Abstract
Acerola (Malpighia emarginata DC) is an exotic fruit that has a high agro-industrial potential. It is known to be rich in ascorbic acid, phenolic compounds, and carotenoid pigments. These nutrients make acerola one of the best sources of natural antioxidants, helping to prevent many conditions and delay aging. Acerola fruit is transformed into concentrate juice then powder to be incorporated into nutritional supplements. The natural ascorbic acid content of juice powders must be between 16 and 17%. Unfortunately, the origin of ascorbic acid in acerola-based products is not always natural. That is to say, some food manufacturers add synthetic ascorbic acid to reach the recommended values (16 to 17%), which can be considered as a falsification of the product. Since a decade, the control of the life cycle and the quality of foodstuffs is an increasingly important concern. In this context, EVEAR Extraction (French company) establishes a high level of traceability of its extracts by combining sourcing, extraction processes and laboratory controls throughout the production process. The determination of the composition of raw material and final products can be determined by spectrometric analysis and more precisely by Nuclear Magnetic Resonance (NMR) spectroscopy. However, spectral analysis remains a tedious and time-consuming task requiring an expert.
In this study, the feasibility of discriminating acerola-based product was investigated using 1H NMR spectroscopy in combination with a supervised classification procedure consisting of several steps: principal component analysis (PCA), a fast Fourier transform (FFT) and a neuronal network classification. A total of 6 classes (Colored Acerola powder, Acerola concentrate, Acerola powder, Ascorbic Acid, Acerola with added ascorbic acid, Other extract) were examined. Following the classical approaches, we opted for a convergent network using hidden layers and a divergent output. The results demonstrate that 1H NMR spectroscopy combined with ANN analysis is an effective tool for verifying the nature of Acerola samples.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Acerola (Malpighia glabra L.) is a small tree that grows in dry deciduous forests. It is native to central and northern South America and has been cultivated in large areas of Brazil [4]. Its red fruit, which resembles the European cherry, contains about 80% juice and a large amount of ascorbic acid (vitamin C), but it is also rich in other nutrients such as carotenes, thiamin, riboflavin, niacin, proteins, and mineral salts, mainly iron, calcium and phosphorus [2, 3]. Acerola’s high vitamin C content makes it one of the best sources of natural antioxidants, helping to prevent many diseases and delay aging [5]. Vitamin C is involved in several biological functions, such as enhancing collagen formation [8] and is considered one of the major vitamins required by the human body because of its antioxidant properties [15]. Indeed, increased antioxidant intake has been associated with a lower risk of cardiovascular disease [7]. As a result, acerola concentrate is used in the manufacture of many dietary supplements and their quality depends on the quantity of key active components and the absence of undesirable materials such as adulterants and residual solvents. Claims of benefit depend on the presence of specific molecules in the extracts, which must therefore be identified and quantified with great precision. Recently, NMR spectroscopy has been widely used as a qualitative and a quantitative tool to characterize plant extracts. NMR spectroscopy-based metabolome analyses can be highly effective in identifying and quantifying novel and known metabolites [6, 14, 18, 20]. However, the spectral analysis remains a tedious and time-consuming task requiring an expert. Proton nuclear magnetic resonance (1H NMR) allows to obtain a metabolomic profile of the analyzed sample but does not allow to detect the addition of synthetic ascorbic acid in acerola products. Moreover, the addition of 1 to 2% ascorbic acid changes the metabolomic profile slightly but it is not possible to see this modification with the human eye. Hence the interest of using artificial intelligence. The idea is to train the model with real acerola concentrates, concentrates transformed into powder (without addition of ascorbic acid) and concentrates transformed into powder with addition of ascorbic acid, then query it to classify spectra of unknown products. To date and to the best of our knowledge, No NMR method coupled with artificial intelligence has been implemented for the classification of acerola products according to their composition in order to detect the addition of synthetic ascorbic acid.
The determination of the nature of the extracts can be summarized as a classification problem. Data classification is the process of analyzing structured or unstructured data and organizing them into categories based on the type and content of the signals. There are several types of classification: unsupervised and supervised (Logistic Regression, SVM, etc.) techniques [12]. Among the methods in the literature, the classification proposed in this work is based on Convolutional Neural Networks (CNNs) [1]. Indeed, CNNs have shown their efficiency in the creation of feature maps. These maps are a strong point for NMR spectrum analysis since they are invariant to the small transformation introduced by the measurement.
To detail this approach, the paper is decomposed as follows: first, a state of the art on spectral analysis methods and on neural network classification is proposed. In Sect. 3, we will explain how the data are collected and how our model is made. Section 4, that details the first results of our approach, is followed by a discussion in the conclusion.
2 State of the Art
2.1 Overview
To ensure the authenticity of herbal extracts or herbal products, the process of standardization is a lengthy one, requiring proper sample preparation, time-consuming analytical method development for the resolution of an analyte peak from the complex natural extract, and more importantly, a pure authentic natural product. As a solution to these issues, the creation of a reliable and simple method is needed as an alternative to standard analyses.
Metabolomic profiling is a discipline that focuses on the detailed description of the metabolite composition of herbal extract. Metabolomics thus focuses on the analysis of metabolites that represent the final phenotype of the extracted herb. These metabolites are low molecular weight molecules (molecular weight < 1500 Da) and can be sugars, amino acids or fatty acids, their levels reflect changes in the genome, transcriptome and proteome. Proton nuclear magnetic resonance (1H NMR) is used in this kind of study because it is a highly reproducible technology that offers information about all metabolites in a herbal extract sample that are over the limit of detection. While artificial intelligence has been widely used in the pre-processing of NMR data, peak identification, peak integration, its use in metabolomics is not as developed as it is in other omics domains like as genomics [16]. In this study, artificial intelligence techniques such as artificial neural networks, genetic algorithms, and genetic programming will be applied to metabolomic data.
2.2 Nuclear Magnetic Resonance (NMR)
Nuclear Magnetic Resonance (NMR) spectroscopy has evolved into a powerful tool for metabolomic analysis of plant extract [6]. it’s non destructive, fast, accurate, quantitative and information-rich analytical method. It is a highly repeatable and reproducible method when compared to mass spectrometry. It is possible to compare, distinguish, or classify samples using NMR spectra. However, because of the high level of signal overlap, especially in one-dimensional NMR spectra, this approach has been limited in its application. Indeed, NMR appears to be a good fit for artificial intelligence techniques because of this. The most typical process in NMR data handling is data pre-processing, which involves converting the free induction decay (FID) to a matrix of chemical shift and intensity, baseline correction, normalization and peak alignment [16]. On an NMR spectrum, each metabolite has characteristic peaks whose position is well defined and whose intensity correlates with the amount of this metabolite.
In this study, NMR spectra of acerola samples under different formulations and of other extracts are recorded. The spectra are processed and transformed into a matrix with the chemical shifts on the x-axis and the intensity on the y-axis. The classification of the spectra of large series is generally done using unsupervised (PCA) or supervised statistical (PLS) methods.
2.3 Classification
In machine learning applications, the goal is to train a model (or enable it to learn) from the data it is given, and thus improve its output (results). Machine learning techniques are generally split into two families: supervised and unsupervised learning methods. Unsupervised models learn from unlabelled data: the trained model works on raw data and looks for patterns in the given information. They are very convenient because the input data does not need to be tagged or labelled. However these techniques can be limited when the task is to distinguishing between many complex classes. Supervised models require labelled data and they usually need more computing capacity (and more data). Given the exact information of the expected class for each data (label), the model is able to discern the specificity of two closely related classes since it knows they must be different. However, when using supervised techniques, one must be careful about over-fitting. When the training of the model is not well established (for instance, when the input data is too specific), it can identify false relations that bias the reasoning, resulting in wrong classifications.
The NMR spectra of some molecules are very close and are quite difficult to differentiate for humans. As a result, the use of unsupervised models seems inconsistent with the high similarity of the data. In our case, supervised model seem the most appropriate. The distinction between two close NMR spectrum seems too complicated to be done by the model itself.
In this context, many algorithms for spectral matching have been developed since the 1970s [9, 11]. Most of these algorithms were developed for mass spectrometry. Their applications quickly extended to vibrational spectroscopy. These conventional spectral matching approaches are iterative techniques. They are based on the identification of the largest similarities between the unknown spectrum and the reference spectrum. These approaches document the tools and techniques that are used to automate this classification through AI. The first results of 1D CNN for spectral data analysis [21] revealed the potential of using machine learning methods for spectral analysis, from classification of a substance to identification of components in a mixture in various scientific fields [13]. The majority of recent publications use 1D CNNs for various spectral applications.
The first results of 1D CNN for spectral data analysis [21] revealed the potential of using machine learning methods for spectral analysis, from classification of a substance to identification of components in a mixture in various scientific fields [13]. The majority of recent publications use 1D CNNs for various spectral applications. However, a few studies have highlighted the need for further research in order to address the problem of not having enough samples compared to the number of features.
3 Material and Methods
3.1 Global Overview
This section provides a description of the dataset used, containing NMR signal recordings of the different samples to be analyzed, as well as the extracted features used to generate the classification model. Then, the methodology used all along the experimentation is described. The design and implementation of the ANN presented in this work relies on Python programming language and on Keras and Tensorflow libraries, that are among the most used for deep learning applications. The key steps in the proposed classification process for sample tractability are: data preparation, network construction, training and evaluation.
3.2 Data
1D 1H NMR spectrum was acquired for each sample. Spectra were acquired on a Bruker Advance-400MHz spectrometer using a 5mm broad-band probe tuned to detect 1H resonances at 400.15 MHz. Data were collected without sample rotation at 300K, as à 64K complex points using a noesygppr pulse sequence with 90\(^\circ \) pulse length and pre-saturation to remove the residual water signal. The number of scan was set at 16. The receiver gain was set to 90.5 and the spectral width was fixed to 20ppm. Obtaind FID were converted to spectra using Topspin 3.5 software. Spectra were processed (phase correction, baseline correction) and the signal of TSP set at 0 ppm was used as an internal reference for chemical shift measurement. Finally, spectra were converted to csv files using Mnova software. The csv files contain the chemical shifts on the x-axis and the corresponding intensities on the y-axis.
The data are distributed in the following way (Table 1).
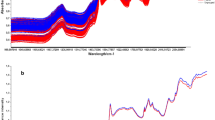
The following figure (Fig. 1) show two spectra that are visually close but belong to two different classes.
3.3 Model
When exploring existing work in Sect. 2, we identified two techniques that could be relevant for our classification problem:
Deep Neural Network (DNNs) are fully connected networks, all neurons in each neuron in each layer are connected to all neurons in the following layer. The input of a neuron is therefore equal to the number of neurons in the previous layer. The classic DNN structure consists of an input layer, several hidden layers and an output layer. When data enters the input layer, the output values are calculated layer by layer in the network. In each hidden layer, after receiving a vector consisting of the output values of each neuron in the previous layer, it is multiplied by the weights associated with each neuron in the current layer to obtain the weighted sum. The activation function of a neuron is specific to its layer. The functions are chosen according to the type of problem. Sigmoid functions are non-linear as opposed to Rectified Linear Unit (ReLu) for example.
Convolutional Neural Networks (CNNs) are a particular architecture of deep networks [10]. They are designed to process data from several sources: 1D for sequences, 2D for images and 3D for videos [17]. They are very suitable for shape or pattern recognition while being insensitive to scale factor, rotation, etc. In opposition to DNNs, convolution networks are not strongly connected. The hidden layers are separated by layers acting like filter. These filters represent weights and biases. In general, the basic structure of CNNs consists of convolution layers, nonlinear layers and pooling layers. To avoid combinatorial explosion, all neurons in a convolution layer share the same filter, i.e. the same weights and biases, in order to reduce the number of training parameters. As with DNNs, the outputs of these filters are then passed through nonlinear layers that typically use the ReLU function. The role of pooling layers is to aggregate semantically similar features to identify complex features by creating maximal or average subsamples in the feature maps. Sometimes pooling layers are also used to avoid network overfitting and improve model generalization. Given the excellent ability of CNNs to analyze spatial information, they can be applied to NMR spectra reconstruction, denoising, and chemical shift prediction. A convolution network is more appropriate in our case of study because it will allow us to build a feature map corresponding to the different parts of the NMR spectrum.
When using Keras library to implement an ANN in Python, it is necessary to specify the type of model to be created. There are two ways to define Keras models: Sequential and Functional. A sequential model refers to the fact that the output of each layer is taken as the input to the next layer, and this is the type of model developed in this work. The objective is to build a feature map representing the different metabolites characteristic of a class of plant extract (Fig. 2).
The built network has 7 layers. The input layer is composed of as many neurons as there are points on the spectrum. It has been reduced to 1300 datas thanks to Evear’s expertise. The following layers reduce little by little the vector to propose an output vector of 6 neurons, as summarized in Table 2.
A network is associated with several metrics:
-
Optimization algorithm: The ANN will use an optimization algorithm to calculate the weight of each neuron. There are several ways to do this. The most common one is to minimize (or maximize) an objective function E(X) which is a mathematical function depending on the internal training parameters of the model where X are features. The result of E(X) is used to compute the objective values Y of the set of training parameters where Y ars labes. The most commonly used optimization algorithms in ANNs are gradient descent. In our case, we use a classical backpropagation proposed by Tensorflow.
-
Loss function: The loss function, also known as the cost function, is a function that measures the quality of the network response. A high result indicates that the ANN is performing poorly and a low result indicates that the ANN is performing positively. This is the function that is optimized or minimized when backpropagation is performed. There are several mathematical functions that can be used, the choice of one of them depends on the problem to be solved. The most suitable function for classification is the cross-entropy function. The cross-entropy loss, or log loss, measures the performance of a classification model whose output, Y, is a probability value P, between 0 and 1, and is calculated using the following equation (Eq. 1). The cross-entropy loss increases as the predicted probability deviates from the actual label. This function is used for classification problems.
$$\begin{aligned} -(y log(p)+(1-y)log(1-p)) \end{aligned}$$(1)
4 Results
Our dataset contains a set of 8960 spectra distributed as describe in Table 1. Following a similar procedure as in traditional approaches [19], the dataset was randomized and then separated into two samples: Training 80% and Validation 20%. The learning (training) phase is a backpropagation repeated thirty times.
The indicators of the training phase are presented in this section. Reviewing learning curves of models during training can help to diagnose problems with learning, such as an underfit or overfit model, and if the training and validation datasets are suitably representative.
We first observe the accuracy of the model (how well it is able to guess the expected class for a given input spectra). This learning curve is calculated from a hold-out validation dataset that gives an idea of how well the model is generalizing.
As illustrated in Fig. 3(a), the accuracy increases with the repetition of the training batch. At the end of this step, we can see that our network reaches an accuracy level of 93% on the validation dataset. On samples that are not part of the original dataset, we observed similar results. The loss curve, depicted in Fig. 1(b), shows that our network is learning well and is converging to an optimum. There would not be much benefit in extending the learning phase.
5 Conclusion and Future Work Direction
The aim of this work was to create a classification model of acerola samples using an NMR spectrum as a data source. Once the data was made usable by the classical techniques of chemistry, we could use it to ensure a better traceability. The advantage of this model is to save time and precision to the Evear-extraction team. Through the introduction of a CNNs, we were able to define and train a model capable of meeting our expectations. The training game was discriminating enough to properly train the 7 layers of our CNN. However, from an industrial point of view, it will be necessary to increase the capabilities of the POC to take into account more complex plant extracts. However, from an industrial point of view, it will be necessary to increase the capabilities of the POC to take into account more complex plant extracts.
However, this work is only a proof of concept. The proposed classification method of plant extracts could be significantly improved by exploring the following directions:
-
using different (and more) classes for the network. Indeed, the acerola is only one example of the plants to be traced. For this, it will be necessary to increase the precision of the network and rework its topology.
-
increasing the reliability of the traceability by being able to define the metabolites and their quantity present in a spectrum.
-
strengthening our knowledge of the spectrum by analyzing the parameters described in the architecture of the neural network or using feature selection techniques for the dataset.
References
Aghdam, H.H., Heravi, E.J.: Guide to convolutional neural networks. New York NY: Springer 10(978–973), 51 (2017)
Albertino, A., Barge, A., Cravotto, G., Genzini, L., Gobetto, R., Vincenti, M.: Natural origin of ascorbic acid: validation by 13C NMR and IRMS. Food Chem. 112(3), 715–720 (2009)
Alves Filho, E., Silva, L.M., Canuto, K.: Metabolomic profiling of acerola clones according to the ripening stage. Food Mesure 15, 416–424 (2021)
Anand, P., Revathy, B.: Acerola, an untapped functional superfruit: a review on latest frontiers. J. Food Sci. Technol. 55, 3373–3384 (2018)
Belwal, T., et al.: Phytopharmacology of acerola (Malpighia spp.) and its potential as functional food. Trends Food Sci. Technol. 74, 99–106 (2018)
Deborde, C., Moing, A., Roch, L., Jacob, D., Rolin, D., Giraudeau, P.: Plant metabolism as studied by NMR spectroscopy. Prog. Nucl. Magn. Reson. Spectrosc. 102–103, 61–97 (2017)
Ellingsen, I., Seljeflot, I., Arnesen, H., Tonstad, S.: Vitamin c consumption is associated with less progression in carotid intima media thickness in elderly men: a 3-year intervention study. Nutr. Metab. Cardiovasc. Dis. 19, 8–14 (2009)
Findik, R., Ilkaya, F., Guresci, S., Guzel, H., Karabulut, S., Karakaya, J.: Effect of vitamin C on collagen structure of cardinal and uterosacral ligaments during pregnancy. Eur. J. Obstet. Gynecol. Reproductive Biol. 201, 31–35 (2016)
Grotch, S.L.: Matching of mass spectra when peak height is encoded to one bit. Anal. Chem. 42(11), 1214–1222 (1970)
Gu, J., et al.: Recent advances in convolutional neural networks. Pattern Recogn. 77, 354–377 (2018)
Knock, B., Smith, I., Wright, D., Ridley, R., Kelly, W.: Compound identification by computer matching of low resolution mass spectra. Anal. Chem. 42(13), 1516–1520 (1970)
Lorena, A.C., Garcia, L.P., Lehmann, J., Souto, M.C., Ho, T.K.: How complex is your classification problem? A survey on measuring classification complexity. ACM Comput. Surv. (CSUR) 52(5), 1–34 (2019)
Lussier, F., Thibault, V., Charron, B., Wallace, G.Q., Masson, J.F.: Deep learning and artificial intelligence methods for Raman and surface-enhanced Raman scattering. TrAC Trends Anal. Chem. 124, 115796 (2020)
Pauli, G., Jaki, B., Lankin, D.: Quantitative 1 h NMR: development and potential of a method for natural products analysis. J. Nat. Prod. 68, 133–149 (2005)
Podmore, I., Griffiths, H., Herbert, K., Mistry, N., Mistry, P., Lunec, J.: Vitamin c exhibits pro-oxidant properties. Nature 392, 559 (1998)
Pomyen, Y., Wanichthanarak, K., Poungsombat, P., Fahrmann, J., Grapov, D., Khoomrung, S.: Deep metabolome: applications of deep learning in metabolomics. Comput. Struct. Biotechnol. J. 18, 2818–2825 (2020)
Shamsaldin, A.S., Fattah, P., Rashid, T.A., Al-Salihi, N.K.: A study of the convolutional neural networks applications. UKH J. Sci. Eng. 3(2), 31–40 (2019)
Smolinska, A., Blanchet, L., Buydens, L., Wijmenga, S.: NMR and pattern recognition methods in metabolomics: from data acquisition to biomarker discovery: a review. Anal. Chim. Acta 750, 82–97 (2012)
Tan, J., Yang, J., Wu, S., Chen, G., Zhao, J.: A critical look at the current train/test split in machine learning. arXiv preprint arXiv:2106.04525 (2021)
Ward, J., Baker, J., Beale, M.: Recent applications of NMR spectroscopy in plant metabolomics. FEBS J. 274, 1126–1131 (2007)
Yang, J., Xu, J., Zhang, X., Wu, C., Lin, T., Ying, Y.: Deep learning for vibrational spectral analysis: recent progress and a practical guide. Anal. Chim. Acta 1081, 6–17 (2019)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 IFIP International Federation for Information Processing
About this paper
Cite this paper
Combis, L. et al. (2023). NMR Spectroscopy and Learning-Based Classification of Acerola Samples. In: Noël, F., Nyffenegger, F., Rivest, L., Bouras, A. (eds) Product Lifecycle Management. PLM in Transition Times: The Place of Humans and Transformative Technologies. PLM 2022. IFIP Advances in Information and Communication Technology, vol 667. Springer, Cham. https://doi.org/10.1007/978-3-031-25182-5_32
Download citation
DOI: https://doi.org/10.1007/978-3-031-25182-5_32
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-25181-8
Online ISBN: 978-3-031-25182-5
eBook Packages: Computer ScienceComputer Science (R0)