Abstract
A high proportion of lung cancer cases are detected at a late cancer stage when they present with symptoms to general practitioners (GP). Early diagnosis is a challenge because many symptoms are also common in other diseases. Therefore, this study aims to assess UK primary care data of patients one, two and three years prior to lung cancer diagnosis to capture trends in clinical features of patients with the goal of early diagnosis and thus potentially curative treatment. This longitudinal study utilises data from the Clinical Practice Research Datalink (CPRD) with linked data from the National Cancer Registration and Analysis Service (NCRAS). A comprehensive list of Read codes is created to select features of interest to establish if a patient has experienced a certain medical condition or not. The comparison of the relative frequencies of the identified predictors associated with cases and controls reveals the importance of the following groups of features: ‘Cough Wheeze’ and ‘Bronchitis unspecified’, ‘Dyspnoea’ and ‘Upper Respiratory Infection’, which are frequent events for lung cancer cases, where a high proportion of cases were also identified using ‘Haemoptysis’ and ‘Peripheral vascular disease’.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Lung cancer is the third most diagnosed cancer and the leading cause of cancer mortality in the United Kingdom (UK) and worldwide [3]. It is estimated that by 2030, lung cancer will be the third-highest cause of death in high-income countries and the fifth-highest cause in middle-income countries [13]. Detecting lung cancer at an early stage remains a major challenge for clinicians, where most of the lung cancer cases are undetectable until an advanced stage. The detection of lung cancer at a late stage of disease progression reduces the chance of disease cure where the disease becomes rapidly fatal, dropping the 5-year survival rate drastically to 10%. Recognition of lung cancer at an early stage can result in better prognosis with a 5-year survival rate, and thus the UK National Health Service (NHS) long-term plan is to boost cancer careFootnote 1.
In the UK, the general practitioners (GPs) play a major role in the detection and management of lung cancer, where a significant percentage of lung cancer cases are detected symptomatically when patients present to the GP with cancer alarm symptoms [14]. However, these symptoms are also quite common in other conditions, posing a challenge for healthcare professionals to determine high-risk symptomatic patients eligible for further analysis and the targeting of screening to people at a high enough risk of lung cancer to benefit, from the other individuals who will not. Currently, the identification of a high-risk target population for lung cancer screening is gaining importance due to evidence that illustrates the ability of Low-Dose Computed Tomography (LDCT) to reduce mortality. The results from NLST [20] and other pilot trials [1, 4, 6, 8, 17, 19, 21] show that lung cancer screening with LDCT can save lives and reduce death from lung cancer by 20% or more in high risk smokers.
GPs record primary care and referral information of patients in Electronic Medical Records (EMRs), where some GPs contribute their EMRs structured data in an anonymised form to data warehouses such as the Clinical Practice Research Datalink (CPRD). Therefore, the CPRD primary care database can be considered a rich source of health data, including demographic information, symptoms, diagnoses, tests, therapies, immunisation and referrals to secondary care. The EMRs records of the CPRD database offer great potential for researchers when conducting epidemiological studies that can address important questions of interest in healthcare. The EMRs of patients collected by GPs can provide a very valuable resource of information: many subjects screened in the past were at relatively low risk and benefited little, and costs were high. To be clinically and cost effective, LDCT screening needs to be offered to people at a high enough risk of lung cancer to benefit.
In this study, we aim to assess UK primary care data of patients one, two and three years prior to lung cancer diagnosis to capture trends in clinical features with the goal of early diagnosis and to identify those at high enough risk to benefit. This longitudinal study uses data from the Clinical Practice Research Datalink (CPRD) with linked data from the National Cancer Registration and Analysis Service (NCRAS). The features were identified for patients with an incident diagnosis of lung cancer in cohorts within the study period (01/01/2000- 31/12/2015). A comprehensive code list of features was created by our lung cancer clinician partners. This study is reliant on Read Codes to establish if a patient has experienced a certain medical symptom or condition or not, and the unstructured text data were inaccessible in this dataset.
2 Methods
2.1 Study Design and Population
CPRD is an ongoing primary care database of coded anonymised information about patients from GPs, including demographics, symptoms, diagnoses, drug prescriptions, immunisation, investigation and test results. Linkages enable follow-up of patients beyond the primary care setting. Data are recorded by GP staff using a hierarchical clinical classification system, called Read codes. Each Read code represents a health-related concept, which is also represented by a Read term (i.e., the plain language description described in the medical dictionary). More details about the CPRD “GOLD” dataset that is drawn from the EMRs software Vision can be found in [5, 18]. Approval for use of data for this project was granted by the CPRD Independent Scientific Advisory Committee (ISAC) (Protocol numbers \(18\_223\) and \(20\_014\)R). The study is a longitudinal case-control study in which data collected within the CPRD are used to compare features of interest between cases (i.e., individuals who later received a diagnosis of lung cancer) and controls (i.e., individuals with no lung cancer record). The initial extraction population from the CPRD GOLD database comprises all cases eligible for data linkage to the NCRAS cancer registry database. Patients are selected from the CPRD database and included in the study according to the following criteria:
-
1.
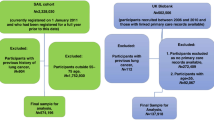
Patients with lung cancer (cases) are identified by the presence of one or more lung cancer diagnostic codes occurring within the study period (01/01/2000- 31/12/2015) and the date of the first lung cancer code was considered as the “index date”. Patients who had a record of lung cancer (within 01/01/1990- 31/12/2015) prior to their index date were excluded. The index date is defined as the date of the first ever record of a lung cancer diagnosis within follow up for the cases and a matched index date for the controls. The start of follow-up is defined as the latest of the patient registration date, the practice Up-to-standard (UTS) date and 01/01/2000. The end of follow-up will be defined as the earliest of the patient transfer out date, the practice last collection date, the CPRD GOLD death date and 31/12/2015. Furthermore, patients who are eligible for linkage to Hospital Episode Statistics (HES), National Cancer Registration and Analysis Service (NCRAS), ONS Death registration and patient level deprivation data are only included. Lung cancer cases were 40 years or older at the index date and had the event within their UTS follow-up. All patients within the CPRD Gold dataset matching these criteria were extracted. 26,701 cases have at least 12 months of follow up prior to their index date, as explained in Fig. 1.
-
2.
Control participants matched cases based on general practice, sex, and year of birth (within ±5 years), and had no lung cancer code anywhere in their patient record (either in CPRD GOLD or in the Cancer Registry). We also ensured that controls had at least 12 months of follow up prior to the index date of their matched case. CPRD used Index date Matching. In this algorithm, the case patient has a specified index date that must fall between the follow-up start and follow-up end dates of the control patient. This can be seen in Fig. 2. The start of follow up for the controls will be amended to ensure they have 12 months UTS follow up prior to the index date of their matched case.
In the final dataset, 26,701 cases were identified in the cancer registry data and CPRD GOLD. Up to 10 matching controls will be provided for each case. Once eligible patients are identified, the entire available coded records for cases and controls are extracted from the data files, as illustrated in Table 1. The data files are: Patients (i.e., 1 file), Consultation (i.e., 8 files), Clinical (i.e., 8 files), Additional clinical (i.e., 2 files), Referral (i.e., 1 file), Immunisation (i.e., 1 file), Test (i.e., 10 files), and Therapy (i.e., 28 files).
2.2 Demographic Characteristics of Cases and Controls
A total of 26,701 patients and 267,010 matched controls meeting the inclusion criteria were included in the analyses. Removing the missing values from the matched controls data (i.e., 388 (0.15%)) resulted in a dataset of 26,701 patient samples and 266,622 matched controls. Gender characteristics of both lung cancer patients and controls are shown in Table 2. Lung cancer patients and matched controls have similar age and sex distributions, as expected given the matching process, as shown in Fig. 3 and Fig. 4.
2.3 Features of Interest
Since EMRs data are recorded as Read codes, the associated data analysis relies mainly on generating code-lists to define features of interest. A code list can be defined as a collection of codes that describe certain medical conditions which can be used by researchers to investigate patient EMRs. Our code list comprises of 1,468 codes based on 17 groups of features, which are: Any Pulmonary Tuberculosis (i.e., 208 codes), Pulmonary Tuberculosis (i.e., 83 codes), Cough Wheeze (i.e., 48 codes), Pneumonia (i.e., 168 codes), Haemoptysis (i.e., 12 codes), Emphysema (i.e., 26 codes), Hypertension (i.e., 74 codes), Acute Myocardial Infarction (i.e., 65 codes), Bronchitis Unspecified (i.e., 95 codes), Dyspnoea (i.e., 65 codes), Cystic fibrosis (i.e., 17 codes), Upper Respiratory Infection (i.e., 310 codes), Idiopathic (i.e., 17 codes), Chronic Kidney Disease (i.e., 147 codes), Acute Nephritis With Lesions (i.e., 7 codes), Peripheral Vascular Disease (i.e., 90 codes), and Congestive Heart Failure (i.e., 34 codes). Read codes are utilised to select those groups of features for lung cancer reported in both cases and controls. This means that patients were identified as having experienced Dyspnoea (for instance) if they had a consultation with a Read code corresponding to that symptom. The identified list of Read codes is utilised to extract lung cancer cases and controls from the created data files (see Table 1). In this study, the relative frequencies of the identified predictors are assessed and compared between the records of cases and controls based on a set of clinical descriptions called medical codes (medcode) found in the clinical, referral, and test files, as explained in Table 3.
Table 4 explains the relative frequencies of the identified features between the clinical records of cases and controls. The group of features ‘Cough Wheeze’, which comprise 48 medcodes seems to be more frequent in the clinical records of cases (i.e., 1.03% ), compared with the controls (i.e., 0.74%) and also in comparison to other features. Furthermore, the group feature ‘Bronchitis Unspecified’ can be considered as a frequent event for lung cancer cases (i.e., 1.01%) compared with controls (i.e., 0.60%), and also in comparison to other subsets of features. The percentage of patients with ‘Dyspnoea’ as well as ‘Upper Respiratory Infection’ seems to be higher in the clinical records of cases (0.46%, 0.52%) compared with the clinical records of controls (0.27%, 0.34%), respectively. Furthermore, a bag of codes model is presented in Fig. 5 for cases and controls to show the frequency of codes in each cohort of clinical records. The medcode ‘92’ (equivalent to the Read code (171..00) that represents the ‘Cough’ symptom) constitutes 14% of the clinical records for both groups of samples, whereas the medcode ‘68’ (which corresponds to the medical concept of ‘Chest infection’ with Read code (H06z011)) constitutes 10% of the clinical records of cases and 9% of the corresponding records of controls. Moreover, the medcodes ‘2581’ (which represents the feature ‘Chest infection NOS’ with Read code (H06z000)) comprises 7% of the clinical records of cases compared with 5% of the records in the control group. Medcodes ‘1273’, and ‘799’ (‘C/O - cough’ and ‘Essential hypertension’) are also frequent events in the clinical records of both groups of samples, as illustrated in Fig. 5.
Table 5 shows the comparison of relative frequencies of the identified predictors between the referral records of cases and controls. As we have seen in the clinical file in Table 4, the percentage of patients with ‘Cough Wheeze’ features is higher in the referral records of lung cancer patients (i.e., 0.60%) compared to the controls (i.e., 0.30%) and in comparison to other lung cancer symptoms. Moreover, of 291,496 cases, 0.30% had ‘Haemoptysis’ in their referral records in comparison to (i.e., 0.06%) controls out of 3,184,693 records in the referral file. The group of features ‘Bronchitis Unspecified’ seems to be more frequent in the referral records of lung cancer cases (i.e., 0.33%) compared with control samples (i.e., 0.16%). The proportions of patients with ‘Dyspnoea’ and ‘Upper Respiratory Infection’ are higher in the referral records of cases (i.e., 0.55%, 0.32%) compared with the negative samples in the control group (i.e., 0.33%, 0.12%) respectively. The group of features ‘Peripheral vascular disease’ is more frequent in the referral records of cases (0.32%) compared to controls (0.15%) and in comparison to other groups of features. Furthermore, a bag of codes model is presented in Fig. 6 for cases and controls to show the frequency of codes in each cohort of referral records. The medcode ‘92’ (equivalent to Read code (171..00) representing the ‘Cough’ symptom) constitutes 14% of the referral records of cases and 12% of the referral records of controls, highlighting the importance of this symptom.‘Shortness of breath’ - (741/(R060800)) is slightly higher in the referral records of controls (10%) than cases (9%). Referring patients to the respiratory physician - (i.e., 10874/(ZL5A500)) is higher for cases (7%) than controls (5%) in the referral file. ‘Intermittent claudication’ (1517/(G73z000)) constitutes 7% of the referral records of cases compared to 4% of the referral records of controls. Moreover, the ‘Haemoptysis’ symptom (2244/(R063.00)), comprises 7% of the referral records of cases in comparison to 3% of the corresponding records of controls.
In the Test file, the group of features ‘Chronic Kidney Disease’ seems to be a frequent event for both groups of samples, where its relative frequency for controls is slightly higher than cases. A bag of codes model is also created for the test records of cases and controls to show the relative frequencies of the features between these groups of samples. The ‘GFR calculated abbreviated MDRD’ (medcode ‘23250’ and Read code ‘451E.00’) comprises 80% of the test records of lung cancer cases and 81% of the test records of controls, as shown in Fig. 7. As a result, the total number of EMRs extracted from clinical, referral, and test files for cases is 1,105,653 compared to 12,620,203 EMRs for control samples, resulting in a dataset of 13,725,856 samples (Table 6).
3 Data Analysis
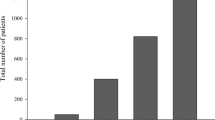
As mentioned previously, the created dataset contains 13,725,856 samples, where the majority are the control samples (i.e., 12,620,203 (91.94%)) and the minority are the lung cancer cases (i.e., 1,105,653 (8.06%)), as shown in Fig. 8 (a). Training a machine learning classification model using a dataset that suffers from an imbalanced class distribution such as this poses a tough challenge for learning algorithms in terms of capturing something meaningful from the minority samples. The issue of imbalanced class distribution simply refers to the challenge that occurs when the number of samples that represent the class of interest is much lower than the other classes, which can be considered a common problem in real-world data. In situations like this, the classifiers are more likely to be biased towards the majority class causing a high-level of miss-classification rate of the minority class as shown in Fig. 9 (b), where the percentage of lung cancer cases that were incorrectly classified is 95.9% compared to 99.8% correctly classified controls. However, if we attempt to quantify the predictive performance of the classification model using the well-known accuracy metric, the outcome is 92.1%, as shown in Fig. 9 (a). Therefore, adopting reliable evaluation measurements, as illustrated in Fig. 9 (b) demonstrates the consequences of feeding the learning models with imbalanced class data.
In our research problem, the dataset can be considered highly imbalanced class data, where the majority are the controls (i.e., 91.94%), due to the fact that we have 10 matched controls for each lung cancer patient defined based on the matching process of age, gender, and GPs, as discussed in Sect. 2. Due to the advent of artificial intelligence based methods in analysing clinical data [2, 7, 10, 15, 16], several methods have been proposed in the literature for tackling imbalanced class issues, including oversampling, undersampling, and hybrid approaches, which integrate oversampling and undersampling techniques [9, 11, 12]. For the work presented in this paper, a particular form of an under-sampling technique was utilised and performed for creating several data samplings from the original dataset, rather than simply eliminating some of the samples from the majority class and losing some potentially very useful information. This undersampling technique has the potential to address the issues caused by imbalanced class data, in which we have one matching control at each file. As a result, we will have 10 matching case-control files. The matching case-control files are: Matching-file1 (i.e., 26700 samples), Matching-file2 (i.e., 26699 samples), Matching-file3 (i.e., 26695 samples), Matching-file4 (i.e., 26693 samples), Matching-file5 (i.e., 26689 samples), Matching-file6 (i.e., 26677 samples), Matching-file7 (i.e., 26663 samples), Matching-file8 (i.e., 26637 samples), Matching-file9 (i.e., 26610 samples), Matching-file9 (i.e., 26559 samples). The difference in the number of samples across Matching-files is due to having 388 missing values distributed in the matching files as follows respectively: (1, 2, 6, 8, 12, 24, 38, 64, 91, 142). For instance, selecting the first data sampling (Matching-file1 for performing the classification task), resulted in a more realistic and reliable accuracy (69%), as was quantified in Fig. 10 - (a). Furthermore, detecting the underlying structure of the data has improved drastically due to having enough representative examples for each class, leading to a dramatic improvement in the True Positive rate (TP), from (4.1%) (to 64.7%), which in turn has improved the capacity of the model to correctly classifying positive patients, as shown in Fig. 10 - (b).
4 Conclusion
In this paper, we emphasise the importance of the groups of features: ‘Cough Wheeze’, ‘Bronchitis unspecified’, ‘Dyspnoea’, and ‘Upper Respiratory Infection’ for the early detection of lung cancer. These symptoms are the commonest symptoms of lung cancer cases based on the utilised medical care dataset, where the percentage of patients defined with those symptoms seems to be higher in the EMRs of cases compared with controls, and also in comparison to other symptoms. We also found that a high percentage of patients identified using ‘Haemoptysis’ and ‘Peripheral vascular disease’ in comparison to other symptoms, highlighting the potential significance of those features. In the context of testing, ‘Chronic Kidney Disease’ is a frequent event in the test records of cases and controls, particularly the GFR calculated abbreviated MDRD (23250/(451E.00)), which constitutes around 80% of the test EMRs of both groups of samples. Currently, in the medical domain, it has been shown that there are still trends in overestimating ‘Haemoptysis’ and underestimating ‘Cough’, ‘Bronchitis unspecified’ and ‘Dyspnoea’, which are demonstrated in our research to be frequent events for lung cancer patients. Therefore, more emphasis should be placed on the symptoms of ‘Cough’, ‘Bronchitis unspecified’ and ‘Dyspnoea’ as for ‘Haemoptysis’.
References
Becker, N., et al.: Randomized study on early detection of lung cancer with MSCT in Germany: results of the first 3 years of follow-up after randomization. J. Thorac. Oncol. 10(6), 890–896 (2015)
Chen, L., Yan, J., Chen, J., Sheng, Y., Xu, Z., Mahmud, M.: An event based topic learning pipeline for neuroimaging literature mining. Brain Inf. 7(1), 1–14 (2020). https://doi.org/10.1186/s40708-020-00121-1
Ferlay, J., et al.: Cancer incidence and mortality patterns in Europe: estimates for 40 countries in 2012. Eur. J. Cancer 49(6), 1374–1403 (2013)
Field, J.K., et al.: The UK lung cancer screening trial: a pilot randomised controlled trial of low-dose computed tomography screening for the early detection of lung cancer. Health Technol. Assess. (Winchester, England) 20(40), 1 (2016)
Herrett, E., et al.: Data resource profile: clinical practice research datalink (CPRD). Int. J. Epidemiol. 44(3), 827–836 (2015)
Infante, M., et al.: Long-term follow-up results of the DANTE trial, a randomized study of lung cancer screening with spiral computed tomography. Am. J. Respir. Crit. Care Med. 191(10), 1166–1175 (2015)
Kaiser, M.S., et al.: iWorksafe: towards healthy workplaces during COVID-19 with an intelligent phealth app for industrial settings. IEEE Access 9, 13814–13828 (2021)
van Klaveren, R.J., et al.: Management of lung nodules detected by volume CT scanning. New England J. Med. 361(23), 2221–2229 (2009)
López, V., Fernández, A., García, S., Palade, V., Herrera, F.: An insight into classification with imbalanced data: empirical results and current trends on using data intrinsic characteristics. Inf. Sci. 250, 113–141 (2013)
Mahmud, M., Kaiser, M.S.: Machine learning in fighting pandemics: a COVID-19 case study. In: Santosh, K.C., Joshi, A. (eds.) COVID-19: Prediction, Decision-Making, and its Impacts. LNDECT, vol. 60, pp. 77–81. Springer, Singapore (2021). https://doi.org/10.1007/978-981-15-9682-7_9
Mahmud, M., Kaiser, M.S., McGinnity, T.M., Hussain, A.: Deep learning in mining biological data. Cogn. Comput. 13(1), 1–33 (2020). https://doi.org/10.1007/s12559-020-09773-x
Mahmud, M., Kaiser, M.S., Hussain, A., Vassanelli, S.: Applications of deep learning and reinforcement learning to biological data. IEEE Trans. Neural Netw. Learn. Syst. 29(6), 2063–2079 (2018)
Mathers, C.D., Loncar, D.: Projections of global mortality and burden of disease from 2002 to 2030. PLoS Med. 3(11), 1 (2006)
McDonald, L., et al.: Suspected cancer symptoms and blood test results in primary care before a diagnosis of lung cancer: a case-control study. Future Oncol. 15(33), 3755–3762 (2019)
Nahian, M.J.A., et al.: Towards an accelerometer-based elderly fall detection system using cross-disciplinary time series features. IEEE Access 9, 39413–39431 (2021)
Noor, M.B.T., et al.: Application of deep learning in detecting neurological disorders from magnetic resonance images: a survey on the detection of Alzheimer’s disease, Parkinson’s disease and schizophrenia. Brain Inf. 7(1), 1–21 (2020)
Paci, E., et al.: Mortality, survival and incidence rates in the ITALUNG randomised lung cancer screening trial. Thorax 72(9), 825–831 (2017)
Padmanabhan, S.: Cprd gold data specification (2015). https://www.ed.ac.uk/files/atoms/files/cprd_gold_full_data_specification. pdf
Sverzellati, N., et al.: Low-dose computed tomography for lung cancer screening: comparison of performance between annual and biennial screen. Eur. Radiol. 26(11), 3821–3829 (2016). https://doi.org/10.1007/s00330-016-4228-3
Team, N.L.S.T.R.: Reduced lung-cancer mortality with low-dose computed tomographic screening. New Engl. J. Med. 365(5), 395–409 (2011)
Wille, M.M., et al.: Results of the randomized Danish lung cancer screening trial with focus on high-risk profiling. Am. J. Respir. Crit. Care Med. 193(5), 542–551 (2016)
Acknowledgement
We would like to thank the Medical Technologies and Advanced Materials Strategic Research Theme at Nottingham Trent University for financial support.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Alzubaidi, A. et al. (2021). Selecting Lung Cancer Patients from UK Primary Care Data: A Longitudinal Study of Feature Trends. In: Mahmud, M., Kaiser, M.S., Kasabov, N., Iftekharuddin, K., Zhong, N. (eds) Applied Intelligence and Informatics. AII 2021. Communications in Computer and Information Science, vol 1435. Springer, Cham. https://doi.org/10.1007/978-3-030-82269-9_4
Download citation
DOI: https://doi.org/10.1007/978-3-030-82269-9_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-82268-2
Online ISBN: 978-3-030-82269-9
eBook Packages: Computer ScienceComputer Science (R0)