Abstract
We report the sequencing of SARS-CoV-2 Omicron variants from 75 patients, using nanopore long-read sequencing chemistry. These data show a range of mutations in spike glycoprotein that are both unique and common to other populations.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Coronavirus disease 2019 (COVID-19) is caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), a betacoronaviridae family member, and has been a primary and urgent concern worldwide [1,2,3]. As of March 4, 2022, over 107 countries had reported infections due to Omicron variants, since the reporting of first case on November 29, 2021 [4]. India saw the first few Omicron cases originating in the state of Karnataka on December 1, 2021 [5], with Delhi reporting a case later from a Tanzania returnee [6]. In this study, we sought to sequence all COVID-19 samples including Omicron variants that were reported in our tertiary care to gain further insights into the mutations occurring in this SARS-CoV-2 variant.
2 Methods
Nasopharyngeal swab samples were collected from 75 patients with a travel history of Africa/Middle East. Here, we randomly analysed samples from 10 representative patients who presented with mild symptoms (fever, cold, cough, sore throat and mild weakness) within 3 days of onset of infection and prior to hospitalization. The samples were used as an input for the ARTIC network “Midnight” protocol (Fig. 15.1) for PCR tiling of SARS-CoV-2, including sequencing with Oxford Nanopore Technologies (ONT) long-read whole-genome sequencing (Rapid Barcoding Kit 96/SQL-RBK-110-96) [7, 8].
Midnight workflow for preparation of SARS-CoV-2 whole-genome sequencing. This method was similar to the ARTIC amplicon sequencing protocol for MinION for SARS-CoV-2 v3 (LoCost) by Josh Quick and the method used in Freed et al. [8]
3 Results and Discussion
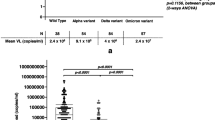
ONT sequencing yielded an average of 25 million reads from all 10 samples, spanning 96.28% of the SARS-CoV-2 genome (20× coverage depth) (Table 15.1). To check the transmissibility associated with the number of mutations in the spike glycoprotein associated with receptor-binding domain (RBD), we compared the 44 common mutations from our samples with the recently emerging mutations of Omicron. Our preliminary analysis indicated that the Omicron variant subcladed with the dominant Delta variant and might have evolved rapidly from multiple mutations (Tables 15.2a, 15.2b, 15.3 and 15.4). A neighbourhood joining tree was constructed using Clustal Omega with the sequences sorted vertically, thereby drawing a circular and unrooted tree (Fig. 15.2a) [9]. We observed that the Indian Omicron variants were clustered together with a root emerging from OL815455, the variant that was first detected from Botswana. The iTOL containing the 75 sequenced samples and Wuhan reference yielded distinct clades in both unrooted and rooted circular tree (data not shown) and the four samples that were claded separately suggested that these were among the first suspected Omicron cases in India (Fig. 15.2a) [10,11,12]. We obtained p.Thr614Ile, p.Thr1822Ile, p.Thr6098Ile and p.Asp155Tyr from LNHD9, p.Ala701Val and p.Val1887Ile from LNHD8 and p.Gly667Ser from LNHD1. However, our preliminary observations indicated that none of these are known to confer detrimental properties to the spike (e.g. changes in transmissibility, severity or immune evasion). Mutations in the spike proteins (Fig. 15.2b(i–iii)) of SARS-CoV-2 variants of concern have also been compared to the parental SARS-CoV-2 isolate B.1 suggesting that the amino acid substitutions are already found in altered positions but with distinct substitutions (Supplementary Tables 15.1 and 15.2).
(a) Circular phylogenetic tree of all 75 samples from India claded with the Wuhan reference genome. The unrooted tree shows a clear dissection of Wuhan from other lineages. All LNHD accessions are labelled. In the Indian sub-population, spike mutations (n = 35) were seen with the nearest residue if in loop/termini region (A67V, V70I(69), T95I, G142V, Y145H(143), N211I, L212I, G339D, R346K, S371L, S373P, S375F, K417N, N440K, G446S, S477N, T478K, E484A, Q493R, G496S, Q498R, N501Y, Y505H T547K, D614G, H655Y, N679K(674), P681H(674), A701V, N764K, D796Y, N856K, Q954H, N969K and L981F). (b) (i) Spike glycoprotein (PDB: 6acc, EM 3.6 Angstrom) with RBD in down conformation. (ii) Multi-Venn diagram of three samples LNHD1, LNHD8 and LNHD9 showing unique and common mutations to all the LNHD series. (iii) Spike glycoprotein (PDB: 6acj, EM 4.2 Angstrom) in complex with host cell receptor ACE2 (green ribbon). (Also see links to Supplementary Tables 15.1 and 15.2)
The limitation of our study is that although the adopted ARTIC sequencing protocol allowed the confirmation of SARS-CoV-2 infections, we did not carry out analyses to determine the probable structural impact of mutations on binding of antibodies produced by existing vaccines or previous SARS-CoV-2 infections, as described by Kannan et al. [13].
In conclusion, our study has demonstrated the utility of nanopore sequencing for SARS-CoV-2 genomes from clinical specimens. We firmly hope that prompt diagnosis and rapid whole-genome analysis would allow a decisive response to the SARS-CoV-2 outbreak that will bring disease control and prevention efforts.
References
Zheng J (2020) SARS-CoV-2: an Emerging Coronavirus that Causes a Global Threat. Int J Biol Sci 16(10):1678–1685
Kaur A, Chopra M, Bhushan M, et al (2021) The Omic Insights on Unfolding Saga of COVID-19. Front Immunol 12:724914. https://doi.org/10.3389/fimmu.2021.724914
Kupferschmidt K, Vogel G (2021) How bad is omicron? Some clues are emerging. Science 374(6573):1304–1305
Bai Y, Du Z, Xu M, et al (2021) International risk of SARS-CoV-2 omicron variant importations originating in South Africa. J Travel Med. 2022 Sep 17;29(6):taac073. https://doi.org/10.1093/jtm/taac073
Hindustan Times; India’s first Omicron cases detected in Karnataka. https://www.hindustantimes.com/india-news/indias-firstomicron-cases-detected-in-karnataka-101638445884205.html. Accessed December 02, 2021
The Hindu; The Hindu Delhi reports first case of Omicron variant. https://www.thehindu.com/news/national/delhi-reports-first-case-of-omicronvariant/article37849305.ece. Accessed December 05, 2021.
Tyson JR, James P, Stoddart D, et al (2020) Improvements to the ARTIC multiplex PCR method for SARS-CoV-2 genome sequencing using nanopore. bioRxiv. https://doi.org/10.1101/2020.09.04.283077
Freed NE, Vlková M, Faisal MB, Silander OK (2020) Rapid and inexpensive whole-genome sequencing of SARS-CoV-2 using 1200 bp tiled amplicons and Oxford Nanopore Rapid Barcoding. Biol Methods Protoc 5(1):bpaa014. https://doi.org/10.1093/biomethods/bpaa014
Sievers F, Wilm A, Dineen D, et al (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539. https://doi.org/10.1038/msb.2011.75
Cov-lineages.org. https://cov-lineages.org/lineage_list.html
Letunic I, Bork P (2021) Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation, Nucleic Acids Res 49(W1):W293–W296
Elbe S, Buckland-Merrett G (2017) Data, disease and diplomacy: GISAID’s innovative contribution to global health. Glob Chall 1(1):33–46
Kannan SA, Austin N. Spratt, et al (2022) Omicron SARS-CoV-2 variant: Unique features and their impact on pre-existing antibodies. J Autoimmun 126:102779. https://doi.org/10.1016/j.jaut.2021.102779
Acknowledgements
The authors gratefully acknowledge Government of Delhi, INSACOG and Ethics Committee of Maulana Azad Medical College, Delhi, India. The consultant’s physicians and laboratory staff members provided initial diagnostic testing of the SARS-CoV-2 samples.
Ethics Statement
Informed consent was judiciously taken before the sample was sequenced. The Institutional Ethics Committee of Maulana Azad Medical College, Delhi, India, had given approval for the study (F.1/IEC/MAMC/85/03/2021).
Data Availability
All Omicron variant samples have been uploaded to NCBI-GenBank with accession IDs ON063241–ON063253 (https://www.ncbi.nlm.nih.gov/nuccore/?term=ON063241:ON063253[accn]) and ON060006–ON060067 (https://www.ncbi.nlm.nih.gov/nuccore/?term=ON060006:ON060067[accn]). The same have also been submitted to GISAID.org with hCoV-19/India/un-LNHDXX/2021 series
-
EPI_ISL_7864703
-
EPI_ISL_7876997
-
EPI_ISL_7877026
-
EPI_ISL_7877006
-
EPI_ISL_7877093
-
EPI_ISL_7877115
-
EPI_ISL_7877191
-
EPI_ISL_7877201
-
EPI_ISL_7877202
-
EPI_ISL_7877203
-
EPI_ISL_7877297
-
EPI_ISL_7889640
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic Supplementary Material
Supplementary information and the INSACOG and SCOG MAMC LNH author list are available at https://zenodo.org/record/6452357#.YlWHbXVBy-o
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Kumar, S. et al. (2023). Amplicon-Based Nanopore Sequencing of Patients Infected by the SARS-CoV-2 Omicron (B.1.1.529) Variant in India. In: Guest , P.C. (eds) Application of Omic Techniques to Identify New Biomarkers and Drug Targets for COVID-19. Advances in Experimental Medicine and Biology(), vol 1412. Springer, Cham. https://doi.org/10.1007/978-3-031-28012-2_15
Download citation
DOI: https://doi.org/10.1007/978-3-031-28012-2_15
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-28011-5
Online ISBN: 978-3-031-28012-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)