Abstract
Skin cancer is the most commonly diagnosed cancer in the United States with over a million cases being detected each year. Fortunately, early detection provides high odds of recovery. The traditional method of detection involves clinical screening, which is prone to false positives, followed by an invasive biopsy. While this provides for a high rate of detection, it is intrusive and costly. Artificial Intelligence for medical image analysis has proved effective in assisting in the diagnosis of many medical maladies, yet fine variations in the appearance of skin lesions has made applications to skin cancer detection difficult. We report that a deep convolutional neural network (CNN) trained over clinically labeled images (pixels) can accurately assist in the diagnosis of early-stage skin cancers. Specifically, we analyze skin lesions using CNN and evaluate its performance on seven dermatologist-certified clinical image types: Actinic keratoses and intraepithelial carcinoma (Bowen’s disease), basal cell carcinoma, benign keratosis-like lesions (solar lentigines, seborrheic keratoses, and lichen-planus-like keratoses), dermatofibroma, melanoma, melanocytic nevi, and vascular lesions (angiomas, angiokeratomas, pyogenic granulomas, and hemorrhage). The model provides significantly high levels of average accuracy, specificity, and sensitivity across these types.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
- Artificial intelligence
- Convolutional neural networks
- Deep learning
- Medical image analysis
- Cancer
- Image processing
1 Introduction
Skin cancer is the leading form of cancer in the United States [1]. It forms when skin cells multiply abnormally and can prove fatal if it is allowed to metastasize to other areas of the body through the lymphatic system. Most skin cancers result from exposure to Ultraviolet (UV) light. When the skin is unprotected, UV radiation damages DNA and can produce genetic mutations, which can subsequently lead to cancerous growths [2]. According to Didona et al. [3] the most common types of skin cancer in Caucasian populations are melanoma and nonmelanoma (i.e., basal and squamous cell carcinoma) skin cancers (NMSC), with melanoma accounting for 4% of all deaths from cancer [4].
Two methods are commonly employed to diagnose whether a skin sample (biopsy) should be taken: Visual examination of the skin by a physician [5, 6]; or dermatoscopy [7] and/or epiluminescence microscopy by a trained clinician [8]. Thus, initial diagnostic efficiency currently depends exclusively on the competence and perceptual capabilities of the practitioner. Perhaps unsurprisingly, both methods have been found to result in suboptimal detection efficacy [9] with false positives abounding. Hence there exists an urgent need for a screening method with increased sensitivity and specificity to be developed.
To address these issues, medical practitioners have increasingly been seeking to employ automated image processing tools that can more effectively diagnose skin cancer [10]. Maier et al. [11] successfully used dermatoscopic images to train an Artificial Neural Network to differentiate deadly melanomas from melanocytic nevi. Although promising, this study, like earlier attempts [12], was hampered by small sample sizes and a lack image variation [13].
Recent increases in data availability, paired with technological advances, have revigorated these efforts. A deep learning approach was successfully employed and returned more accurate diagnoses than most trained experts [14, 15]. Gautman et al. [16] issued an automation challenge to modelers and reported the top submission had an accuracy of 85.5% for disease classification. More recently, a deep convolutional neural network (CNN) model known as MobileNetV2 using a transfer learning method classified benign versus malignant lesions with an accuracy of 91.33% [17].
The objective of the current research is to expand earlier efforts in developing automated skin cancer detection systems by producing a model capable of accurately classifying seven different types of skin lesions. Stakeholders in this endeavor are patients with skin lesions and practitioners. Prefacing our findings, we demonstrate that our approach can provide a high degree of accuracy (95%) in the early diagnosis of skin cancer(s). Importantly, because human perception is not required, we argue this approach should greatly minimize the negative impact of human factors.
2 Dataset
The dataset was compiled by the Medical University of Vienna [18] and includes 10,015 images of pigmented skin lesions. Images were sampled equally from male and female patients with an average age of 51. Images were collected from different parts of the body (e.g., face, ear, and neck) and captured in resolutions ranging from 8 × 8 pixels to 450 × 600 pixels. Figure 1 displays a sample of the images used in the study. Images fall into seven different classifications:
-
Melanoma (mel): The most dangerous form of skin cancer which generally develops from pigment-containing cells known as melanocytes [19].
-
Basal cell carcinoma (bcc): This cancer affects the basal cells which are responsible for the production of new skins. While it rarely metastasizes it does spread easily [20].
-
Actinic keratosis (akiec): This “pre-cancer” indicator appears as a scaly patch resulting from accumulated UV exposure [21].
-
Benign keratosis-like lesions (bkl): A benign, painless skin disorder which is mostly associated with aging and exposure to UV light [22].
-
Vascular lesions (vasc): Common birthmarks that can be flat or raised [23].
-
Dermatofibroma (df): Superficial benign fibrous histiocytoma which primarily occur in women [24].
-
Melanocytic nevi (nv): Benign birthmarks and moles that resemble melanoma [25].
A lower extremity sample for a 50-year-old male diagnosed with melanocytic nevei (upper left). A lesion sample from a 60-year-old male diagnosed with melanoma (upper right). A face lesion sample from a 70-year-old female diagnosed with basal cell carcinoma (lower left) and a lesion sample from a 50-year-old female diagnosed with benign keratosis-like lesions (upper right)
Importantly, these classes are not unique. Thus, some patients may present with more than one type of lesion. More than 50% of lesion images were confirmed by pathology, while the ground truth for the rest of were either follow-up, expert consensus, or confirmation by in vivo confocal microscopy.
3 Methodology
CNNs are the state of the art in deep learning for image classification [26], and there are numerous applications for CNN medical image analysis [27].
3.1 Image Preprocessing
Images were preprocessed using normalization techniques, for example, scaling image intensity to the range of [0, 1]. To increase processing speed each image was down sampled to 50 × 50 × 3 pixels. Images were unevenly distributed across classes. Thus, to remove potential bias subsets were created by randomly sampling evenly from the seven categories that ensured the complete population was considered. One aspect of preparation of the images was to be assure that no repeated image should be appeared in training dataset, to address this issue, a chi-square distance measurement technique was used [28].
Data augmentation techniques were employed to increase the number of images available to train on. Images were rotated, zoomed, and flipped.
3.2 CNN
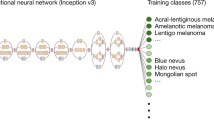
The architecture of CNN model employed is shown in Fig. 2. It consists of two convolutional parts: First, two convolutional layers followed by a pooling layer with a dropout rate of 0.25; second, two convolutional layers, a pooling layer and dropout rate of 0.30, trailed by a flattening of densely connected layers. The convolutional, also known as pooling steps, condense information. The lowest resolution images did not provide enough information to allow for the second convolutional/pooling layers and were omitted. Sometimes results from the first convolutional models were good, besting more complex models. This pattern was particularly true for medium-resolution images. It appears that medium-resolution images can run out of the information required by more complex models, and performance begins to suffer.
3.3 VGG-Net
The last set of models considered employed the VGG16 algorithm [29]. The general structure of this network is a 16-layer CNN that uses 3 × 3 filters with stride and pad of 1, along with 2 × 2 max pooling layers with stride of 2. The convolutional layers have 16, 64, 128, 256, 512 nodes successively. As the spatial size of the input volumes at each layer decrease, the result of the convolutional and pool layers, the depth of the volumes increases as the number of filters increases, doubling after each maxpool layer. The flattened layers consist of 1098, 4098, and 7 nodes. The final layer employs a SoftMax activation function.
4 Results
In the current analysis the CNN was found to produce an accuracy of 93% and minimal test loss of 0.18%. However, it did not sufficiently address the issue of overfitting. This can be seen clearly in the wide gaps between training and test set performance in Fig. 3.
When compared to the CNN without data augmentation, the model including augmented data did improve accuracy to 94% and decreased loss to 0.14% (shown in Fig. 4). The problem of overfitting was not ideal.
The VGG16 had an average accuracy of 93.67%, sensitivity of 95.66%, and specificity of 80.43%. A ten-fold cross validation was used to estimate the efficiency of the model. The learning curve of this topology is shown in Fig. 5. The learning curves indicate that the training loss decreases to a point of stability, and the small gap with training loss suggest overfitting was mostly resolved.
Metrics for each k-fold of the model are shown in Table 1. From the table, the ability of the model to correctly identify those with cancer (i.e., true positive rate) is as high as 96% and never lower than 94%, while the ability to correctly identify those without cancer (i.e., true negative rate) is as high as 83% and no lower than 70%. Notably higher than those provided by experts [9].
Table 2 indicates how accurately each of the seven classes of skin lesions are predicted. The most common type of skin cancer (bcc) is predicted with an accuracy of 95.61%. The deadliest skin cancer (mel) is predicted with an accuracy greater than 90%. Thus, the model performs well in diagnosing the most serious cases.
5 Conclusions
A deep learning approach to diagnosing different types of skin lesions ranging from potentially deadly skin cancers to benign age spots was employed. Results indicate that an automated approach can be used to effectively diagnose the etiology of lesions, detecting skin cancers more accurately than human experts [9]. This is an important finding with high pragmatic value. An approach to skin cancer screening will greatly improve health outcomes for patients while reducing resource expenditures. For instance, patients will be able to obtain accurate preliminary diagnosis from their primary care physician without seeking out a specialist, rural patients would be able to obtain a diagnosis through telemedicine, and laboratories would likely see a reduction in the number of unnecessary biopsies needing to be processed. While further testing and refinement is required, we believe the current results can help healthcare providers to make more accurate decisions.
References
Melanoma of the Skin Statistics, 9 April 2020. [Online]. Available: https://www.cdc.gov/cancer/skin/statistics/index.htm. Accessed 23 April 2020
D.L. Narayanan, R.N. Saladi, J.L. Fox, Ultraviolet radiation and skin cancer. Int. J. Dermatol. 9, 978–986 (2010)
D. Didona, G. Paolino, U. Bottoni, C. Cantisani, Non-melanoma skin cancer pathogenesis overview. Biomedicine 6(1), 6 (2018)
E. Losina, R.P. Walensky, A. Geller, F.C. Bedingfield, L.L. Wolf, B.A. Gilchrest, K.A. Freedberg, Visual screening for malignant melanoma: a cost-effectiveness analysis. Arch. Dermatol. 143(1), 21–28 (2007)
A.I. Riker, N. Zea, T. Trinh, The epidemiology, prevention, and detection of melanoma. J. Oschner 10(2), 56–65 (2010)
L. Brochez, E. Verhaeghe, L. Bleyen, J.M. Naeyaert, Diagnostic ability of general practitioners and dermatologists in discriminating pigmented skin lesions. J. Am. Acad. Dermatol. 44(6), 979–986 (2001)
A. Herschorn, Dermoscopy for melanoma detection in family practice. Can. Fam. Physician 58(7), 740–745 (2012)
A. Steiner, M. Binder, M. Schemper, K. Wolff, H. Pehamberger, Statistical evaluation of epiluminescence microscopy criteria for melanocytic pigmented skin lesions. J. Am. Acad. Dermatol. 29(4), 581–588 (1993)
J.K. Winkler, F. Christine, T. Ferdinand, et al., Association between surgical skin markings in Dermoscopic images and diagnostic performance of a deep learning convolutional neural network for melanoma recognition. JAMA Dermatol. 155(10), 1135–1141 (2019)
A. Marka, J.B. Carter, E. Toto, S. Hassanpour, Automated detection of nonmelanoma skin cancer using digital images: a systematic review. BMC Med. Imaging 19(21) (2019)
H. Maier, M. Schemper, B. Ortel, M. Binder, A. Tanew, H. Hönigsmann, Skin tumors in Photochemotherapy for psoriasis: a single-center follow-up of 496 patients. Dermatology 193, 185–191 (1996)
P. Rubegni, M. Burroni, B. Perotti, M. Fimiani, L. Andreassi, G. Cevenini, Digital Dermoscopy analysis and artificial neural network for the differentiation of clinically atypical pigmented skin lesions: a retrospective study. J. Investig. Dermatol. 119(2), 417–474 (2001)
P. Tschandl, T. Wiesner, Advances in the diagnosis of pigmented skin lesions. Br. J. Dermatol. 178, 9–11 (2018)
T.J. Brinker, A. Hekler, et al., Deep learning outperformed 136 of 157 dermatologists in a head-to-head dermoscopic melanoma image classification task. Eur. J. Cancer 113, 47–54 (2019)
A. Dascalu, E.O. Davis, Skin cancer detection by deep learning and sound analysis algorithms: a prospective clinical study of an elementary dermoscope. EBio Med. 43, 107–113 (2019)
D. Gautman, N. C. Codella, E. Celebi, B. Helba, M. Marchetti, N. Mishra, A. Halpern, Skin lesion analysis toward melanoma detection: a challenge at the International Symposium on Biomedical Imaging (ISBI). International Symposium on Biomedical Imaging, 2016
A. Cherif, M. Misbhauddin, M. Ech-Cherif, Deep neural network based mobile dermoscopy application for triaging skin cancer detection. Information security, ICCAIS, 2019
P. Tschandl, C. Rosendahl, H. Kittler, The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci. Data 161, 1–6 (2018)
Y. Liu, M. Saeed Sheikh, Melanoma: molecular pathogenesis and therapeutic management. Mol. Cell. Pharmacol. 6(3), 228 (2014)
J. Lanoue, G. Goldenberg, A comprehensive review of existing and emerging nonsurgical therapies. J. Clin. Aesth. Dermatol. 9(5), 26–36 (2016)
J.Q. Del Rosso, L. Kircik, G. Goldenberg, B. Berman, Comprehensive management of actinic keratoses. J. Clin. Aesth. Dermatol. 7(9), 2–12 (2014)
U. Wollina, Seborrheic Keratoses – the most common benign skin tumor of humans. Clinical presentation and an update on pathogenesis and treatment options. Open Access J. Med. Sci. 6(11) (2018)
S.C. Nair, Vascular anomalies of the head and neck region. J. Maxillofac Oral Surg. 17(1), 1–12 (2018)
J. Alves, D.M. Matos, H.F. Barreiros, E. Bártolo, Variants of dermatofibroma - a histopathological study. An. Bras. Dermatol. 89(3), 472–477 (2014)
M.R. Roh, P. Eliades, S. Gupta, H. Tsao, Genetics of melanocytic nevi. Pigment Cell Melanoma Res. 28(6), 661–672 (2015)
A. Krizhevsky, I. Sutskever, G.E. Hinton, ImageNet classification with deep convolutional. Adv. Neural Inf. Proces. Syst., 1097–1105 (2012)
O. Faust, Y. Hagiwara, T.J. Hong, O.S. Lih, U.R. Acharya, Deep learning for healthcare applications based on physiological signals: a review. Comput. Methods Prog. Biomed. 161, 1–13 (2018)
S. Hadipour, S. Aram, R. Sadeghian, Similar multi-modal image detection in multi-source dermatoscopic images of cancerous pigmented skin lesions. Submitted to The 2020 World Congress in Computer Science, Computer Engineering, & Applied Computing, 2020
K. Simonyan, A. Zisserman, Very deep convolutional networks for large-scale image recognition. arXiv preprint 1409, 1556 (2014)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Srivastava, R., Rahamathullah, M.J., Aram, S., Ashby, N., Sadeghian, R. (2021). A Deep Learning Approach to Diagnose Skin Cancer Using Image Processing. In: Arabnia, H.R., Ferens, K., de la Fuente, D., Kozerenko, E.B., Olivas Varela, J.A., Tinetti, F.G. (eds) Advances in Artificial Intelligence and Applied Cognitive Computing. Transactions on Computational Science and Computational Intelligence. Springer, Cham. https://doi.org/10.1007/978-3-030-70296-0_12
Download citation
DOI: https://doi.org/10.1007/978-3-030-70296-0_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-70295-3
Online ISBN: 978-3-030-70296-0
eBook Packages: Computer ScienceComputer Science (R0)