Abstract
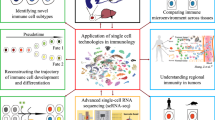
The immune system functions to protect the host from diverse pathogenic microorganisms as well as cancers, where transcriptional regulations in various immune cells are stringently controlled to ensure the appropriate activities of the system. Therefore, the transcriptional landscapes in different types of immune cells are distinctive to achieve respective roles in the immune system. The transcriptome analysis in bulk or a population of cells provides essential information which promotes a better understanding of gene regulations and molecular mechanisms shaping respective cell types. Furthermore, given the substantial heterogeneity in immune cells, transcriptome analysis at a single cell resolution would be contributive for identifying disease-relevant cells and genes, which facilitate us to understand the underlying mechanisms of immune disorders. In this chapter, we discuss the relevance of transcriptional profiling of immune cells including single-cell transcriptomes which have great potential to dissect the function of the immune system.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Immune system
- Transcriptome analysis
- Single cell resolution
- Single-cell RNA sequencing
- mTECs
- RNA Sequencing (RNA-seq)
- Microarrays
- Infections
- Autoimmune disorders
- Immune dysregulation
- Peripheral blood mononuclear cells (PBMCs)
- Single-cell transcriptomics
- Differentially expressed genes (DEGs)
- Thymus
- Thymic epithelial cells (TECs)
- Central tolerance
- Autoimmune regulator (AIRE)
- APECED
- mTEC
- cTEC
- Post-Aire mTEC
- Corneocyte-like mTEC
- mTEC IV
- Thymic tuft cells
Immunology, which is the study for the immune system, started in the late nineteenth century beginning with two significant discoveries. One was the phagocytosis by macrophages which plays a critical host-defense mechanism against invading pathogens found by Elie Metchnikoff (1845–1916). The other one was an antibody which can neutralize microbial toxins discovered by Emil von Behring (1854–1917) and Paul Ehrlich (1854–1915) (Kaufmann 2017). Since then, immunology has been a field of intensive biomedical research and contributed to society by providing pivotal knowledge on both basic science and clinical applications along with its development. While the classical and authentic function of the immune system is to protect our bodies from diverse pathogenic microorganisms, including bacterias, viruses, and parasites, recent immunological studies revealed different parts of the immune system in eliminating cancer cells and regulating physiologic processes in diverse tissues such as the nervous system function, metabolic state, thermogenesis, and tissue repair (Chaplin 2010; Rankin and Artis 2018; Rouse and Sehrawat 2010). Now, we recognize the immune system is a multifunctional biological system and vital for our health and survival.
The immune system is composed of different types of immune cells, which do not form a single organ like the brain and heart but are spread throughout the body to achieve rapid responses to invading pathogens. As transcriptional regulation plays a crucial role in shaping these immune cells of diverse differentiation and activation status, various immune cells were examined at the transcriptional level. These profiling analyses effectively yielded relevant insights of immune cells regarding respective regulatory mechanisms and crucial factors involved in cell differentiation and activations (Amit et al. 2011; Lara-Astiaso et al. 2014; Mostafavi et al. 2016; Smale and Fisher 2002; Uhlen et al. 2019). Furthermore, recent single-cell transcriptome analyses by single-cell RNA sequencing (scRNA-seq) provide unprecedented high-resolution insights of immune cells which cannot be captured by studies in bulk and are expected to promote our understanding of the nature of immune cells in both physiological and pathological contexts (Proserpio and Mahata 2016; Roy 2019; Seumois and Vijayanand 2019; Stubbington et al. 2017; Xie et al. 2021). We introduce general features of the immune system and discuss the transcriptome analysis applied to explore the immune system.
1 The Immune System and Immune Cells
The immune system not only protects us from diverse infections, including bacterias, viruses, and parasites, but also eliminates cancer cells and healing wounds. The efficiency of the immune activity relies on the orchestrated functions of a set of different types of immune cells, which are responsible for the diverse steps of the process. In the case of infections, for example, these include pathogen recognition, the cascade to recruit and activate effector cells, and the final clearance by other immune cells. Identifications of different types of cells involved in the immune process have been a keen target of immunological research for decades, and accordingly, types of immune cells have been expanded, which contributed to the dissection of the immune functions. It was started from the discovery of white blood cells in 1843 by Gabriel Andral (1797–1876) and William Addison (1802–1881). Then, different types of immune cells have been progressively identified along with the development of technologies such as flow cytometry in the 1960s and monoclonal antibodies in the 1970s, which were collectively employed to specify CD4+ T cells and CD8+ T cells, for instance (Hajdu 2003; Jayasinghe 2020; Packer 2021). The major populations of immune cells include granulocytes and macrophages with innate ability to phagocytose bacteria, antibody-producing B cells which were discovered before the 1990s, and more than 80 immune cell populations are recognized to date (Fig. 10.1) (Ackerman 1964; Hayakawa et al. 1983; Maecker et al. 2012; Stein et al. 1992).
Overview of immune cell populations
Arrows indicate schematic representation of the standard model for hematopoietic stem cell differentiation. Self-reactive T cells are eliminated by negative selection after the CD4/CD8 double-positive (DP) stage in the thymic medulla with the help of mTECs. ILC innate lymphoid cell, NK natural killer cell, gdT gamma delta (γδ) T cell, Treg regulatory T cell, DC dendritic cell
While different immune cells possess distinctive functions, essentially all immune cells develop from a hematopoietic stem cell in the bone marrow and share the same genome except for rearranged genes (i.e., T cell receptor and immunoglobulin). Through the differentiation pathways that can be parsed up to as many as ten successive steps, immune cells acquire their divergent capabilities, which are established by correspondent transcriptional landscapes (Hardy and Hayakawa 2001; Rothenberg 2014). As such, transcriptional regulations are the most fundamental mechanisms controlling immune cells and the immune system. Thus, it is not surprising that the recent development of single-cell transcriptomics is promising to confirm existing populations and unveil new populations efficiently in an unbiased manner.
2 Transcriptome Analysis of Different Subsets in Bulk
While more than 80 immune cell subsets are recognized throughout our body, many subsets residing in lymph nodes, tissues, and organs, immunocytes in peripheral blood are the most feasible cells to be examined for research and clinical diagnostics (Chou and Li 2018; Maecker et al. 2012; Novershtern et al. 2011). A few large transcriptomic studies have been done on different immune cell populations in blood. For example, 13 immune cell types in peripheral blood were examined by Schmiedel et al. (2018) 29 immune cell types by Monaco et al. (2019), and 18 immune cell types by Uhlen et al. (2019). In these studies, they isolated immune cells in blood including such as monocytes, natural killer (NK) cells, neutrophils, basophils, B cells, CD4+ and CD8+ T cells , as well as dendritic cells (DCs) employing known markers and fluorescence-activated cell sorters (FACS), and profiled whole transcriptomes by RNA sequencing (RNA-seq) or microarrays. These studies have revealed the distinctive global expression profiles of various immune cells where granulocyte cell types (neutrophils, basophils, and eosinophils) are discrete from others, all lymphocytes make a cluster, including T cells, NK cells, and B cells. In contrast, monocytes are closely related to DCs. According to the study by Uhlen et al., among ~16,000 genes detected in 18 immune cell types, ~10,000 were detected in single-cell types, which were almost comparable to genes detected in cell lines (~9,500 genes per cell line). Of these, 5,934 genes were seen across all immune cells and 1,713 genes in a single-cell type: 9,939 genes showed low specificity for cell types. The sets of differentially expressed and co-expressed genes were served to deduce the functional modules of genes with the aid of bioinformatics such as enrichment analysis using the gene ontology (GO). Furthermore, these transcriptome atlases in each cell type are valuable to promote the understanding of primary immunodeficiency diseases (PID). PID are a large group of over 400 different diseases caused by quantitative and functional changes in the various mechanisms involved in immune response and associated with complications including infections, autoimmune disorders, immune dysregulation with lymphoproliferation, inflammatory disorders, lymphomas, and other types of cancers (Amaya-Uribe et al. 2019; Sánchez-Ramón et al. 2019). While PID are caused by genetic disorders and 354 diseases were listed as consequences of monogenic defects in genes associated with the immune system involving 224 identified genes, the mechanism of disease is often incompletely understood (Uhlen et al. 2019) (https://www.omim.org). Uhlen et al. hypothesized that an analysis of cellular expression of identified genes could help generate a better mechanistic investigation and analyzed 224 PID genes across their 18 immune cell populations. They divided these PID genes into seven clusters according to the shared expression pattern among cell populations and found some PID genes are expressed explicitly in restricted populations. These included the CEBPE gene in which mutations can cause specific granule deficiency 1 (SG1) highly expressed in eosinophils. Although SG1 has been considered a neutrophil-granule deficiency associated with recurrent pyogenic infections, CEBPE’s expression in eosinophils suggested that eosinophil deficiency might also be involved in SG1.
It is also worth noting that the variable mRNA abundance in different immune cells was carefully examined in the study by Monaco et al., and they developed an enhanced method for normalization. The normalization for mRNA abundance can become essential for differential expression analyses. For example, if the analysis is done with two cell types of essentially different total mRNA amounts per cell (e.g., same 10 gene X mRNA molecules are expressed, but cell type A is expressing 100 mRNA molecules in total, and cell type B is expressing 1,000 mRNA molecules in total), this can lead to the misleading that the gene X is downregulated in cell type B. Indeed, existing normalization methods for transcriptome profiling such as the UQ, TMM, and RLE cannot correctly identify transcriptomes in which the overall transcriptional activity is suppressed or enhanced (Anders and Huber 2010; Bullard et al. 2010; Robinson and Oshlack 2010). Normalizing mRNA abundance also becomes relevant to analyzing the transcriptomes from cells of heterogeneous populations such as peripheral blood mononuclear cells (PBMCs) by employing a deconvolution method. Deconvolution processing computationally estimates the proportions of distinctive cell types in a heterogeneous sample utilizing the normalized abundance of mRNA in each cell type as references It is an effective solution to determine the composition of each immune cell type in PBMCs (Abbas et al. 2009; Shen-Orr and Gaujoux 2013). Considering that the proportion of immune cell subsets in PBMCs can be dynamically affected by the disease, age, or interventions (e.g., vaccines and drugs), the composition of immune cell populations needs to be carefully evaluated. Otherwise, it is not always possible to accurately determine which immune cell types are responsible for any given transcriptomic changes in PBMCs. The transcriptome profiling can contribute to the results that are inconclusive or difficult to interpret. Hence, the appropriate normalization method is crucial for differential expression analyses and deconvolution approaches. Monaco et al. developed an advanced and robust normalization method that can be applied for future transcriptome analyses of PBMCs by taking advantage of the breadth and granularity of the datasets from 29 isolated immune cell types.
In summary, transcriptome analyses of isolated immune cells from peripheral blood elucidate individual immune cell population’ divergent gene expression patterns, which promote our understanding of diseases related to the immune system. The transcriptome analyses of isolated immune cells are also critical as the resource for analyzing transcriptomes obtained from whole peripheral blood. Furthermore, large transcriptomic studies of isolated immune cells provide opportunities to develop and validate analysis pipelines which would be impractical from heterogeneous samples.
Another point to mention here is that the immune cells have been exploited to investigate the regulatory mechanisms of gene expressions. The immune system serves as an excellent model to explore the gene regulations along changing cell states, as discrete cell populations can be readily purified by well-established markers along differentiation and activation pathways that have been carefully characterized by persuasive studies. We took advantage of the breadth and granularity of immune cells to study the dynamic epigenetic landscapes associated with the target gene expression (Yoshida et al. 2019). The study provided a deep insight to understand immunological differentiation and function and the broad relevance of gene regulatory elements on the genome, such as a profound dichotomy within mammalian gene regulation by enhancers and promoters.
3 Transcriptomes Analyses Employing Whole Blood and PBMC
Blood is an invaluable source to examine our health not only because of the easy accessibility and minimal invasiveness during sampling but also because of the breadth of information it can provide (Sohn 2017). Transcriptomes in PBMCs have also been investigated intensively for scientific research (Corkum et al. 2015; Mello et al. 2012), as well as in medical contexts such as ischemic stroke (Baird et al. 2015), ulcerative colitis (Miao et al. 2013), epilepsy (Karsten et al. 2011), and sepsis (Davenport et al. 2016) to characterize diseases, and epidemiological contexts including aging (Peters et al. 2015), obesity (Homuth et al. 2015), and lifestyle factors such as smoking, drinking, and nutrition (Burton et al. 2018; Dumeaux et al. 2010).
Since PBMCs can include variable naïve and activated immune cells recirculating throughout the body, PBMCs transcriptome analyses are expected to promote the characterization of the whole immune system. However, due to the heterogeneity and dynamics of the components of immune cell types in PMBCs, cell population-level resolution is not successfully achieved so far even with the cutting-edge approaches such as deconvolutions mentioned and thus straight immunological interpretations (e.g., a specific immune cell population is enlarged in donor A than donor B, or a set of genes are more activated in immune cell population X in donor A than B) are readily possible. Accordingly, different approaches employing systems biology are preferentially applied for analyzing transcriptomes from PBMCs (Chaussabel 2015).
Systems biology is an approach in the biomedical research field to understand the larger picture hidden in the biological system by putting pieces of information from the system together. A hypothesis being constructed based on all observed parameters associated with a given biological system, systems biology is compatible with high throughput technologies called “omics” such as genomics, transcriptomics, proteomics, and metabolomics by which a biology system is comprehensively profiled (Aizat et al. 2018; Veenstra 2021). In omics, the parameters are not chosen in advance like in more traditional assays, and these approaches are inherently unbiased. Importantly, as the potency of systems biology intrinsically relies on the variability of observed parameters , the size and heterogeneity of a dataset are crucial for the analyses employing systems biology, and thus more informative results can be expected from the larger dataset (Koumakis 2020; McCue and McCoy 2017; Qin et al. 2015). Schmidt et al. reported the analysis of blood transcriptomes of 3,388 adult individuals (mean age = 58 years), together with phenotypic attributes including disease history, medication status, lifestyle factors, and body mass index (BMI) (Schmidt et al. 2020). Although there were preceding studies analyzing blood transcriptomics, studies were composed of relatively smaller sample sizes related to specific diseases, which restricted the analytical power due to the limited variability in the transcriptomic states and health conditions. Schmidt et al. demonstrated the diversity of blood transcriptomes with modules of co-expressed genes linking to different biological functions. They visualized the molecular heterogeneity of transcriptomes combining with different phenotypic statuses by employing state-of-the-art machine learning methods. The results include two major transcriptomic types, one relating to inflammation enhanced in male, elderly, and overweighted people, and the other one to activated immune responses in female, younger, and ordinary weighted people. They also found that transcriptome signatures are associated with immune response and the increase of inflammatory processes are shared among multiple diseases, aging, and obesity, indicating common underlying mechanisms.
Together, transcriptome analyses employing blood or PBMC is not straightforward to elucidate biological processes at the cell population level and characterize specific immune processes. However, they can provide an unprecedented opportunity to evaluate various diseases and lifestyle factors. They will be applicable for medical diagnostics and molecular and epidemiological research, which will contribute to the promotion of the personalized medicine .
4 Transcriptome Analyses: From Bulk to Single Cells
As mentioned earlier , transcriptome analyses using isolated immune cells as well as blood cells are beneficial to promote our understanding of the immune system by shedding light on disease pathogenesis and global immunity. However, as these are averaged profiles of immune cells and the transcriptomes of minor cells are masked by other major cells, it is not feasible to detect its relevance if rare subsets of cells are responsible for an immune phenotype. The heterogeneity is evident in blood cells and PMBC. Still, FACS-isolated cells according to their markers can also be heterogeneous because immune cell types are too heterogeneous to be entirely separated by known markers. Furthermore, immune cells can be activated by various stimuli such as pathogens and secreted proteins from other cell types (i.e., cytokines) temporarily in an unsynchronized manner. The heterogeneity should also be considered when rare subsets of cells (e.g., antigen-specific T cells or B cells) drive the immune responses by temporal activation (Chattopadhyay et al. 2014; Mostafavi et al. 2016). Hence, single-cell analysis is most anticipated when seeking rare distinctive subsets of cells relating to biological outcomes, for example, when rare cells are essential for conferring protection or inducing pathologic status.
5 Recent Development of Single-Cell Transcriptomics
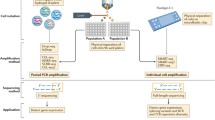
Before the transcriptome analyses from single cells became possible, cDNA synthesis and amplification from a single cell were first succeeded by Iscove in 1990 and Coleman in 1992 (Brady et al. 1990; Eberwine et al. 1992). The cDNA was analyzed using DNA microarrays in the early 2000s and subsequently combined with next-generation sequencing (NGS) technology for single-cell RNA sequencing (scRNA-seq) around 2010 (Bengtsson et al. 2005; Islam et al. 2011; Klein et al. 2002; Kurimoto et al. 2006; Tang et al. 2009). There have been various scRNA-seq methods developed ranging from relatively lower throughput but more detailed full-length transcriptomic data from individual cells to higher throughput with focused coverage on the 3′ terminal of the transcript (Jaitin et al. 2014; Klein et al. 2015; Macosko et al. 2015; Picelli et al. 2013). Currently, scRNA-seq employing a commercial kit from 10× Genomics (Pleasanton, CA) is presumably the most popular. They allow us to profile up to 10,000 cells at a time, and have been used in more than 1,000 publications (Daniloski et al. 2021; Stewart et al. 2020).
6 Applying scRNA-seq for Immune Cells
The heterogeneity in immune cells mirrors the unusual flexibility of the immune system and is essential to protect our bodies efficiently from diverse pathogens. It was recognized in the 1970s and successively confirmed along with the identifications of the cluster of differentiation (CD) antigens using monoclonal antibodies (Engel et al. 2015; Talal 1973). For example, a type of T cells marked by CD4 glycoprotein molecule on the cell surface was identified around 1980. Then subtypes including Th1, Th2, Th17, and the regulatory T cells (Tregs) were identified later (Engleman et al. 1981; Harrington et al. 2005; Mosmann et al. 1986; Park et al. 2005; Sakaguchi et al. 1995). However, given that these distinctions between subtypes are defined by the expression of a few specific markers, these classifications might be a very simplified categorization. Indeed, Teichmann and colleagues demonstrated a subpopulation in Th2 cells which produces the steroid pregnenolone by employing the scRNA-seq approach (Mahata et al. 2014). Importantly, as the comprehensive transcriptome analysis was accomplished by scRNA-seq, they could identify co-regulated genes in the subpopulation, which facilitated the characterization of the cells. Shalek et al. also reported the transcriptomic heterogeneity within bone-marrow-derived dendritic cells (BMDCs) which were seemingly homogenous using scRNA-seq (Shalek et al. 2013). They found hundreds of key immune genes, including genes very highly expressed at the population level, are bimodally expressed across cells. While these pioneering researches employed mouse cells, scRNA-seq approaches were also effectively applied to human cells later .
Karamitros et al. employed scRNA-seq to investigate the transcriptomic differences between progenitor populations in human cord blood (i.e., lymphoid-primed multipotential progenitors: LMPPs, granulocyte-macrophage progenitors: GMPs and multi-lymphoid progenitors: MLPs which were FACS-isolated according to known markers) (Karamitros et al. 2018). They revealed these progenitors were transcriptionally distinct and heterogeneous at the single-cell level, with cells from different progenitor populations showing a transcriptional continuum. Combining with the results from functional assays , they argued a continuum of progenitors executed lymphoid and myeloid differentiation, rather than progenitors downstream of stem cells are uni-lineage. Considering that functional assays can only demonstrate the potential rather than actual cell fate in vivo, and a failure to display functional potential might reflect the assay’s problem, transcriptome analysis adequately contributed to declaring progenitor’s fate in vivo. Recently, Xie et al. profiled 7,551 human blood cells isolated from 21 healthy donors (Xie et al. 2021). They isolated 32 immunophenotypic cell types by FACS and measured transcriptomes in single cells by scRNA-seq. These cells include hematopoietic stem cells, progenitors , and mature immune cells, representing the whole-blood system. The transcriptomic profiles from these 7,551 cells constitute a comprehensive atlas for hematopoietic cells at single-cell resolution. Besides they identified putative long non-coding RNAs (lncRNAs) and transcription factors regulating the differentiation of immune cells, the atlas is also valuable as a resource. It will be utilized by the community to understand the transcriptomic regulations underlying hematopoiesis and immune cell differentiation.
7 scRNA-seq Analysis in Diseases
Measuring the transcriptomes at single-cell resolution by scRNA-seq is innovating our understanding of immune cells in a physiological setting, as mentioned above. In addition, this approach has afforded new options to study the immune response in pathological conditions. What types of cells are responsible for the dysregulated immune response in diseases? By employing comprehensive transcriptome profiling at single-cell resolution, it is possible to examine whether new pathogenic cell subsets developed in disease and the expansion (or contraction) of physiological cell subsets are accompanied. For example, Golumbeanu et al. employed the scRNA-seq approach for dissecting HIV-infected primary CD4+ T cells (Golumbeanu et al. 2018). HIV can persist in latently infected cells despite the effective treatments, which hampers HIV eradication. Hence strategies so-called “shock and kill“ have been developed aiming at reactivating HIV production from the latent cells, so as these cells will die due to virus-mediated cytotoxicity and be killed by cytotoxic CD8+ T cells. However, reactivations of HIV expression are limited to a fraction of latent cells, and the heterogeneity of latently infected cells was suggested. Golumbeanu et al. identified two major cell subpopulations characterized by a set of 134 differentially expressed genes (DEGs) by employing scRNA-seq. Gene ontology analysis revealed enrichment of viral processes, translational regulation, RNA and protein metabolism as well as cell activation genes among these DEGs, which indicates different HIV reactivation potentials for each cluster. They argue that these DEGs are valuable to facilitate the identification of successful reactivations and to identify potential biomarkers of inducible cells.
The composition of the tumor microenvironment (TME) is known to affect the prognosis of cancer patients. For example, higher infiltrates of cytotoxic and memory CD8+ T cells, Th1 CD4+ cells, and NK cells are usually associated with better outcomes, whereas Th2 and Th17 CD4+ cells and Tregs with poor prognosis in several cancers (Fridman et al. 2012). Indeed, while immunotherapies for lung cancer can significantly improve the prognosis for patients, their efficacy varies and depends on in part the number and properties of tumor-infiltrating T cells. Guo et al. investigated the heterogeneity within the tumor-infiltrating T cell by scRNA-seq (Guo et al. 2018). They performed scRNA-seq for 12,346 T cells from 14 untreated non-small-cell lung cancer (NSCLC) patients to comprehensively understand the infiltrating T cells regarding composition, lineage and functional status, and demonstrated the heterogeneity within exhausted CD8+ T cells and Tregs. T-cell exhaustion was originally identified in mice during chronic infection and was later observed in cancer patients (Jiang et al. 2015; Pauken et al. 2016). Exhausted T cells in TME are hyporesponsive states expressing increased inhibitory receptors and decreased effector cytokines, which provoke the failure of cancer elimination. Reinvigorating T-cell exhaustion by such as anti-CTLA-4 (ipilimumab) and anti-PD-1 (nivolumab and pembrolizumab) represents a promising strategy to treat cancer. Since scRNA-seq facilitates trajectory inference or so-called pseudo-time ordering which estimates the cellular identity along with a consecutive differentiation without prior knowledge, they could analyze CD8+ T cells undergoing exhaustion in TME and anticipate two clusters of cells preceding exhaustion, including their transcriptome signatures (Saelens et al. 2019). They employed the transcriptome datasets from TCGA LUAD (The Cancer Genome Atlas Lung Adenocarcinoma) and demonstrated that a high ratio of pre-exhausted to exhausted T cells was associated with a better prognosis. Furthermore, they identified heterogeneity within Tregs in TME, marked by the bimodal expression pattern of TNFRSF9 which is a known activation marker for Tregs. They found a set of 260 genes, including REL and LAYN which are associated with immunosuppressive functions, are highly expressed in TNFRSF9+ Tregs compared to TNFRSF9− Tregs. Importantly, survival analysis employing the TCGA LUAD dataset indicated that higher expressions of these 260 genes were predictive of a worse prognosis. These results represent the efficacy of an approach using scRNA-seq to reveal the heterogeneity in immune cell populations and identify potential clinical biomarkers.
Compared with bulk RNA-seq, scRNA-seq detects the transcriptome nuance in single cells that contribute to revealing the heterogeneity in a seemingly single population. With state-of-the-art machine learning and big data analytics, scRNA-seq has been becoming valuable to identify unknown subpopulations and their transcriptome signatures that affect the biological process and disease diagnosis. However, it is worth noting that scRNA-seq also has limitations compared with bulk RNA-seq, which include relatively low sensitivity, the bias of the transcriptome coverage, and overall cost. Hence, we anticipate that bulk RNA-seq will not lose its value.
8 The Paradigm of Self vs. Non-self from a Transcriptomic Viewpoint
Heterogeneity in the immune cells includes diversity at the DNA level besides the RNA and protein levels which establish the heterogeneity on the population level as discussed. At the DNA level, the number of T-cell receptors (TCRs) and the B-cell receptors (BCR) are estimated to be in the order of 107 whereas the human genome contains roughly 30,000 genes (Fugmann et al. 2000; Nikolich-Zugich et al. 2004). These are produced by somatic DNA recombination called V(D)J recombination in developing lymphocytes during the early stages of T and B cell differentiation. The exceptional divergency endows the immune system with potent effector mechanisms to destroy and eliminate a broad range of pathogenic microorganisms. As the recombination is nearly random, which appropriate to achieve the reactivity to targets of essentially unlimited diversities, it also causes the possibility of self-reactivity at the same time. Therefore, it is critical for the immune system to have mechanisms discriminating self from non-self to avoid destroying the host’s own tissues. The capability of the immune system to avoid damaging the host’s tissues is known as self-tolerance . As the failure of self-tolerance is associated with various autoimmune diseases, this mechanism has been broadly studied in immunology (Besnard et al. 2021; Klein et al. 2014; Sakaguchi et al. 2020).
One of the pivotal roles of T cells is to recognize and kill host cells infected by microbes which otherwise serve as factories for producing replicated microbes. This is managed by a mechanism where infected cells present a molecular complex of microbe antigens and Major Histocompatibility Complex (MHC) class I molecules on the cell surface, which are recognized and killed by T cells with a compatible TCR . As MHC molecules also present normal self-peptides on the cell surface, it is crucial for T cells to maintain self-tolerance. Negative selection of self-reactive T cells is an important process in the thymus where developing T cells of self-reactivity are eliminated if their TCRs react to self-peptides on MHC molecules . Intriguingly, essentially all protein-coding genes are expressed in sets of cells in the thymus. Negative selection functions effectively and comprehensively in the thymus where thymic epithelial cells (TECs) play a pivotal role. In the final section, we discuss how the establishment of self-tolerance in the thymus has been studied using transcriptomic data obtained by novel technologies.
8.1 Thymic Epithelial Cells (TECs) in the Thymic Stroma
The thymus is a highly specialized organ for the establishment of self-tolerance, which is characterized by the “education” of immature T cells. Thymus’ key function is to provide diverse competent T cells that can recognize and eliminate foreign antigens, while they are tolerant to self-components. This complicated process is mainly orchestrated by TECs that form reticular structures in the thymus. TECs are divided into two major subsets by their localization, molecular characteristics, and functions: cortical TECs (cTECs) and medullary TECs (mTECs). Specifically, cTECs are responsible for T-cell lineage commitment and positive selection, while mTECs contribute to the negative selection of self-reactive T cells and/or their cell-fate diversion into Treg lineages (Kyewski and Klein 2006; Matsumoto et al. 2019). These incomparable roles in mTECs are achieved by the expression and presentation of diverse self-antigens complexed with MHC molecule on the surface of mTECs. Notably, to effectively screen for considerable self-reactive thymocyte clones, mTECs are equipped with a unique capacity to express almost 90% of the coding genome, including thousands of tissue-restricted antigens (TRAs) (Kadouri et al. 2020). As expected, the impairment of this “central tolerance” machinery can result in various autoimmune diseases. However, most autoimmune diseases are multifactorial , making it difficult to elucidate their pathogenesis. In this regard, autoimmune regulator (AIRE) and forkhead box P3 (FOXP3), both of which work as transcription factors, are very characteristic genes that cause severe autoimmunity by a single gene mutation. Considering its intimacy with TECs, we focus on and review Aire, an intriguing transcriptional regulator .
8.2 Aire in mTEC
The human AIRE gene was first cloned as the causative gene for autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy (APECED) (Finnish-German APECED Consortium 1997; Nagamine et al. 1997). APECED shows autosomal recessive inheritance and patients have been preferentially reported in certain populations such as Finns, Norwegians, Sardinians, and Iranian Jews (Myhre et al. 2001). The human AIRE gene is composed of 14 exons and is located in the region q22.3 of chromosome 21, encoding a 545 amino-acid protein with a molecular weight of 57.5 kDa (Pitkanen et al. 2000). Importantly, Aire is almost exclusively expressed in mTECs in the thymus.
APECED patients’ symptoms are characterized by a variable combination of (i) failure of the endocrine organs, (ii) chronic mucocutaneous candidiasis, and (iii) dystrophy of the ectoderm-derived tissues (Ahonen et al. 1990). The “hypoparathyroidism,” “adrenal insufficiency (Addison disease),” and “chronic mucocutaneous candidiasis” are regarded as the triad of APECED. Notably, APECED patients have high levels of serum autoantibodies reacting specifically with components in the affected organs, like antibodies against steroidogenic enzymes of the P450 superfamily (e.g., P450c21 and P450c17) in the adrenal cortex (Peterson and Peltonen 2005). Furthermore, unique neutralizing autoantibodies to type I IFN and Th17-related cytokines are frequently detected in patients and these antibodies had been considered to be responsible for the development of chronic mucocutaneous candidiasis (Kisand et al. 2010; Puel et al. 2010). However, this long-standing hypothesis was recently challenged by another group, arguing that aberrantly enhanced type 1 immunity in the patients promotes candida infection susceptibility (Break et al. 2021).
Following the identification of the human AIRE gene, Aire-knockout (Aire-KO) mice (B6 genetic background) were generated to elucidate the mechanisms underlying the Aire deficiency and breakdown of self-tolerance (Anderson et al. 2002). Although the Aire-KO mice showed a rather milder phenotype than APECED patients, they developed lymphocytic infiltrates in several organs with the production of several autoantibodies. Remarkably, Aire-deficient mTECs showed considerable reduction in TRAs, raising a model wherein Aire functions predominantly as a direct transcriptional activator of TRA genes, and reduced TRAs is the cause of autoimmunity in Aire-KO mice (Anderson et al. 2002). This story seems perspicuous and reasonable, but some questions remain. Kuroda et al. reported that although mRNA levels of α-fodrin in mTECs were not reduced, autoantibodies against this molecule were produced in their Aire-deficient mouse model (Kuroda et al. 2005). Another example came from the Aire-KO mice on NOD (non-obese diabetic) background that developed severe autoimmune pancreatitis attacking acinar cells in parallel with a production of autoantibodies against pancreas-specific protein disulfide isomerase (PDIp), despite that the expression of PDIp was retained in Aire-deficient mTECs (Niki et al. 2006). Although further study is required, it is possible that Aire-dependent TRA reduction may not be the sole factor for the breakdown of self-tolerance in Aire-KO mice. In this regard, the role of Aire in the maturation program of mTECs has been proposed (Matsumoto 2011). Interestingly, each TRA protein is expressed only in a few mTECs, considered to be 1–3% of total mTECs with ordered stochasticity (Derbinski et al. 2008). The complete expression of all TRAs by the total mTEC population must be owing to the summation of mosaic expression of TRAs by individual mTECs (Kadouri et al. 2020).
8.3 The Molecular Function of Aire
Aire protein is localized in the nucleus as the shape of nuclear dots, resembling promyelocytic leukemia (PML) nuclear bodies, but they were revealed largely not to be colocalized (Akiyoshi et al. 2004). Considering its localization and structure, the Aire protein appears to be a putative transcriptional regulator, consisting of two plant homeodomain-type zinc-fingers (PHD-fingers), a DNA-binding domain (SAND), and four nuclear receptor binding LXXLL motifs (Kumar et al. 2001). These structural and functional domains are well conserved across phyla (Saltis et al. 2008). Many studies argue about the transcriptional role in Aire, but several unique features that differ from conventional transcription factors have been reported. Aire is apparently involved in the regulation of its target loci in collaboration with lots of partner proteins, forming a large multimolecular complex (Mathis and Benoist 2009). For example, CREB-binding protein (CBP) was the first identified Aire’s partner (Pitkanen et al. 2000). It has also been reported that Aire recruits p-TEFb for transcriptional elongation of target genes (Oven et al. 2007), followed by a study arguing that bromodomain-containing protein, Brd4, bridges Aire and p-TEFb (Yoshida et al. 2015). Furthermore, a broad screen for Aire-targeted coimmunoprecipitation followed by high-throughput mass spectrometry newly identified putative Aire-interacting proteins involved in multiple biological pathways, including nuclear transport, chromatin structure, binding to the transcription machinery, and pre-mRNA processing (Abramson et al. 2010).
Aire’s extraordinary broad transcriptional effect seems to be achieved by activating ectopic transcription, not through specific recognition of TRA gene promoters or enhancer motif. Instead, Aire appears to bind to the repressive chromatin mark H3K4me0 with its PHD1 finger domain (Koh et al. 2008; Org et al. 2008), and release RNA polymerase II paused just downstream of transcriptional start site (TSS) (Giraud et al. 2012). Moreover, recent bioinformatics revealed that Aire-containing complexes are predominantly located on mTEC super-enhancers, which are chromatin stretches enclosing TSS of Aire-dependent genes (Bansal et al. 2017).
8.4 mTEC Heterogeneity Defined by the Single-Cell Approach
As described above, TECs have been divided into cTECs (EpCAM+Ly51+UEA1−) and mTECs (EpCAM+Ly51−UEA1+), histologically and cytologically. Referring to their ontogeny, the evidence regarding the bipotent progenitor cells that give rise to both mTEC and cTEC lineages is emerging in the fetal and early neonatal thymus (Bleul et al. 2006; Rossi et al. 2006), characterized by cTEC-like molecular markers (Baik et al. 2013; Ohigashi et al. 2013). In contrast, it is still controversial about the existence and molecular characteristics of corresponding progenitors in the adult thymus (Ulyanchenko et al. 2016; Wong et al. 2014).
Depending on their molecular characteristics, mTECs were previously categorized as mTEClow (Aire−CD80lowMHC-IIlow) and mTEChigh (Aire+CD80highMHC-IIhigh). “Central tolerance” is primarily achieved by the effective expression and presentation of TRAs from mTEChigh to the developing thymocytes. mTEChigh are differentiated from a part of mTEClow and require RANK and CD40 signals for the development (Akiyama et al. 2008). In comparison with mTEChigh, mTEClow fraction appeared to contain multiple subsets as studied in the past several years: (i) developing stage of mTEC lineage (recently categorized as “Ccl21+ mTEC”) and (ii) terminally differentiated stage of mTECs, called “post-Aire mTEC” (Nishikawa et al. 2010) or “corneocyte-like mTEC ” (Kadouri et al. 2020) (Table 10.1). Post-Aire mTECs lose their nuclei as they form Hassall’s corpuscles. Notably, Aire-deficient mice have reduced numbers of Krt10+ post-Aire mTECs and impaired formation of Hassall’s corpuscles in their thymi, which suggests that Aire may control the differentiation program of mTECs (Matsumoto 2011; Yano et al. 2008).
Furthermore, recent high-throughput scRNA-seq revealed that TECs, especially mTECs, consist of more heterogeneous groups than previously appreciated (Bornstein et al. 2018; Dhalla et al. 2020; Miller et al. 2018; Miragaia et al. 2018). Bornstein et al. categorized mTECs into four subsets as follows: (i) mTEC I (Ccl21+ mTEC), (ii) mTEC II (previous “mTEChigh”), (iii) mTEC III (previous “post-Aire mTEC” or “corneocyte-like mTEC”), and (iv) a newly identified mTEC IV (called “thymic tuft cells”). The existence of the thymic tuft cells, which are considered to establish an immune microenvironment in the thymus, was simultaneously reported by two groups (Bornstein et al. 2018; Miller et al. 2018). Thymic tuft cells are remarkably similar to peripheral tuft cells existing at mucosal barriers in that they express canonical taste transduction pathway molecules and IL-25, whereas the expression of MHC-II and CD74 is characteristic to thymic tuft cells (Miller et al. 2018). Moreover, Dhalla et al. identified a “proliferating mTEC” cluster that exhibited upregulation of Mki67 with Aire, but its biology is still controversial (Ishikawa et al. 2021).
Recently, two groups have reported scRNA-seq studies focusing on human TECs (Bautista et al. 2021; Park et al. 2020). Importantly, human TECs have been revealed to contain similar subsets to mouse TECs (i.e., mTEC I-IV), and the expression of TRA genes and APECED relevant genes are enriched in the AIRE-expressing mTEChigh cluster (Bautista et al. 2021). Bautista et al. also reported the existence of immature TECs, which express canonical TEC genes but lacking characteristic genes of cTECs and mTECs, from both datasets. Moreover, some unique TEC subsets that are specific to humans were identified. Both groups reported the existence of (i) MYOD1 and MYOG expressing myoid cells, and (ii) NEUROD1, NEUROG, and CHGA expressing neuroendocrine cells. Bautista et al. further identified (iii) SOX10 and MPZ expressing myelin cells. Interestingly, expressions of myasthenia gravis relevant genes (i.e., CHRNA1, TTN, and MUSK) were predominantly found in the myoid, and neuroendocrine subsets (Bautista et al. 2021). It evokes the possibility that these unique AIRE− populations also participate in the induction of immune tolerance, while these cells may not directly present antigens due to their low levels of MHC (HLA) expression. In summary, recent transcriptome analysis at single-cell resolution revealed that the thymus orchestrates the establishment of self-tolerance by the coordination of quite heterogenous TEC subsets, collaborating with unique transcriptional machineries .
9 Conclusion
In this chapter, we have described the recent advances in transcriptome analyses especially focusing on the bulk RNA-seq and scRNA-seq approaches that helped our understanding of the immune system more globally. In the last part of the chapter, we touched on how these techniques have now been bringing a new paradigm for self vs. non-self-discrimination in the thymus. The study on Aire deficiency, a monogenic autoimmune disease, has underscored the importance of the advent of new technologies to draw a whole picture of transcriptional control of the immune system. We are hoping that the complete picture of the transcripts of each immune cell type and the integration of this knowledge will pave the way to a comprehensive understanding of the immune system from a novel viewpoint.
References
Abbas AR, Wolslegel K, Seshasayee D, Modrusan Z, Clark HF (2009) Deconvolution of blood microarray data identifies cellular activation patterns in systemic lupus erythematosus. PLoS One 4:e6098
Abramson J, Giraud M, Benoist C, Mathis D (2010) Aire's partners in the molecular control of immunological tolerance. Cell 140:123–135
Ackerman GA (1964) Histochemical differentiation during neutrophil development and maturation. Ann N Y Acad Sci 113:537–565
Ahonen P, Myllarniemi S, Sipila I, Perheentupa J (1990) Clinical variation of autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy (APECED) in a series of 68 patients. N Engl J Med 322:1829–1836
Aizat WM, Ismail I, Noor NM (2018) Recent development in omics studies. Adv Exp Med Biol 1102:1–9
Akiyama T, Shimo Y, Yanai H, Qin J, Ohshima D, Maruyama Y, Asaumi Y, Kitazawa J, Takayanagi H, Penninger JM et al (2008) The tumor necrosis factor family receptors RANK and CD40 cooperatively establish the thymic medullary microenvironment and self-tolerance. Immunity 29:423–437
Akiyoshi H, Hatakeyama S, Pitkanen J, Mouri Y, Doucas V, Kudoh J, Tsurugaya K, Uchida D, Matsushima A, Oshikawa K et al (2004) Subcellular expression of autoimmune regulator (AIRE) is organized in a spatiotemporal manner. J Biol Chem 279:33984–33991
Amaya-Uribe L, Rojas M, Azizi G, Anaya J-M, Gershwin ME (2019) Primary immunodeficiency and autoimmunity: a comprehensive review. J Autoimmun 99:52–72
Amit I, Regev A, Hacohen N (2011) Strategies to discover regulatory circuits of the mammalian immune system. Nat Rev Immunol 11:873–880
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Nature Precedings
Anderson MS, Venanzi ES, Klein L, Chen Z, Berzins SP, Turley SJ, von Boehmer H, Bronson R, Dierich A, Benoist C, Mathis D (2002) Projection of an immunological self shadow within the thymus by the aire protein. Science 298:1395–1401
Baik S, Jenkinson EJ, Lane PJ, Anderson G, Jenkinson WE (2013) Generation of both cortical and Aire(+) medullary thymic epithelial compartments from CD205(+) progenitors. Eur J Immunol 43:589–594
Baird AE, Soper SA, Pullagurla SR, Adamski MG (2015) Recent and near-future advances in nucleic acid-based diagnosis of stroke. Expert Rev Mol Diagn 15:665–679
Bansal K, Yoshida H, Benoist C, Mathis D (2017) The transcriptional regulator Aire binds to and activates super-enhancers. Nat Immunol 18:263–273
Bautista JL, Cramer NT, Miller CN, Chavez J, Berrios DI, Byrnes LE, Germino J, Ntranos V, Sneddon JB, Burt TD et al (2021) Single-cell transcriptional profiling of human thymic stroma uncovers novel cellular heterogeneity in the thymic medulla. Nat Commun 12:1096
Bengtsson M, Ståhlberg A, Rorsman P, Kubista M (2005) Gene expression profiling in single cells from the pancreatic islets of Langerhans reveals lognormal distribution of mRNA levels. Genome Res 15:1388–1392
Besnard M, Padonou F, Provin N, Giraud M, Guillonneau C (2021) AIRE deficiency, from preclinical models to human APECED disease. Dis Model Mech 14:dmm046359
Bleul CC, Corbeaux T, Reuter A, Fisch P, Monting JS, Boehm T (2006) Formation of a functional thymus initiated by a postnatal epithelial progenitor cell. Nature 441:992–996
Bornstein C, Nevo S, Giladi A, Kadouri N, Pouzolles M, Gerbe F, David E, Machado A, Chuprin A, Toth B et al (2018) Single-cell mapping of the thymic stroma identifies IL-25-producing tuft epithelial cells. Nature 559:622–626
Brady G, Barbara M, Iscove NN (1990) Representative in vitro cDNA amplification from individual hemopoietic cells and colonies. Methods Mol Cell Biol 2:17–25
Break TJ, Oikonomou V, Dutzan N, Desai JV, Swidergall M, Freiwald T, Chauss D, Harrison OJ, Alejo J, Williams DW et al (2021) Aberrant type 1 immunity drives susceptibility to mucosal fungal infections. Science 371:eaay5731
Bullard JH, Purdom E, Hansen KD, Dudoit S (2010) Evaluation of statistical methods for normalization and differential expression in mRNA-Seq experiments. BMC Bioinformatics 11:94
Burton KJ, Pimentel G, Zangger N, Vionnet N, Drai J, McTernan PG, Pralong FP, Delorenzi M, Vergères G (2018) Modulation of the peripheral blood transcriptome by the ingestion of probiotic yoghurt and acidified milk in healthy, young men. PLoS One 13:e0192947
Chaplin DD (2010) Overview of the immune response. J Allergy Clin Immunol 125:S3–S23
Chattopadhyay PK, Gierahn TM, Roederer M, Love JC (2014) Single-cell technologies for monitoring immune systems. Nat Immunol 15:128–135
Chaussabel D (2015) Assessment of immune status using blood transcriptomics and potential implications for global health. Semin Immunol 27:58–66
Chou C, Li MO (2018) Tissue-resident lymphocytes across innate and adaptive lineages. Front Immunol 9:2104
Corkum CP, Ings DP, Burgess C, Karwowska S, Kroll W, Michalak TI (2015) Immune cell subsets and their gene expression profiles from human PBMC isolated by vacutainer Cell Preparation Tube (CPT™) and standard density gradient. BMC Immunol 16:48
Daniloski Z, Jordan TX, Wessels H-H, Hoagland DA, Kasela S, Legut M, Maniatis S, Mimitou EP, Lu L, Geller E et al (2021) Identification of required host factors for SARS-CoV-2 infection in human cells. Cell 184:92–105.e116
Davenport EE, Burnham KL, Radhakrishnan J, Humburg P, Hutton P, Mills TC, Rautanen A, Gordon AC, Garrard C, Hill AVS et al (2016) Genomic landscape of the individual host response and outcomes in sepsis: a prospective cohort study. Lancet Respir Med 4:259–271
Derbinski J, Pinto S, Rosch S, Hexel K, Kyewski B (2008) Promiscuous gene expression patterns in single medullary thymic epithelial cells argue for a stochastic mechanism. Proc Natl Acad Sci U S A 105:657–662
Dhalla F, Baran-Gale J, Maio S, Chappell L, Hollander GA, Ponting CP (2020) Biologically indeterminate yet ordered promiscuous gene expression in single medullary thymic epithelial cells. EMBO J 39:e101828
Dumeaux V, Olsen KS, Nuel G, Paulssen RH, Børresen-Dale A-L, Lund E (2010) Deciphering normal blood gene expression variation – the NOWAC postgenome study. PLoS Genet 6:e1000873
Eberwine J, Yeh H, Miyashiro K, Cao Y, Nair S, Finnell R, Zettel M, Coleman P (1992) Analysis of gene expression in single live neurons. Proc Natl Acad Sci U S A 89:3010–3014
Engel P, Boumsell L, Balderas R, Bensussan A, Gattei V, Horejsi V, Jin B-Q, Malavasi F, Mortari F, Schwartz-Albiez R et al (2015) CD nomenclature 2015: human leukocyte differentiation antigen workshops as a driving force in immunology. J Immunol (Baltimore, Md : 1950) 195:4555–4563
Engleman EG, Benike CJ, Grumet FC, Evans RL (1981) Activation of human T lymphocyte subsets: helper and suppressor/cytotoxic T cells recognize and respond to distinct histocompatibility antigens. J Immunol (Baltimore, Md : 1950) 127:2124–2129
Finnish-German APECED Consortium (1997) An autoimmune disease, APECED, caused by mutations in a novel gene featuring two PHD-type zinc-finger domains. Nat Genet 17:399–403
Fridman WH, Pagès F, Sautès-Fridman C, Galon J (2012) The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer 12:298–306
Fugmann SD, Lee AI, Shockett PE, Villey IJ, Schatz DG (2000) The RAG proteins and V(D)J recombination: complexes, ends, and transposition. Annu Rev Immunol 18:495–527
Giraud M, Yoshida H, Abramson J, Rahl PB, Young RA, Mathis D, Benoist C (2012) Aire unleashes stalled RNA polymerase to induce ectopic gene expression in thymic epithelial cells. Proc Natl Acad Sci U S A 109:535–540
Golumbeanu M, Cristinelli S, Rato S, Munoz M, Cavassini M, Beerenwinkel N, Ciuffi A (2018) Single-cell RNA-seq reveals transcriptional heterogeneity in latent and reactivated HIV-infected cells. Cell Rep 23:942–950
Guo X, Zhang Y, Zheng L, Zheng C, Song J, Zhang Q, Kang B, Liu Z, Jin L, Xing R et al (2018) Global characterization of T cells in non-small-cell lung cancer by single-cell sequencing. Nat Med 24:978–985
Hajdu SI (2003) A note from history: the discovery of blood cells. Ann Clin Lab Sci 33:237–238
Hardy RR, Hayakawa K (2001) B cell development pathways. Annu Rev Immunol 19:595–621
Harrington LE, Hatton RD, Mangan PR, Turner H, Murphy TL, Murphy KM, Weaver CT (2005) Interleukin 17-producing CD4+ effector T cells develop via a lineage distinct from the T helper type 1 and 2 lineages. Nat Immunol 6:1123–1132
Hayakawa K, Hardy RR, Parks DR, Herzenberg LA (1983) The "Ly-1 B" cell subpopulation in normal immunodefective, and autoimmune mice. J Exp Med 157:202–218
Homuth G, Wahl S, Müller C, Schurmann C, Mäder U, Blankenberg S, Carstensen M, Dörr M, Endlich K, Englbrecht C et al (2015) Extensive alterations of the whole-blood transcriptome are associated with body mass index: results of an mRNA profiling study involving two large population-based cohorts. BMC Med Genet 8:65
Ishikawa T, Akiyama N, Akiyama T (2021) In pursuit of adult progenitors of thymic epithelial cells. Front Immunol 12:621824
Islam S, Kjällquist U, Moliner A, Zajac P, Fan J-B, Lönnerberg P, Linnarsson S (2011) Characterization of the single-cell transcriptional landscape by highly multiplex RNA-seq. Genome Res 21:1160–1167
Jaitin DA, Kenigsberg E, Keren-Shaul H, Elefant N, Paul F, Zaretsky I, Mildner A, Cohen N, Jung S, Tanay A, Amit I (2014) Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types, vol 343. Science (New York, N.Y.), pp 776–779
Jayasinghe SN (2020) Reimagining flow cytometric cell sorting. Adv Biosyst 4:e2000019
Jiang Y, Li Y, Zhu B (2015) T-cell exhaustion in the tumor microenvironment. Cell Death Dis 6:e1792
Kadouri N, Nevo S, Goldfarb Y, Abramson J (2020) Thymic epithelial cell heterogeneity: TEC by TEC. Nat Rev Immunol 20:239–253
Karamitros D, Stoilova B, Aboukhalil Z, Hamey F, Reinisch A, Samitsch M, Quek L, Otto G, Repapi E, Doondeea J et al (2018) Single-cell analysis reveals the continuum of human lympho-myeloid progenitor cells. Nat Immunol 19:85–97
Karsten SL, Kudo LC, Bragin AJ (2011) Use of peripheral blood transcriptome biomarkers for epilepsy prediction. Neurosci Lett 497:213–217
Kaufmann SHE (2017) Emil von Behring: translational medicine at the dawn of immunology. Nat Rev Immunol 17:341–343
Kisand K, Boe Wolff AS, Podkrajsek KT, Tserel L, Link M, Kisand KV, Ersvaer E, Perheentupa J, Erichsen MM, Bratanic N et al (2010) Chronic mucocutaneous candidiasis in APECED or thymoma patients correlates with autoimmunity to Th17-associated cytokines. J Exp Med 207:299–308
Klein CA, Seidl S, Petat-Dutter K, Offner S, Geigl JB, Schmidt-Kittler O, Wendler N, Passlick B, Huber RM, Schlimok G et al (2002) Combined transcriptome and genome analysis of single micrometastatic cells. Nat Biotechnol 20:387–392
Klein L, Kyewski B, Allen PM, Hogquist KA (2014) Positive and negative selection of the T cell repertoire: what thymocytes see (and don't see). Nat Rev Immunol 14:377–391
Klein AM, Mazutis L, Akartuna I, Tallapragada N, Veres A, Li V, Peshkin L, Weitz DA, Kirschner MW (2015) Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161:1187–1201
Koh AS, Kuo AJ, Park SY, Cheung P, Abramson J, Bua D, Carney D, Shoelson SE, Gozani O, Kingston RE et al (2008) Aire employs a histone-binding module to mediate immunological tolerance, linking chromatin regulation with organ-specific autoimmunity. Proc Natl Acad Sci U S A 105:15878–15883
Koumakis L (2020) Deep learning models in genomics; are we there yet? Comput Struct Biotechnol J 18:1466–1473
Kumar PG, Laloraya M, Wang CY, Ruan QG, Davoodi-Semiromi A, Kao KJ, She JX (2001) The autoimmune regulator (AIRE) is a DNA-binding protein. J Biol Chem 276:41357–41364
Kurimoto K, Yabuta Y, Ohinata Y, Ono Y, Uno KD, Yamada RG, Ueda HR, Saitou M (2006) An improved single-cell cDNA amplification method for efficient high-density oligonucleotide microarray analysis. Nucleic Acids Res 34:e42
Kuroda N, Mitani T, Takeda N, Ishimaru N, Arakaki R, Hayashi Y, Bando Y, Izumi K, Takahashi T, Nomura T et al (2005) Development of autoimmunity against transcriptionally unrepressed target antigen in the thymus of Aire-deficient mice. J Immunol 174:1862–1870
Kyewski B, Klein L (2006) A central role for central tolerance. Annu Rev Immunol 24:571–606
Lara-Astiaso D, Weiner A, Lorenzo-Vivas E, Zaretsky I, Jaitin DA, David E, Keren-Shaul H, Mildner A, Winter D, Jung S et al (2014) Immunogenetics. Chromatin state dynamics during blood formation, vol 345. Science (New York, N.Y.), pp 943–949
Macosko EZ, Basu A, Satija R, Nemesh J, Shekhar K, Goldman M, Tirosh I, Bialas AR, Kamitaki N, Martersteck EM et al (2015) Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161:1202–1214
Maecker HT, McCoy JP, Nussenblatt R (2012) Standardizing immunophenotyping for the human immunology project. Nat Rev Immunol 12:191–200
Mahata B, Zhang X, Kolodziejczyk AA, Proserpio V, Haim-Vilmovsky L, Taylor AE, Hebenstreit D, Dingler FA, Moignard V, Göttgens B et al (2014) Single-cell RNA sequencing reveals T helper cells synthesizing steroids de novo to contribute to immune homeostasis. Cell Rep 7:1130–1142
Mathis D, Benoist C (2009) Aire. Annu Rev Immunol 27:287–312
Matsumoto M (2011) Contrasting models for the roles of Aire in the differentiation program of epithelial cells in the thymic medulla. Eur J Immunol 41:12–17
Matsumoto M, Rodrigues PM, Sousa L, Tsuneyama K, Matsumoto M, Alves NL (2019) The ins and outs of thymic epithelial cell differentiation and function. In: Passos GA (ed) Thymus transcriptome and cell biology. Springer, pp 35–66
McCue ME, McCoy AM (2017) The scope of big data in one medicine: unprecedented opportunities and challenges. Front Vet Sci 4:194
Mello VDF, Kolehmanien M, Schwab U, Pulkkinen L, Uusitupa M (2012) Gene expression of peripheral blood mononuclear cells as a tool in dietary intervention studies: what do we know so far? Mol Nutr Food Res 56:1160–1172
Miao Y-L, Xiao Y-L, Du Y, Duan L-P (2013) Gene expression profiles in peripheral blood mononuclear cells of ulcerative colitis patients. World J Gastroenterol 19:3339–3346
Miller CN, Proekt I, von Moltke J, Wells KL, Rajpurkar AR, Wang H, Rattay K, Khan IS, Metzger TC, Pollack JL et al (2018) Thymic tuft cells promote an IL-4-enriched medulla and shape thymocyte development. Nature 559:627–631
Miragaia RJ, Zhang X, Gomes T, Svensson V, Ilicic T, Henriksson J, Kar G, Lonnberg T (2018) Single-cell RNA-sequencing resolves self-antigen expression during mTEC development. Sci Rep 8:685
Monaco G, Lee B, Xu W, Mustafah S, Hwang YY, Carré C, Burdin N, Visan L, Ceccarelli M, Poidinger M et al (2019) RNA-seq signatures normalized by mRNA abundance allow absolute deconvolution of human immune cell types. Cell Rep 26:1627–1640.e1627
Mosmann TR, Cherwinski H, Bond MW, Giedlin MA, Coffman RL (1986) Two types of murine helper T cell clone. I. Definition according to profiles of lymphokine activities and secreted proteins. J Immunol (Baltimore, Md : 1950) 136:2348–2357
Mostafavi S, Yoshida H, Moodley D, LeBoité H, Rothamel K, Raj T, Ye CJ, Chevrier N, Zhang S-Y, Feng T et al (2016) Parsing the interferon transcriptional network and its disease associations. Cell 164:564–578
Myhre AG, Halonen M, Eskelin P, Ekwall O, Hedstrand H, Rorsman F, Kampe O, Husebye ES (2001) Autoimmune polyendocrine syndrome type 1 (APS I) in Norway. Clin Endocrinol 54:211–217
Nagamine K, Peterson P, Scott HS, Kudoh J, Minoshima S, Heino M, Krohn KJ, Lalioti MD, Mullis PE, Antonarakis SE et al (1997) Positional cloning of the APECED gene. Nat Genet 17:393–398
Niki S, Oshikawa K, Mouri Y, Hirota F, Matsushima A, Yano M, Han H, Bando Y, Izumi K, Matsumoto M et al (2006) Alteration of intra-pancreatic target-organ specificity by abrogation of Aire in NOD mice. J Clin Invest 116:1292–1301
Nikolich-Zugich J, Slifka MK, Messaoudi I (2004) The many important facets of T-cell repertoire diversity. Nat Rev Immunol 4:123–132
Nishikawa Y, Hirota F, Yano M, Kitajima H, Miyazaki J, Kawamoto H, Mouri Y, Matsumoto M (2010) Biphasic Aire expression in early embryos and in medullary thymic epithelial cells before end-stage terminal differentiation. J Exp Med 207:963–971
Novershtern N, Subramanian A, Lawton LN, Mak RH, Haining WN, McConkey ME, Habib N, Yosef N, Chang CY, Shay T et al (2011) Densely interconnected transcriptional circuits control cell states in human hematopoiesis. Cell 144:296–309
Ohigashi I, Zuklys S, Sakata M, Mayer CE, Zhanybekova S, Murata S, Tanaka K, Hollander GA, Takahama Y (2013) Aire-expressing thymic medullary epithelial cells originate from beta5t-expressing progenitor cells. Proc Natl Acad Sci U S A 110:9885–9890
Org T, Chignola F, Hetenyi C, Gaetani M, Rebane A, Liiv I, Maran U, Mollica L, Bottomley MJ, Musco G, Peterson P (2008) The autoimmune regulator PHD finger binds to non-methylated histone H3K4 to activate gene expression. EMBO Rep 9:370–376
Oven I, Brdickova N, Kohoutek J, Vaupotic T, Narat M, Peterlin BM (2007) AIRE recruits P-TEFb for transcriptional elongation of target genes in medullary thymic epithelial cells. Mol Cell Biol 27:8815–8823
Packer D (2021) The history of the antibody as a tool. Acta Histochem 123:151710
Park H, Li Z, Yang XO, Chang SH, Nurieva R, Wang Y-H, Wang Y, Hood L, Zhu Z, Tian Q, Dong C (2005) A distinct lineage of CD4 T cells regulates tissue inflammation by producing interleukin 17. Nat Immunol 6:1133–1141
Park JE, Botting RA, Dominguez Conde C, Popescu DM, Lavaert M, Kunz DJ, Goh I, Stephenson E, Ragazzini R, Tuck E et al (2020) A cell atlas of human thymic development defines T cell repertoire formation. Science 367:eaay3224
Pauken KE, Sammons MA, Odorizzi PM, Manne S, Godec J, Khan O, Drake AM, Chen Z, Sen DR, Kurachi M et al (2016) Epigenetic stability of exhausted T cells limits durability of reinvigoration by PD-1 blockade. Science (New York, NY) 354:1160–1165
Peters MJ, Joehanes R, Pilling LC, Schurmann C, Conneely KN, Powell J, Reinmaa E, Sutphin GL, Zhernakova A, Schramm K et al (2015) The transcriptional landscape of age in human peripheral blood. Nat Commun 6:8570
Peterson P, Peltonen L (2005) Autoimmune polyendocrinopathy syndrome type 1 (APS1) and AIRE gene: new views on molecular basis of autoimmunity. J Autoimmun 25(Suppl):49–55
Picelli S, Björklund ÅK, Faridani OR, Sagasser S, Winberg G, Sandberg R (2013) Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat Methods 10:1096–1098
Pitkanen J, Doucas V, Sternsdorf T, Nakajima T, Aratani S, Jensen K, Will H, Vahamurto P, Ollila J, Vihinen M et al (2000) The autoimmune regulator protein has transcriptional transactivating properties and interacts with the common coactivator CREB-binding protein. J Biol Chem 275:16802–16809
Proserpio V, Mahata B (2016) Single-cell technologies to study the immune system. Immunology 147:133–140
Puel A, Doffinger R, Natividad A, Chrabieh M, Barcenas-Morales G, Picard C, Cobat A, Ouachee-Chardin M, Toulon A, Bustamante J et al (2010) Autoantibodies against IL-17A, IL-17F, and IL-22 in patients with chronic mucocutaneous candidiasis and autoimmune polyendocrine syndrome type I. J Exp Med 207:291–297
Qin Y, Yalamanchili HK, Qin J, Yan B, Wang J (2015) The current status and challenges in computational analysis of genomic big data. Big Data Res 2:12–18
Rankin LC, Artis D (2018) Beyond host defense: emerging functions of the immune system in regulating complex tissue physiology. Cell 173:554–567
Robinson MD, Oshlack A (2010) A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol 11:R25
Rossi SW, Jenkinson WE, Anderson G, Jenkinson EJ (2006) Clonal analysis reveals a common progenitor for thymic cortical and medullary epithelium. Nature 441:988–991
Rothenberg EV (2014) Transcriptional control of early T and B cell developmental choices. Annu Rev Immunol 32:283–321
Rouse BT, Sehrawat S (2010) Immunity and immunopathology to viruses: what decides the outcome? Nat Rev Immunol 10:514–526
Roy AL (2019) Transcriptional regulation in the immune system: one cell at a time. Front Immunol 10:1355
Saelens W, Cannoodt R, Todorov H, Saeys Y (2019) A comparison of single-cell trajectory inference methods. Nat Biotechnol 37:547–554
Sakaguchi S, Sakaguchi N, Asano M, Itoh M, Toda M (1995) Immunologic self-tolerance maintained by activated T cells expressing IL-2 receptor alpha-chains (CD25). Breakdown of a single mechanism of self-tolerance causes various autoimmune diseases. J Immunol (Baltimore, Md : 1950) 155:1151–1164
Sakaguchi S, Mikami N, Wing JB, Tanaka A, Ichiyama K, Ohkura N (2020) Regulatory T cells and human disease. Annu Rev Immunol 38:541–566
Saltis M, Criscitiello MF, Ohta Y, Keefe M, Trede NS, Goitsuka R, Flajnik MF (2008) Evolutionarily conserved and divergent regions of the autoimmune regulator (Aire) gene: a comparative analysis. Immunogenetics 60:105–114
Sánchez-Ramón S, Bermúdez A, González-Granado LI, Rodríguez-Gallego C, Sastre A, Soler-Palacín P (2019) Primary and secondary immunodeficiency diseases in oncohaematology: warning signs, diagnosis, and management. Front Immunol 10:586
Schmidt M, Hopp L, Arakelyan A, Kirsten H, Engel C, Wirkner K, Krohn K, Burkhardt R, Thiery J, Loeffler M et al (2020) The human blood transcriptome in a large population cohort and its relation to aging and health. Front Big Data 3:548873
Schmiedel BJ, Singh D, Madrigal A, Valdovino-Gonzalez AG, White BM, Zapardiel-Gonzalo J, Ha B, Altay G, Greenbaum JA, McVicker G et al (2018) Impact of genetic polymorphisms on human immune cell gene expression. Cell 175:1701–1715.e1716
Seumois G, Vijayanand P (2019) Single-cell analysis to understand the diversity of immune cell types that drive disease pathogenesis. J Allergy Clin Immunol 144:1150–1153
Shalek AK, Satija R, Adiconis X, Gertner RS, Gaublomme JT, Raychowdhury R, Schwartz S, Yosef N, Malboeuf C, Lu D et al (2013) Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498:236–240
Shen-Orr SS, Gaujoux R (2013) Computational deconvolution: extracting cell type-specific information from heterogeneous samples. Curr Opin Immunol 25:571–578
Smale ST, Fisher AG (2002) Chromatin structure and gene regulation in the immune system. Annu Rev Immunol 20:427–462
Sohn E (2017) Diagnosis: frontiers in blood testing. Nature 549:S16–S18
Stein M, Keshav S, Harris N, Gordon S (1992) Interleukin 4 potently enhances murine macrophage mannose receptor activity: a marker of alternative immunologic macrophage activation. J Exp Med 176:287–292
Stewart CA, Gay CM, Xi Y, Sivajothi S, Sivakamasundari V, Fujimoto J, Bolisetty M, Hartsfield PM, Balasubramaniyan V, Chalishazar MD et al (2020) Single-cell analyses reveal increased intratumoral heterogeneity after the onset of therapy resistance in small-cell lung cancer. Nat Can 1:423–436
Stubbington MJT, Rozenblatt-Rosen O, Regev A, Teichmann SA (2017) Single-cell transcriptomics to explore the immune system in health and disease. Science (New York, NY) 358:58–63
Talal N (1973) Lymphocyte heterogeneity and function. Arthritis Rheum 16:422–425
Tang F, Barbacioru C, Wang Y, Nordman E, Lee C, Xu N, Wang X, Bodeau J, Tuch BB, Siddiqui A et al (2009) mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods 6:377–382
Uhlen M, Karlsson MJ, Zhong W, Tebani A, Pou C, Mikes J, Lakshmikanth T, Forsström B, Edfors F, Odeberg J et al (2019) A genome-wide transcriptomic analysis of protein-coding genes in human blood cells. Science (New York, NY) 366:eaax9198
Ulyanchenko S, O'Neill KE, Medley T, Farley AM, Vaidya HJ, Cook AM, Blair NF, Blackburn CC (2016) Identification of a bipotent epithelial progenitor population in the adult thymus. Cell Rep 14:2819–2832
Veenstra TD (2021) Omics in systems biology: current progress and future outlook. Proteomics 21:e2000235
Wong K, Lister NL, Barsanti M, Lim JM, Hammett MV, Khong DM, Siatskas C, Gray DH, Boyd RL, Chidgey AP (2014) Multilineage potential and self-renewal define an epithelial progenitor cell population in the adult thymus. Cell Rep 8:1198–1209
Xie X, Liu M, Zhang Y, Wang B, Zhu C, Wang C, Li Q, Huo Y, Guo J, Xu C et al (2021) Single-cell transcriptomic landscape of human blood cells. Natl Sci Rev 8:nwaa180
Yano M, Kuroda N, Han H, Meguro-Horike M, Nishikawa Y, Kiyonari H, Maemura K, Yanagawa Y, Obata K, Takahashi S et al (2008) Aire controls the differentiation program of thymic epithelial cells in the medulla for the establishment of self-tolerance. J Exp Med 205:2827–2838
Yoshida H, Bansal K, Schaefer U, Chapman T, Rioja I, Proekt I, Anderson MS, Prinjha RK, Tarakhovsky A, Benoist C, Mathis D (2015) Brd4 bridges the transcriptional regulators, Aire and P-TEFb, to promote elongation of peripheral-tissue antigen transcripts in thymic stromal cells. Proc Natl Acad Sci U S A 112:E4448–E4457
Yoshida H, Lareau CA, Ramirez RN, Rose SA, Maier B, Wroblewska A, Desland F, Chudnovskiy A, Mortha A, Dominguez C et al (2019) The cis-regulatory atlas of the mouse immune system. Cell 176(897-912):e820
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Yoshida, H., Matsumoto, M., Matsumoto, M. (2022). Transcriptomics to Dissect the Immune System. In: Passos, G.A. (eds) Transcriptomics in Health and Disease. Springer, Cham. https://doi.org/10.1007/978-3-030-87821-4_10
Download citation
DOI: https://doi.org/10.1007/978-3-030-87821-4_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-87820-7
Online ISBN: 978-3-030-87821-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)