Abstract
Diabetic Retinopathy (DR) is a very common retinal disease in the world, which can affect vision and even cause blindness. Early diagnosis can effectively prevent the disease, or at least delay the progression of DR. However, most methods are based on regular single-view images, which would lack complete information of lesions. In this paper, a novel method is proposed to achieve DR classification using ultra-widefield images (UWF). The proposed network includes a dual-branch network, an efficient channel attention (ECA) module, a spatial attention (SA) module, and an atrous spatial pyramid pooling (ASPP) module. Specifically, the dual-branch network uses ResNet-34 model as the backbone. The ASPP module enlarges the receptive field to extract rich feature information by setting different dilated rates. To emphasize the useful information and suppress the useless information, the ECA and SA modules are utilized to extract important channel information and spatial information respectively. To reduce the parameters of the network, we use a global average pooling (GAP) layer to compress the features. The experimental results on the UWF images collected by a local hospital show that our model performs very well.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
- Diabetic retinopathy
- Dual-branch network
- Efficient channel and spatial attention
- Atrous spatial pyramid pooling
1 Introduction
Diabetic retinopathy (DR) originates from diabetes and is a relatively common fundus disease that can cause blindness. It is estimated that about 93 million people worldwide suffer from DR [1], and this number continues to rise. Since DR can be detected by the morphological changes of the retina, many researchers are devoted to the research of disease detection based on fundus images to assist ophthalmologists for diagnosis. However, most methods are based on single-view fundus images, which lead to incomplete information of lesions. So some researchers chose ultra-widefield (UWF) images as their studying objects [2,3,4]. In clinical, compared with conventional single-view scanning images, the UWF images have a vision of 180–200° and contain more region information, which is beneficial for the accurate diagnosis. For instance, Nagasato et al. proposed a central retinal vein occlusion detection method based on UWF fundus images [5]. Pellegrini et al. presented an method for artery/vein classification using UWF images [6].
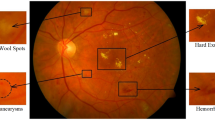
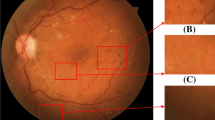
The UWF fundus images of patients with DR and those of normal people are shown in Fig. 1. It shows that there is little contrast between the lesion area and the normal area, with differences in lesion size and inhomogeneous distribution, which brings great challenges to the processing and analysis of the UWF images.
Deep learning performs well in medical image processing and analysis and many researchers have applied it to automatically diagnose various diseases. For example, Brown et al. designed an automatic detection method for retinopathy of prematurity using deep convolutional network [7]. Li et al. used a deep learning method to detect DR and diabetic macular oedema [8]. Diaz-Pinto et al. presented a method based on deep learning to detect and classify glaucoma [9]. Xie et al. used a cross-attention network for fundus diseases classification [10]. However, compared to the background area, some lesion areas are small and the background area is more obvious than the target area (especially in UWF images). Some researchers tried to use more complex models or design multiple networks to extract the discriminative features. For instance, Hamwood et al. used a fully convolutional network to determine the positions of cone photoreceptors [11]. Ruan et al. achieved kidney tumor segmentation on CT images by using a multi-branch feature sharing network [12]. However, these networks have advantages in extracting global features, but easily overlook the important local information, which is disadvantageous for the expression of discriminative detailed features. To address this situation, some researchers have turned to attention mechanism [13,14,15,16]. The attention mechanism can obtain more detailed information of the target, which is very helpful for the detection and discrimination of lesion.
In this paper, a dual-branch network with ResNet-34 [17] as the backbone is proposed to extract features. We use atrous spatial pyramid pooling module (ASPP) to enlarges the receptive field and obtain rich semantic information. According to the dependence of spatial and channel information between the features, the efficient channel attention (ECA) and spatial attention (SA) modules are utilized to emphasize the useful information after multiple convolutional layers of the network. Finally, we use a global average pooling (GAP) layer to compress the features and concatenate the features from two branches to complete the final classification task.
2 Methodology
The architecture of our proposed method is shown in Fig. 2. We propose a dual-branch network to extract features and enlarge the receptive field by ASPP to obtain richer semantic information. At the same time, the ECA and SA modules are employed to emphasize some important information. After obtaining discriminative features by GAP, the features from the two branches are concatenated and used for the prediction. The detailed information of modules will be described below.
2.1 Dual-Branch Network
Inspired by [9], we propose a dual-branch network to extract depth features. Each branch of the model is based on the ResNet-34 model, which solves the gradient disappearance problem in deep networks through residual connection. It is worth noting that the first three layers of the two branch networks share parameters, so the scale of the model parameters is reduced. We use the UWF images as the input of the dual-branch network, and the extracted rich deep semantic features can be used in the subsequent modules.
2.2 Atrous Spatial Pyramid Pooling Module
General neural networks use pooling operation to enlarge the receptive field to capture more spatial information while reducing the resolution, but this process will have some loss of detailed information. To solve this issue, some researchers used atrous convolution (AC) instead of pooling operation. We can obtain the context information of different scales by setting different dilated rates. At the same time, spatial pyramid pooling (SPP) [18] can detect complex objects in multiple fields of view and extract features from different angles and then aggregate them by 1 × 1 convolution, which improves detection accuracy. Combining the advantages of AC and SPP, an atrous spatial pyramid pooling (ASPP) module is added to a branch of the network to extract more context features, which can further improves the classification performance on UWF images without increasing parameters. Specifically, we use three different dilated rates to perform atrous convolution operation and all the extracted features are integrated by a 1 × 1 convolution layer. The structure of ASPP module is demonstrated in Fig. 3 in which we set the dilated rate to 1, 2, and 3 respectively. Because the extraction of information with a large dilated rate may only have an obvious effect on the detection of some large objects, while the focus area of most fundus diseases is relatively small compared with the global background area.
The structure of ECA and SA modules. After a GAP layer, ECA generates channel attention weights matrix by a one-dimensional convolution of size K, where K is adaptively determined by the number of channels. The channel attention map is obtained by multiplying the channel attention weight matrix with the original feature map. After AP and MP, we use a convolutional layer and an sigmoid function to integrate the concatenated features as the spatial attention weight matrix. Finally, the channel-refined map is multiplied by the spatial attention weights to get the final output.
2.3 Efficient Channel Attention Module and Spatial Attention Module
Inspired by [19,20,21], we use the attention mechanism of efficient channel attention (ECA) and spatial attention (SA) to enhance the feature representation ability. The ECA and SA modules are shown in Fig. 4. We perform a GAP operation on the feature map and capture the cross-channel information from each channel and its K neighbors, which can be achieved by a one-dimensional convolution of size K. Then use a matrix multiplication operation to integrate the output of the previous step with the original feature map. After that, we perform average pooling (AP) operation and max pooling (MP) operation, and the spatial attention coefficients matrix are obtained by a simple convolutional layer and sigmoid operation. Finally, all features are integrated and become the output of this module. Moreover, an adaptive selection method of one-dimensional convolution kernel size in ECA is employed to find the suitable receptive region of kernel. The formula is described as follows:
where K is the kernel size of one-dimensional convolution, C is the number of channels, b and γ are manually parameters, which are set as 1 and 2 respectively.
3 Experiments
3.1 Dataset and Implementation Details
The experiment performs on ultra-widefield scanning laser ophthalmoscopy images which is called UWF dataset. The UWF is obtained from a local hospital and includes four categories of images. We choose the DR and the Normal images here. Among them, we have 398 images of DR and 948 images of Normal, which are divided into training data and testing data at 3:1. The specific divisions of the training data and testing data are shown in Table 1. In addition, the images in the UWF dataset have a high resolution of 2600 × 2048 which would be difficult to put the original images directly into the network for training. So we resize the images to 448 × 448. Meanwhile, random vertical and horizontal flipping are used for data augmentation in the training set to prevent network overfitting. Accuracy, Precision, Recall, and F1-score are used to evaluate the network classification performance.
We implement our work with PyTorch and use GPUs to accelerate the training process. The pre-trained ResNet-34 model is used for the first three layers of the network, and the second two layers are initialized randomly. During training, we choose the Adam with default values as the optimizer. We set the max epoch to 80, set the size of every training batch to 8 and set the size of every testing batch to 1. In addition, the learning rate is 0.0001 and decays by 10% per 50 epochs.
3.2 Experimental Results
The experimental results of different methods is shown in Table 2. We choose VGG16 [22], ResNet-34 [17], ResNet-50 [17], InceptionV3 [23], and DenseNet121 [24] for comparison. The VGG16, ResNet-34, ResNet-50, and DenseNet121 contain the characteristics of deep networks, while InceptionV3 has the characteristics of multiple branches and great feature extraction ability. So these networks are good baselines for research.
From Table 2, we can see that among the backbone networks, InceptionV3 has the best performance, followed by ResNet-34. So we know the multi-branch has an impact in the stage of feature extraction. From line 2, line 3, and line 5, it shows that the deep networks perform not as good as other networks. Generally, the deeper networks are easily overfitting and difficult to train. The proposed method extracts multi-scale features and uses the attention mechanism to strengthen the degree of attention to some important information. From Table 2, our model performs well compared with the other backbone networks and achieves the highest accuracy of 98.82%.
Ablation experiments are conducted to estimate the modules in our model. Specifically, we select the following networks for comparison: ResNet-34 model (ResNet34), dual-branch model (DB), dual-branch model with ASPP (DB-ASPP), dual-branch model with attention modules (DB-AM), and dual-branch network with all complete modules (Proposed). The experimental results are shown in Table 3. We can know that adding corresponding modules to the backbone can slightly strengthen the classification performance. From the experimental results, the score of dual-branch model with ECA and SA on Recall is lower than backbone, because both ECA and SA modules operate with deep features. Without strengthening the global feature extraction capability, the use of ECA and SA alone will produce biased errors in the classification results. So the result has a slight decrease in Recall.
To more intuitively evaluate our method, we plot the receiver operating characteristic (ROC) curves and the area under curve (AUC) is used as the evaluation metric. The ROC curves are shown in Fig. 5 in which we can observe our model has very good performance.
4 Conclusions
We propose a novel dual-branch network for DR classification. Two branches use ResNet-34 as the backbone, which can strengthen the extraction ability. ASPP enlarges the receptive field and integrates more features. ECA and SA emphasize the important information of feature space to obtain more discriminative features. The experimental results show our method performs better than other methods on the UWF dataset.
References
Yau, J.W., et al.: Global prevalence and major risk factors of diabetic retinopathy. Diabetes Care 35(3), 556–564 (2012)
Webb, R.H., Hughes, G.W.: Scanning laser ophthalmoscope. IEEE Trans. Biomed. Eng. 7, 488–492 (1981)
Haleem, M.S., Han, L., van Hemert, J., Li, B., Fleming, A.: Retinal area detector from scanning laser ophthalmoscope (SLO) images for diagnosing retinal diseases. IEEE J. Biomed. Health Inform. 19(4), 1472–1482 (2014)
Ashok, V., Murugesan, G.: Detection of retinal area from scanning laser ophthalmoscope images (SLO) using deep neural network. Int. J. Biomed. Eng. Technol. 23(2–4), 303–314 (2017)
Nagasato, D., et al.: Deep neural network-based method for detecting central retinal vein occlusion using ultrawide-field fundus ophthalmoscopy. Ophthalmology 2018 (2018)
Pellegrini, E., et al.: A graph cut approach to artery/vein classification in ultra-widefield scanning laser ophthalmoscopy. IEEE Trans. Med. Imag. 37(2), 516–526 (2017)
Brown, J.M., et al.: Automated diagnosis of plus disease in retinopathy of prematurity using deep convolutional neural networks. JAMA Ophthalmol. 136(7), 803–810 (2018)
Li, F., et al.: Deep learning-based automated detection for diabetic retinopathy and diabetic macular oedema in retinal fundus photographs. Eye, 1–9 (2021)
Diaz-Pinto, A., et al.: Retinal image synthesis and semi-supervised learning for glaucoma assessment. IEEE Trans. Med. Imag. 38(9), 2211–2218 (2019)
Xie, H., et al.: Cross-attention multi-branch network for fundus diseases classification using SLO images. Med. Image Anal. 71, 102031 (2021)
Hamwood, J., et al.: Automatic detection of cone photoreceptors with fully convolutional networks. Transl. Vis. Sci. Technol. 8(6), 10 (2019)
Ruan, Y., et al.: MB-FSGAN: joint segmentation and quantification of kidney tumor on CT by the multi-branch feature sharing generative adversarial network. Med. Image Anal. 64, 101721 (2020)
Mou, L., et al.: CS-Net: Channel and Spatial Attention Network for Curvilinear Structure Segmentation. In: Shen, D., et al. (eds.) MICCAI 2019. LNCS, vol. 11764, pp. 721–730. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-32239-7_80
Nie, D., Wang, L., Xiang, L., Zhou, S., Adeli, E., Shen, D.: Difficulty-aware attention network with confidence learning for medical image segmentation. In: AAAI, pp. 1085–1092 (2019)
Shaikh, M., Kollerathu, V.A., Krishnamurthi, G.: Recurrent attention mechanism networks for enhanced classification of biomedical images. In: IEEE ISBI, pp. 1260–1264 (2019)
Zhang, J., Xie, Y., Xia, Y., Shen, C.: Attention residual learning for skin lesion classification. IEEE Trans. Med. Imag. 38(9), 2092–2103 (2019)
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition. In: CVPR, pp. 770–778 (2016)
Gu, Z., et al.: DeepDisc: optic disc segmentation based on atrous convolution and spatial pyramid pooling. In: Stoyanov, D., et al. (eds.) OMIA/COMPAY 2018, LNCS, vol. 11039, pp. 253–260. Springer, Cham (2018)
Schlemper, J., et al.: Attention gated networks: learning to leverage salient regions in medical images. Med. Image Anal. 53, 197–207 (2019)
Xu, K., et al.: Show, attend and tell: neural image caption generation with visual attention. In: Bach, F.R., Blei, D.M. (eds.) International Conference on Machine Learning 2015, PMLR, vol. 37, pp. 2048–2057. PMLR, Lille (2015)
Wang, Q., et al.: ECA-Net: efficient channel attention for deep convolutional neural networks. In: CVPR (2020)
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition. In: ICLR (2015)
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J., Wojna, Z.: Rethinking the inception architecture for computer vision. In: CVPR, pp. 2818–2826 (2016)
Huang, G., et al.: Densely connected convolutional networks. In: CVPR, pp. 4700–4708 (2017)
Acknowledgements
This work was supported partly by National Natural Science Foundation of China (Nos. 61871274, 61801305 and 81571758), National Natural Science Foundation of Guangdong Province (No. 2020A1515010649 and No. 2019A1515111205), Guangdong Province Key Laboratory of Popular High Performance Computers (No. 2017B030314073), Guangdong Laboratory of Artificial-Intelligence and Cyber-Economics (SZ), the China Postdoctoral Science Foundation (2021M692196), Shenzhen Peacock Plan (Nos. KQTD2016053112051497 and KQTD2015033016104926), Shenzhen Key Basic Research Project (Nos. JCYJ20190808165209410, 20190808145011259, JCYJ20180507184647636, GJHZ20190822095414576 and JCYJ20170302153337765, JCYJ20170302150411789, JCYJ20170302142515949, GCZX2017040715180580, GJHZ20180418190529516, and JSGG20180507183215520), NTUT-SZU Joint Research Program (No. 2020003), Special Project in Key Areas of Ordinary Universities of Guangdong Province (No. 2019KZDZX1015), Shenzhen Key Medical Discipline Construction Fund (No. SZXK038), Shenzhen Fund for Guangdong Provincial High-level Clinical Key Speciaties (No. SZGSP014), Shenzhen-Hong Kong Co-financing Project (No. SGDX20190920110403741).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Tian, Z. et al. (2021). Dual-Branch Attention Network and Atrous Spatial Pyramid Pooling for Diabetic Retinopathy Classification Using Ultra-Widefield Images. In: Fu, H., Garvin, M.K., MacGillivray, T., Xu, Y., Zheng, Y. (eds) Ophthalmic Medical Image Analysis. OMIA 2021. Lecture Notes in Computer Science(), vol 12970. Springer, Cham. https://doi.org/10.1007/978-3-030-87000-3_13
Download citation
DOI: https://doi.org/10.1007/978-3-030-87000-3_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-86999-1
Online ISBN: 978-3-030-87000-3
eBook Packages: Computer ScienceComputer Science (R0)