Abstract
Alterations in transcriptional programs are a fundamental feature of prostate (PCa) and breast cancer (BrCa), and frequently target the actions of the principal steroidal nuclear receptors (NRs), namely the androgen receptor (AR) and the estrogen receptor alpha (ERα), respectively. Indeed, the functions of AR and ERα are central to both prostate and mammary gland biology. The genomic interactions of these NRs become highly distorted in part by changing how they functionally interact with a cohort of non-steroidal Type II NRs, which are by contrast relatively understudied compared to their steroidal cousins. For example, the AR cistrome overlaps with cistromes of different Type II NRs, which suggests a high potential for integrated NR functions to tailor transcriptional signals. Over recent years the cistromes of these Type II NRs, including HNF4s, RARs, PPARs and VDR, have been studied in PCa and BrCa revealing convergence and functional consequences, and are reviewed in the current chapter.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Breast cancer
- Prostate cancer
- Non-steroidal nuclear receptors
- Cistrome
- Transcriptome
- Bookmarking
- Epigenetics
1 Nuclear Receptor Genomic Interactions Are Highly Integrated and Sense a Wide Variety of Inputs
The collective transcriptional actions of nuclear receptors (NRs) form a central conduit for hormonal, dietary and environmental compounds to signal to the genome. Specifically, NRs act as sensors that respond to both the presence and absence of a diverse array of ligands and in turn initiate and fine-tune transcriptional events. The impact of NR gene regulatory complexes is evident in development, metabolism, circadian rhythm and cell fate decisions including differentiation phenotypes. Reflecting this widespread importance, there is clear evidence for their disruption acting as disease drivers for various syndromes including cancer [1,2,3,4,5].
The classical sex steroids bind cognate receptors with high affinity; estradiol binds estrogen receptor, NR3A1/ERα, and dihydrotestosterone binds the androgen receptor, NR3C4/AR. Beyond these ligands seco-steroids, retinoid derivatives and bioactive dietary-derived factors such as fatty acids, oxysterols, heme, and bile acids act as ligands and regulate the genomic interactions of a broader group of the NRs. More broadly, these integrated and environmentally-driven NR-genomic interactions are central to concepts such as nutrigenomics and provide the rationale for positioning a wider panel of NRs as promising therapeutic targets in cancer [6,7,8]. Finally, other NRs, without known ligands, have also been identified, known as orphan receptors [9]. Collectively, the interaction of all these NRs allows for the highly dexterous transcriptional outputs, underpinned by the dynamic and mobile NR-genomic interactions, known as NR cistromes. In turn, the NR cistrome gene-regulatory functions are regulated by NR-associated coregulators including coactivators, corepressors and other transcription factors (TFs) and thereby provide a further level of control to regulate transcription [10,11,12,13].
NRs are classified based on mode of action as Type I, II, III, or IV [14]. Steroid NRs are Type I and in the absence of ligand these receptors are often largely cytoplasmic associated with heat shock proteins. Ligand binding results in their dissociation from heat shock proteins and NR homo-dimerization and translocation to the nucleus. Type II NRs, in contrast, reside in the nucleus as heterodimers (for example with RXRs) and bound to genome even in the absence of ligands [15]. Types III and Type IV are orphan receptors, for which ligands are unknown, or possibly don’t exist, and are also generally located in the nucleus and bind DNA as homodimers (Type III) or monomers (Type IV).
The impact of NRs is highly evident across many high-profile and impactful hormone-dependent cancers, including not only prostate cancer (PCa) and breast cancer (BrCa), but also other cancers including ovarian, endometrium, testis, thyroid, and pancreas. An appreciation of the relationship between steroids and cancers of the reproductive system was pioneered by the work of Sir George Beatson in the nineteenth century, who began to define the relationship between estrogen and BrCa risk [16]. Subsequently, in the 1940s this concept was echoed by the work of Dr. Charles Huggins and colleagues who established the endocrine synthesis of androgens and the relationship to PCa [17]. As a result, the genomic functions of AR and ERα in PCa and BrCa, respectively, are highly studied and these are well understood TFs. Additionally, there is a parallel and, in many cases, emerging appreciation of how these cancers are impacted by non-steroidal NRs, and the potential for the genomic cross-talk between steroidal and non-steroidal NRs. For example, there are physiological and gene regulatory studies that strongly support the concept that Type I and Type II NRs function in a range of cooperative and antagonist cross-talk signaling mechanisms, for example centered around AR [18,19,20,21,22,23], and ERα [24,25,26,27,28,29].
The focus of the current chapter is to summarize genomic insights into the Type II NRs in hormone-dependent cancer including the vitamin D receptor (NR1I1VDR), retinoic acid receptors (NR1B1/RARα, NR1B2/RARβ, and NR1B3/RARγ), and peroxisome proliferator-activated receptors (NR1C1/PPARα, NR1C2/PPARδ, and NR1C3/PPARγ) [9], summarized in Table 13.1. Clearly, orphan receptors, given they have no identified ligands, also fall under the classification of non-steroidal receptors. In parallel, the understanding of adopted nuclear orphans and orphan NRs is evolving, and reveal further insights into NR functions in terms of genomic distribution and cross-talk with signaling pathways including those that are key targets for pharmacological pathways [30].
2 Genomic Interactions of Non-steroidal Nuclear Receptors in PCa and BrCa
2.1 The Vitamin D Receptor
Supporting an anti-tumorigenic role for the VDR men whose prostate tumors have higher VDR expression have significantly lower prostate-specific antigen, lower Gleason score and less advanced tumor stage [31]. The circulating pre-hormone vitamin D3 is the precursor to the active hormone calcitriol (1alpha,25dihydroxyvitaminD3 (1α,25(OH)2D3)) that binds to the VDR. Epidemiological approaches have identified relationships between low circulating vitamin D3 and cancer incidence, and that 1α,25(OH)2D3 suppresses early prostate carcinogenesis by regulating genes involved in proliferation, differentiation and apoptosis [32]. Underscoring the potential importance of this signaling axis, genomic studies in murine VDR knockout cells as well as human studies have suggested that 1α,25(OH)2D3 can regulate as much as 3% of the mouse or human genome directly and/or indirectly [33].
Several studies have assessed the VDR cistrome in PCa [34, 35] by VDR chromatin immunoprecipitation sequencing (ChIP-Seq). Work by Fleet et al [34] identified binding at ~3400 protein-coding genes, ~680 long noncoding RNAs, and ~ 470 miRNAs. This included VDR-bound peaks at known VDR target genes including CYP24A1 and IGFBP3. Peak distribution was evenly divided between intergenic and intronic regions, supporting both long-range and proximal regulation. These studies also suggested that 1α,25(OH)2D3 amplifies signals mediated through other TFs including NF-Kappa-B Inhibitor Alpha (NFKBIA) and FOXO1, and some peaks near immune response related genes (e.g., L1R2) hint towards VDR regulation of immune processes.
A further VDR-ChIP Seq study in non-malignant prostate cells (PrEc) [35] identified ~5000 VDR binding sites, again including well-known targets (e.g., CYP24A1) and, interestingly, ligand activation led to a significant decrease in the number of VDR-ChIP peaks, reflecting perhaps an active role for the basal VDR in gene expression. Sites with loss of peaks include aminoacyl tRNA synthetase genes, which in turn leads to decreased proliferation. VDR also binds near genes regulating neural differentiation, which raises a possibility that itmay also be linked to neuroendocrine trans differentiation in PCa.
Finally, a recent study from our lab [doi.org/10.1101/2022.01.31.478573] addressed VDR function in the context of PCa health disparities by examining a panel of European American (EA) (HPr1-AR and LNCaP) and African American (AA) cell lines (RC43N, RC43T, RC77N and RC77T). These analyses lent strong evidence to the concept that the VDR is a significantly more potent transcriptional regulator in AA than EA prostate cells, and that in PCa this signaling is distorted and suppressed. In non-malignant RC43N cells, VDR ChIP-Seq identified significant basal and 1α,25(OH)2D3 dependent VDR binding sites, with ~1300 in total associated with transcriptional responses enriched for circadian rhythm and inflammation networks. In parallel, 1α,25(OH)2D3-dependent ATAC-Seq also revealed the greatest impact on chromatin accessibility in RC43N cells, with significant gain of nucleosome-free regions at enhancers. By contrast, in malignant EA and AA cell models 1α,25(OH)2D3 led to a loss of VDR binding. Motif prediction identified a diverse set of enriched motifs within peaks, including the VDR motif and other NRs including the AR and RARs. The suppressed transcriptional responses in AA PCa cells associated with reduced expression of Bromodomain adjacent to zinc finger domain protein 1A (BAZ1A), a component of the human SWI/SNF complex, and restored expression of this protein led to significantly enhanced 1α,25(OH)2D3-regulated transcriptome.
There are also equally compelling epidemiological associations between vitamin D3 and breast cancer incidence. For example, the Vdr −/− mouse [36, 37] displays a range of mammary gland phenotypes in terms of disrupted development of the gland, and then changing sensitives to the control of programmed cell death within epithelial cells. In parallel there are a wide range of pre-clinical studies which all support a potentially anti-tumorigenic role in BrCa [38].
Two studies have examined VDR genomic interactions which revealed that in MCF-7 BrCa cells, VDR has ~2300 VDR-binding sites in the absence of 1,25(OH)2D3, and ~7,400 sites following ligand stimulation (4 h). Out of these, ~700 sites remained unchanged in both presence and absence of ligand. A significant numbers of VDR-binding sites were detected in intergenic regions, and distal from promoters, and VDR-bound enhancers were enriched in apoptotic and metabolic pathways. In a series of comprehensive studies led by Kevin White and coworkers [39, 40] multi-cistrome analyses were undertaken for a range of more than 20 NRs including non-steroidal ones in BrCa cancer cell lines [39,40,41]. Within these studies VDR binding was analyzed in MCF-7 cells and also reported ~7000 binding regions, which were more distal to TSS regions than many of the other NRs, and in terms of network topology demonstrated lower interconnectedness compared to NRs such as the retinoic acid receptors. These workers were able to undertake integrative regions.
Together these data strongly support the VDR playing an important role in the biology of the prostate and mammary glands, and suggest disruption of VDR signalling is carcinogenic by disrupting a wide number of gene regulatory mechanisms including overlap with other NRs.
2.2 Retinoic Acid Receptors
The NR1B1/RARα represents one of the earliest examples of targeted cancer therapy, involving all-trans retinoic acid in acute promyelocytic leukemia [42, 43]. This was a major catalyst for the development of the field of differentiation therapy, whereby compounds such as retinoic acid would be in cancers to limit their proliferation and induce either differentiation or programmed cell death [44, 45]. In part, these actions were the motivations for cistromic studies on the VDR, RARs and multiple NRs in PCa, and BrCa [44, 45].
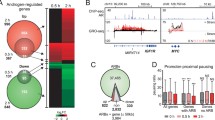
In the prostate, retinoic acid regulates normal differentiation and the Rarγ knockout mouse exhibits prostate metaplasia [46, 47], both suggesting the receptor plays a role in control of cell growth. Reflecting this, NR1B3/RARγ is commonly down-regulated in PCa3, for example because of up-regulated miR-96-5p, and this leads to significant changes to AR signaling [48]. In a non-malignant prostate cell line, RWPE-1, under basal conditions the RARγ cistrome is ~1250 peaks and interestingly the addition of a RARγ-selective ligand (CD437) restricts the number of peaks to ~350, which are mostly shared with the basal state (only ~50 appear unique). These data also revealed that RARγ significantly enhanced AR function, and regulation of AR target genes, and that the RARγ cistrome significantly overlapped with AR binding at active enhancers. In turn, reduced expression genes that were annotated RARγ binding was associated with aggressive PCa [48].
In MCF-7 BrCa cells, RARα/γ and ERα form a genomic antagonism [40] in a so-called “Yin and Yang” manner to regulate proliferation and survival. These NRs balance expression of shared gene targets in part because RARs overlaps significantly with ERα binding in a genome wide fashion. These co-occupied regions are in the vicinity of genes for which estrogen and retinoic acid regulate antagonistically. The number of peaks in the presence of selective RARα (AM580) and RARγ (CD437) ligands was ~7300 for RARα and ~ 3200 for RARγ sites, and using a generous distance cut-off of 1 kb between the center of the peaks there was a significant overlap of sites; it is unclear how many of the peaks actually overlap as opposed to being closely adjacent. This therefore suggests convergence at the level of gene-regulatory actions rather than perhaps direct chromatin-accessibility [40]. Together, these data suggest significant genomic interactions between RARs and both AR and ERα in PCa and BrCa.
Interestingly, the related paralog, RARβ, appears to be a bona fide tumor suppressor in BrCa and PCa. For example, methylation patterns of the CpG islands associated with the RARβ promoter are exploited in algorithms to predict tumor grade and progression risks in these tumors [49,50,51,52]. Against this backdrop it is perhaps surprising that there are no cistrome data for this receptor in these cancers, although it has been undertaken in brain tissues [53].
2.3 RAR Related Orphan Receptor C
NR1F3/RORC encodes RORγ and is amplified and upregulated in metastatic recurrent PCa tumors following androgen deprivation therapy. It acts as an upstream regulator of AR and appears to drive AR expression, as well as to facilitate recruitment of coactivators such as Nuclear Receptor Coactivator 1 and 3 (NCOA1/3, SRC1/3). Furthermore, pharmacological targeting with an antagonist to RORγ reduces expression of AR as well as the oncogenic AR splice variant 7 and reduces AR genomic binding, and as a result reduced expression of various AR target genes. This regulation appears to be a targeted AR event, as inhibiting RORγ does not alter genome-wide histone modifications associated with chromatin accessibility [54].
Studies on RORγ in BrCa have suggested that its function is an essential activator of the cholesterol-biosynthesis program, as it binds to cholesterol-biosynthesis genes, and it facilitates the genomic recruitment of Sterol regulatory element-binding protein 2 (SREBP2) in Triple-negative BrCa [55]. From a genome-wide perspective there appear to be a massive number of RORγ binding sites in the HCC70 BrCa cell line, in excess of 30,000, and these are highly shared with SREBP2 binding sites. Again, similarly to PCa, a RORγ antagonist very potently inhibits BCa tumor growth in vitro and in xenografts [55]. Similarly, the related RORα is also a potential tumor suppressor and a therapeutic target for BrCa [56, 57] but as yet cistromic studies have not been undertaken and so the extent of genomic cooperation between these two receptors remains unknown. RORγ therefore plays a paramount role in regulating cholesterol-biosynthesis through its own genomic binding leads to the recruitment of SREBP2 at the gene targets to stimulate the cholesterol-biosynthesis.
2.4 Peroxisome Proliferator-Activated Receptors
PPARs regulate energy production, lipid metabolism, and inflammation [58]. In triple negative BrCa MDA-MB-231 cells, ChIP-Seq and transcriptomic analyses identified ~500 PPARδ peaks and, amongst these, the hormone ANGPTL4 was a significant PPARδ target [59]. In another study, using a transformed variant of the non-malignant breast epithelial cell, MCF10A-NeuT cells, PPARγ binds to a large number of sites and regulates genes and notably EphA-Amphiregulin as well as genes involved in chemokine signaling [60]. Similarly, PPARα, PPARδ and PPARγ bind to ~2230, ~3250 and ~ 6300 genomic regions respectively in MCF7 cells with PPARγ binding as sites at a greater distal distance to TSS [39] than the other PPARs. Interestingly, the PPARδ cistrome shared a significant proportion (~70%) of its binding sites with RARα and RARγ, and in part this led to the concept of high occupancy target (HOT) regions in the genome. Specifically, these are regions that are significantly shared by multiple NRs and other TFs, and appear to be found disproportionately associated with genes associated with cancer development and progression. The functional significance of these sites is illustrated by shared PPARδ and RARs binding sites at target genes, which in turn are associated with poor prognosis in BrCa. More widely these genomic findings also support a concept of selectively targeting RARs and PPARδ to inhibit synergistically BrCa growth.
Set against these interesting data, to date there are no cistromic studies of PPARs in PCa. This is all the more striking given that there is a considerable literature on PPARs [61,62,63,64,65] and the PPAR coregulator PPARGC1α [66,67,68] playing significant roles in PCa carcinogenesis. Such studies would also be able to address the concept of HOT regions in PCa, and how these cistromic patterns impact AR signaling.
2.5 Hepatocyte Nuclear Factor 4 α and γ
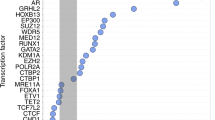
In PCa, NR2A1/HNF4γ appears to function as a pioneer factor that generates and maintains enhancer landscape at lineage genes, for example those associated with neuronal lineages, and which impacts AR signaling in a more nuanced manner. For example, restoring HNF4γ expression reduces AR sensitivity towards androgen deprivation therapy [69], and increased HNF4γ expression does not alter the AR cistrome or AR signaling directly, but increased FOXA1 binding at a subset of HNF4γ sites. Approximately 35% of HNF4γ peaks share binding FOXA1, and a smaller proportion of HNF4γ peaks directly overlap with AR peaks. Therefore, HNF4γ binding sites appear to cooperate with FOXA1 to establish and maintain enhancers that facilitate lineage-specific transcriptomes in the prostate; this is potentially corrupted in PCa progression [69]. Similarly, NR2A2/HNF4α appears to exert a tumor suppressor function and has reduced expression in PCa tissues, cell lines, and xenografts of androgen deprivation therapy recurrent PCa [70] through epigenetic mechanisms. For example, HNF4α binds constitutively to binding sites in the promoter of CDKN1A, which guides AR to bind upon dihydrotestosterone stimulation. Indeed, the motifs of HNF4α are over-represented within unique AR-binding loci, and the cistrome shows significant overlap with AR-binding sites [71]. Again, given these potent cooperative actions between HNF4 receptors with a principal steroid hormone receptor, it is perhaps surprising that similar studies haven’t yet been undertaken in BrCa.
2.6 COUP Transcription Factor I and II
NR2F1/COUP-TF I is one of the earliest cloned NRs, first being identified in the late 1980s [72], and subsequently led to the discovery of NR2F2/COUP-TF II [73]. Several studies [39, 74, 75] have analyzed the COUP-TF II cistrome in BrCa. High expression of COUP-TF II is related with better survival in ERα-positive BrCa patients but not in ERα-negative patients, and COUP-TF II cooperates with pioneer factors such as FOXA1 and GATA3 to promote ERα function [74, 75]. These findings suggest a cooperativity between ERα and COUP-TF-11, and although estradiol is not required for COUP-TF II binding, inhibition of COUP-TF II decreases ERα binding, chromatin accessibility (ATAC-Seq peaks were reduced by 70% after COUP-TF II depletion), and estradiol-dependent cell growth suggesting a protein-protein interaction. Together, these data suggest a complex interdependency between estradiol, ERα and COUP-TF II. In MCF-7 cells, approximately, 40% of ERα binding sites overlap with FOXA1, 60% with COUP-TF II and 70% with GATA3, and there is evidence for shared binding at super-enhancers on a wide-spread scale which directly leads to high de novo transcription. Indeed, this integration also impacts other NRs downstream, including RARβ [76]. These roles for COUP-TF II in regulating ERα-mediated transcription make it an interesting potential therapeutic target in BCa. In parallel studies COUP-TF I-specific agonists suppress metastasis supporting a wider role for COUP-TFs to interact with ERα and to regulate anticancer actions [77].
2.7 NUR77
NR4A1/NUR77 is an orphan NR that acts in a ligand-independent manner. In a recent study [78], NUR77 was reported to regulate immediate early genes, suppressing replication stress in BCa and acting as a master regulator through a transcriptional processing checkpoint. Genome-wide analyses revealed that NUR77 binds the gene body and 3’ UTR of immediate early genes, inhibits transcriptional elongation, generating R-loops and accessible chromatin domains. Under stress, dissociation of NUR77 leads to a burst of expression of these transcriptionally poised genes thereby suggesting a role for NUR77 in governing transcriptional responses to chronic replication stress. Although there are no genome-wide cistrome studies of NUR77 in PCa, there is strong evidence for it regulating programmed cell death in this cancer [79, 80].
3 Mechanisms of NR Cooperation: Bookmarking Functions by Non-steroidal NRs
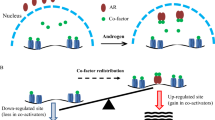
Mitotic bookmarking functions to retain epigenetic states throughout the cell cycle at gene loci that are poised for immediate reactivation post-mitotically (Fig. 13.1). This involves the retention of histone variants, regulatory proteins and modifications, and some selected TFs. Bookmarking mechanisms prevent the spreading of heterochromatin into genomic regions which are pre-marked for TF future actions. In this manner, these epigenetic mechanisms regulate genes that coordinately control cell growth and lineage maintenance following mitosis. Furthermore, it is clear these mechanisms are corrupted in carcinogenesis and tumor maintenance leading to deregulated proliferation and compromised control of differentiation [81,82,83,84,85,86].
A model for bookmarking function by Type II nuclear receptors for Type I nuclear receptors. As cells go through the cell cycle and division chromatin assumes different conformations, becoming most compacted during metaphase of mitosis. Prior to this, many proteins to chromatin associations are lost as a result of degradation and displacement. However, a number of transcription factors are retained such that transcription, or the marking of sites for transcription, can be activated rapidly in G1. This function is termed bookmarking and there is evidence that Type II nuclear receptors that are nuclear resident in both the presence and absence of ligand (mitosis, purple open symbols) can serve this function for other nuclear receptors (G1, purple solid symbol)
Several Type II NRs have been reported to have bookmarking properties independent of ligand exposure, again reflecting their predominant nuclear location. NR1I2/PXR remains constitutively associated with mitotic chromatin specifically at the CYP3A4 promoter during mitosis [88]. A region of PXR contains a ‘mitotic chromatin binding-determining region’ which exerts these functions. The bookmarking property of PXR is impeded by direct interaction with the orphan NR small heterodimer partner (SHP) perhaps underscoring the importance of this function [89]. Other examples of NRs appearing to play a bookmarking function include NR3B2/ESRBB, which is a major pluripotency TF that remains bound to key regulatory regions during mitosis [90]; it is bound widely with at least 10,000 binding sites and maintains nucleosome positioning during mitosis to ensure the rapid post-mitotic re-establishment of functional regulatory complexes at selected enhancers and promoters [91]. Similarly, HNF4α bookmarks specific genomic regions and keeps them competent for future activation during liver development [92].
The raises an interesting question of whether this bookmarking property is a generalized feature of NRs, and specifically those NRs that are nuclear resident independent of ligand exposure maybe retaining enhancer access through mitosis for other NRs. This concept is supported by examples above of Type II and Type I NR co-regulation of gene expression programs. Given that non-steroidal NRs in PCa and BrCa are frequently disrupted for example with decreased expression (e.g., RARγ), this may suggest that Type II NRs bookmark and regulate the actions of AR and ERα. However, there is also evidence of ligand activated (and therefore nuclear resident) AR and ERα being associated with mitotic chromatin although it is unclear if these complexes are the cause or consequence of other NRs/TFs serving as bookmarking factors [87].
More generally, there are clear examples of AR and ERα being genomically relocated to other sites during cancer initiation and progression, and in response to NR-targeted therapies. For example, the AR is reprogrammed specifically to genomic sites that are normally regulated in development only in the transition to metastatic PCa by reactivating latent regulatory elements active in fetal prostate organogenesis [93]. It is a tantalizing prospect that the interactions between Type II and Type I NRs is in part underpinned by Type II NR bookmarking enhancers and regulatory regions that are regulated by Type I NR binding to promote cell fate decisions such as differentiation. Furthermore, disruption of these Type II NR complexes potentially disrupts these functions.
4 Genomic Approaches to Defining Type I and II NR Cistromes and Interactions
Methods to map histone and TF genomic interactions emerged in the 1990s with the development of ChIP approaches [94,95,96], and became genome-wide with the advent of microarray technologies giving rise to so-called ChIP–chip [97] approaches, and then subsequently ChIP-Seq [98]. This key technology has been profoundly improved and diversified to tackle limitations such as protein abundance, cross-linking efficiency and antibody availability and specificity. For example, Cleavage Under Targets & Release Using Nuclease (CUT&RUN) and CUT&Tag has made it easier to study TF binding and histone modifications at genome scale [99, 100]. Similarly, the development of ATAC-Seq (Transposase Accessible Chromatin followed by high-throughput sequencing) [101] has enabled the measurement of chromatin accessibility and has also been refined to address single cells and to improve accuracy. More widely, genomic approaches are advancing rapidly to encompass single cell resolution, which allows ever more complex biological questions to be addressed [102]. In parallel, CRISPR technologies are enabling the tagging of proteins, and DNA and epigenome editing, to more establish conditional cell contexts with which to test NR functions more accurately [103, 104].
Matching these wet-lab advancements has been an equally explosive growth in the dry-lab to develop and refine the analyses of cistromic data and combine it with parallel transcriptomic data. This challenge of integrating cistrome to transcriptome data is surprisingly complex. For example, defining NR:enhancer:gene interactions that are driven by NRs is challenging because of the large number of NR and coregulator interactions, which are altered by diverse and interdependent genetic and epigenetic mechanisms, and are further controlled by the 3-D genome [105,106,107]. Thus, NR:enhancer:gene relationships are dynamic and non-linear, with each gene regulated by multiple enhancers in a time- and signal-dependent manner [108, 109], and occur over large genomic distances [110].
Defining the statistical significance of NR:gene relationships, or even NR:NR:gene relationships, is a question of whether a NR signal-to-gene-expression relationship is occurring more than predicted by chance, which in turn requires defining the background of NR:gene relationships. Random sampling methods such as bootstrapping can be used to simulate the distribution of NR:gene relationships changes across the genome for statistical comparison [111, 112], and parsimonious annotation of the genome, for example with the ChromHMM algorithm [113] to define epigenetic states, or the ROSE algorithm to define super-enhancers [114] [115] can refine these statistical challenges. Furthermore, testing the overlap of target NR ChIP-Seq data with comprehensive data sets, such as contained in Cistrome DB [116], allows co-enrichment testing of hundreds of TF and histone modification ChIP-Seq datasets to reveal the extent of enrichment with other NRs and their coregulators. RNA-Seq undertaken in parallel treatments can be matched with these highly annotated cistromic data to define cistrome-transcriptome relationships and test their phenotypic associations for example using Kolmogorov–Smirnov tests to examine differences in cumulative distribution plots for cistrome binding sites with respect to nearest gene, and again using bootstrapping approaches to measure how the specific cistrome-relationships associate with gene expression patterns [117].
Thus, there are many routes through testing NR:gene relationships and this most likely underpins the frequently divergent findings in the literature. On top of this there are multiple methods for cistrome [118, 119] or transciptome [120, 121] analyses and as yet there are few commonly accepted protocol standards, in contrast, for example to the MIAME-compliant protocols for microarray analyses [122]. Therefore, it is unsurprising that for a given NR there is little consensus on the number of significant binding sites, what motifs are most enriched, what the genomic distribution is and how it relates to transcription.
5 Conclusion
Non-steroidal NRs have been somewhat neglected from a genomic perspective, although it is clear their actions and interactions with steroidal NRs are biologically impactful. In this chapter we attempted to provide a broad overview of the advances in understanding non-steroidal nuclear receptor cistromes and their interaction with other AR and ERα in PCa and BrCa and highlighted the expanding impact of the genome wide studies in NR biology. These NRs are potential therapeutic targets in cancer and may be exploited to augment traditional therapeutic approaches. Cistromic studies are rapidly advancing and revealing unprecedented insights into the interactions between Type I and Type II NRs, even with some methodological ambiguities.
References
De Bosscher K, Desmet SJ, Clarisse D, Estebanez-Perpina E, Brunsveld L (2020) Nuclear receptor crosstalk – defining the mechanisms for therapeutic innovation. Nat Rev Endocrinol 16:363–377
Zhao L, Zhou S, Gustafsson JA (2019) Nuclear receptors: recent drug discovery for cancer therapies. Endocr Rev 40:1207–1249
Long MD, Campbell MJ (2015) Pan-cancer analyses of the nuclear receptor superfamily. Nucl Receptor Res 2
Duez H, Staels B (2010) Nuclear receptors linking circadian rhythms and cardiometabolic control. Arterioscler Thromb Vasc Biol 30:1529–1534
Sonoda J, Pei L, Evans RM (2008) Nuclear receptors: decoding metabolic disease. FEBS Lett 582:2–9
Takahashi N, Goto T, Hirai S, Uemura T, Kawada T (2009) Genome science of lipid metabolism and obesity. Forum Nutr 61:25–38
Ruskovska T et al (2021) Systematic Bioinformatic analyses of nutrigenomic modifications by polyphenols associated with Cardiometabolic health in humans-evidence from targeted nutrigenomic studies. Nutrients 13
Bravo-Ruiz I, Medina MA, Martinez-Poveda B (2021) From food to genes: transcriptional regulation of metabolism by lipids and carbohydrates. Nutrients 13
Tao LJ, Seo DE, Jackson B, Ivanova NB, Santori FR (2020) Nuclear hormone receptors and their ligands: metabolites in control of transcription. Cell 9
Fuller PJ, Yang J, Young MJ (2017) 30 years of the mineralocorticoid receptor: Coregulators as mediators of mineralocorticoid receptor signalling diversity. J Endocrinol 234:T23–T34
Schnyder S, Kupr B, Handschin C (2017) Coregulator-mediated control of skeletal muscle plasticity - a mini-review. Biochimie 136:49–54
Obeid JP, Zafar N, El Hokayem J (2016) Steroid hormone receptor Coregulators in endocrine cancers. IUBMB Life 68:504–515
Dasgupta S, Lonard DM, O’Malley BW (2014) Nuclear receptor coactivators: master regulators of human health and disease. Annu Rev Med 65:279–292
Weikum ER, Liu X, Ortlund EA (2018) The nuclear receptor superfamily: a structural perspective. Protein Sci 27:1876–1892
Evans RM, Mangelsdorf DJ (2014) Nuclear receptors, RXR, and the big bang. Cell 157:255–266
Paul J (1981) Sir George Beatson and the Royal Beatson Memorial Hospital. Med Hist 25:200–201
Huggins C, Stevens R, Hodges CV (1941) Studies on prostatic cancer: II. The effects of castration on advanced carcinoma of the prostate gland. Arch Surg 43:209–223
Smith KW, Thompson PD, Rodriguez EP, Mackay L, Cobice DF (2019) Effects of vitamin D as a regulator of androgen intracrinology in LNCAP prostate cancer cells. Biochem Biophys Res Commun 519:579–584
Olokpa E, Bolden A, Stewart LV (2016) The androgen receptor regulates PPARgamma expression and activity in human prostate cancer cells. J Cell Physiol 231:2664–2672
Eskra JN, Kuiper JW, Walden PD, Bosland MC, Ozten N (2017) Interactive effects of 9-cis-retinoic acid and androgen on proliferation, differentiation, and apoptosis of LNCaP prostate cancer cells. Eur J Cancer Prev 26:71–77
Wang JH, Tuohimaa P (2008) Calcitriol and TO-901317 interact in human prostate cancer LNCaP cells. Gene Regul Syst Bio 2:97–105
Cai Y et al (2003) Cytochrome P450 genes are differentially expressed in female and male hepatocyte retinoid X receptor alpha-deficient mice. Endocrinology 144:2311–2318
Lareyre JJ et al (2000) Characterization of an androgen-specific response region within the 5′ flanking region of the murine epididymal retinoic acid binding protein gene. Biol Reprod 63:1881–1892
Yetkin D, Balli E, Ayaz F (2021) Antiproliferative activity of tamoxifen, vitamin D3 and their concomitant treatment. EXCLI J 20:1394–1406
Yang S et al (2021) Differential effects of estrogen receptor alpha and beta on endogenous ligands of peroxisome proliferator-activated receptor gamma in papillary thyroid cancer. Front Endocrinol (Lausanne) 12:708248
Tanaka N et al (2021) Pemafibrate, a novel selective PPARalpha modulator, attenuates tamoxifen-induced fatty liver disease. Clin J Gastroenterol 14:846–851
Verma A, Cohen DJ, Jacobs TW, Boyan BD, Schwartz Z (2021) The relative expression of ERalpha isoforms ERalpha66 and ERalpha36 controls the cellular response to 24R,25-Dihydroxyvitamin D3 in breast cancer. Mol Cancer Res 19:99–111
Hasan N, Sonnenschein C, Soto AM (2019) Vitamin D3 constrains estrogen’s effects and influences mammary epithelial organization in 3D cultures. Sci Rep 9:7423
Fan P, Abderrahman B, Chai TS, Yerrum S, Jordan VC (2018) Targeting peroxisome proliferator-activated receptor gamma to increase estrogen-induced apoptosis in estrogen-deprived breast cancer cells. Mol Cancer Ther 17:2732–2745
de Vera IMS (2018) Advances in orphan nuclear receptor pharmacology: a new era in drug discovery. ACS Pharmacol Transl Sci 1:134–137
Hendrickson WK et al (2011) Vitamin D receptor protein expression in tumor tissue and prostate cancer progression. J Clin Oncol 29:2378–2385
Feldman D, Krishnan AV, Swami S, Giovannucci E, Feldman BJ (2014) The role of vitamin D in reducing cancer risk and progression. Nat Rev Cancer 14:342–357
Bouillon R et al (2008) Vitamin D and human health: lessons from vitamin D receptor null mice. Endocr Rev 29:726–776
Fleet JC et al (2019) Vitamin D signaling suppresses early prostate carcinogenesis in TgAPT121 mice. Cancer Prev Res (Phila) 12:343–356
Baumann B, Lugli G, Gao S, Zenner M, Nonn L (2019) High levels of PIWI-interacting RNAs are present in the small RNA landscape of prostate epithelium from vitamin D clinical trial specimens. Prostate 79:840–855
Tavera-Mendoza LE et al (2017) Vitamin D receptor regulates autophagy in the normal mammary gland and in luminal breast cancer cells. Proc Natl Acad Sci U S A 114:E2186–E2194
Zinser G, Packman K, Welsh J (2002) Vitamin D(3) receptor ablation alters mammary gland morphogenesis. Development 129:3067–3076
Vanhevel J, Verlinden L, Doms S, Wildiers H, Verstuyf A (2022) The role of vitamin D in breast cancer risk and progression. Endocr Relat Cancer 29:R33–R55
Kittler R et al (2013) A comprehensive nuclear receptor network for breast cancer cells. Cell Rep 3:538–551
Hua S, Kittler R, White KP (2009) Genomic antagonism between retinoic acid and estrogen signaling in breast cancer. Cell 137:1259–1271
Al-Dhaheri M et al (2011) CARM1 is an important determinant of ERalpha-dependent breast cancer cell differentiation and proliferation in breast cancer cells. Cancer Res 71:2118–2128
Agadir A et al (1994) Retinoic acid receptors: involvement in acute promyelocytic leukemia. Cell Mol Biol (Noisy-le-Grand) 40:263–274
Lin RJ et al (1998) Role of the histone deacetylase complex in acute promyelocytic leukaemia. Nature 391:811–814
de The H (2018) Differentiation therapy revisited. Nat Rev Cancer 18:117–127
Spira AI, Carducci MA (2003) Differentiation therapy. Curr Opin Pharmacol 3:338–343
Aboseif SR, Dahiya R, Narayan P, Cunha GR (1997) Effect of retinoic acid on prostatic development. Prostate 31:161–167
Lohnes D et al (1993) Function of retinoic acid receptor gamma in the mouse. Cell 73:643–658
Long MD et al (2019) The miR-96 and RARgamma signaling axis governs androgen signaling and prostate cancer progression. Oncogene 38:421–444
de Ruijter TC et al (2020) Prognostic DNA methylation markers for hormone receptor breast cancer: a systematic review. Breast Cancer Res 22:13
Bakavicius A et al (2019) Urinary DNA methylation biomarkers for prediction of prostate cancer upgrading and upstaging. Clin Epigenetics 11:115
Liu X, Giguere V (2014) Inactivation of RARbeta inhibits Wnt1-induced mammary tumorigenesis by suppressing epithelial-mesenchymal transitions. Nucl Recept Signal 12:e004
Jiang D et al (2014) Meta-analyses of methylation markers for prostate cancer. Tumour Biol 35:10449–10455
Niewiadomska-Cimicka A et al (2017) Genome-wide analysis of RARbeta transcriptional targets in mouse striatum links retinoic acid signaling with Huntington’s disease and other neurodegenerative disorders. Mol Neurobiol 54:3859–3878
Wang J et al (2016) ROR-gamma drives androgen receptor expression and represents a therapeutic target in castration-resistant prostate cancer. Nat Med 22:488–496
Cai D et al (2019) RORgamma is a targetable master regulator of cholesterol biosynthesis in a cancer subtype. Nat Commun 10:4621
Du J, Xu R (2012) RORalpha, a potential tumor suppressor and therapeutic target of breast cancer. Int J Mol Sci 13:15755–15766
Mao W et al (2021) RORalpha suppresses cancer-associated inflammation by repressing respiratory complex I-dependent ROS generation. Int J Mol Sci 22
Decara J et al (2020) Peroxisome proliferator-activated receptors: experimental targeting for the treatment of inflammatory bowel diseases. Front Pharmacol 11:730
Adhikary T et al (2013) Inverse PPARbeta/delta agonists suppress oncogenic signaling to the ANGPTL4 gene and inhibit cancer cell invasion. Oncogene 32:5241–5252
Jiao X et al (2021) Ppargamma1 facilitates ErbB2-mammary adenocarcinoma in mice. Cancers (Basel) 13
Galbraith LCA et al (2021) PPAR-gamma induced AKT3 expression increases levels of mitochondrial biogenesis driving prostate cancer. Oncogene 40:2355–2366
Liu S et al (2014) Differential roles of PPARgamma vs TR4 in prostate cancer and metabolic diseases. Endocr Relat Cancer 21:R279–R300
Kaikkonen S, Paakinaho V, Sutinen P, Levonen AL, Palvimo JJ (2013) Prostaglandin 15d-PGJ(2) inhibits androgen receptor signaling in prostate cancer cells. Mol Endocrinol 27:212–223
Govindarajan R et al (2007) Thiazolidinediones and the risk of lung, prostate, and colon cancer in patients with diabetes. J Clin Oncol 25:1476–1481
Butler R, Mitchell SH, Tindall DJ, Young CY (2000) Nonapoptotic cell death associated with S-phase arrest of prostate cancer cells via the peroxisome proliferator-activated receptor gamma ligand, 15-deoxy-delta12,14-prostaglandin J2. Cell Growth Differ 11:49–61
Zheng K, Chen S, Hu X (2022) Peroxisome proliferator activated receptor gamma Coactivator-1 alpha: a double-edged sword in prostate cancer. Curr Cancer Drug Targets
Siddappa M et al (2020) Identification of transcription factor co-regulators that drive prostate cancer progression. Sci Rep 10:20332
Torrano V et al (2016) The metabolic co-regulator PGC1alpha suppresses prostate cancer metastasis. Nat Cell Biol 18:645–656
Shukla S et al (2017) Aberrant activation of a gastrointestinal transcriptional circuit in prostate cancer mediates castration resistance. Cancer Cell 32:792–806 e797
Wang Z et al (2020) Nuclear receptor HNF4alpha performs a tumor suppressor function in prostate cancer via its induction of p21-driven cellular senescence. Oncogene 39:1572–1589
Pihlajamaa P et al (2014) Tissue-specific pioneer factors associate with androgen receptor cistromes and transcription programs. EMBO J 33:312–326
Miyajima N et al (1988) Identification of two novel members of erbA superfamily by molecular cloning: the gene products of the two are highly related to each other. Nucleic Acids Res 16:11057–11074
Ladias JA, Karathanasis SK (1991) Regulation of the apolipoprotein AI gene by ARP-1, a novel member of the steroid receptor superfamily. Science 251:561–565
Erdos E, Balint BL (2020) NR2F2 orphan nuclear receptor is involved in estrogen receptor alpha-mediated transcriptional regulation in luminal a breast cancer cells. Int J Mol Sci 21
Jiang G et al (2019) Cooperativity of co-factor NR2F2 with Pioneer factors GATA3, FOXA1 in promoting ERalpha function. Theranostics 9:6501–6516
Sosa MS et al (2015) NR2F1 controls tumour cell dormancy via SOX9- and RARbeta-driven quiescence programmes. Nat Commun 6:6170
Khalil BD et al (2022) An NR2F1-specific agonist suppresses metastasis by inducing cancer cell dormancy. J Exp Med 219
Guo H et al (2021) NR4A1 regulates expression of immediate early genes, suppressing replication stress in cancer. Mol Cell 81:4041–4058 e4015
Agostini-Dreyer A, Jetzt AE, Stires H, Cohick WS (2015) Endogenous IGFBP-3 mediates intrinsic apoptosis through modulation of Nur77 phosphorylation and nuclear export. Endocrinology 156:4141–4151
Lee KW et al (2007) Contribution of the orphan nuclear receptor Nur77 to the apoptotic action of IGFBP-3. Carcinogenesis 28:1653–1658
Bellec M et al (2022) The control of transcriptional memory by stable mitotic bookmarking. Nat Commun 13:1176
Caravaca JM et al (2013) Bookmarking by specific and nonspecific binding of FoxA1 pioneer factor to mitotic chromosomes. Genes Dev 27:251–260
Kadauke S et al (2012) Tissue-specific mitotic bookmarking by hematopoietic transcription factor GATA1. Cell 150:725–737
Zaidi SK et al (2010) Mitotic bookmarking of genes: a novel dimension to epigenetic control. Nat Rev Genet 11:583–589
Kouskouti A, Talianidis I (2005) Histone modifications defining active genes persist after transcriptional and mitotic inactivation. EMBO J 24:347–357
Michelotti EF, Sanford S, Levens D (1997) Marking of active genes on mitotic chromosomes. Nature 388:895–899
Kumar S, Chaturvedi NK, Kumar S, Tyagi RK (2008) Agonist-mediated docking of androgen receptor onto the mitotic chromatin platform discriminates intrinsic mode of action of prostate cancer drugs. Biochim Biophys Acta 1783:59–73
Rana M, Dash AK, Ponnusamy K, Tyagi RK (2018) Nuclear localization signal region in nuclear receptor PXR governs the receptor association with mitotic chromatin. Chromosom Res 26:255–276
Kumar S, Vijayan R, Dash AK, Gourinath S, Tyagi RK (2021) Nuclear receptor SHP dampens transcription function and abrogates mitotic chromatin association of PXR and ERalpha via intermolecular interactions. Biochim Biophys Acta Gene Regul Mech 1864:194683
Festuccia N et al (2016) Mitotic binding of Esrrb marks key regulatory regions of the pluripotency network. Nat Cell Biol 18:1139–1148
Festuccia N et al (2019) Transcription factor activity and nucleosome organization in mitosis. Genome Res 29:250–260
Karagianni P, Moulos P, Schmidt D, Odom DT, Talianidis I (2020) Bookmarking by non-pioneer transcription factors during liver development establishes competence for future gene activation. Cell Rep 30:1319–1328 e1316
Pomerantz MM et al (2020) Prostate cancer reactivates developmental epigenomic programs during metastatic progression. Nat Genet 52:790–799
Franke A et al (1992) Polycomb and polyhomeotic are constituents of a multimeric protein complex in chromatin of Drosophila melanogaster. EMBO J 11:2941–2950
Dedon PC, Soults JA, Allis CD, Gorovsky MA (1991) A simplified formaldehyde fixation and immunoprecipitation technique for studying protein-DNA interactions. Anal Biochem 197:83–90
Orlando V, Paro R (1995) Chromatin multiprotein complexes involved in the maintenance of transcription patterns. Curr Opin Genet Dev 5:174–179
Buck MJ, Lieb JD (2004) ChIP-chip: considerations for the design, analysis, and application of genome-wide chromatin immunoprecipitation experiments. Genomics 83:349–360
Johnson DS, Mortazavi A, Myers RM, Wold B (2007) Genome-wide mapping of in vivo protein-DNA interactions. Science 316:1497–1502
Kaya-Okur HS et al (2019) CUT&tag for efficient epigenomic profiling of small samples and single cells. Nat Commun 10:1930
Skene PJ, Henikoff S (2017) An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. elife 6
Buenrostro JD, Giresi PG, Zaba LC, Chang HY, Greenleaf WJ (2013) Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods 10:1213–1218
King HW et al (2021) Integrated single-cell transcriptomics and epigenomics reveals strong germinal center-associated etiology of autoimmune risk loci. Sci Immunol 6:eabh3768
Chakravarti R et al (2022) A review on CRISPR-mediated epigenome editing: a future directive for therapeutic management of cancer. Curr Drug Targets
Liu G, Lin Q, Jin S, Gao C (2022) The CRISPR-Cas toolbox and gene editing technologies. Mol Cell 82:333–347
Guo Y et al (2018) CRISPR-mediated deletion of prostate cancer risk-associated CTCF loop anchors identifies repressive chromatin loops. Genome Biol 19:160
Rhie SK et al (2019) A high-resolution 3D epigenomic map reveals insights into the creation of the prostate cancer transcriptome. Nat Commun 10:4154
Taberlay PC et al (2016) Three-dimensional disorganization of the cancer genome occurs coincident with long-range genetic and epigenetic alterations. Genome Res 26:719–731
Roadmap Epigenomics C et al (2015) Integrative analysis of 111 reference human epigenomes. Nature 518:317–330
Thurman RE et al (2012) The accessible chromatin landscape of the human genome. Nature 489:75–82
Jung I et al (2019) A compendium of promoter-centered long-range chromatin interactions in the human genome. Nat Genet 51:1442–1449
Franke J, Neumann MH (2000) Bootstrapping neural networks. Neural Comput 12:1929–1949
Long MD et al (2015) Integrative genomic analysis in K562 chronic myelogenous leukemia cells reveals that proximal NCOR1 binding positively regulates genes that govern erythroid differentiation and Imatinib sensitivity. Nucleic Acids Res 43:7330–7348
Ernst J, Kellis M (2017) Chromatin-state discovery and genome annotation with ChromHMM. Nat Protoc 12:2478–2492
Kim DH et al (2021) BET Bromodomain inhibition blocks an AR-repressed, E2F1-activated treatment-emergent neuroendocrine prostate cancer lineage plasticity program. Clin Cancer Res 27:4923–4936
Whyte WA et al (2013) Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153:307–319
Layer RM et al (2018) GIGGLE: a search engine for large-scale integrated genome analysis. Nat Methods 15:123–126
Long MD, van Den B, Singh P, Battaglia S, Campbell MJ (2015) Integrative genomic analysis in K562 chronic myelogenous leukemia cells reveals that proximal NCOR1 binding positively regulates genes that govern erythroid differentiation and Imatinib sensitivity. Nucleic Acids Res 43:7330–7348
Feng J, Liu T, Qin B, Zhang Y, Liu XS (2012) Identifying ChIP-seq enrichment using MACS. Nat Protoc 7:1728–1740
Lun AT, Smyth GK (2016) Csaw: a Bioconductor package for differential binding analysis of ChIP-seq data using sliding windows. Nucleic Acids Res 44:e45
Law CW et al (2016) RNA-seq analysis is easy as 1-2-3 with limma, Glimma and edgeR. F1000Res 5
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C (2017) Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods 14:417–419
Brazma A et al (2001) Minimum information about a microarray experiment (MIAME)-toward standards for microarray data. Nat Genet 29:365–371
Funding
MJC acknowledge support in part from the Prostate program of the Department of Defense Congressionally Directed Medical Research Programs [W81XWH-20-1-0373; W81XWH-21-1-0850]; the Breast program of the Department of Defense Congressionally Directed Medical Research Programs [W81XWH-21-1-0555]; Prostate Cancer UK [RIA18-ST2–022]. MJC also acknowledges National Institute of Health Cancer Center Support Grant (P30CA016058) to the OSUCCC The James.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Wani, S.A., Campbell, M.J. (2022). Genomic Insights into Non-steroidal Nuclear Receptors in Prostate and Breast Cancer. In: Campbell, M.J., Bevan, C.L. (eds) Nuclear Receptors in Human Health and Disease. Advances in Experimental Medicine and Biology, vol 1390. Springer, Cham. https://doi.org/10.1007/978-3-031-11836-4_13
Download citation
DOI: https://doi.org/10.1007/978-3-031-11836-4_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-11835-7
Online ISBN: 978-3-031-11836-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)