Abstract
Plants produce a variety of secondary metabolites which are being used as a source of medicine since the beginning of mankind, albeit most of them are synthesized in low concentrations. The developments in the field of ‘omics’ techniques help in the identification of genes of these metabolites having complex regulatory networks. Genetic engineering helps in manipulating the pathway which in turn increases the metabolite content and RNA interference (RNAi) is one such tool being used for the same. It is a homology dependent gene silencing technology in which the expression of pathway gene/promoter can be regulated by the introduction of double-stranded RNA (dsRNA) as it degrades the target mRNA. Since its discovery, this tool has been useful in manipulating the biosynthetic flux toward desired metabolite(s) by down-regulation of the competing pathway. In this chapter we discuss about RNAi as a tool to manipulate secondary metabolite pathways in plants.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Biosynthetic pathway
- Medicinal plants
- Metabolic engineering
- RNA interference (RNAi)
- Secondary metabolites
9.1 Introduction
Plants produce around 2,00,000 types of secondary metabolites as a defense response and they are useful sources of drugs, fragrances, pigments, food additives, and pesticides for mankind (Dixon and Strack 2003; Kutchan and Dixon 2005). It is estimated that 70–80% of the people worldwide rely mainly on herbal medicines for their primary healthcare (Canter et al. 2005). Reports document that out of 50,000–70,000 plants that are used worldwide for medicinal purposes, nearly 10,000 plants have become endangered (Brouwer et al. 2002; Edward 2004). World Health Organization (WHO) estimated that the market of herbal medicine will grow up to US$5 trillion by the year 2050 with an annual growth of 5–15% (Kumar and Gupta 2008). Due to complex chemical structures of the metabolites, they are difficult to synthesize chemically, and metabolites such as ajmalicine, ajmaline, artemisinin, berberine, colchicines, digoxin, ginsenosides, morphine, quinine, shikonin, taxol, vincristine, vinblastine, etc., are still extracted from plants (Rao and Ravishankar 2002). However plants synthesize metabolites in low concentrations and are restricted to a particular species or genus (Verpoorte et al. 2002). Thus to fulfill the demand, a large number of plants are collected from the wild which depletes the plants from natural habitat. Another problem faced by industries is the requirement of a large quantity of material for extraction of metabolites e.g., 2.5 kg of taxol requires 27,000 tons of Taxus brevifolia bark and thus the availability of plants for herbal medicines becomes a major problem (Rates 2001).
Synthesis of metabolites is under the control of different genes that are expressed in a particular tissue or cell type (Pichersky and Gang 2000). The plant genome contains 20,000–60,000 genes of which around 15–25% are involved in the synthesis of secondary metabolites (Bevan et al. 1998; Somerville and Somerville 1999). Metabolic engineering of pathways has key applications in alleviating the demands for limited natural resources (Lau et al. 2014). The secondary metabolite pathways are chain reactions catalyzed by enzymes that convert substrates into products with one or more branched points (Farré et al. 2014). Thus main challenge in manipulating the pathways is their complex nature which involves many regulatory factors (Kooke and Keurentjes 2012). Different strategies like blocking a competitive pathway, over-expressing regulatory genes/transcription factors, or inhibiting the catabolism of molecules can be used for the enhancement of metabolites (Koffas et al. 1999; Gomez-Galera et al. 2007).
9.2 Metabolic Engineering
Metabolic engineering is defined as the ‘directed improvement of product formation or cellular properties through the modification of specific biochemical reactions or the introduction of new genes with the use of recombinant DNA technology’ (Stephanopoulos 1999). The main aim of this technique is to redirect the precursor pool toward the synthesis of the desired compound(s) through alteration in the gene expression, and it is done either in positive (over-expression) or negative (down-regulation) manner (Pickens et al. 2011; Farré et al. 2014). The metabolic flux of the pathways can be regulated by the metabolites themselves, which in turn influences the activity of enzymes, transcription factors, and signaling proteins. The chemical diversity mainly arises through alkaloid, phenylpropanoid, and terpenoid pathways, thus number of studies have been carried out for identification of their regulatory genes and transcription factors (Wu and Chappell 2008; Nagegowda 2010). High throughput ‘omics’ technologies like genomics, transcriptomics, proteomics, and metabolomics are being used for elucidation of the pathways (Vemuri and Aristidou 2005; Caspi et al. 2013). In non-model plants where whole genome sequencing is not available, gene identification is done by a comparatively cheaper technique like expressed sequence tags (ESTs) (Joshi and Pathak 2019). Thus, the process of metabolic engineering in medicinal plants research is divided into three steps: (i) selection of plant species and elucidation of the pathways through ‘omics’ technology, (ii) targeting the gene of interest through genetic engineering tool, and (iii) screening the plants for metabolite content (Lau et al. 2014) (Fig. 9.1).
One of the key ways to reduce the levels of undesirable metabolites is recessive gene disruption and dominant gene silencing (Tang and Galili 2004). But the latter is a more promising approach to decrease the synthesis of undesirable compounds by suppression of branch-point gene which redirects the enzymatic reactions to increase the metabolite(s) of interest (DellaPenna 2001). Silencing the expression of a particular gene can be done in three different ways: (i) transcriptional gene silencing (TGS), (ii) post-transcriptional gene silencing (PTGS), and (iii) translation inhibition (Hamilton and Baulcombe 1999; Mansoor et al. 2006). But the central dogma of life suggests that if mRNA is silenced, further synthesis of secondary metabolites will be stopped (Abdurakhmonov 2016). RNA interference (RNAi) also known as post-transcriptional gene silencing (PTGS) is frequently used for gene down-regulation and thus known as the ‘knock-down’ method (Tang and Galili 2004).
9.3 RNA Interference (RNAi)
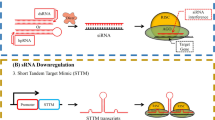
RNAi is a quick, easy, and sequence-specific homology-based tool to down-regulate the expression of targeted mRNA (Small 2007). Initially it was thought to function as a part of the defense mechanism against viruses when discovered in plants (Mansoor et al. 2006). The history of RNAi is nearly three decades old where Napoli and co-workers in 1990 transformed petunia plants with chalcone synthase (CHS) gene and the flower color changed from dark purple to white/chimeric, and this phenomenon was named as co-suppression. After five years, Guo and Kemphues (1995) reported knock-down of par-1 gene expression in Caenorhabditis elegans through both sense and antisense RNA. The reason behind gene silencing remained unknown till Andrew Fire and Craig Mello reported that potent and specific genetic interference can be done by double-stranded RNA (dsRNA) in C. elegans which triggered the silencing of genes as it had identical sequences to the mRNA. This type of gene silencing was termed as ‘RNA interference (RNAi)’ (Fire et al. 1998) and in 2006 Fire and Mello received the Nobel Prize for discovering it (Allen et al. 2004). At the same time similar phenomenon was also reported in plants by Waterhouse et al. (1998) where dsRNA induced gene silencing which was more efficient than either sense or antisense RNA. RNAi technology suppresses the expression of enzymes that are expressed in the number of tissues at different developmental stages, whereas sense or antisense RNA fails to block the activity of enzymes that are encoded by multigene family (Larkin et al. 2007). Wesley et al. (2001) compared the silencing efficiency of hpRNA (dsRNA) and antisense RNA, and reported that hpRNA increases gene silencing by 90–100%. Thus it was confirmed that RNAi became the most promising tool for the suppression of dominant gene expression (Smith et al. 2000). One advantage of this tool is its dominant nature and the silenced gene is passed on in the T1 generation which created new opportunities in agriculture and production of metabolites (Lessard et al. 2002; Verpoorte et al. 2002). Many researchers use in vitro cultures to down-regulate the gene as it reduces the risk of contaminating food sources and environment, and provides a platform to test a metabolic engineering strategy that will be utilized for large scale production of metabolites (Wu and Chappell 2008). The main aims of RNAi technology for engineering secondary metabolites synthesis is given in Fig. 9.2.
9.3.1 Mechanism
Micro RNA (miRNA), short interfering RNA (siRNA), and small hairpin RNA (hpRNA) are types of small non-coding RNAs that are mainly involved in RNAi mechanism (Aukerman and Sakai 2003; Palatnik et al. 2003). Artificial microRNA (amiRNA)-based vectors have also proved to be effective for gene silencing since the last decade (Warthmann et al. 2008). Smith et al. (2000) suggested that a more feasible approach is to clone both sense and antisense sequences separated by an intron region which forms a hairpin RNA (hpRNA) molecule upon transcription and triggers gene silencing. Aberrant single-stranded RNA (ssRNA) with an intron-hairpin construction triggers the generation of dsRNA by RNA-dependent RNA polymerase (RdRP) and activates the RNAi pathway (Waterhouse et al. 2001). Dicer, a ribonuclease III-type enzyme, is activated by ATP which recognizes dsRNA and cuts them into smaller segments of 21–25 bp. These small RNAs are then incorporated into a nuclease complex known as the ‘RNA-induced silencing complex’ (RISC) which contains argonaute protein (AGO). Then one of the strands of siRNA (guide strand) becomes stably associated with AGO and the other strand (passenger strand) is degraded. The guide strand then leads RISC to its target mRNA and AGO protein binds the guide strand to the target sequence for complementary base pairing. Successful docking of the RISC-siRNA complex with mRNA will then either block the translation or degrade mRNA using exonucleases (Kusaba 2004). Reports suggest that the directionality of dsRNA processing and the target RNA cleavage sites are predefined, and the sequence complementary to the guide siRNA will be recognized and cleave the target mRNA in the central region which is 10-12 nt from the 5’ end of siRNA (Elbashir et al. 2001). Lastly, the siRNA molecules are amplified via RdRp on the target mRNA and these siRNAs will, in turn, induce a secondary RNA interference i.e., transitive RNAi (Denli and Hannon 2003).
9.3.2 Vector and Transformation Methods
Different vectors are used to suppress gene expression in plants and the vector-based RNAi technology was improved by using an intron as the linker (Waterhouse et al. 1998; Smith et al. 2000). These RNAi vectors are specifically designed to generate long dsRNA with the same sequence as the target genes. Similarly, vectors designed to express hairpin RNAs (hpRNAs) are also successfully applied to silence the corresponding target genes (Wesley et al. 2003). Nowadays biotechnology companies are developing specialized vector constructs for RNA interference in plants (see table), which after transformation into host plant converts into dsRNAs and triggers efficient silencing.
One of the major issues in plant genetic transformation is to obtain a stably transformed plant which depends on the transformation methods. The first choice is Gram-negative, soil-borne pathogen Agrobacterium spp., which is also known as ‘natural genetic engineer’ is commonly used to transform numerous dicotyledonous plants (Zupan et al. 2000). But the wild-type Ti plasmid is very large (200 kb) and difficult to manipulate, which was overcome by the development of binary vectors (Bevan 1984). In such a system, the Ti plasmid of Agrobacterium has been disarmed by removing the T-DNA and keeping vir regions intact. Simultaneously, a separate binary vector is constructed which carries an origin of replication that is compatible with the Ti plasmid of Agrobacterium. When the binary vector is introduced into Agrobacterium the vir genes of Ti plasmid will act in trans to transfer the recombinant T-DNA from the binary vector to the host plant cell. As the binary vectors are smaller and comparatively easier to construct than wild-type Ti plasmids, the Agrobacterium-mediated transformation is considered as a reliable technique (Lessard et al. 2002).
Transient gene expression in majority of the plant species can be done via particle bombardment and electroporation. These techniques are useful especially when long term expression is not required for e.g., to test the effectiveness of various gene constructs before stable transformation (Lessard et al. 2002). One of the advantages of this method is high transformation frequency, which resulted in the successful transformation of plastids in tobacco and tomato (Maliga 2001). But these methods require the use of tissue culture protocols to regenerate transgenic plants/callus whereas Agrobacterium-mediated transformation overcomes this limitation by directly transforming germ-line cells or seeds and is one of the first choice for RNA interference in plants (Tague 2001).
RNAi is a promising way to manipulate the metabolite pathway (Borgio 2009) and it was first used by Mahmoud and Croteau (2001) in Mentha x piperita to reduce the level of menthofuran through antisense suppression of the mfs gene which codes for the cytochrome P450 (+) menthofuran synthase, which in turn increased the content of essential oils in plants. Later on many studies documented that the content of various volatiles can be increased in Mentha spp. by silencing different genes or transcription factors (Mahmoud et al. 2004; Wang et al. 2016; Reddy et al. 2017). Since the beginning of this technique, berberine bridge enzyme (BBE) is the gene of interest for RNAi research as many scientists knock-down the expression of this gene to study its effect on the content of different alkaloids, especially benzophenanthridine type in many plant species (Park et al. 2002; Frick et al. 2004; Fujii et al. 2007). Waterhouse et al. (1998) documented that this technology can be useful to alter the flower colors as compared to conventional breeding and genetic transformation. RNAi has been applied to suppress the genes of anthocyanin biosynthesis like anthocyanidin synthase (ANS) in Torenia spp. which changed the flower color in transgenic plants (Nagira et al. 2006; Nakamura et al. 2006). Similarly, other genes of flavonoid pathways like isoflavone synthase (IFS), flavone synthase II (FNSII), flavonol synthase (FLS), flavanone 3-hydroxylase (F3H), flavonoid 3’-hydroxylase (F3’H), flavonoid 3’,5’-hydroxylase (F3’5’H), flavone 6-hydroxylase (CYP82D1.1), flavone 8-hydroxylase (CYP82D2), chalcone isomerase (CHI), chalcone synthase (CHS), etc., were silenced and their effect on flavonoids was reported by many workers (Subramanian et al. 2005; Nakatsuka et al. 2007; Seitz et al. 2007; Park et al. 2011; Jiang et al. 2014; Zhang et al. 2015; Zhao et al. 2018). Recently Hu et al. (2020) reported that the down-regulation of one of the flavonoid biosynthetic pathway gene laccase gene (Lac1) affects the cotton fiber development. Whereas Liu et al. (2002) down-regulated the expression of two fatty acid desaturase genes i.e., stearoyl-acyl-carrier protein Δ9-desaturase (SAD) and oleoyl-phosphatidylcholine ω6-desaturase (FAD) in cotton seeds, which increased the content of stearic acid and oleic acid for better oil quality. Similarly, the content of different types of ginsenosides was increased or decreased in different species of Panax (P. ginseng, P. notoginseng and P. quinquefolium) by RNAi technique to identify the roles of different genes in ginsenoside biosynthetic pathway (Han et al. 2006; Zhao et al. 2015; Wang et al. 2017). This strategy has been used for commercial-scale production of desired plant products e.g., decaffeinated Coffea arabica and Coffea canephora plants were produced by silencing theobromine synthase gene using RNAi (Ogita et al. 2003, 2004). Table 9.1 depicts the plant species in which RNAi has been used to silence the secondary metabolite genes as well as the vector and transformation methods used for the same.
9.4 Conclusion
RNAi is the choice of present-day researchers to manipulate the genes synthesizing secondary metabolites. Since RNAi is a sequence-specific process, this requires the selection of a unique or conserved region of the target gene which ensures that the multiple gene families can be silenced. But the major bottleneck is that the complete information about the genomes of many non-model plants for secondary metabolite synthesis is lacking. The major drawback of RNAi tool is its unintended targets as 21–25 nt homology is required to suppress the gene function, even then it is still being used for identifying the gene functions and to increase the content of the desired metabolite.
References
Abdurakhmonov IY (ed) (2016) RNA interference—a hallmark of cellular function and gene manipulation. Intech Publications, Croatia, p 456
Ali A, Ahmad MM, Khan MA, Saxena P, Khan S, Abdin MZ (2017) RNAi‑mediated modulation of squalene synthase gene expression in Artemisia annua L. and its impact on artemisinin biosynthesis. Rend Fis Acc Lincei 28(4):731–741
Allen RS, Millgate AG, Chitty JA, Thisleton J, Miller JA, Fist AJ, Gerlach WL, Larkin PJ (2004) RNAi-mediated replacement of morphine with the nonnarcotic alkaloid reticuline in opium poppy. Nat Biotechnol 22(12):1559–1566
Allen RS, Miller JA, Chitty JA, Fist AJ, Gerlach WL, Larkin PJ (2008) Metabolic engineering of morphinan alkaloids by over-expression and RNAi suppression of salutaridinol 7-O-acetyltransferase in opium poppy. Plant Biotechnol J 6(1):22–30
Aukerman MJ, Sakai H (2003) Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 15(11):2730–2741
Bai Z, Li W, Jia Y, Yue Z, Jiao J, Huang W, Xia P, Liang Z (2018) The ethylene response factor SmERF6 co-regulates the transcription of SmCPS1 and SmKSL1 and is involved in tanshinone biosynthesis in Salvia miltiorrhiza hairy roots. Planta 248(1):243–255
Bevan M (1984) Binary Agrobacterium vectors for plant transformation. Nucleic Acids Res 12(22):8711–8721
Bevan M, Bancroft I, Bent E, Love K, Goodman H, Dean C, Bergkamp R, Dirkse W, Van Staveren M, Stiekema W et al (1998) Analysis of 1.9 Mb of contiguous sequence from chromosome 4 of Arabidopsis thaliana. Nature 391(6666):485–488
Borgio JF (2009) RNA interference (RNAi) technology: a promising tool for medicinal plant research. J Med Plants Res 3(13):1176–1183
Brouwer C, Bruce W, Maddock S, Avramova Z, Bowen B (2002) Suppression of transgene silencing by matrix attachment regions in maize: a dual role for the maize 5’ ADH1 matrix attachment region. Plant Cell 14(9):2251–2264
Cankar K, Jongedijk E, Klompmaker M, Majdic T, Mumm R, Bouwmeester H, Bosch D, Beekwilder J (2015) (+)-Valencene production in Nicotiana benthamiana is increased by downregulation of competing pathways. Biotechnol J 10(1):180–189
Canter PH, Thomas H, Ernst E (2005) Bringing medicinal plants into cultivation: opportunities and challenges for biotechnology. Trends Biotechnol 23(4):180–185
Caspi R, Dreher K, Karp PD (2013) The challenge of constructing, classifying and representing metabolic pathways. FEMS Microbiol Lett 345(2):85–93
Cheng Q, Su P, Hu Y, He Y, Gao W, Huang L (2014) RNA interference-mediated repression of SmCPS (copalyldiphosphate synthase) expression in hairy roots of Salvia miltiorrhiza causes a decrease of tanshinones and sheds light on the functional role of SmCPS. Biotechnol Lett 36(2):363–369
Chintapakorn Y, Hamill JD (2003) Antisense-mediated down-regulation of putrescine N-methyl transferase activity in transgenic Nicotiana tabacum L. can lead to elevated levels of anabatine at the expense of nicotine. Plant Mol Biol 53(1–2):87–105
Coleman HD, Park JY, Nair R, Chapple C, Mansfield SD (2008) RNAi-mediated suppression of p-coumaroyl-CoA 3’-hydroxylase in hybrid poplar impacts lignin deposition and soluble secondary metabolism. Proc Natl Acad Sci USA 105(11):4501–4506
DeBoer KD, Dalton HL, Edward FJ, Ryan SM, Hamill JD (2013) RNAi-mediated down-regulation of ornithine decarboxylase (ODC) impedes wound-stress stimulation of anabasine synthesis in Nicotiana glauca. Phytochem 86:21–28
DellaPenna D (2001) Plant metabolic engineering. Plant Physiol 125(1):160–163
Denli AM, Hannon GJ (2003) RNAi: an ever-growing puzzle. Trends Biochem Sci 28(4):196–201
Dexter R, Qualley A, Kish CM, Ma CJ, Koeduka T, Nagegowda DA, Dudareva N, Pichersky E, Clark D (2007) Characterization of a petunia acetyltransferase involved in the biosynthesis of the floral volatile isoeugenol. Plant J 49(2):265–275
Dhar N, Sarangapani S, Reddy VA, Kumar N, Panicker D, Jin J, Chua N-H, Sarojam R (2020) Characterization of a sweet basil acyltransferase involved in eugenol biosynthesis. J Exp Bot. https://doi.org/10.1093/jxb/eraa142
Dixon RA, Strack D (2003) Phytochemistry meets genome analysis and beyond. Phytochem 62:815–816
Edward R (2004) No remedy in insight for herbal ransack. New Sci 181:10–11
Elbashir SM, Lendeckel W, Tuschl T (2001) RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev 15(2):188–200
Fang R, Zou A, Zhao H, Wu F, Zhu Y, Zhao H, Liao Y, Tang RJ, Pang Y, Yang R, Wang X, Qi J, Lu G, Yang Y (2016) Transgenic studies reveal the positive role of LeEIL-1 in regulating shikonin biosynthesis in Lithospermum erythrorhizon hairy roots. BMC Plant Biol 16(1):121
Farré G, Blancquaert D, Capell T, Van Der Straeten D, Christou P, Zhu C (2014) Engineering complex metabolic pathways in plants. Annu Rev Plant Biol 65:187–223
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391(6696):806–811
Floss DS, Schliemann W, Schmidt J, Strack D, Walter MH (2008) RNA interference-mediated repression of MtCCD1 in mycorrhizal roots of Medicago truncatula causes accumulation of C27 apocarotenoids, shedding light on the functional role of CCD11[W][OA]. Plant Physiol 148(3):1267–1282
Frick S, Chitty JA, Kramell R, Schmidt J, Allen RS, Larkin PJ, Kutchan TM (2004) Transformation of opium poppy (Papaver somniferum L.) with antisense berberine bridge enzyme gene (anti-bbe) via somatic embryogenesis results in an altered ratio of alkaloids in latex but not in roots. Transgenic Res 13(6):607–613
Fu X, Shi P, He Q, Shen Q, Tang Y, Pan Q, Ma Y, Yan T, Chen M, Hao X, Liu P, Li L, Wang Y, Sun X, Tang K (2017) AaPDR3, a PDR transporter 3, is involved in sesquiterpene β-caryophyllene transport in Artemisia annua. Front Plant Sci 8:723
Fujii N, Inui T, Iwasa K, Morishige T, Sato F (2007) Knockdown of berberine bridge enzyme by RNAi accumulates (S)-reticuline and activates a silent pathway in cultured California poppy cells. Transgenic Res 16(3):363–375
Gao W, Xu F-C, Long L, Li Y, Zhang J-L, Chong L, Botella JR, Song C-P (2020) The gland localized CGP1 controls gland pigmentation and gossypol accumulation in cotton. Plant Biotechnol J. https://doi.org/10.1111/pbi.13323
Gavilano LB, Coleman NP, Burnley LE, Bowman ML, Kalengamaliro NE, Hayes A, Bush L, Siminszky B (2006) Genetic engineering of Nicotiana tabacum for reduced nornicotine content. J Agric Food Chem 54(24):9071–9078
Gomez-Galera S, Pelacho AM, Gene A, Capell T, Christou P (2007) The genetic manipulation of medicinal and aromatic plants. Plant Cell Rep 26(10):1689–1715
Guo S, Kemphues KJ (1995) par-1, a gene required for establishing polarity in C. elegans embryos, encodes a putative Ser/Thr kinase that is asymmetrically distributed. Cell 81(4):611–620
Hamilton AJ, Baulcombe DC (1999) A species of small antisense RNA in posttranscriptional gene silencing in plants. Science 286(5441):950–952
Han JY, Kwon YS, Yang DC, Jung YR, Choi YE (2006) Expression and RNA interference-induced silencing of the dammarenediol synthase gene in Panax ginseng. Plant Cell Physiol 47(12):1653–1662
Han JY, In JG, Kwon YS, Choi YE (2010) Regulation of ginsenoside and phytosterol biosynthesis by RNA interferences of squalene epoxidase gene in Panax ginseng. Phytochem 71(1):36–46
Han JY, Kim MJ, Ban YW, Hwang HS, Choi YE (2013) The involvement of β-amyrin 28-oxidase (CYP716A52v2) in oleanane-type ginsenoside biosynthesis in Panax ginseng. Plant Cell Physiol 54(12):2034–2046
Han J, Liu H-tao, Wang S-chang, Wang C-run, Miao G (2020) A Class I TGA transcription factor from Tripterygium wilfordii Hook.f. modulates the biosynthesis of secondary metabolites in both native and heterologous hosts. Plant Sci. https://doi.org/10.1016/j.plantsci.2019.110293
Hao X, Zhong Y, Nutzmann H-W, Fu X, Yan T, Shen Q, Chen M, Ma Y, Zhao J, Osbourn A et al (2019) Light-induced artemisinin biosynthesis is regulated by the bZIP transcription factor AaHY5 in Artemisia annua. Plant Cell Physiol 60(8):1747–1760
Harding SA, Frost CJ, Tsai C-J (2020) Defoliation‐induced compensatory transpiration is compromised in SUT4‐RNAi Populus. bioRxiv. https://doi.org/10.1101/2020.01.13.905406
Hoffmann L, Besseau S, Geoffroy P, Ritzenthaler C, Meyer D, Lapierre C, Pollet B, Legrand M (2004) Silencing of hydroxycinnamoyl-coenzyme A shikimate/quinate hydroxycinnamoyltransferase affects phenylpropanoid biosynthesis. Plant Cell 16(6):1446–1465
Hu Q, Xiao S, Guan Q, Tu L, Sheng F, Du X, Zhang X (2020) The laccase gene GhLac1 modulates fiber initiation and elongation by coordinating jasmonic acid and flavonoid metabolism. Crop J. https://doi.org/10.1016/j.cj.2019.11.006
Jiang Y, Hu Y, Wang B, Wu T (2014) Bivalent RNA interference to increase isoflavone biosynthesis in soybean (Glycine max). Braz Arch Biol Technol 57(2):163–170
Jiang J, Liao X, Jin X, Tan L, Lu Q, Yuan C, Xue Y, Yin N, Lin N, Chai Y (2020) MYB43 in oilseed rape (Brassica napus) positively regulates vascular lignification, plant morphology and yield potential but negatively affects resistance to Sclerotinia sclerotiorum. Genes 11:581
Jørgensen K, Bak S, Busk PK, Sørensen C, Olsen CE, Puonti-Kaerlas J, Møller BL (2005) Cassava plants with a depleted cyanogenic glucoside content in leaves and tubers. Distribution of cyanogenic glucosides, their site of synthesis and transport, and blockage of the biosynthesis by RNA interference technology. Plant Physiol 139(1):363–374
Joshi AG, Pathak AR (2019) EST (Expressed Sequence Tag): A technique for identification of plant secondary metabolite genes. In: Ozturk M, Hakeem KR (eds) Plant and human health, vol 2. phytochemistry and molecular aspects. Springer, Sawitzerland, pp 207–225
Kaminaga Y, Schnepp J, Peel G, Kish CM, Ben-Nissan G, Weiss D, Orlova I, Lavie O, Rhodes D, Wood K, Porterfield DM, Cooper AJ, Schloss JV, Pichersky E, Vainstein A, Dudareva N (2006) Plant phenylacetaldehyde synthase is a bifunctional homotetrameric enzyme that catalyzed phenylalanine decarboxylation and oxidation. J Biol Chem 281(33):23357–23366
Katsumoto Y, Fukuchi-Mizutani M, Fukui Y, Brugliera F, Holton TA, Karan M, Nakamura N, Yonekura-Sakakibara K, Togami J, Pigeaire A, Tao GQ, Nehra NS, Lu CY, Dyson BK, Tsuda S, Ashikari T, Kusumi T, Mason JG, Tanaka Y (2007) Engineering of the rose flavonoid biosynthetic pathway successfully generated blue-hued flowers accumulating delphinidin. Plant Cell Physiol 48(11):1589–1600
Kempe K, Higashi Y, Frick S, Sabarna K, Kutchan TM (2009) RNAi suppression of the morphine biosynthetic gene salAT and evidence of association of pathway enzymes. Phytochem 70(5):579–589
Kim HJ, Ono E, Morimoto K, Yamagaki T, Okazawa A, Kobayashi A, Satake H (2009) Metabolic engineering of lignan biosynthesis in Forsythia cell culture. Plant Cell Physiol 50(12):2200–2209
Koffas M, Roberge C, Lee K, Stephanopoulos G (1999) Metabolic engineering. Annu Rev Biomed Eng 1(1):535–557
Kooke R, Keurentjes JJ (2012) Multi-dimensional regulation of metabolic networks shaping plant development and performance. J Exp Bot 63(9):3353–3365
Kruse LH, Stegemann T, Jensen-Kroll J, Engelhardt A, Wesseling AM, Lippert A, Ludwig-Müller J, Ober D (2019) Reduction of pyrrolizidine alkaloid levels in comfrey (Symphytum officinale) hairy roots by RNAi silencing of homospermidine synthase. Planta Med. https://doi.org/10.1055/a-0998-5125
Kumar J, Gupta P (2008) Molecular approaches for improvement of medicinal and aromatic plants. Plant Biotechnol Rep 2(2):93–112
Kumar R, Vashisth D, Misra A, Akhtar MQ, Jalil SU, Shanker K, Gupta MM, Rout PK, Gupta AK, Shasany AK (2016) RNAi down-regulation of cinnamate-4-hydroxylase increases artemisinin biosynthesis in Artemisia annua. Sci Rep 6:26458
Kusaba M (2004) RNA interference in crop plants. Curr Opin Biotechnol 15(2):139–143
Kutchan T, Dixon RA (2005) Secondary metabolism: nature, chemical reservoir under deconvolution. Curr Opin Plant Biol 8(3):227–229
Larkin PJ, Miller JAC, Allen RS, Chitty JA, Gerlach WL, Kutchan SFTM, Fist AJ (2007) Increasing morphinan alkaloid production by over-expressing codeinone reductase in transgenic Papaver somniferum. Plant Biotechnol J 5(1):26–37
Lau W, Fischbach MA, Osbourn A, Sattely ES (2014) Key applications of plant metabolic engineering. PLoS Biol 12(6):e1001879
Lessard PA, Kulaveerasingam H, York GM, Strong A, Sinskey AJ (2002) Manipulating gene expression for the metabolic engineering of plants. Metab Eng 4(1):67–79
Li S, Wu Y, Kuang J, Wang H, Du T, Huang Y, Zhang Y, Cao X, Wang Z (2018) SmMYB111 is a key factor to phenolic acid biosynthesis and interacts with both SmTTG1 and SmbHLH51 in Salvia miltiorrhiza. J Agric Food Chem 66(30):8069–8078
Liu Q, Singh SP, Green AG (2002) High-stearic and high-oleic cottonseed oils produced by hairpin RNA-mediated post-transcriptional gene silencing. Plant Physiol 129(4):1732–1743
Liu Y, Zhao Z, Xue Z, Wang L, Cai Y, Wang P, Wei T, Gong J, Liu Z, Li JJ, Li S, Xiang F (2016) An intronless β-amyrin synthase gene is more efficient in oleanolic acid accumulation than its paralog in Gentiana straminea. Sci Rep 6:33364
Liu Q, Liu Y, Xu Y, Yao L, Liu Z, Cheng H, Ma M, Wud J, Wang W, Ninga W (2018) Overexpression of and RNA interference with hydroxycinnamoyl-CoA quinate hydroxycinnamoyl transferase affect the chlorogenic acid metabolic pathway and enhance salt tolerance in Taraxacum antungense Kitag. Phytochem Lett 28:116–123
Liu Q, Yao L, Xu Y, Cheng H, Wang W, Liu Z, Liu J, Cui X, Zhou Y, Ning W (2019) In vitro evaluation of hydroxycinnamoyl CoA:quinate hydroxycinnamoyl transferase expression and regulation in Taraxacum antungense in relation to 5-caffeoylquinic acid production. Phytochem 162:148–156
Liu L, Yang D, Xing B, Zhang C, Liang Z (2020) SmMYB98b positive regulation to tanshinones in Salvia miltiorrhiza Bunge hairy roots. Plant Cell Tiss Organ Cult 140:459–467
Lu C, Zhao SJ, Wang XS (2017a) Functional regulation of a UDP-glucosyltransferase gene (Pq3-O-UGT1) by RNA interference and overexpression in Panax quinquefolius. Plant Cell Tiss Organ Cult 129(3):445–456
Lu C, Zhao SJ, Wei G, Zhao H, Qu Q (2017b) Functional regulation of ginsenoside biosynthesis by RNA interferences of a UDP-glycosyltransferase gene in Panax ginseng and Panax quinquefolius. Plant Physiol Biochem 111:67–76
Ma Y, Ma XH, Meng FY, Zhan ZL, Guo J, Huang LQ (2016) RNA interference targeting CYP76AH1 in hairy roots of Salvia miltiorrhiza reveals its key role in the biosynthetic pathway of tanshinones. Biochem Biophys Res Commun 477(2):155–160
Ma K, Jiang Y, Yu ZY, Huang YT, Zhan YG, Fan GZ (2019) H2S-induced NO/SNO positively promotes betulin production in Betula platyphylla. Ind Crops Prod. https://doi.org/10.1016/j.indcrop.2019.111608
Mahmoud SS, Croteau RB (2001) Metabolic engineering of essential oil yield and composition in mint by altering expression of deoxyxylulose phosphate reductoisomerase and menthofuran synthase. Proc Natl Acad Sci USA 98(15):8915–8920
Mahmoud SS, Williams M, Croteau R (2004) Cosuppression of limonene-3-hydroxylase in peppermint promotes accumulation of limonene in the essential oil. Phytochem 65(5):547–554
Maliga P (2001) Plastid engineering bears fruit. Nat Biotechnol 19(9):826–827
Mansoor S, Amin I, Hussain M, Zafar Y, Briddon RW (2006) Engineering novel traits in plants through RNA interference. Trends Plant Science 11(11):559–565
Martinez DH, Payyavula RS, Kudithipudi C, Shen Y, Xu D, Warek U, Strickland JA, Melis A (2020) Genetic attenuation of alkaloids and nicotine content in tobacco (Nicotiana tabacum). Planta 251:92
Mishra S, Bansal S, Mishra B, Sangwan RS, Asha Jadaun JS, Sangwan NS (2016) RNAi and homologous over-expression based functional approaches reveal triterpenoid synthase gene- cycloartenol synthase is involved in downstream withanolide biosynthesis in Withania somnifera. PLoS ONE 11(2):e0149691
Nagegowda DA (2010) Plant volatile terpenoid metabolism: Biosynthetic genes, transcriptional regulation and subcellular compartmentation. FEBS Lett 584(14):2965–2973
Nagira Y, Shimamura K, Hirai S, Shimanuki M, Kodama H, Ozeki Y (2006) Identification and characterization of genes induced for anthocyanin synthesis and chlorophyll degradation in regenerated torenia shoots using suppression subtractive hybridization, cDNA microarrays, and RNAi techniques. J Plant Res 119(3):217–230
Nakamura N, Fukuchi-Mizutani M, Miyazaki K, Suzuki K, Tanaka Y (2006) RNAi suppression of the anthocyanidin synthase gene in Torenia hybrida yields white flowers with higher frequency and better stability than antisense and sense suppression. Plant Biotechnol 23(1):13–17
Nakatsuka T, Abe Y, Kakizaki Y, Yamamura S, Nishihara M (2007) Production of red-flowered plants by genetic engineering of multiple flavonoid biosynthetic genes. Plant Cell Rep 26(11):1951–1959
Napoli C, Lemieux C, Jorgensen R (1990) Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 2:279–289
Niephaus E, Muller B, van Deenen N, Lassowskat I, Bonin M, Finkemeier I, Prufer D, Gronover CS (2019) Uncovering mechanisms of rubber biosynthesis in Taraxacum koksaghyz—role of cis-prenyltransferase-like 1 protein. Plant J 100:591–609
Nishihara M, Nakatsuka T, Yamamura S (2005) Flavonoid components and flower color change in transgenic tobacco plants by suppression of chalcone isomerase gene. FEBS Lett 579(27):6074–6078
Ogita S, Uefuji H, Yamaguchi Y, Koizumi N, Sano H (2003) Producing decaffeinated coffee plants. Nature 423(6942):823
Ogita S, Uefuji H, Morimoto M, Sano H (2004) Application of RNAi to confirm theobromine as the major intermediate for caffeine biosynthesis of coffee plants with potential for construction of decaffeinated varieties. Plant Mol Biol 54(6):931–941
Ono E, Fukuchi-Mizutani M, Nakamura N, Fukui Y, Yonekura-Sakakibara K, Yamaguchi M, Nakayama T, Tanaka T, Kusumi T, Tanaka Y (2006) Yellow flowers generated by expression of the aurone biosynthetic pathway. Proc Natl Acad Sci USA 103(29):11075–11080
Orlova I, Marshall-Colón A, Schnepp J, Wood B, Varbanova M, Fridman E, Blakeslee JJ, Peer WA, Murphy AS, Rhodes D, Pichersky E, Dudareva N (2006) Reduction of benzenoid synthesis in petunia flowers reveals multiple pathways to benzoic acid and enhancement in auxin transport. Plant Cell 18(12):3458–4375
Pal S, Rastogi S, Nagegowda DA, Gupta MM, Shasany AK, Chanotiya CS (2019) RNAi of sterol methyl transferase1 reveals its direct role in diverting intermediates towards withanolide/phytosterol biosynthesis in Withania somnifera. Plant Cell Physiol 60(3):672–686
Palatnik JF, Allen E, Wu X, Schommer C, Schwab R, Carrington JC, Weigel D (2003) Control of leaf morphogenesis by microRNAs. Nature 425(6955):257–263
Park SU, Yu M, Facchini PJ (2002) Antisense RNA-mediated suppression of benzophenanthridine alkaloid biosynthesis in transgenic cell cultures of California poppy. Plant Physiol 128(2):696–706
Park S, Yu M, Facchini PJ (2003) Modulation of berberine bridge enzyme levels in transgenic root cultures of California poppy alters the accumulation of benzophenanthridine alkaloids. Plant Mol Biol 51(2):153–164
Park NI, Xu H, Li X, Kim SJ, Park SU (2011) Enhancement of flavone levels through overexpression of chalcone isomerase in hairy root cultures of Scutellaria baicalensis. Funct Integr Genomics 11(3):491–496
Park SB, Chun JH, Ban YW, Han JY, Choi YE (2016) Alteration of Panax ginseng saponin composition by overexpression and RNA interference of the protopanaxadiol 6-hydroxylase gene (CYP716A53v2). J Ginseng Res 40(1):47–54
Pichersky E, Gang DR (2000) Genetics and biochemistry of secondary metabolites in plants: an evolutionary perspective. Trends Plant Sci 5(10):439–445
Pickens LB, Tang Y, Chooi YH (2011) Metabolic engineering for the production of natural products. Annu Rev Chem Biomol Eng 2(1):211–236
Qiu F, Zeng J, Wang J, Huang J-P, Zhou W, Yang C, Lan X, Chen M, Huang S-X, Kai G et al (2020) Functional genomics analysis reveals two novel genes required for littorine biosynthesis. New Phytol 225:1906–1914
Rao SR, Ravishankar GA (2002) Plant cell cultures: chemical factories of secondary metabolites. Biotechnol Adv 20(2):101–153
Rates SMK (2001) Plants as source of drugs. Toxicon 39(5):603–613
Reddy VA, Wang Q, Dhar N, Kumar N, Venkatesh PN, Rajan C, Panicker D, Sridhar V, Mao HZ, Sarojam R (2017) Spearmint R2R3-MYB transcription factor MsMYB negatively regulates monoterpene production and suppresses the expression of geranyl diphosphate synthase large subunit (MsGPPS.LSU). Plant Biotechnol J 15(9):1105–1119
Saema S, ur Rahman L, Niranjan A, Ahmad IZ, Misra P (2015) RNAi-mediated gene silencing of WsSGTL1 in W. somnifera affects growth and glycosylation pattern. Plant Signal Behav 10(12):e1078064
Seitz C, Vitten M, Steinbach P, Hartl S, Hirsche J, Rathje W, Treutter D, Forkmann G (2007) Redirection of anthocyanin synthesis on Osteospermum hybrida by a two-enzyme manipulation strategy. Phytochem 68(6):824–833
Shen Q, Lu X, Yan T, Fu X, Lv Z, Zhang F, Pan Q, Wang G, Sun X, Tang K (2016) The jasmonate-responsive AaMYC2 transcription factor positively regulates artemisinin biosynthesis in Artemisia annua. New Phytol 210(4):1269–1281
Singh S, Pal S, Shanker K, Chanotiya CS, Gupta MM, Dwivedi UN, Shasany AK (2014) Sterol partitioning by HMGR and DXR for routing intermediates toward withanolide biosynthesis. Physiol Plant 152(4):617–633
Skirycz A, Jozefczuk S, Stobiecki M, Muth D, Zanor MI, Witt I, Mueller-Roeber B (2007) Transcription factor AtDOF4;2 affects phenylpropanoid metabolism in Arabidopsis thaliana. New Phytol 175(3):425–438
Small I (2007) RNAi for revealing and engineering plant gene functions. Curr Opin Biotechnol 18(2):148–153
Smith NA, Singh SP, Wang MB, Stoutjesdijk PA, Green AG, Waterhouse PM (2000) Total silencing by intron-spliced hairpin RNAs. Nature 407(6802):319–320
Somerville C, Somerville S (1999) Plant functional genomics. Science 285(5426):380–383
Song J, Wang Z (2011) RNAi-mediated suppression of the phenylalanine ammonia-lyase gene in Salvia miltiorrhiza causes abnormal phenotypes and a reduction in rosmarinic acid biosynthesis. J Plant Res 124(1):183–192
Stanley LE, Ding B, Sun W, Mou F, Hill C, Chen S, Yuan Y-W (2020) A tetratricopeptide repeat protein regulates carotenoid biosynthesis and chromoplast development in Monkeyflowers (Mimulus). Plant Cell 32:1536–1555
Stephanopoulos G (1999) Metabolic fluxes and metabolic engineering. Metab Eng 1(1):1–11
Su P, Guan H, Zhao Y, Tong Y, Xu M, Zhang Y, Hu T, Yang J, Cheng Q, Gao L, Liu Y, Zhou J, Peters RJ, Huang L, Gao W (2018) Identification and functional characterization of diterpene synthases for triptolide biosynthesis from Tripterygium wilfordii. Plant J 93(1):50–65
Subramanian S, Graham MY, Yu O, Graham TL (2005) RNA interference of soybean isoflavone synthase genes leads to silencing in tissues distal to the transformation site and to enhanced susceptibility to Phytophthora sojae. Plant Physiol 137(4):1345–1353
Sujeeth N, Mehterov N, Gupta S, Qureshi MK, Fischer A, Proost S, Omidbakhshfard MA, Obata T, Benina M, Staykov N et al (2020) A novel seed plants gene regulates oxidative stress tolerance in Arabidopsis thaliana. Cell Mol Life Sci 77:705–718
Sun Y, Zhao SJ, Liang YL, Le W, Cao HJ (2013) Regulation and differential expression of protopanaxadiol synthase in Asian and American ginseng ginsenoside biosynthesis by RNA interferences. Plant Growth Regul 71(3):207–217
Tague BW (2001) Germ-line transformation of Arabidopsis lasiocarpa. Transgenic Res 10(3):259–267
Takagi K, Nishizawa K, Hirose A, Kita A, Ishimoto M (2011) Manipulation of saponin biosynthesis by RNA interference-mediated silencing of β-amyrin synthase gene expression in soybean. Plant Cell Rep 30(10):1835–1846
Tang G, Galili G (2004) Using RNAi to improve plant nutritional value: from mechanism to application. Trends Biotechnol 22(9):463–469
Todd AT, Liu E, Polvi SL, Pammett RT, Page JE (2010) A functional genomics screen identifies diverse transcription factors that regulate alkaloid biosynthesis in Nicotiana benthamiana. Plant J 62(4):589–600
Tsuro M, Tomomatsu K, Inukai C, Tujii S, Asada S (2019) RNAi targeting the gene for 1,8-cineole synthase induces recomposition of leaf essential oil in lavandin (Lavandula × intermedia Emeric). Vitro Cell Dev Biol-Plant 55:165–171
Underwood BA, Tieman DM, Shibuya K, Dexter RJ, Loucas HM, Simkin AJ, Sims CA, Schmelz EA, Klee HJ, Clark DG (2005) Ethylene-regulated floral volatile synthesis in petunia corollas. Plant Physiol 138:255–266
Vemuri GN, Aristidou AA (2005) Metabolic engineering in the -omics era: elucidating and modulating regulatory networks. Microbiol Mol Biol Rev 69(2):197–216
Verdonk JC, Haring MA, van Tunen AJ, Schuurink RC (2005) ODORANT1 regulates fragrance biosynthesis in petunia flowers. Plant Cell 17(5):1612–1624
Verpoorte R, Contin A, Memelink J (2002) Biotechnology for the production of plant secondary metabolites. Phytochem 1(1):13–25
Wagner A, Ralph J, Akiyama T, Flint H, Phillips L, Torr K, Nanayakkara B, Te Kiri L (2007) Exploring lignification in conifers by silencing hydroxycinnamoyl-CoA: shikimate hydroxycinnamoyltransferase in Pinus radiate. Proc Natl Acad Sci USA 104(28):11856–11861
Wang E, Wagner GJ (2003) Elucidation of the functions of genes central to diterpene metabolism in tobacco trichomes using PTGS. Planta 216(4):686–691
Wang Z, Cui L, Chen C, Liu X, Yan Y, Wang Z (2012) Downregulation of cinnamoyl CoA reductase affects lignin and phenolic acids biosynthesis in Salvia miltiorrhiza Bunge. Plant Mol Biol Rep 30(4):1229–1236
Wang L, Zhao SJ, Liang YL, Sun Y, Cao HJ, Han Y (2014) Identification of the protopanaxatriol synthase gene CYP6H for ginsenoside biosynthesis in Panax quinquefolius. Funct Integr Genomics 14(3):559–570
Wang Q, Reddy VA, Panicker D, Mao HZ, Kumar N, Rajan C, Venkatesh PN, Chua NH, Sarojam R (2016) Metabolic engineering of terpene biosynthesis in plants using a trichome-specific transcription factor MsYABBY5 from spearmint (Mentha spicata). Plant Biotechnol J 14(7):1619–1632
Wang Z, Ge Q, Chen C, Jin X, Cao X, Wang Z (2017) Function analysis of caffeoyl-CoA O-methyltransferase for biosynthesis of lignin and phenolic acid in Salvia miltiorrhiza. Appl Biochem Biotechnol 181(2):562–572
Wang CH, Lei XY, Xia J, Wang JW (2018) Effect of down-regulating 1-deoxy-d-xylulose-5-phosphate reductoisomerase by RNAi on growth and artemisinin biosynthesis in Artemisia annua L. Plant Growth Regul 84(3):549–559
Wang Z, Wang S, Xiao Y, Li Z, Wu M, Xie X, Li H, Mu W, Li F, Liu P et al (2020) Functional characterization of a HD-ZIP IV transcription factor NtHDG2 in regulating flavonols biosynthesis in Nicotiana tabacum. Plant Physiol Biochem 146:259–268
Warthmann N, Chen H, Ossowski S, Weigel D, Herve P (2008) Highly specific gene silencing by artificial miRNAs in rice. PLoS ONE 3(3):e1829
Waterhouse PM, Graham MW, Wang MB (1998) Virus resistance and gene silencing in plants can be induced by simultaneous expression of sense and antisense RNA. Proc Natl Acad Sci USA 95(23):13959–13964
Waterhouse PM, Ming-Bo W, Lough T (2001) Gene silencing as an adaptive defence against viruses. Nature 411(6839):834–842
Wesley SV, Helliwell CA, Smith NA, Wang MB, Rouse DT, Liu Q, Gooding PS, Singh SP, Abbott D, Stoutjesdijk PA, Robinson SP, Gleave AP, Green AG, Waterhouse PM (2001) Constructs for efficient, effective and high throughput gene silencing in plants. Plant J 27(6):581–590
Wesley SV, Liu Q, Wielopolska A, Ellacott G, Smith N, Singh S, Helliwell C (2003) Custom knock-outs with hairpin RNA-mediated gene silencing. Methods Mol Biol 236:273–286
Wróbel-Kwiatkowska M, Starzycki M, Zebrowski J, Oszmiański J, Szopa J (2007) Lignin deficiency in transgenic flax resulted in plants with improved mechanical properties. J Biotechnol 128(4):919–934
Wu SQ, Chappell J (2008) Metabolic engineering of natural products in plants; tools of the trade and challenges for the future. Curr Opin Biotechnol 19(2):145–152
Wu Y, Zhang Y, Li L, Guo X, Wang B, Cao X, Wang Z (2018) AtPAP1 interacts with and activates SmbHLH51, a positive regulator to phenolic acids biosynthesis in Salvia miltiorrhiza. Front Plant Sci 9:1687
Wu Z, Wang N, Hisano H, Cao Y, Wu F, Liu W, Bao Y, Wang Z-Y, Fu C (2019) Simultaneous regulation of F5H in COMT-RNAi transgenic switchgrass alters effects of COMT suppression on syringyl lignin biosynthesis. Plant Biotechnol J 17:836–845
Xiao Y, Feng J, Li Q, Zhou Y, Bu Q, Zhou J, Tan H, Yang Y, Zhang L, Chen W (2020) IiWRKY34 positively regulates yield, lignin biosynthesis and stress tolerance in Isatis indigotica. Acta Pharma Sin B. https://doi.org/10.1016/j.apsb.2019.12.020
Yang Y, Ge F, Sun Y, Liu D, Chen C (2017) Strengthening triterpene saponins biosynthesis by over-expression of farnesyl pyrophosphate synthase gene and RNA interference of cycloartenol synthase gene in Panax notoginseng cells. Molecules 22(4):E581
Yang Y, Zhang Z, Li R, Yi Y, Yang H, Wang C, Wang Z, Liu Y (2020) RgC3H involves in the biosynthesis of allelopathic phenolic acids and alters their release amount in Rehmannia glutinosa roots. Plants 9:567
Yin J, Yang J, Ma H, Liang T, Li Y, Xiao J, Tian H, Xu Z, Zhan Y (2020) Expression characteristics and function of CAS and a new beta-amyrin synthase in triterpenoid synthesis in birch (Betula platyphylla Suk.). Plant Sci. https://doi.org/10.1016/j.plantsci.2020.110433
Zhang S, Ma P, Yang D, Li W, Liang Z, Liu Y, Liu F (2013) Cloning and characterization of a putative R2R3 MYB transcriptional repressor of the rosmarinic acid biosynthetic pathway from Salvia miltiorrhiza. PLoS ONE 8(9):e73259
Zhang Y, Yan YP, Wu YC, Hua WP, Chen C, Ge Q, Wang ZZ (2014) Pathway engineering for phenolic acid accumulations in Salvia miltiorrhiza by combinational genetic manipulation. Metab Eng 21:71–80
Zhang S, Li H, Liang X, Yan Y, Xia P, Jia Y, Liang Z (2015) Enhanced production of phenolic acids in Salvia miltiorrhiza hairy root cultures by combing the RNAi-mediated silencing of chalcone synthase gene with salicylic acid treatment. Biochem Eng J 103:185–192
Zhang Y, Zhao Y, Wang J, Hu T, Tong Y, Zhou J, Song Y, Gao W, Huang L (2018) Overexpression and RNA interference of TwDXR regulate the accumulation of terpenoid active ingredients in Tripterygium wilfordii. Biotechnol Lett 40(2):419–425
Zhang B, Huo Y, Zhang J, Zhang X, Zhu C (2019a) Agrobacterium rhizogenes-mediated RNAi of Tripterygium wilfordii and application for functional study of terpenoid biosynthesis pathway genes. Ind Crop Prod. https://doi.org/10.1016/j.indcrop.2019.111509
Zhang Y, Ji A, Xu Z, Luo H, Song J (2019b) The AP2/ERF transcription factor SmERF128 positively regulates diterpenoid biosynthesis in Salvia miltiorrhiza. Plant Mol Biol 100(1–2):83–93
Zhang J-H, Lv H-Z, Liu W-J, Ji A-J, Zhang X, Song J-Y, Luo H-M, Chen S-L (2020) bHLH transcription factor SmbHLH92 negatively regulates biosynthesis of phenolic acids and tanshinones in Salvia miltiorrhiza. Chin Herb Med. https://doi.org/10.1016/j.chmed.2020.04.001
Zhao C, Xu T, Liang Y, Zhao S, Ren L, Wang Q, Dou B (2015) Functional analysis of β-amyrin synthase gene in ginsenoside biosynthesis by RNA interference. Plant Cell Rep 34(8):1307–1315
Zhao Q, Zhang Y, Wang G, Hill L, Weng JK, Chen XY, Xue H, Martin C (2016) A specialized flavone biosynthetic pathway has evolved in the medicinal plant, Scutellaria baicalensis. Sci Adv 2(4):e1501780
Zhao Q, Cui MY, Levsh O, Yang D, Liu J, Li J, Hill L, Yang L, Hu Y, Weng JK, Chen XY, Martin C (2018) Two CYP82D enzymes function as flavone hydroxylases in the biosynthesis of root-specific 40-deoxyflavones in Scutellaria baicalensis. Mol Plant 11(1):135–148
Zhao Q, Yang J, Cui MY, Liu J, Fang Y, Yan M, Qiu W, Shang H, Xu Z, Yidiresi R, Weng JK, Pluskal T, Vigouroux M, Steuernagel B, Wei Y, Yang L, Hu Y, Chen XY, Martin C (2019) The reference genome sequence of Scutellaria baicalensis provides insights into the evolution of wogonin biosynthesis. Mol Plant 12(7):935–950
Zhao T, Li S, Wang J, Zhou Q, Yang C, Bai F, Lan X, Chen M, Liao Z (2020) Engineering tropane alkaloid production based on metabolic characterization of ornithine decarboxylase in Atropa belladonna. ACS Synth Biol 9(2):437–448
Zhu Y, Chu SJ, Luo YL, Fu JY, Tang CY, Lu GH, Pang YJ, Wang XM, Yang RW, Qi JL, Yang YH (2018) Involvement of LeMRP, an ATP-binding cassette transporter, in shikonin transport and biosynthesis in Lithospermum erythrorhizon. Plant Biol 20(2):365–373
Zupan J, Muth TR, Draper O, Zambryski P (2000) The transfer of DNA from Agrobacterium tumefaciens into plants: A feast of fundamental insights. Plant J 23(1):11–28
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Pathak, A.R., Patel, S.R., Joshi, A.G. (2021). RNA Interference (RNAi): A Genetic Tool to Manipulate Plant Secondary Metabolite Pathways. In: Tang, G., Teotia, S., Tang, X., Singh, D. (eds) RNA-Based Technologies for Functional Genomics in Plants. Concepts and Strategies in Plant Sciences. Springer, Cham. https://doi.org/10.1007/978-3-030-64994-4_9
Download citation
DOI: https://doi.org/10.1007/978-3-030-64994-4_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-64993-7
Online ISBN: 978-3-030-64994-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)