Abstract
The electrocardiogram (ECG), a recording of the electrical activity of the heart, is commonly used for cardiac analysis, but lack of abnormal ECG signal data restricts the development of high quality automatic auxiliary diagnosis. In this paper, we introduce an LSTM and GAN based ECG abnormal signal generator to alleviate the issue. By training with a small set of real abnormal signals, the proposed generator can learn and produce high quality fake abnormal signals. The fake signals are then combined with real signals to train abnormal ECG classifiers. We show that our method can significantly improve the ability of classifiers in recognizing the uncommon case with a low proportion in the database.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
The electrocardiogram (ECG) is a record of the heart muscle movement and has been widely used in detection and treatment for cardiac diseases. Different from normal ECG signals, arrhythmia signals can be harbingers of some dangerous heart diseases. Early diagnosis helps early detection of chronic disease and starts treatment as soon as possible. For emergency heart attack such as ventricular fibrillation, timely detection can significantly improve the survival rate [1]. Therefore, efficient and effective ECG arrhythmia detection and classification is very important. However, arrhythmia beats usually occur sporadically and unexpected, which makes them extremely difficult to record. The rarity of arrhythmia data limits the training quality of classifiers and becomes a hindrance on the road to a comprehensive diagnosis system [2].
To enable auto-diagnosis with limited real arrhythmia data, an efficient model to generate ECG signals with high confidence and quality, especially the abnormal ECG signals, is necessary. To build a system that can mimic arrhythmia signal, we choose long short-term memory (LSTM) and generative adversarial nets (GANs) as main components. The GANs were first published in 2003 by Goodfellow et al. [3]. In the system, there is a generator (G) that will continuously produce fake signals to approach real database, while there is a discriminator (D) that can determine whether the input is real or from the generator. Both G and D will be trained at the same time until the discriminator cannot define the true label. Instead of using the original ECG signal, we use the feature got from an LSTM encoder to train the generator.
LSTM is a special recurrent neural network model, which can selectively remember the important information of the input. It is good at dealing with correlated data sequence and commonly used on speech recognition [4] and sequence translation [5]. In our system, the LSTM encoder and decoder are trained to find the commonality of a group of data and exclude the effort from differences among individuals. To learn and mimic arrhythmia signals, a GAN model [3] is included between LSTM encoder and decoder to learn the commonality within hidden states. Previous study shows that GANs can generate similar output based on input efficiently [6]. So we choose GANs as the generative model to synthesize the commonality given by LSTM encoder. After enough training, it can generate fake states and pass them to LSTM decoder to produce high quality fake signals.
Our main contributions are listed below: (1) We proposed an LSTM and GAN based arrhythmia generator, which can learn from a small data set and produce high quality arrhythmia signals. (2) We optimize the system performance by studying the correlation between training iteration and training samples. (3) We verify the effectiveness of our method on MIT-BIH data set with random forest classifiers. Compared to classifier trained with only real data, the same classifier that is trained with both real and fake data achieves an accuracy boost from 84.24 to 95.46%.
The remaining of this paper is organized as follows: Sect. 2 is the related work about both mathematical and machine learning based ECG models. In Sect. 3, our method will be discussed in detail. Section 4 describes the classifier in order to test our method. And the result is shown in Sect. 5. Discussion about future work is given in Sect. 6. The paper is concluded in Sect. 7.
2 Related Work
2.1 Mathematical ECG Models
Mathematical ECG models use a set of dynamic equations to fit the ECG behavior. In [7], a realistic synthetic ECG generator is built by using a set of state equations to generate a 3D ECG trajectory in a 3D state space. ECG signal is represented by a sum of Gaussian functions with amplitudes, angular spreads, and locations controlled by the Gaussian kernel parameters as in Eq. 1. In [8], a sum of Gaussian kernels are fitted to a normal ECG signal and then used to generate abnormal signal. Switching between normal and abnormal beat types is achieved by using a first-order Markov chain. The state probability vector (P) is trained to be changed based on factor such as R–R time series. According to the P and the ECG morphology parameters, the next ECG beat type can be determined.
In [9] to apply the model to filtering arrhythmia, a new wave-based ECG dynamic model (WEDM) by separating different events of the ECG has been introduced. In this model, the ECG signal was separated into three events as the P, C, and the T that represent the P-wave, the QRS complex, and the T-wave, respectively. Beside the (P,Q,R,S,T) events, there are also parameters needed for R-R process used in the wave-based synthetic ECG generation, which are mean heart rate, the standard deviation of the heart rate, the mean normalized low frequency, the standard deviation of the normalized low frequency, the mean respiratory rate, the standard deviation of the respiratory rate, and the low frequency to high frequency ratio. By controlling the set of parameters, it is possible to generate abnormal signals.
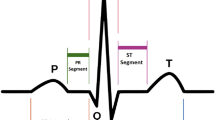
Mathematical models can generate high quality ECG signal with enough data to extract the parameters. These parameters are often more than just standard parameters P, Q, R, S, T as marked in Fig. 1. More parameters, such as mean heart rate, the standard deviation of heart rate, the mean normalized low frequency, etc., are often necessary to configure the models. However, lack of enough ECG data for each type of abnormality causes extreme difficulty on parameter extractions and makes mathematical models inappropriate for general abnormal ECG signal generation.
2.2 Machine Learning ECG Models
The generative adversarial networks (GANs) [3], based on the mini–max two-player game theory, show a great superiority in generating high quality artificial data. In [10], a GAN based inverse mapping method has been produced. Instead of cooperation between the constructed and original image space, the latent space has been used to update the generator. The similarity of the reconstructed image to original image is around 0.8266, which is higher than direct training. In [11], personalized GANs (PGANs) are developed for patient-specific ECG classification. The PGANs learn how to synthesize patient-specific signals by dealing with the P, Q, R, S, T events. The results are used as additional training data to get higher classification rate on personal ECG data. Three types of arrhythmia are considered in the work with an average accuracy rate about 93%. However, the models are only trained to produce personalized ECG signals. As sporadic types of arrhythmia are even harder to collect for each individual, it is impractical to train GANs for personalized abnormal ECG signal generation.
Long short-term memory (LSTM) can solve the vanishing gradient problem caused by the gradual reduction of the back propagation. It has high performance when dealing with time series related issues such as speech recognition, machine translation, and encoding/decoding [4]. For ECG signal classification, LSTM encoder/decoder classifier can achieve 99.39% for ECG signals [12]. However, it only studies common arrhythmia that is equally distributed and ignores rare cases.
Although those models can generate both normal and abnormal signals, there are still limitations. For the mathematical model, there are too many parameters needed for computation. On the other hand, the input real data need to be analyzed before it can be used for calculating. For the VAE and PGAN models, the models are only trained to produce personalized ECG signals. However, for those sporadic types of arrhythmia, it is hard to record and cannot be used for training. Therefore, a general purposed ECG signal generator is needed.
3 Methodology
Figure 2 illustrates the overall flow of our LSTM and GAN based ECG signal generator. Instead of using LSTM as classifier, we use two LSTMs, one is as encoder to translate ECG signal data x into hidden states h, and another is as decoder to convert h back to x. Not only more data, but also more iterations can help improve the quality of the LSTM encoder and decoder. To generate fake ECG signals, we insert GANs between LSTM encoder and decoder and train it to generate fake latent vector hf that is similar to h. The LSTM encoder, decoder, and GANs are trained separately for each type of abnormal signals. For abnormal signal type that has less than 1000 beats, the iteration is increased to improve performance. After enough fake ECG data are generated, a random forest classifier is trained with both real and fake ECG data.
3.1 LSTM Encoder and Decoder
In this section, we describe the LSTM unit and how to train LSTM encoder and decoder. Figure 2b shows the general diagram of LSTM encoder and decoder. Each unit represents an LSTM cell. It takes the input signal x(t), the previous unit output h(t), the previous cell state C(t − 1), and the bias vector b as inputs. Inside a unit, three gates (f,i,o) work together to calculate the hidden state h. The gates’ activation value vectors are calculated to determine whether the input information should be used. The W is the separate weight matrices for each input. The activation of three gates is calculated in the following way: the forget gate (f) first determinates whether the previous memory should be used. The total input of the cell passes through the input gate (i) and sigmoid function (tanh) and then multiplies with the activation value of the input gate. After adding input value to the cell state C, the final output hidden state h is calculated by multiplying the output gate activation value (o) and tanh(C). h is used to update W for each gate.
The decoder is formed as an inverse process of the encoder, and it works by taking the final hidden state h as the first input. The conditional decoder takes the last generated output as one of the inputs that is represented as the dot line in Fig. 3b. The final output of the decoder is an ECG signal restored based on the hidden state h. All parameters including weight W and bias b are the same as the encoder unit but in reverse order.
3.2 GANs for Fake Hidden State Generation
The GANs are used to generate fake hidden state that can pass to the LSTM decoder to produce fake ECG signal waveform. The basic idea of the GANs is a mini–max game of two players: the generator (G) and the discriminator (D). D exterminates the samples to determine whether they are valid (real hidden states) or invalid (fake hidden states), while G keeps creating samples with similar distribution to real hidden states. We implemented the generator and discriminator in the following way: Generator(G): The generator is a 2-layer feed-forward fully connected ANN with ReLu. It takes a random noise as input and gives the fake hidden state hf as output. Discriminator(D): The discriminator is a 4-layer feed-forward fully connected ANN with ReLu between each layer. It is trained to distinguish the real and fake states.
Algorithm 1 summarizes the training of GANs. The output of the function D(x) is the predicted label from the discriminator of input x, while the \(MSE(Y,\hat {Y}) = \frac {1}{n} \sum _{i=1}^{n} (Y_i-\hat {Y}_i)^{2}\), Y is the predicted label, and the \(\hat {Y}\) is the ground truth. A mini-batch of real state and a mini-batch of fake state made from the former step variables are chosen. The training is based on two simultaneous stochastic gradient descending processes. At each step, mini-loss functions G loss and D loss are all given by D. D loss shows the ability of D in different real and fake states. A 0 D loss means it can perfectly distinguish the inputs, and the higher D loss is the worse situation it will be. The G loss represents how much the G can “cheat” D. G loss equals to 0 means all the fake states given by G are considered as real, the higher, the worse. A learning rate of 0.0002 is chosen [6].
4 Random Forest Arrhythmia Classifier
The random forest (RF) classifier used to classify different types of abnormal ECG signals. The RF is a method that brunch of decision trees work together. Each tree votes independently, and the final decision is made with the class that gets most votes. The decision tree is built by randomly choosing n features from the total m characteristics of the samples, where \((n=\sqrt {m})\). The child nodes are decided by using those n features until the current n features have been used in its parent node. Multiple decision trees are built independently based on randomly picked samples of n features. Combination of all the decision trees forms a random forest. The contribution of RF works as follows:
-
(1)
Randomly pick N samples from the original data set with replacement.
-
(2)
Train the root node of the decision node based on the N samples.
-
(3)
Randomly choose m features from the total \(M(m=\sqrt {M})\) characteristics of the samples. Use the m features as the split nodes until the current m features have been used in the parent node.
-
(4)
Independently build multiple decision trees following steps 1–3, while all trees are built without pruning.
-
(5)
Combine those decision trees with counters.
The feature for RF classifier in our system is the signal itself. The classifier is trained based on real data only to classify each arrhythmia type, and then another classifier is trained to prove the effect of real and fake data used together.
5 Result
5.1 ECG Database
In experiments we use MIT-BIH arrhythmia database [13]. In the MIT-BIH database, total 23 types of annotations have been record. The 4 types we used are: left bundle branch block beat (LB) with 8011 beats, right bundle branch block beat (RB) with 6425 beats, aberrated atrial premature beat (AA) with 6548 beats, and the atrial fibrillation (AF) with only 310 beats as the rare ECG type.
5.2 Result for LSTM and GANs
Figure 4 shows the output of decoder for type AF with different iterations. As iteration number increases, LSTM decoder can better capture the commonality of AF signals. Figure 5 gives the error vs. iteration curves for all four types. Figure 6 shows the fake hidden states generated by GANs and its fake ECG signal generated by the LSTM decoder. We can see that GANs+LSTM decoder can produce similar but not exactly the same abnormal ECG signals, which are suitable for the later training of the RF classifier.
5.3 Classification with Fake ECG Signal
Here we compare the qualities of training based on real data only vs. training on both real and fake data. Table 1 summarizes the training result with or without fake data and also comparisons to other existing methods. RF stands for RF classifier trained with only real data. It achieves an average 84.24% accuracy in total 21,298 testing samples. And for each abnormal ECG type, the classifications rates are: 95.33% for LB, 61.34% for RB, 96.29% for AA, and 0.18% for AF. RLG stands for RF classifier trained with both real and fake data. Fake signals are added to data sets of each type to an equal number of 8200. After training, the average classification accuracy rate increases to 95.46%. Moreover, for each type, we have 95.28% for LB, 95.39% for RB, 95.85% for AA, and 93.55% for AF. Note that testing is only done on real data for a fair comparison, and we can see a significant boost in accuracy especially for AF. The huge difference between with or without fake data on AF classification shows the importance of data balance for training as well as verifies the effectiveness of our approach.
6 Future Work
Although ECG based arrhythmia test is attracting more and more attentions and more and more ECG databases are created, the arrhythmia beats included in are still not comparable to normal beats. In the MIT-BIH database [13], only 16 beats of 110,000 are recorded as atrial escape beat. On the other hand, all the annotations are given by two cardiologists artificially, and some of the public databases such as [16] do not include arrhythmia annotation. Therefore, mathematical models can be used as auxil. In [17], new models based on probability density function are created to make more detailed divisions about ECG signals. In that case, neural network can participate in adjust parameters.
There are research that show that ECG signals vary from person to person [18], and arrhythmia ECG models can also be created to adapt to different persons. With the development of wearable ECG detect device [19], personalized ECG signal can be collected easily. Since our system can generate ECG beat from few input real signals, we can build a personalized classification system within few periods of testing.
Hardware based ECG application is another field with good development prespects. In [20] a 65-nm CMOS is used for personal cardiac monitoring. A classifier is trained based on the MIT-BIH database and then tested with connected ECG processor. With the hardware based neural network implementation method [21] and database extended by our generator, a faster arrhythmia can be trained to get higher accuracy.
Therefore, we will consider in future the feasibility of this hardware–software co-operated application. We want to create a wearable device with pre-installed arrhythmia classifier that can be adjusted by users’ ECG signal to finish the personal cardiac monitoring.
7 Conclusion
In this work, we present an LSTM and GAN based ECG signal generator to improve abnormal ECG classification especially for rare types. Our LSTM encoder can extract the hidden states that represent the commonality between individuals of the same ECG type. With fake states from GANs, our LSTM decoder can produce high quality fake signals for later detection and classification. We implemented a random forest classifier to verify the effectiveness of our approach. With the help of fake ECG signals, the average classification rate improves from 84.24 to 95.46%, with classification accuracy for rare ECG types (AF) boosted from 0.18 to 93.55%
References
V. Fuster, L.E. Rydén, D.S. Cannom, H.J. Crijns, A.B. Curtis, K.A. Ellenbogen, J.L. Halperin, J.-Y. Le Heuzey, G.N. Kay, J.E. Lowe, et al., ACC/AHA/ESC 2006 guidelines for the management of patients with atrial fibrillation: a report of the American college of cardiology/American heart association task force on practice guidelines and the European society of cardiology committee for practice guidelines (writing committee to revise the 2001 guidelines for the management of patients with atrial fibrillation) developed in collaboration with the European heart rhythm association and the heart rhythm society. J. Am. College Cardiol. 48(4), e149–e246 (2006)
E.J.d.S. Luz, W.R. Schwartz, G. Cámara-Chávez, D. Menotti, ECG-based heartbeat classification for arrhythmia detection: a survey. Comput. Methods Programs Biomed. 127, 144–164 (2016)
I. Goodfellow, J. Pouget-Abadie, M. Mirza, B. Xu, D. Warde-Farley, S. Ozair, A. Courville, Y. Bengio, Generative adversarial nets, in Advances in Neural Information Processing Systems (2014), pp. 2672–2680
A. Graves, N. Jaitly, A.-r. Mohamed, Hybrid speech recognition with deep bidirectional LSTM, in 2013 IEEE Workshop on Automatic Speech Recognition and Understanding (IEEE, Piscataway, 2013), pp. 273–278
I. Sutskever, O. Vinyals, Q.V. Le, Sequence to sequence learning with neural networks, in Advances in Neural Information Processing Systems (2014), pp. 3104–3112
I. Goodfellow, Nips 2016 tutorial: Generative adversarial networks (2016). Preprint arXiv:1701.00160
P.E. McSharry, G.D. Clifford, L. Tarassenko, L.A. Smith, A dynamical model for generating synthetic electrocardiogram signals. IEEE Trans. Biomed. Eng. 50(3), 289–294 (2003)
J. Lee, D.D. McManus, P. Bourrell, L. Sörnmo, K.H. Chon, Atrial flutter and atrial tachycardia detection using Bayesian approach with high resolution time–frequency spectrum from ECG recordings. Biomed. Signal Proc. Control 8(6), 992–999 (2013)
G. Clifford, A. Shoeb, P. McSharry, B. Janz, Model-based filtering, compression and classification of the ECG. Int. J. Bioelectro. 7(1), 158–161 (2005)
G. Perarnau, J. Van De Weijer, B. Raducanu, J.M. Álvarez, Invertible conditional GANs for image editing 2016. Preprint arXiv:1611.06355
T. Golany, K. Radinsky, PGANs: Personalized generative adversarial networks for ECG synthesis to improve patient-specific deep ECG classification, in Proceedings of the AAAI Conference on Artificial Intelligence (2019)
Ö. Yildirim, A novel wavelet sequence based on deep bidirectional LSTM network model for ECG signal classification. Comput. Biol. Med. 96, 189–202 (2018)
G.B. Moody, R.G. Mark, The impact of the MIT-BIH arrhythmia database. IEEE Eng. Medicine Biology Mag. 20(3), 45–50 (2001)
R.G. Kumar, Y. Kumaraswamy, et al., Investigating cardiac arrhythmia in ECG using random forest classification. Int. J. Comput. Appl. 37(4), 31–34 (2012)
Z. Zhang, J. Dong, X. Luo, K.-S. Choi, X. Wu, Heartbeat classification using disease-specific feature selection. Comput. Biol. Med. 46, 79–89 (2014)
S. Pouryayevali, S. Wahabi, S. Hari, D. Hatzinakos, On establishing evaluation standards for ECG biometrics, in 2014 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP) (IEEE, Piscataway, 2014), pp. 3774–3778
J.P. do Vale Madeiro, J.A.L. Marques, T. Han, R.C. Pedrosa, Evaluation of mathematical models for QRS feature extraction and QRS morphology classification in ECG signals. Measurement 156, 107580 (2020)
S.K. Berkaya, A.K. Uysal, E.S. Gunal, S. Ergin, S. Gunal, M.B. Gulmezoglu, A survey on ECG analysis. Biomed. Signal Proc. Control 43, 216–235 (2018)
N. Glazkova, T. Podladchikova, R. Gerzer, D. Stepanova, Non-invasive wearable ECG-patch system for astronauts and patients on earth. Acta Astronaut. 166, 613–618 (2020)
S. Yin, M. Kim, D. Kadetotad, Y. Liu, C. Bae, S.J. Kim, Y. Cao, J.-s. Seo, A 1.06-μw smart ECG processor in 65-nm CMOS for real-time biometric authentication and personal cardiac monitoring. IEEE J. Solid-State Circ. 54(8), 2316–2326 (2019)
F. Zhang, M. Hu, Mitigate parasitic resistance in resistive crossbar-based convolutional neural networks (2019).Preprint arXiv:1912.08716
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Sun, H., Zhang, F., Zhang, Y. (2021). An LSTM and GAN Based ECG Abnormal Signal Generator. In: Arabnia, H.R., Ferens, K., de la Fuente, D., Kozerenko, E.B., Olivas Varela, J.A., Tinetti, F.G. (eds) Advances in Artificial Intelligence and Applied Cognitive Computing. Transactions on Computational Science and Computational Intelligence. Springer, Cham. https://doi.org/10.1007/978-3-030-70296-0_54
Download citation
DOI: https://doi.org/10.1007/978-3-030-70296-0_54
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-70295-3
Online ISBN: 978-3-030-70296-0
eBook Packages: Computer ScienceComputer Science (R0)