Abstract
Organic contaminants are among the main pollutants of ecosystems because of their presence in domestic, agricultural, or industrial effluents. Indeed, many organic xenobiotics such as aromatic hydrocarbons, pesticides, synthetic dyes, etc., are not easily biodegradable in the environment and can therefore accumulate in ecosystems causing various toxic symptoms in exposed organisms, including humans. Yeast-assisted biological treatment has emerged as a promising new strategy for the biodegradation of such hazardous contaminants. Firstly, this chapter provides an overview of the applications of yeast in the biodegradation of organic contaminants. Subsequently, synthetic dyes were chosen as a model of organic pollutants to highlight the enzymes involved in their biodegradation process using various yeast strains. Indeed, the main oxidases involved are laccase, tyrosinase, lignin peroxidase, and manganese peroxidase. While the main reductases are Azoreductase, NADH-DCIP reductase, and malachite green reductase. The last section highlights the effects of physicochemical conditions on the effectiveness of mycoremediation.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
5.1 Introduction
Yeasts are a group of polyphyletic fungi composed of basidiomycetes and ascomycetes species that have the particularity of living in a single-celled state. The environmental role that yeasts play is identical to that of other fungi. In fact, they act as saprophytes that transform plant and animal organic matter into biomass and its by-products (Kutty and Philip 2008). These ecological properties have been exploited by man since antiquity and continue to be developed even in the present day in a range of applications such as the fermentation processes (Beer, Wine, Sake, Soy Sauce), in food and feed ingredients (enzymes, flavors, pigments, amino acids, organic acids), the biocatalysis (pharmaceuticals, chiral chemical intermediates, and biotransformation), the biocontrol (food and feed safety, crop protection, and probiotics), as well as in fundamental research in biology and biomedical (Molecular biology, pathway engineering, systems biology mechanisms, drug metabolism and resistance, etc.) (Johnson and Echavarri-erasun 2011).
In addition to these applications, the exploitation of this ecological principle of yeasts in the biodegradation of different organic materials has prompted scientists to evaluate their capacity in the biodegradation of different carbon-based xenobiotics such as organic solvents, humic substances, phenolic compounds, petroleum, surfactants, pesticides, pharmaceuticals, and dyes, etc. (Aksu 2005). Numerous studies have reported the ability of yeast to biodegrade a variety of hazardous contaminants, including aromatic hydrocarbons (Deeba et al. 2018), phenol compounds (Filipowicz et al. 2020), pesticides (Han et al. 2019; Isia et al. 2019), fungicides (Kucharska et al. 2020), insecticide (Chen et al. 2012) and herbicides (States and States 2011) and synthetic dyes (Danouche et al. 2021c, 2022).
In this chapter, we first reviewed existing studies dealing with the topic of biodegradation of organic pollutants using halotolerant yeast strains. Then, we approached a comprehensive analysis of the enzymatic process involved by various yeast strains in the biodegradation of synthetic dyes as a model of organic pollution. Finally, we discussed the involvement of physicochemical factors in enhancing the mycoremediation capacity of yeast species.
5.2 Biodegradation of Organic Pollutants by Yeast
Biodegradation process is defined as an energy-dependent mechanism by which organic substances are decomposed into simpler and smaller by-products through the action of various enzymes (Kaushik and Malik 2009). This bioprocess is called mineralization when the products of the biodegradation are more straightforward elements, such as H2O, CO2, NH3, CH4, H2S, or PO3. This same process is defined as biotransformation when the organic compounds are not completely mineralized (Danouche et al. 2021b). Numerous halophilic microorganisms belonging to bacteria, fungi, and microalgae have shown the ability to decompose a wide variety of organic hazardous substances under high salt conditions (Castillo-carvajal et al. 2014). In recent years, the biotechnological application of fungi (mycoremediation) has become a model example for bioremoval of organic contaminants. It has been reported that various species of fungal could be used for the biodegradation of organic chemicals including aromatic and aliphatic hydrocarbons, industrial dyes, and other organic contaminants, released into the aquatic environment from various industrial and agricultural sectors (Aleu and Collado 2009; Sen et al. 2016). Despite the advantages of yeast strains over other species of the fungal kingdom, including rapid growth, high plasticity, and the ability to adapt to adverse growth conditions (Jafari et al. 2014; Sen et al. 2016), only a few studies have been conducted on the use of halotolerant yeast strains for the biodegradation of organic pollutants.
Aliphatic and Aromatic Hydrocarbons
The yeast species described as hydrocarbon degraders are mostly from the genera of Yarrowia, Candida, Pichia, Debaryomyces, Sporidiobolus, Metschnikowia, Lodderomyces, Rhodosporidium, Leucosporidium, Rhodotorula, Stephanoascus, Sporobolomyces, Trichosporon, and Cryptococcus (Csutak et al. 2010; Kumari and Abraham 2011; Jain and Bajpai 2012; Gargouri et al. 2015; Deeba et al. 2018). Recently, Hashem et al. (2018) reported that other yeast strains of Meyerozyma guilliermondii KKUY-0214, Yamadazyma mexicana KKUY-0160, R. taiwanensis KKUY-0162, P. kluyveri KKUY- 0163, R. ingeniosa KKUY-0170, and C. pseudointermedia KKUY-0192 were approved for their ability to degrade both aromatic and aliphatic hydrocarbon.
Phenolic Compounds
Pollution with phenolic compounds can occur in the soil as well as in water bodies, due to their presence in discharges from industrial, agricultural, or domestic activities (Anku et al. 2017). Species of the genus Candida are documented as yeast strains with the highest capacity to decompose a diverse range of phenolic compounds. For example, a yeast strain of C. rugopelliculosa was reported to be able to decompose various phenolic compounds such as phenol, bisphenol A, nonylphenol, 4-methylphenol, 4-ethylphenol, 4-tert-butylphenol, 4-tert-OP, 4-tert-, and isooctane (Huang et al. 2017). The phenol was also reported to be degraded by other strains of C. tropicalis (Gong et al. 2021), C. tropicalis PHB5 (Basak et al. 2019), C. subhashii A011, C. oregonensis B021, Schizoblastosporion starkeyi-henricii L012 (Filipowicz et al. 2020), and R. kratochvilovae HIMPA1 (Patel et al. 2017).
Pesticides, Fungicides, Insecticide and Herbicides
Various agricultural practices lead to the release of organic contaminants into the soil and surface or ground water. Selected yeast strains can be used as biodegradation agents for these xenobiotics. The pesticides like diazinon or pendimethalin can be biodegraded with Saccharomyces cerevisiae (Ehrampoush et al. 2017) or Clavispora lusitaniae (Han et al. 2019) respectively. The biodegradation of fungicides such as propiconazole was illustrated through the use of yeast strains of Aureobasidium pullulans, Rhodotorula glutinis, and Cryptococcus sp. (Kucharska et al. 2020). Regarding the insecticide, Chen et al. (2012) showed that C. pelliculosa was efficient in the biodegradation of Bifenthrin. Also, C. xestobii was documented to have a high biodegradation capacity of Metolachlor and Alachlor herbicides (States and States 2011).
Synthetic Dyes
Biodegradation of synthetic dyes by yeast has also been documented in the literature using different yeast strains (Danouche et al. 2021b). The most commonly used yeast for the degradation of synthetic dyes are strains belonging to the phylum Ascomycetes, such as Saccharomyces, Candida, and Pichia species. While only a few studies have involved basidiomycetous yeast strains, namely Trichosporon and Pseudozyma (Pajot et al. 2014). In the following section, we will focus on synthetic dyes as a model of organic pollutants because of their different chemical characteristics that make them resistant to biodegradation in natural ecosystems, as well as because of their toxicity toward exposed organisms, including humans (Danouche et al. 2021a). In the remainder of this chapter, an in-depth review of the enzymatic mechanisms involved in the biodegradation of these chemicals by yeast is presented in detail.
5.3 Yeast Enzymes Implications in the Biodegradation of Synthetic Dyes
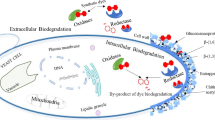
Synthetic dyes can be degraded enzymatically by yeast cells in either the extracellular or in the intracellular compartment. The most studied enzymes for the biodegradation of dyes by yeast are the oxidases, which are the class of enzymes that use oxygen (O2) as an electron acceptor to catalyze the redox reaction, generating H2O or H2O2 as products. They contain a metal or a Flavin-type coenzyme on the active site (Phale et al. 2019). In addition, it has been shown that some reductases are also involved by some yeast strains. The reductases indicated for the biodegradation of synthetic dyes are Azoreductase, NADH-DCIP reductase, and Malachite green reductase (Danouche et al. 2021b).
Laccase (Lac: EC 1.10.3.2)
Synthetic dyes biocatalysis with Lac can be achieved by direct biodegradation of the dye molecule by a nonspecific radical mechanism. This enzymatic pathway has the advantage of avoiding the formation of toxic by-products, such as aromatic amines, which are usually obtained as a result of specific cleavage of the azo bond of various dyes via reductases or chemical catalytic processes (Dave et al. 2015). Several studies have documented the involvement of Lac in the biodegradation of the synthetic dye by yeast strains of Sterigmatomyces halophilus SSA-1575 (Al-Tohamy et al. 2020), Galactomyces geotrichum GG (Guo et al. 2019), Cyberlindnera fabianii (Danouche et al. 2021c), T. akiyoshidainum HP2023 (Martorell et al. 2017a), T. multisporum, and T. laibachii (Pajot et al. 2007).
Tyrosinase (Tyr: E.C. 1.14.18.1): referred also to as monophenol monooxygenase or polyphenol oxidase. It is an oxidase with copper coenzyme, which can be employed for the detoxification of wastewater containing phenol or other organic pollutants (Kim and Uyama 2005). The catalytic reaction of the dyes with Tyr occurs in two successive steps, a first catalyzing reaction is the o-hydroxylation of the monophenols to the corresponding catechols (monophenolase activity), next a second oxidation of monophenols to the corresponding o-quinones (diphenolase activity) (Duckworth and Coleman 1970). On their involvement in the biodegradation of synthetic dyes by yeast cells, it has been identified in only a few yeast species. (Danouche et al. 2021c) reported their involvement in the biodegradation of the azo dye Acid Red 14 with a yeast strain of C. fabianii, in addition to other yeast strains including G. geotrichum MTCC (Waghmode et al. 2012a, b), S. cerevisiae MTCC 463 (Jadhav et al. 2007), C. krusei strains (Charumathi and Das 2011), Candida sp. MM 4035, T. porosum MM 4037, C. satwnus MM 4034, Barnettozyma californica MM 4018 (Martorell et al. 2012).
Lignin Peroxidase (LiP: EC 1.11.1.14)
LiP is an extracellular enzyme whose enzymatic substrate is nonspecific, this particularity confers it the capacity to degrade various aromatic phenolic and non-phenolic compounds (Chowdhary et al. 2018). For example, the biodegradation of sulfonated azo dye with LiP can be accomplished in two consecutive one-electron oxidations of the oxidized LiP forms by H2O2 in the phenolic ring, where the corresponding carbonium ion bearing the azo bond contributes to the formation of quinone and phenyldiazine by nucleophilic attack by H2O. The phenyldiazine product is then oxidized by O2 to a phenyl radical and the azo bond is removed as N2, and then the phenyl radical extracts hydrogen from its surroundings to produce a stable aromatic compound (Chivukula et al. 1995). The catalytic activity of LiP was investigated during the biodegradation of various synthetic dyes by basidiomycota yeast strains of T. laibachii and T. multisporum (Pajot et al. 2007), as well as, by ascomycota yeast strains of S. halophilus SSA-1575 (Al-Tohamy et al. 2020), G. geotrichum (Guo et al. 2019), P. occidentalis (Song et al. 2018a), S. cerevisiae (Jadhav et al. 2007), C. krusei (Charumathi and Das 2011), Diutina rugosa (Bankole et al. 2017), and C. samutprakarnensis (Song et al. 2018b).
Manganese Peroxidase (EC 1.11.1.13)
MnP is a substrate-specific oxidase that oxidizes Mn2 + to Mn3+ from the surface of the enzyme and subsequently oxidizes phenolic substrates such as model lignin compounds or other organic contaminants (Zhou et al. 2013). Yeast used MnP for the biodegradation of synthetic dyes as well, it was revealed in yeast strains of C. fabianii (Danouche et al. 2021c), P. occidentalis (Song et al. 2018a), D. polymorphus, C. tropicalis (Yang et al. 2008), T. multisporum, and T. laibachii (Pajot et al. 2007).
Azoreductase (AzoR: EC 1.7.1.6): are a class of enzymes that catalyze the reduction reaction such as the reduction of azo bonds (-N=N-) of azo dyes and nitroaromatic and azoic drugs (Misal and Gawai 2018). The AzoR can be classified according to their structures, or according to flavin dependence. The flavin-dependent class of AzoR can also be divided based on their coenzymes like NADH, NADPH (Saratale et al. 2011; Solís et al. 2012). The involvement of AzoR in dye biodegradation by yeast has been reported in some research employing yeast strains of Issatchenkia occidentalis (Ramalho et al. 2004), C. fabianii (Danouche et al. 2021c), C. krusei (Charumathi and Das 2011), S. cerevisiae MTCC 463 (Jadhav et al. 2007), and T. beigelii NCIM-3326 (Saratale et al. 2009a).
NADH-Preferring 2,6-Dichloroindophenol Reductase (NADH-DCIP: EC 1.6.99.3)
NADH-DCIP reductase is an oxidoreductase that reduces 2,6-dichloroindo-phenol (DCIP) with NADH as an electron donor (Nishiya and Yamamoto 2007). Some studies have shown an increase in the NADH-DCIP reductase activity during the biodegradation of various azo dyes with strains of P. occidentalis G1 (Song et al. 2018a), C. samutprakarnensis (Song et al. 2018b), D. rugosa (Bankole et al. 2017), P. kudriavzevii CR-Y103 (Rosu et al. 2018), and T. beigelii NCIM-3326 (Saratale et al. 2009a).
Malachite Green Reductase
NADH is used as an electron donor by this reductase to transform malachite green into leucomalachite green. It has been first reported by Jadhav and Govindwar (2006) that in S. cerevisiae MTCC 463 was used for the biodegradation of green malachite. Next, Jadhav et al. (2008b) demonstrated their implication in the biodegradation of methyl red by G. geotrichum MTCC 1360. Also, Charumathi and Das (2011) reported the increase of MG-reductase activity in C. krusei used for the biodegradation of Basic Violet 3, as well as, in S. cerevisiae used for the biodegradation of Malachite green (Biradar et al. 2016).
Regardless of the mechanisms involved, the performance of the microorganisms in the bioremediation of organic pollutants can be influenced by various environmental factors, including nutrients, pH, temperature, etc. It is therefore critical to emphasize the impact of these parameters on the yeast’s ability to eliminate such contaminants.
5.4 Factors Controlling Mycoremediation Performance
Yeast cells are sensitive to the environmental conditions in which they grow. Determining the impact of these physiochemical factors on the efficiency of the removal of synthetic dyes is therefore crucial in order to make the mycoremediation process faster, more efficient, and more practical for large-scale applications. There are two ways to perform this optimization, either using a single-factor optimization approach or based on statistical methods of optimization by the design of the experiment (Gönen and Aksu 2009; Mahmoud 2016).
Carbon and Nitrogen Sources
The effect of carbon and nitrogen sources on the bioaccumulation or the biodegradation capacity of dye by yeast strains has been the subject of several studies. It has been reported that at constant sucrose content, the concentrations of both Remazol Black B and Remazol Blue dyes inhibited the growth of C. tropicalis, with constant dye concentration, the growth efficiency and the bioaccumulation capacity increased with sucrose concentration up to 15 g L−1 (Aksu and Dönmez 2005). This combined effect was also analyzed using a statistical approach of response surface methodology. Okur et al. (2014) found that the optimal values for dye uptake by C. tropicalis correspond to sugar concentration of 5.1 g L−1 and a dye concentration of 499 mg L−1 of initial dye concentration. Also, Gönen and Aksu (2009) found that the optimum combination predicted by response surface methodology (RSM) confirmed that C. utilis was able to bioaccumulate Remazol turquoise blue-G with a maximum uptake yield of 82.0% in 15 g L−1 sucrose and 50 mg L−1 dye concentration. Additionally, (Das et al. 2010) confirmed that P. fermentans MTCC 189 was able to accumulate Basic violet 3 up to 69.8% in 10 mg L−1 of dye-containing medium and 24 g L−1 sugar extracted from sugarcane bagasse through RSM analysis. Concerning the carbon sources effect on the biodegradation efficiency of dye by yeast cells, it has been reported that the addition of carbon sources, especially glucose at a certain level, stimulates the biodegradation of dyes (Chang et al. 2000; Waghmode et al. 2011). Indeed, glucose is an essential element in several mechanisms, it provides an energy source for yeast growth, it regenerates the redox mediators NADH and FADH, and also it serves as a substrate for the production of H2O2, which acts in turn as a co-substrate for MnP and LiP (Swamy and Ramsay 1999; Jafari et al. 2014). In the same way, the supplementation of the medium with nitrogen sources such as peptone, yeast extract, urea, or others favors the regeneration of NADH which is used as an electron donor for the reduction of azo dyes by the different enzymes (Bras et al. 2001).
Dye Concentration
The initial dye concentration can significantly influence their removal efficiency. A higher dye concentration gradually decreases the percentage of their decolorization. This can be attributed to the toxicity of the dyes toward the yeast cells, or to a lower biomass production making the decolorization operation inefficient (Saratale et al. 2011). It has been reported by Das et al. (2010) that the bioaccumulation of Acid Blue 93, Direct Red 28, and Basic Violet 3 by P. fermentans MTCC 189 decreased as the initial concentration of these dyes increased from 10 to 30 mg L−1. Likewise, Dönmez (2002) and Aksu (2003) described that the increase in the initial dye concentration inhibited the growth and caused a long lag period of S. cerevisiae and C. tropicalis. Other studies indicate that the presence of dyes at high concentrations may inhibit azoreductase activity during enzymatic biodegradation of the synthetic dye, due to the binding of dye molecules to the active site of enzymes (Jadhav et al. 2008a; Saratale et al. 2009a).
Temperature
Temperature is one of the crucial factors that influence the metabolic pathways, the enzymatic activity, and the physicochemical interaction of dye molecules with the cell wall. According to the available literature, no study has been devoted to the question of the effect of temperature on the bioaccumulation of dyes using yeast cells. Therefore, it is very important to consider this factor as a research question for future studies. Regarding the effect of temperature on the ability of yeast to biodegrade dyes, many studies have been conducted on the activation energy of the involved enzymes (Chequer et al. 2013; Miranda et al. 2013). The performance of decolorization increased with increasing temperature to the optimum temperature, then a reduction in activity occurs at higher temperatures (Tan et al. 2013, 2014, 2016). The decrease in the ability of yeast strains to remove the dye molecules can be attributed to the denaturation of the enzymes involved or to the resulting loss of cell viability (Saratale et al. 2009b).
pH
pH of solutions is one of the most influential factors on the ability of yeast cells to bioremediate various organic pollutants, including synthetic dyes, it can modify the physicochemical properties of the dye molecules, as well as the physicochemical properties of the yeast surface where the cell-dye molecule interaction initially occurs (Fu and Viraraghavan 2002). At low pH, the surface of yeast cells becomes protonated with a positive charge, which promotes the binding of anionic dyes. On the other hand, at higher pH values, the cell surface acquires a negative charge, resulting in the electrostatic attraction of cationic dyes (Charumathi and Das 2012). The pH may have an inhibitory effect on the transport of dye molecules across the cell membrane, which is considered the first stage of the intracellular biodegradation or bioaccumulation process (Khan et al. 2013). In addition, the initial pH can affect the physiology of the yeast cells and the enzymatic activity. Most yeast strains show better decolorization efficiency under neutral or acidic conditions. The optimal value for the bioaccumulation of Remazol Blue, Reactive Red, and Reactive Black by the yeast strain C. tropicalis (Dönmez and Aksu 2002) as well as for Remazol Red RB, Remazol Black B, and Remazol Blue by S. Cerevisiae (Aksu 2003) was observed was 3.0. Likewise, Das et al. (2010) investigated that the maximum bioaccumulation rate of Basic Violet3, Direct Red 28, and Acid Blue 93 by growing cells of P. fermentans MTCC 189 was at pH 5.0.
Shaking
Agitation allows a uniform distribution of oxygen and nutrients in the medium. It also facilitates the exchange of gases produced during the fermentation process of dyes by yeast cells (Yu and Wen 2005; Pajot et al. 2007; Yang et al. 2008; Martorell et al. 2017b). Differing opinions have been expressed regarding the effect of oxygen on the decolorization process, with some researchers considering that during the dye reduction reaction by the reductase, oxygen behaves as a stronger electron acceptor than the dye molecule, thus preventing the azo dye reduction reaction (Kalyani et al. 2008). On the other hand, other research indicates that oxidizing enzymes, such as laccases, require oxygen to oxidize aromatic molecules, including dyes (Thurston 1994). Therefore, it is necessary to optimize the level of agitation that regulates the concentration of dissolved oxygen in the medium throughout the yeast decolorization process to ensure an effective treatment.
5.5 Conclusions
On the basis of the research discussed in this chapter, it is appropriate to consider mycoremediation as a cost-effective, eco-friendly, and efficient approach for the biodegradation of organic contaminants. However, an efficient mycoremediation process requires the optimization of physicochemical conditions, notably the initial concentration of the pollutant, the supply of nitrogen or carbon sources, as well as growth conditions such as pH, temperature, and agitation. On the other hand, the effectiveness of this kind of bio-processes should not be limited to emphasizing the degradation of the considered pollutant molecule; instead, it requires characterization and evaluation of the toxicity of the obtained by-products, since they can have more harmful effects than the original molecule. Lastly, we strongly recommend the application of this emerging biotechnology in wastewater treatment at pilot or large scale in order to prove its potential application.
References
Aksu Z (2003) Reactive dye bioaccumulation by Saccharomyces cerevisiae. Process Biochem 38:1437–1444. https://doi.org/10.1016/S0032-9592(03)00034-7
Aksu Z (2005) Application of biosorption for the removal of organic pollutants: a review. Process Biochem 40:997–1026. https://doi.org/10.1016/j.procbio.2004.04.008
Aksu Z, Dönmez G (2005) Combined effects of molasses sucrose and reactive dye on the growth and dye bioaccumulation properties of Candida tropicalis. Process Biochem 40:2443–2454. https://doi.org/10.1016/j.procbio.2004.09.013
Aleu J, Collado IG (2009) Pollutants biodegradation by fungi. Curr Org Chem 13:1194. https://doi.org/10.2174/138527209788921774
Al-Tohamy R, Kenawy ER, Sun J, Ali SS (2020) Performance of a newly isolated salt-tolerant yeast strain Sterigmatomyces halophilus SSA-1575 for azo dye decolorization and detoxification. Front Microbiol 11:1–19. https://doi.org/10.3389/fmicb.2020.01163
Anku WW, Mamo MA, Mamo MA, Penny PW, Govender PP (2017) Phenolic compounds compounds in in water : water : sources, phenolic toxicity and treatment methods toxicity and treatment methods, in: Phenolic compounds—natural sources, importance and applications. pp. 419–443. https://doi.org/10.5772/66927
Bankole PO, Adekunle AA, Obidi OF, Olukanni OD, Govindwar SP (2017) Degradation of indigo dye by a newly isolated yeast, Diutina rugosa from dye wastewater polluted soil. J Environ Chem Eng 5:4639–4648. https://doi.org/10.1016/j.jece.2017.08.050
Basak B, Jeon B, Kurade MB, Saratale GD, Bhunia B, Chatterjee PK, Dey A (2019) Biodegradation of high concentration phenol using sugarcane bagasse immobilized Candida tropicalis PHB5 in a packed-bed column reactor. Ecotoxicol Environ Saf 180:317–325. https://doi.org/10.1016/j.ecoenv.2019.05.020
Biradar SP, Rane NR, Patil TS, Khandare RV, Govindwar SP, Pawar PK (2016) Herbal augmentation enhances malachite green biodegradation efficacy of Saccharomyces cerevisiae. Biol 71:475–483. https://doi.org/10.1515/biolog-2016-0069
Bras R, Ferra MIA, Pinheir HM, Goncalves IC (2001) Batch tests for assessing decolourisation of azo dyes by methanogenic and mixed cultures. J Biotechnol 89:155. https://doi.org/10.1016/s0168-1656(01)00312-1
Castillo-carvajal LC, Sanz-martín JL, Barragán-huerta BE (2014) Biodegradation of organic pollutants in saline wastewater by halophilic microorganisms : a review. Environ Sci Pollut Res. https://doi.org/10.1007/s11356-014-3036-z
Chang J, Kuo T, Chao Y, Ho J, Lin P (2000) Azo dye decolorization with a mutant Escherichia coli strain. Biotechnol Lett 22:807–812. https://doi.org/10.1023/A:1005624707777
Charumathi D, Das N (2011) Biodegradation of Basic Violet 3 by Candida krusei isolated from textile wastewater. Biodegradation 22:1169–1180. https://doi.org/10.1007/s10532-011-9472-2
Charumathi D, Das N (2012) Packed bed column studies for the removal of synthetic dyes from textile wastewater using immobilised dead C. tropicalis. Desalination 285:22–30. https://doi.org/10.1016/j.desal.2011.09.023
Chen S, Luo J, Hu M, Geng P, Zhang Y (2012) Microbial detoxification of bifenthrin by a novel yeast and its potential for contaminated soils treatment. PLoS One 7:30862. https://doi.org/10.1371/journal.pone.0030862
Chequer FD, de Oliveira GAR, Ferraz EA, Cardoso JC, Zanoni MB, de Oliveira DP (2013) Textile dyes: dyeing process and environmental impact, in: In eco-friendly textile dyeing and finishing. IntechOpen. 268. https://doi.org/10.5772/3436
Chivukula M, Spadaro JT, Renganathan V (1995) Lignin peroxidase-catalyzed oxidation of sulfonated azo dyes generates novel Sulfophenyl hydroperoxides. Biochemistry 34:7765–7772. https://doi.org/10.1021/bi00023a024
Chowdhary P, More N, Yadav A, Bharagava RN (2018) Ligninolytic enzymes: an introduction and applications in the food industry, enzymes in food biotechnology: production, applications, and future prospects. Elsevier Inc. https://doi.org/10.1016/B978-0-12-813280-7.00012-8
Csutak O, Stoica I, Ghindea R, Tanase AM, Vassu T (2010) Insights on yeast bioremediation processes. Rom Biotechnol Lett 15:5066–5071
Danouche M, El Arroussi H, Bahafid W, El Ghachtouli N, El Arroussi H, Bahafid W, An NEG (2021a) An overview of the biosorption mechanism for the bioremediation of synthetic dyes using yeast cells. Environ Technol Rev 10:58–76. https://doi.org/10.1080/21622515.2020.1869839
Danouche M, El Arroussi H, El N (2021b) Mycoremediation of synthetic dyes by yeast cells: a sustainable biodegradation approach. Environ Sustain 1–18. https://doi.org/10.1007/s42398-020-00150-w
Danouche M, Ferioun M, Bahafid W, El Ghachtouli N (2021c) Mycoremediation of azo dyes using Cyberlindnera fabianii yeast strain: application of designs of experiments for decolorization optimization. Water Environ Res:1–15. https://doi.org/10.1002/wer.1499
Danouche M, El Arroussi H, El Ghachtouli N (2022) Bioremoval of Acid Red 14 dye by Wickerhamomyces anomalus biomass: kinetic and thermodynamic study, characterization of physicochemical interactions, and statistical optimization of the biosorption process. Biomass Conv Bioref. https://doi.org/10.1007/s13399-022-02711-x
Das D, Charumathi D, Das N (2010) Combined effects of sugarcane bagasse extract and synthetic dyes on the growth and bioaccumulation properties of Pichia fermentans MTCC 189. J Hazard Mater 183:497–505. https://doi.org/10.1016/j.jhazmat.2010.07.051
Dave SR, Patel TL, Tipre DR (2015) Bacterial degradation of azo dye containing wastes. In: Microbial degradation of synthetic dyes in wastewaters, environmental science and engineering. Cham, pp 57–83. https://doi.org/10.1007/978-3-319-10942-8_3
Deeba F, Pruthi V, Negi YS (2018) Aromatic hydrocarbon biodegradation activates neutral lipid biosynthesis in oleaginous yeast. Bioresour Technol 255:273–280. https://doi.org/10.1016/j.biortech.2018.01.096
Dönmez G (2002) Bioaccumulation of the reactive textile dyes by Candida tropicalis growing in molasses medium. Enzym Microb Technol 30:363–366. https://doi.org/10.1016/S0141-0229(01)00511-7
Dönmez G, Aksu Z (2002) Removal of chromium(VI) from saline wastewaters by Dunaliella species. Process Biochem 38:751–762. https://doi.org/10.1016/S0032-9592(02)00204-2
Duckworth HW, Coleman JE (1970) Physicochemical and kinetic properties of mushroom tyrosinase. J Biol Chem 245:1613–1625. https://doi.org/10.1016/S0031-9422(00)82247-5
Ehrampoush MH, Sadeghi A, Ghaneian MT, Bonyadi Z (2017) Optimization of diazinon biodegradation from aqueous solutions by Saccharomyces cerevisiae using response surface methodology. AMB Express 1–6. https://doi.org/10.1186/s13568-017-0366-5
Filipowicz N, Momotko M, Boczkaj G, Cieśliński H (2020) Determination of phenol biodegradation pathways in three psychrotolerant. Enzyme Microb Technol J 141:0141–0229. https://doi.org/10.1016/j.enzmictec.2020.109663
Fu Y, Viraraghavan T (2002) Removal of Congo Red from an aqueous solution by fungus Aspergillus niger. Adv Environ Res 7:239–247. https://doi.org/10.1016/S1093-0191(01)00123-X
Gargouri B, Mhiri N, Karray F, Aloui F, Sayadi S (2015) Isolation and characterization of hydrocarbon-degrading yeast strains from petroleum contaminated industrial wastewater. Biomed Res Int 2015:11. https://doi.org/10.1155/2015/929424
Gönen F, Aksu Z (2009) Predictive expressions of growth and Remazol Turquoise Blue-G reactive dye bioaccumulation properties of Candida utilis. Enzym Microb Technol 45:15–21. https://doi.org/10.1016/j.enzmictec.2009.03.006
Gong Y, Ding P, Xu M, Zhang C, Xing K, Qin S (2021) Biodegradation of phenol by a halotolerant versatile yeast Candida tropicalis SDP-1 in wastewater and soil under high salinity conditions. J Environ Manag 289:112525. https://doi.org/10.1016/j.jenvman.2021.112525
Guo G, Tian F, Zhao Y, Tang M, Liu W, Liu C, Xue S, Kong W, Sun Y, Wang S (2019) Aerobic decolorization and detoxification of Acid Scarlet GR by a newly isolated salt-tolerant yeast strain Galactomyces geotrichum GG. Int Biodeterior Biodegrad 145:104818. https://doi.org/10.1016/j.ibiod.2019.104818
Han Y, Tang Z, Bao H, Wu D, Deng X (2019) Degradation of pendimethalin by the yeast YC2 and determination of its two main metabolites †. RSC Adv 9:491–497. https://doi.org/10.1039/c8ra07872f
Hashem M, Alamri SA, Al-zomyh SSAA, Alrumman SA (2018) Biodegradation and detoxi fi cation of aliphatic and aromatic hydrocarbons by new yeast strains. Ecotoxicol Environ Saf 151:28–34. https://doi.org/10.1016/j.ecoenv.2017.12.064
Huang S, Lin C, Kirschner R (2017) Biodegradation of the endocrine disrupter 4-tert-octylphenol by the yeast strain Candida rugopelliculosa RRKY5 via phenolic ring hydroxylation and alkyl chain oxidation pathways Ranjith. Bioresour Technol 226:55–64. https://doi.org/10.1016/j.biortech.2016.11.129
Isia I, Hadibarata T, Sari AA, Al Khulaifi MM (2019) Potential use of a pathogenic yeast Pichia kluyveri FM012 for degradation of dichlorodiphenyltrichloroethane ( DDT ). Water Air Soil Pollut 230:1–11
Jadhav JP, Govindwar SP (2006) Biotransformation of malachite green by Saccharomyces cerevisiae MTCC 463. Yeast 23:315–323. https://doi.org/10.1002/yea.1356
Jadhav JP, Parshetti GK, Kalme SD, Govindwar SP (2007) Decolourization of azo dye methyl red by Saccharomyces cerevisiae MTCC 463. Chemosphere 68:394–400. https://doi.org/10.1016/j.chemosphere.2006.12.087
Jadhav SU, Jadhav MU, Kagalkar AN, Govindwar SP (2008a) Decolorization of brilliant blue G dye mediated by degradation of the microbial consortium of Galactomyces geotrichum and Bacillus sp. J Chinese Inst Chem Eng 39:563–570. https://doi.org/10.1016/j.jcice.2008.06.003
Jadhav SU, Kalme SD, Govindwar SP (2008b) Biodegradation of Methyl red by Galactomyces geotrichum MTCC 1360. Int Biodeterior Biodegrad 62:135–142. https://doi.org/10.1016/j.ibiod.2007.12.010
Jafari N, Soudi MR, Kasra-Kermanshahi R (2014) Biodegradation perspectives of azo dyes by yeasts. Microbiology 83:484–497. https://doi.org/10.1134/s0026261714050130
Jain PK, Bajpai V (2012) Biotechnology of bioremediation—a review. Int J Environ 3:535–549. https://doi.org/10.6088/ijes.2012030131053
Johnson EA, Echavarri-erasun C (2011) Yeast biotechnology. In: The yeasts. Elsevier B.V, pp 21–44. https://doi.org/10.1016/B978-0-444-52149-1.00003-3
Kalyani DC, Patil PS, Jadhav JP, Govindwar SP (2008) Biodegradation of reactive textile dye red BLI by an isolated bacterium Pseudomonas sp. SUK1. Bioresour Technol 99:4635–4641. https://doi.org/10.1016/j.biortech.2007.06.058
Kaushik P, Malik A (2009) Fungal dye decolourization: recent advances and future potential. Environ Int 35:127–141. https://doi.org/10.1016/j.envint.2008.05.010
Khan R, Bhawana P, Fulekar MH (2013) Microbial decolorization and degradation of synthetic dyes: a review. Rev Environ Sci Biotechnol 12:75–97. https://doi.org/10.1007/s11157-012-9287-6
Kim YJ, Uyama H (2005) Tyrosinase inhibitors from natural and synthetic sources: structure, inhibition mechanism and perspective for the future. Cell Mol Life Sci 62:1707–1723. https://doi.org/10.1007/s00018-005-5054-y
Kucharska K, Wachowska U, Czaplicki S (2020) Wheat phyllosphere yeasts degrade propiconazole. BMC Microbiol 20:1–14. https://doi.org/10.1186/s12866-020-01885-6
Kumari M, Abraham J (2011) Biodegradation of diesel oil using yeast Rhodosporidium toruloides. Res J Environ Toxicol 5:369–377. https://doi.org/10.3923/rjet.2011.369.377
Kutty SN, Philip R (2008) Marine yeasts—a review. Yeast 465–483. https://doi.org/10.1002/yea
Mahmoud MS (2016) Decolorization of certain reactive dye from aqueous solution using Baker’s yeast (Saccharomyces cerevisiae) strain. HBRC J 12:88–98. https://doi.org/10.1016/j.hbrcj.2014.07.005
Martorell MM, Pajot HF, Ahmed PM, de Figueroa LIC (2017a) Biodecoloration of Reactive Black 5 by the methylotrophic yeast Candida boidinii MM 4035. J Environ Sci (China) 53:78–87. https://doi.org/10.1016/j.jes.2016.01.033
Martorell MM, Pajot HF, de Figueroa LIC (2012) Dye-decolourizing yeasts isolated from Las Yungas rainforest. Dye assimilation and removal used as selection criteria. Int Biodeterior Biodegrad 66:25–32. https://doi.org/10.1016/j.ibiod.2011.10.005
Martorell MM, Pajot HF, Figueroa LICD (2017b) Biological degradation of Reactive Black 5 dye by yeast Trichosporon akiyoshidainum. J Environ Chem Eng 5:5987–5993. https://doi.org/10.1016/j.jece.2017.11.012
Miranda RDCMD, Gomes EDB, Pereira N, Marin-Morales MA, Machado KMG, de Gusmão NB (2013) Biotreatment of textile effluent in static bioreactor by Curvularia lunata URM 6179 and Phanerochaete chrysosporium URM 6181. Bioresour Technol 142:361–367. https://doi.org/10.1016/j.biortech.2013.05.066
Misal SA, Gawai KR (2018) Azoreductase: a key player of xenobiotic metabolism. Bioresour Bioprocess. https://doi.org/10.1186/s40643-018-0206-8
Nishiya Y, Yamamoto Y (2007) Characterization of a NADH:dichloroindophenol oxidoreductase from Bacillus subtilis. Biosci Biotechnol Biochem 71:611–614. https://doi.org/10.1271/bbb.60548
Okur M, Saraçoglu N, Aksu Z (2014) Use of response surface methodology for the bioaccumulation of Violet 90 metal-complex dye by Candida tropicalis. Turkish J Eng Environ Sci 38:217–230. https://doi.org/10.3906/muh-1409-2
Pajot HF, de Figueroa LIC, Fariña JI (2007) Dye-decolorizing activity in isolated yeasts from the ecoregion of Las Yungas (Tucumán, Argentina). Enzym Microb Technol 40:1503–1511. https://doi.org/10.1016/j.enzmictec.2006.10.038
Pajot HF, Martorell MM, de Figueroa LIC (2014) Ecology of dye decolorizing yeasts. In: Alvarez A, Polti MA (eds) Bioremediation in Latin America: current research and perspectives. Springer, pp 223–240. https://doi.org/10.1007/978-3-319-05738-5_14
Patel A, Sartaj K, Arora N, Pruthi V, Pruthi PA (2017) Biodegradation of phenol via meta cleavage pathway triggers de novo TAG biosynthesis pathway in oleaginous yeast. J Hazard Mater 340:47–56. https://doi.org/10.1016/j.jhazmat.2017.07.013
Phale PS, Sharma A, Gautam K (2019) Microbial degradation of xenobiotics like aromatic pollutants from the terrestrial environments, Pharmaceuticals and Personal Care Products: Waste Management and Treatment Technology Emerging Contaminants and Micro Pollutants. Elsevier Inc. https://doi.org/10.1016/B978-0-12-816189-0.00011-1
Ramalho PA, Cardoso MH, Ramalho MT (2004) Characterization of azo reduction activity in a novel ascomycete yeast strain characterization of azo reduction activity in a novel ascomycete yeast strain. Appl Environ Microbiol 70:2279–2288. https://doi.org/10.1128/AEM.70.4.2279
Rosu CM, Avadanei M, Gherghel D, Mihasan M, Mihai C, Trifan A, Miron A, Vochita G (2018) Biodegradation and detoxification efficiency of azo-dye reactive Orange 16 by Pichia kudriavzevii CR-Y103. Water Air Soil Pollut 229. https://doi.org/10.1007/s11270-017-3668-y
Saratale RG, Saratale GD, Chang JS, Govindwar SP (2009a) Decolorization and biodegradation of textile dye Navy blue HER by Trichosporon beigelii NCIM-3326. J Hazard Mater 166:1421–1428. https://doi.org/10.1016/j.jhazmat.2008.12.068
Saratale RG, Saratale GD, Chang JS, Govindwar SP (2009b) Ecofriendly degradation of sulfonated diazo dye C.I. Reactive Green 19A using Micrococcus glutamicus NCIM-2168. Bioresour Technol 100:3897–3905. https://doi.org/10.1016/j.biortech.2009.03.051
Saratale RG, Saratale GD, Chang JS, Govindwar SP (2011) Bacterial decolorization and degradation of azo dyes: a review. J Taiwan Inst Chem Eng 42:138–157. https://doi.org/10.1016/j.jtice.2010.06.006
Sen SK, Raut S, Bandyopadhyay P, Raut S (2016) Fungal decolouration and degradation of azo dyes: a review. Fungal Biol Rev 30:112–133. https://doi.org/10.1016/j.fbr.2016.06.003
Solís M, Solís A, Pérez HI, Manjarrez N, Flores M (2012) Microbial decolouration of azo dyes: a review. Process Biochem 47:1723–1748. https://doi.org/10.1016/j.procbio.2012.08.014
Song L, Shao Y, Ning S, Tan L (2018a) Performance of a newly isolated salt-tolerant yeast strain Pichia occidentalis G1 for degrading and detoxifying azo dyes. Bioresour Technol 233:21–29. https://doi.org/10.1016/j.biortech.2017.02.065
Song Z, Song L, Shao Y, Tan L (2018b) Degradation and detoxification of azo dyes by a salt-tolerant yeast Cyberlindnera samutprakarnensis S4 under high-salt conditions. World J Microbiol Biotechnol 34. https://doi.org/10.1007/s11274-018-2515-7
States U, States U (2011) Biodegradation and mineralization of metolachlor and alachlor by Candida xestobii. J Agric Food Chem 619–627. https://doi.org/10.1021/jf103508w
Swamy J, Ramsay JA (1999) Effects of glucose and NH 4+ concentrations on sequential dye decoloration by Trametes versicolor. Enzym Microb Technol 25:278–284. https://doi.org/10.1016/S0141-0229(99)00058-7
Tan L, He M, Song L, Fu X, Shi S (2016) Bioresource technology aerobic decolorization, degradation and detoxification of azo dyes by a newly isolated salt-tolerant yeast Scheffersomyces spartinae TLHS-SF1. Bioresour Technol 203:287–294. https://doi.org/10.1016/j.biortech.2015.12.058
Tan L, Li H, Ning S, Xu B (2014) Aerobic decolorization and degradation of azo dyes by suspended growing cells and immobilized cells of a newly isolated yeast Magnusiomyces ingens LH-F1. Bioresour Technol 158:321–328. https://doi.org/10.1016/j.biortech.2014.02.063
Tan L, Ning S, Zhang X, Shi S (2013) Aerobic decolorization and degradation of azo dyes by growing cells of a newly isolated yeast Candida tropicalis TL-F1. Bioresour Technol 138:307–313. https://doi.org/10.1016/j.biortech.2013.03.183
Thurston CF (1994) The structure and function of fungal laccases. Microbiology 140:19–26
Waghmode TR, Kurade MB, Govindwar SP (2011) Time dependent degradation of mixture of structurally different azo and non azo dyes by using Galactomyces geotrichum MTCC 1360. Int Biodeterior Biodegrad 65:479–486. https://doi.org/10.1016/j.ibiod.2011.01.010
Waghmode TR, Kurade MB, Kabra AN, Govindwar SP (2012a) Degradation of remazol red dye by Galactomyces geotrichum MTCC 1360 leading to increased iron uptake in Sorghum vulgare and Phaseolus mungo from soil. Biotechnol Bioprocess Eng 17:117–126. https://doi.org/10.1007/s12257-011-0307-0
Waghmode TR, Kurade MB, Kabra AN, Govindwar SP (2012b) Biodegradation of Rubine GFL by Galactomyces geotrichum MTCC 1360 and subsequent toxicological analysis by using cytotoxicity, genotoxicity and oxidative stress studies. Microbiol (United Kingdom) 158:2344–2352. https://doi.org/10.1099/mic.0.060467-0
Yang Q, Tao L, Yang M, Zhang H (2008) Effects of glucose on the decolorization of Reactive Black 5 by yeast isolates. J Environ Sci 20:105–108. https://doi.org/10.1016/S1001-0742(08)60016-9
Yu Z, Wen X (2005) Screening and identification of yeasts for decolorizing synthetic dyes in industrial wastewater. Int Biodeterior Biodegrad 56:109–114. https://doi.org/10.1016/j.ibiod.2005.05.006
Zhou XW, Cong WR, Su KQ, Zhang YM (2013) Ligninolytic enzymes from Ganoderma spp: current status and potential applications. Crit Rev Microbiol 39:416–426. https://doi.org/10.3109/1040841X.2012.722606
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Mohammed, D., Hicham, E.A., Naima, E.G. (2023). Biodegradation of Environmental Pollutants by Marine Yeasts. In: Encarnação, T., Canelas Pais, A. (eds) Marine Organisms: A Solution to Environmental Pollution?. Environmental Challenges and Solutions. Springer, Cham. https://doi.org/10.1007/978-3-031-17226-7_5
Download citation
DOI: https://doi.org/10.1007/978-3-031-17226-7_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-17225-0
Online ISBN: 978-3-031-17226-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)