Abstract
Lymphoma is a cancer of the lymphatic system, and it can affect many organs throughout the body. Positron emission tomography (PET)/computed tomography (CT) are primary imaging methods to assess lymphoma types and monitor their treatment, where PET is sensitive to identify lymphoma regions while CT preserves anatomical structures. Combining PET and CT is thus useful for lymphoma segmentation because it helps to identify lymphoma types and evaluate treatment effects. However, lymphoma segmentation suffers many challenges, including substantial lymphoma size and shape variance, numerous types, limited PET/CT data for lymphoma, and similar PET signals with adjacent organs. To address these challenges, we integrate label guidance, patch sampling, and negative data augmentation to achieve multi-modal lymphoma segmentation. The training data consist of positive and negative patch samples. These samples are purposely extracted from the original scans with the guidance of lymphoma labels. Negative samples are further supplemented from the PET/CT scans of non-lymphoma patients to better discriminate lymphoma from adjacent organs. The proposed method was validated on the PET/CT scans from 28 patients. Experimental results revealed that the Dice coefficient was improved from 0.11 to 0.43 in comparison with a baseline method the 3D-residual U-Net method. Patch-based strategy is also computational undemanding. These results suggest that the proposed method could be an efficient means to segment lymphoma and possibly used for identifying lymphoma types and assessing their treatment.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Lymphoma is a hematopoietic malignancy with numerous types, and it can affect people of all ages [1]. Lymphoma treatment response is highly dependent on the measurement of tumor burden, which often requires accurate identification of lymphoma regions. Positron emission tomography (PET)/computed tomography (CT) [2, 4, 8] are primary imaging methods to assess lymphoma and monitor treatment response. Figure 1 illustrates an example of PET/CT scans on lymphoma patient. Organs such as kidney and liver are well depicted, but lymphoma is difficult to identify in the CT scan (Left Image). In contrast, the standardized uptake value (SUV) is used to measure fluorodeoxyglucose positron emission tomography uptake or glucose metabolism of the tumor regions in the PET scan. For this reason, lymphoma is visually represented as bright regions in the PET scan (yellow arrows) while organs are hard to delineate. These observations motivate us to develop a multi-modal lymphoma segmentation method as it is useful for lymphoma treatment.
Example of lymphoma distributed on the paired PET-CT scans. Left column: organs are preserved in the CT scan; Center column: lymphoma is highlighted with bright regions in the PET scan (yellow arrows), in which it is randomly spread to the whole body; Right column: PET scan with overlayed lymphoma labels, from which we can observe that lymphomas are outside organs (a1, a2), inside organs (b1, b2), small spots (c1, c2). Kidneys (d1, d2) and bladder (e1, e2) could also have bright normal regions similar to lymphoma. All these challenges are attributed to the difficulty of lymphoma segmentation.
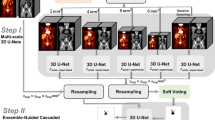
However, lymphoma segmentation is a challenging task because it can randomly spread throughout the body (Fig. 1). It could be either outside organs (a1, a2) or inside organs (b1, b2). Lymphoma also has a wide range of shapes and sizes, such as tiny spots (c1, c2). High SUV values at kidneys (d1, d2) and bladder (e1, e2) are also similar to those at lymphoma. All these difficulties hinder lymphoma segmentation, and only a limited number of methods have been developed for lymphoma segmentation. An ensemble model from DeepMedic was developed for pediatric lymphoma PET/CT scans [11]. DenseX-Net was also developed to segment lymphoma on the whole-body PET/CT scans [7]. However, the input of these methods is 2D slice, which potentially lose the spatial coherence among slices. Another 3D segmentation method based on the belief function was used to segment lymphoma [3], which integrated a feature extraction module and an evidential segmentation (ES) module. Although it achieved decent segmentation results, it has not considered the multi-scale and patch-based framework to further extract the useful information from the details of the PET and CT scans. This paper aims to develop a deep learning-based approach to segment lymphoma on multimodal PET/CT scans. Our method combines Label guided Patch sampling for Multi-Modalities, and negative sample augmentation (LPMM-nsa) to serve the segmentation purpose. The training data are composed of a set of local image patches, and positive (green boxes, Fig. 2), and negative patches (red boxes, Fig. 2) are extracted according to the likelihood of lymphoma regions or non-lymphoma regions. In other words, positive samples are more likely from the lymphoma regions and negative samples are from non-lymphoma regions, which could help to create high-quality data for training. Negative samples are further enhanced from PET/CT scans of non-lymphoma patientsFootnote 1 to better discriminate the lymphoma from organs. Since our method is patch based, the proposed method is naturally computationally undemanding and GPU memory efficient, which is suitable for clinical applications. A validation dataset with 28 lymphoma patients is also created to evaluate the segmentation accuracy, in which the lymphoma size changes drastically, and they are more close to the real clinical practice.
2 Method
For demonstration of the effectiveness of the proposed methods, we choose to use the widely validated 3D-residual U-Net as the back-bone structure to develop our own modules (Fig. 2). It is noted that the proposed methods can be extended to other more advanced structure in simple plug-in fashion such as [3].
2.1 Notation and Formation
Let us first give some notations and formations to improve readability. Multi-modal PET/CT dataset \(\{ \textbf{X}_i \}^n_{i=1}\) where \(\textbf{X}_i\) is a sample in the dataset, which channel-wise concatenates the PET and CT modalities. Training and testing datasets are defined as \(\{ \textbf{X}_i^t \}^k_{i=1}\) and \(\{ \textbf{X}_i^v \}^l_{i=1}\) respectively. Their corresponding labels are, thus, given as \(\{ \textbf{Y}_i^t \}^k_{i=1}\) and \(\{ \textbf{Y}_i^v \}^l_{i=1}\). Each element y in \(\textbf{Y}_i^t\) and \(\textbf{Y}_i^v\) belongs to the set \(\{0,1\}\) denoting lymphoma and non-lymphoma regions, respectively. Let us denote the 3D residual U-Net as \(\textbf{E}\) and the loss function as \(\mathcal {L}\). Then the classic segmentation framework is as follows:
where \(\theta (\textbf{E})\) stands for the trainable parameters of the network and \(\theta ^*\) is the optimized parameters of the network.
2.2 Label-Guided Patch Sampling
To further extract the useful information from the details of the PET/CT scans and decrease the computing resource demanding issue of the huge 3D volumetric data, we utilize the strategies of label-guided patch sampling. Training samples are extracted based on the guidance of data label. The sampling function \({\text {S()}}\) extracts patches non-homogeneously in terms of probability of the presence of lymphoma guided by the label map \(\textbf{Y}_i^t\) because lymphoma regions and their adjacent regions should be highlighted. The sampling process is thus given by:
where the \(\hat{\textbf{X}}^t_{ij}\) represents the j-th patch sampled from the scan dataset \(\textbf{X}^t_i\). m indicates the number of patches. The \(\mathbf {\textbf{f}}\) Bernoulli distribution to sample the images from the lymphoma and non-lymphoma regions according to the \(\textbf{Y}_i^t\):
where the probability p is set as 0.6 in this work, and \(y=1\) if it is a positive sample and \(y=0\) if it is a negative sample. Using patch sampling leads (1) to:
where \(\hat{\textbf{Y}}^t_{ij}\) is the patch label of \(\hat{\textbf{X}}^t_{ij}\).
2.3 Negative Sample Augmentation
Negative sample augmentation is another efficient strategy to enhance the training data. PET/CT scans of the non-lymphoma patients were used for discriminating organs from lymphoma. Therefore, we denote the extra patch dataset as \((\{\hat{\textbf{X}}^e_{ij}\}_{j=1}^s, \{ \textbf{0} \}_{j=1}^s)\), where the negative patches \(\{\hat{\textbf{X}}^e_{ij}\}_{j=1}^s\) are randomly sampled from the extra negative samples \(\{ \textbf{X}_i^e \}^q_{i=1}\). The symbol \(\textbf{0}\) means the all 0 label tensor for the negative sample \(\hat{\textbf{X}}^e_{ij}\). Finally, our framework is expressed as:
Optimizing (5) yields the trained network \(\textbf{E}^*\). During inference, the prediction of the patches of a PET/CT scans is given by \(\hat{\textbf{Y}}^*_{ij} = \textbf{E}^*(\hat{\textbf{X}}^v_{ij})\). These predicted patches are stitched based on the aggregation function:
where the \(\alpha \) represents the parameters for the patch aggregation, which includes patch size, overlap margin, and \(\textbf{Y}^*_i\) is the final lymphoma segmentation.
The ADAM optimizer [6] with weight decay is used for training. The learning rate is set to \(10^{{-3}}\). The proposed method is implemented in PyTorch. Both training and testing are performed on the Nvidia DGX station equipped with a Tesla A100 graphics card with 40 GB GPU memory.
2.4 Data Collection and Validation Methods
Twenty-two lymphoma patients underwent whole body (WB) PET and CT examinations between 2010–2021 were collected, and Research Consortium for Medical Image Analysis (RECOMIA) AI tool was used to initially label the lymphoma regions. These labeled results were then reviewed and manually corrected by an experienced radiology residence. Labeled results were eventually confirmed by a nuclear medicine physician, which generates our lymphoma labels.
All twenty-two PET/CT scans were resampled to \(500\times 500\times 850\) pixels through cropping and padding operations. The intensity is normalized to [0, 1] using the window range of \([-1000, 800]\) on the CT scan and the SUV window range of [0, 40] inspired by [10] on PET scan. We empirically set the patch size as \(64\times 64\times 64\). For the SUV computation, we use the SUV normalized by the body weight (SUVbw) [5]Footnote 2. More specifically, the computation method is listed as follows:
PET image pixels and injected dose are decay corrected to the start of scan. After the conversion, the pet image pixels are in units (g/ml). Three metrics are used for validation, including Dice score (dsc), average symmetric surface distance (assd), and sensitivity. Two experiments were conducted in this work. The first is the comparison between the baseline 3D residual U-Net and the proposed method with different settings, including the proposed label guided patch sampling for multi-modal data (LPMM), and its further improved version with negative sample augmentation (LPMM-nsa). The second is the comparison among single modality (PET or CT only) and multi-modal (PET/CT).
3 Experiments
The comparison of segmentation results using different methods is reported in Table 1. It reveals that the combination of label guidance, patch sampling and negative data augmentation (LPMM-nsa) achieves the highest segmentation accuracy. It also suggested that the input of multi-modal data is another key component to improve the segmentation accuracy as the dice-coefficient is only 0.11 using PET scan only. In contrast, the proposed method can achieve 0.43.
Comparison results in Fig. 3 also supported these findings because the baseline 3D residual U-Net is prone to over-segmenting lymphoma (second row). In contrast, over-segmentation is substantially reduced after using label guidance and patch sampling. However, it could induce the issue of under-segmentation, which was further improved by the addition of negative sample augmentation. Figure 4 also proves the importance of multi-modal input. All lymphomas are missed from the model trained on CT scans only because lymphoma is non-trivial to identify on CT scans. Some lymphoma were segmented using the model with PET scans only, and the segmentation results were vastly improved with both modalities. Since CT and PET concentrate on the different parts of lymphoma patients, they might contribute to each other to more accurately identify lymphoma regions.
To further demonstrate the advantage of our methods, we illustrate several results from each multi-modal method in details in Fig. 3 from axial.
Comparison of lymphoma segmentation results using different segmentation models. The three columns show slices from three scans respectively. First row: ground truth; second row: segmentation results from the baseline 3D-residual U-Net method where lymphomas are over-segmented; third row: results from the segmentation model enhanced with label guided patch sampling where over-segmentation is substantially reduced but with some lymphoma under-segmentated; fourth row: results with the addition of negative sample augmentation, in which lymphomas are accurately segmented.
Comparison of lymphoma segmentation using different image modalities. The three columns show slices from three scans respectively. First row: ground truth, second row: segmentation results using CT only, third row: results using PET only, and fourth row: results using both CT and PET. No lymphomas are segmented on CT scans only, and some lymphomas are found in the results with PET only. Almost all lymphomas are segmented using both modalities
4 Conclusion and Future Work
In this paper, we developed a multi-modal lymphoma segmentation method on PET/CT scans. Three key components were integrated to improve the segmentation accuracy, including label guidance, patch sampling, and negative sample augmentation. Label guidance helps to create effective training samples that are more focused on both lymphoma and non-lymphoma regions. Patch samples not only reduces computational cost, but also avoid over-segmentation from the baseline 3D residual U-Net (third row, Fig. 3). Negative sample augmentation could further reduce the issue of under-segmentation raised by path sampling (fourth row, Fig. 3). Comparing with the segmentation models from single modal, multi-modal is another important property to the segmentation accuracy (Fig. 4. The validation results in Table 1 also proved that the proposed method utilized all effective means to achieve the highest segmentation accuracy.
In the future, we would like to explore more about the multiple modality fusion methods, such as graph-based methods [9], multimodal transformers [12], as well as incorporate additional modal of clinical reports to continuously improve segmentation accuracy. Nevertheless, the proposed method shows the promising results to accurately segment lymphoma on PET/CT scans.
References
A predictive model for aggressive Non-Hodgkin’s lymphoma. N. Engl. J. Med. 329(14), 987–994 (1993). https://doi.org/10.1056/NEJM199309303291402
Czernin, J., Allen-Auerbach, M., Nathanson, D., Herrmann, K.: PET/CT in oncology: current status and perspectives. Curr. Radiol. Rep. 1(3), 177–190 (2013)
Huang, L., Ruan, S., Decazes, P., Denœux, T.: Evidential segmentation of 3D PET/CT images. In: Denœux, T., Lefèvre, E., Liu, Z., Pichon, F. (eds.) BELIEF 2021. LNCS (LNAI), vol. 12915, pp. 159–167. Springer, Cham (2021). https://doi.org/10.1007/978-3-030-88601-1_16
Juweid, M.E., Cheson, B.D.: Positron-emission tomography and assessment of cancer therapy. N. Engl. J. Med. 354(5), 496–507 (2006)
Kim, C.K., Gupta, N.C., Chandramouli, B., Alavi, A.: Standardized uptake values of FDG: body surface area correction is preferable to body weight correction. J. Nucl. Med. 35(1), 164–167 (1994)
Kingma, D.P., Ba, J.: Adam: a method for stochastic optimization. In: Bengio, Y., LeCun, Y. (eds.) 3rd International Conference on Learning Representations, ICLR 2015, San Diego, CA, USA, 7–9 May 2015, Conference Track Proceedings (2015). http://arxiv.org/abs/1412.6980
Li, H., et al.: DenseX-Net: an end-to-end model for lymphoma segmentation in whole-body PET/CT images. IEEE Access 8, 8004–8018 (2019)
Li, J., Xiao, Y.: Application of FDG-PET/CT in radiation oncology. Front. Oncol. 3, 80 (2013)
Liu, L., Nie, F., Wiliem, A., Li, Z., Zhang, T., Lovell, B.C.: Multi-modal joint clustering with application for unsupervised attribute discovery. IEEE Trans. Image Process. 27(9), 4345–4356 (2018)
Noy, A., et al.: The majority of transformed lymphomas have high standardized uptake values (SUVs) on positron emission tomography (PET) scanning similar to diffuse large b-cell lymphoma (DLBCL). Ann. Oncol. 20(3), 508–512 (2009)
Weisman, A.J., et al.: Automated quantification of baseline imaging pet metrics on FDG PET/CT images of pediatric Hodgkin lymphoma patients. EJNMMI Phys. 7(1), 1–12 (2020)
Xu, P., Zhu, X., Clifton, D.A.: Multimodal learning with transformers: a survey. arXiv preprint arXiv:2206.06488 (2022)
Acknowledgements
This research was supported by the National Institutes of Health, Clinical Center and by a Cooperative Research and Development Agreement with Ping An.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Liu, L. et al. (2022). Improved Multi-modal Patch Based Lymphoma Segmentation with Negative Sample Augmentation and Label Guidance on PET/CT Scans. In: Li, X., Lv, J., Huo, Y., Dong, B., Leahy, R.M., Li, Q. (eds) Multiscale Multimodal Medical Imaging. MMMI 2022. Lecture Notes in Computer Science, vol 13594. Springer, Cham. https://doi.org/10.1007/978-3-031-18814-5_12
Download citation
DOI: https://doi.org/10.1007/978-3-031-18814-5_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-18813-8
Online ISBN: 978-3-031-18814-5
eBook Packages: Computer ScienceComputer Science (R0)