Abstract
For cell growth and proliferation, synthesis of cellular components such as DNA, proteins, and lipids is a prerequisite. Cells have to rewire and reprogram their metabolism to allow the production of required cellular materials. Oncogenic signaling pathways such as those driven by KRAS mutations in pancreatic cancer induce metabolic reprogramming. Metabolic dependency can on the other hand be a critical vulnerability so that identifying tumor dependencies on metabolism would provide options for novel targeting strategies. This chapter summarizes the research evidence on metabolic reprogramming of cancer focusing on glucose, glutamine, acetate, and lipid metabolism. Further, studies with inhibitors and drugs targeting cancer metabolism are discussed. Understanding reprogramming strategies of cancer, as well as visualization of metabolism reprogramming networks, could uncover novel therapeutic options.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Metabolic reprogramming
- Glucose metabolism
- Pentose phosphate pathway
- Glutamine metabolism
- Acetate metabolism
- Lipid metabolism
- Pancreatic ductal adenocarcinoma
Understanding Metabolic Reprogramming to Improve Therapeutic Strategies in Pancreatic Cancer
Pancreatic ductal adenocarcinoma (PDAC) is a devastating disease with an unfavorable outcome and is projected to become the second deadliest cancer by 2030, and currently the overall 5-year survival rate is less than 7% [1, 2]. PDAC arises through multistage genetic and histological progression from precursors such as pancreatic intraepithelial neoplasia (PanIN). Activating mutations in the KRAS oncogene are observed in over 90% of PDAC patients, and using genetically engineered mouse models, it has been demonstrated that KRAS mutations influence tumor initiation, progression, and maintenance [3, 4]. Tumorigenesis is dependent on the reprogramming of cellular metabolism which can be a consequence of oncogenic mutations. In line with this, a profound rewiring of metabolic pathways involved in, e.g., glucose, glutamine, and lipid metabolisms, is activated downstream of oncogenic KRAS [5]. In general, metabolic reprogramming has now been recognized as a hallmark of cancer [6]. Cancer cells manipulate metabolisms to keep generating their own cellular components such as DNA, proteins, and lipids for maintaining rapid cell growth. Understanding and identification of metabolic reprogramming strategies of individual cancers could uncover novel potential personalized targets. This chapter provides a background of cancer metabolism focusing on glucose, glutamine, acetate, and lipid metabolism and targeting strategies for modulating enzymes/factors involved in key metabolic pathways.
Glucose Metabolism and Pentose Phosphate Pathway in Pancreatic Cancer
Warburg Effect and Reprogramming of Glucose Metabolism: The Role of Gene Mutations
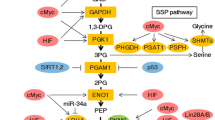
A pioneer of the study of cancer metabolic reprogramming, Otto Heinrich Warburg made a striking discovery known as Warburg effect that many cancer cells preferentially convert glucose into lactate (fermentation) rather than respiration – transporting pyruvate into mitochondria and converted it into acetyl-CoA for subsequent ATP production via the citric acid cycle and electron transport chain – even in the presence of oxygen [7,8,9,10]. Glycolysis, a metabolic pathway that converts glucose into pyruvate (and lactate) in the cytoplasm, is a sequence of ten enzyme-catalyzed reactions (Fig. 13.1). The three reactions converting glucose into glucose 6-phosphate by hexokinase (HK), fructose 6-phosphate into fructose 1,6-bisphosphate by phosphofructokinase (PFK), and phosphoenolpyruvate into pyruvate by pyruvate kinase are key steps. Oncogenic KRASG12D plays a role in upregulating gene expression of the glucose transporter (GLUT) Slc2a1 (SLC: solute carrier), Hk1, Hk2, and Pfk1 as well as Ldha coding lactate dehydrogenase (LDH) , an enzyme for converting pyruvate to lactate. Concomitantly oncogenic KRAS enhances glucose uptake and lactate production in a pancreatic cancer mouse model [11]. The transcription factor p53 is recognized as a key tumor suppressor and also frequently mutated in human tumors. Missense mutations such as R175H, R248Q, and R273H not only result in loss of the tumor suppressive function of p53 but also in oncogenic functions that promote invasion, metastasis, proliferation, and cell survival [12]. Mutation of p53 also enhances glucose uptake by GLUT1 translocation, glycolytic rate, and lactate production in R172H mutant-expressing p53 in murine cancer cells or fibroblasts (R172H is equivalent to human R175H) [13]. Deficiency of another tumor suppressor gene, SMAD4, increases GLUT1 levels and lactate production in cancer cells [14]. KRAS, TP53, CDKN2A, and SMAD4 are the most prevalent genetic mutations in pancreatic cancer [1]; yet these genes are currently not druggable . However, targeting glucose metabolic reprogramming may provide a selective mechanism for eliminating cancer cells.
Regulation of glycolysis, pentose phosphate pathway, and nucleotide biosynthesis. AMPK AMP-activated protein kinase, G6PD glucose 6-phosphate dehydrogenase, HIF hypoxia-inducible factor, HK hexokinase, LDH lactate dehydrogenase, mKRAS mutant KRAS, mp53 mutant p53, MPC mitochondria pyruvate carrier, PFK phosphofructokinase, PGD 6-phosphogluconate dehydrogenase PK pyruvate kinase, PPP pentose phosphate pathway, PRPS phosphoribosylpyrophosphate synthetase, RPE ribose 5-phosphate-3-epimerase, RPIA ribose 5-phosphate isomerase A, SLC solute carrier
Targeting Enzymes and Factors Involved in Glucose Metabolism
Inhibition of GLUT, especially GLUT1 expression, can be an option to halt the proliferation of cancers. A small-molecule GLUT1 inhibitor WZB117 has been shown to block glucose uptake and tumor growth in a tumor xenograft model [15, 16]. Furthermore, WZB117 administration inhibits tumor initiation after implantation of cancer stemlike cells derived from pancreatic cancer cells without causing adverse events in host mice [17]. Overexpression of GLUT1 correlates with poor overall survival of several solid tumors [18], and high GLUT1 expression is also suggested to predict shorter overall survival in patients with pancreatic cancer [19].
In the mammalian glycolytic pathway, PFK1 is rate-limiting and the most important control element. When PFK1 is inactive, the concentration of fructose 6-phosphate rises, and in equilibrium, the level of glucose 6-phosphate also rises. Hexokinase, another key enzyme in the glycolytic pathway, is allosterically inhibited by glucose 6-phosphate; therefore, PFK1 inhibition leads to the inhibition of hexokinase. Activity of PFK1 is stimulated by fructose 2,6-bisphosphate, which is derived from fructose 6-phosphate catalyzed by PFK2. There are four PFK2 isoforms (PFKFB1–4), and PFKFB3 is highly expressed in many types of human cancer including pancreatic cancer [20]. Expression of PFKFB3 can also be regulated by hypoxia [21]. PFKFB3 also regulates angiogenesis and vessel branching [22] and can be an emerging anticancer target. In this line, KAN0438757 has been considered as a selective PFKFB3 inhibitor, and treatment with this inhibitor radiosensitizes cancer cells [23]. Another PFKFB3 blocker 3-(3-pyridinyl)-1-(4-pyridinyl)-2-propen-1-one (3PO) reduces orthotopically implanted pancreatic cancer cell development [24], suggesting that targeting PFKFB3 can be an option for pancreatic cancer treatment.
In the late step in glycolysis, pyruvate kinase plays an important role as catalyzing the last physiological irreversible reaction to produce pyruvate. In mammals, there are four pyruvate kinase isoforms encoded by two genes: isoforms PKL and PKR are derived from the PKLR gene, and PKM1 and PKM2 are derived from the PKM gene through alternative splicing. The amino acid differences in PKM2 result in a fructose 1,6-bisphosphate-binding pocket for positive allosteric regulation [25]. Activation of PFK1 (for producing fructose 1,6-bisphosphate) can therefore not only regulate hexokinase activity but also PKM2 activity. PKM2 is expressed during embryogenesis, regeneration processes, and in cancer, suggesting that PKM2 activity is important in actively proliferating cells [25]. Orthotopically implanted cancer cells expressing PKM2 support tumorigenesis, whereas cells expressing PKM1 reduce tumorigenicity, suggesting that the PKM2 splice isoform is important for cancer metabolism and tumor growth [26]. On the contrary, in some studies activation of PKM2 can inhibit cancer cell proliferation [27, 28]. Furthermore, conditional deletion of PKM2 in a pancreatic cancer mouse model (oncogenic KRASG12D expression and p53 deletion) does not affect mouse survival, tumor weight, or tumor histology [29]. Therefore, targeting PKM2 might not be suitable for pancreatic cancer treatment and needs further investigation.
Pyruvate is a key metabolite in the network of metabolic pathways. Pyruvate in the cytoplasm can be converted into alanine by alanine aminotransferase (ALT) or transported into the mitochondria via mitochondria pyruvate carrier (MPC) and converted there into oxaloacetate by pyruvate carboxylase for gluconeogenesis or converted into acetyl-CoA by the pyruvate dehydrogenase (PDH) complex for the citric acid cycle. Pyruvate decarboxylase catalyzes a reaction converting pyruvate into acetaldehyde in the cytoplasm and mitochondria. Cancer cells however preferentially convert pyruvate into lactate, which is catalyzed by LDH. LDH is a tetramer of two subunits LDHA and LDHB, which assemble into five different combinations [30]. LDHA has a higher affinity for pyruvate than LDHB, and elevated levels of LDHA are a hallmark of many cancer types; hence targeting LDHA can be a promising strategy for cancer therapeutics. Consistently, FX11 (3-dihydroxy-6-methyl-7-(phenylmethyl)-4-propylnaphthalene-1-carboxylic acid), a small-molecule inhibitor of LDHA, inhibits the progression of pancreatic cancer xenografts [31]. Interestingly, the inhibitory effect of FX11 requires mutant p53, and FX11 treatment does not inhibit tumor progression of patient-derived PDAC xenografts without p53 mutation [32], suggesting that targeting LDHA in pancreatic cancer can be an attractive stratification option since drug responsiveness in PDAC patients may depend on the genetic status.
Pentose Phosphate Pathway: Helper of Cancer’s Anabolic Demands
The pentose phosphate pathway (PPP) is another pathway in the cytoplasm for glucose catabolism starting from glucose 6-phosphate. The major function of the PPP is not energy production, but generating extramitochondrial nicotinamide adenine dinucleotide phosphate (NADPH), which is required for fatty acid synthesis and for scavenging reactive oxygen species (ROS). The PPP also supports the synthesis of ribonucleotides. The PPP is divided into two parts, namely, the oxidative arm and non-oxidative arm. The oxidative arm is initiated by conversion of glucose 6-phosphate to 6-phosphogluconolactone by glucose 6-phosphate dehydrogenase (G6PD), which is converted into 6-phosphogluconate by gluconolactonase and further converted into ribulose 5-phosphate by 6-phosphogluconate dehydrogenase (PGD). In the non-oxidative phase of the PPP, ribulose 5-phosphate is either reversibly catalyzed by ribose 5-phosphate isomerase A (RPIA) for producing ribose 5- phosphate or reversibly catalyzed by ribose 5-phosphate-3-epimerase (RPE) for producing xylulose 5-phosphate [33]. Ribose 5-phosphate is converted by phosphoribosylpyrophosphate synthetase (PRPS) to phosphoribosyl pyrophosphate, which serves as the backbone for nucleotide synthesis. Oncogenic KRASG12D upregulates RPIA and RPE gene expression in murine primary cells of a pancreatic cancer model with oncogenic KRASG12D and p53 deficiency. Knockdown of Rpia or Rpe genes in primary cells reduces the flux of glucose into DNA/RNA synthesis and xenograft pancreatic tumor growth [11], and knockdown of Rpia gene inhibits human PDAC cell growth [34]. Ribose 5-phosphate and xylulose 5-phosphate in the non-oxidative branch of the PPP can also be reversibly catalyzed by transketolase and aldolase to fructose 6-phosphate or glyceraldehyde 3-phosphate, which can be utilized in the glycolysis [33]. Vice versa, fructose 6-phosphate and glyceraldehyde 3-phosphate in the glycolytic pathway can be incorporated into the PPP pathway, and many cancer cells generate ribose 5-phosphate through the non-oxidative branch of the PPP for de novo nucleotide biosynthesis [35]. Fructose induces transketolase flux and proliferation of pancreatic cancer cells [36]. High fructose intake has been suggested to be associated with increased pancreatic cancer risk [37]. A key regulator of the non-oxidative branch of the PPP is hypoxia-inducible factor (HIF)-1α which increases the carbon flux into the PPP [35], and HIF-1α directly regulates transketolase gene expression [38]. Taken together, the PPP especially the non-oxidative arm plays an important role in de novo nucleotide biosynthesis, and directly or indirectly targeting enzymes and factors involved in the PPP is a promising therapeutic strategy against pancreatic cancer.
Targeting Enzymes and Factors Involved in the Pentose Phosphate Pathway and Nucleotide Synthesis
Oncogenic KRASG12D reprograms metabolism of the PPP in PDAC through MAPK and Myc pathways [11, 34]. Myc has been further shown to control PRPS2, but not PRPS1, and functional loss of PRPS2 delays Myc-dependent tumor initiation [39]. Since KRAS and Myc are currently not druggable, targeting RPIA, RPE of the non-oxidative branch of the PPP, as well as targeting PRPS2 in the nucleotide biosynthesis pathway can be considered as therapeutic options. Inhibitors of RPIA, RPE, or PRPS remain largely undiscovered. Especially selective PRPS2 inhibitors are challenging to identify, since PRPS2 shares more than 97% amino acid identity with the PRPS1 [40]. So far, pharmacological inhibitors of effector pathways on cancer metabolism have been used. For example, treatment with the MEK inhibitor AZD8330 decreases Rpia gene expression in murine primary cells of a pancreatic cancer model with oncogenic KRASG12D and p53 deficiency [11]. AMP-activated protein kinase (AMPK) phosphorylation leads to conversion of PRPS hexamer to monomer resulting in inhibition of nucleotide synthesis in cancer cells (AMPK activator: A-769662) [41]. Digoxin is an HIF-1α synthesis inhibitor [42], and targeting HIF-1α leads to reduction of transketolase gene expression and improved gemcitabine sensitivity in pancreatic cancer cells [38]. MEK/MAPK, AMPK, and HIF-1α regulate not only the PPP and/or nucleotide biosynthesis. However, reprogramming the reprogrammed metabolism of the PPP and nucleotide biosynthesis in cancer by modulating effectors is also a promising targeting strategy.
Lipid Metabolism in Pancreatic Cancer
Fatty Acid Synthesis as an Entrance of Lipid Metabolism and Critical for Cancer Cell Proliferation
The most prominent metabolic alteration is known as the Warburg effect. However, cancer cells manipulate many other metabolic pathways for building up their own cellular components. Especially, activating lipid synthesis is highly important for cancer cells, because lipids such as phospholipid bilayers are fundamental structural components enabling cellular proliferation. It has been shown that extracellular lipids can sufficiently stimulate pancreatic cancer cell proliferation [43]. However, in a wide variety of tumors, de novo synthesis of fatty acids (FAs) is activated irrespective of the levels of circulating lipids. In contrast to normal cells, cancer cells may gain more than 93% of triacylglycerol FAs via de novo synthesis [44]. In the first step of FA synthesis, cytoplasmic acetyl-CoA is generated from citrate by ATP-citrate lyase (ACLY) and then converted into malonyl-CoA by acetyl-CoA carboxylase (ACC). Malonyl-CoA and acetyl-CoA are coupled to the acyl-carrier protein (ACP) domain of the multienzyme protein fatty acid synthase (FASN) (Fig. 13.2). Via repeated condensations of acetyl groups by the FASN in an NADPH-dependent manner, a basic 16-carbon saturated FA called palmitic acid is generated [45]. In cancer cells, expression of ACLY and ACC is also markedly increased [44]. Furthermore, serum FASN levels are higher in patients with PDAC, in patients with intraductal papillary mucinous neoplasm (IPMN), and in patients with chronic pancreatitis in comparison to healthy controls [46]. Pancreatic cancer patients with high FASN expression in the pancreas show a shorter overall survival than patients with low FASN expression [47]. Furthermore, FASN expression is correlated with poor response to gemcitabine therapy in pancreatic cancer cells [47, 48]. Increased Fasn gene expression is also observed in a pancreatic cancer mouse model with oncogenic KRASG12D and p53 R172H mutation [47], suggesting that enzymes involved in fatty acid synthesis can be important targets.
Regulation of fatty acid synthesis, cholesterol synthesis, fatty acid desaturation, and SREBP translocation. ACC acetyl-CoA carboxylase, ACLY ATP-citrate lyase, ER endoplasmic reticulum, FASN fatty acid synthase, HMG-CoA 3-hydroxy-3-methylglutaryl-CoA or β-hydroxy-β-methylglutaryl-CoA, SCD Δ9-stearoyl-CoA desaturase, SREBP sterol regulatory element-binding protein
Targeting Fatty Acid Synthesis in Cancer
For targeting fatty acid synthesis, several inhibitors for ACLY, ACC, and FASN blockade have been proposed. SB-204990 is an ACLY inhibitor which inhibits lipid synthesis. Intraperitoneal administration of SB-204990 leads to reduced tumor growth in mice carrying xenografts of primary mouse PDAC lines generated from oncogenic KRASG12D with or without p53 R172H mutation [49]. For inhibiting ACC, soraphen A and TOFA (5-(tetradecyloxy)-2-furoic acid) have been shown to block cancer cell growth [50], and treatment with TOFA suppresses the proliferation of pancreatic cancer cells [51]. In a mouse xenograft model, it has been demonstrated that intraperitoneally administered TOFA reduces human ovarian cancer cell development [52]. Oral administration of another ACC inhibitor ND-646 suppresses FA synthesis and tumor growth in lung cancer mouse models where tumors are induced by oncogenic KRASG12D with p53 deficiency or by oncogenic KRASG12D with Stk11 knockout [53]. Serine/threonine kinase 11, also known as liver kinase B1 (LKB1), activates AMPK for ACC inhibition. BAY ACC022 (another ACC inhibitor) attenuates growth of pancreatic cancer cell xenograft in mice [54]. These observations suggest that inhibiting the first step of FA synthesis is an attractive strategy for cancer therapy.
Targeting FASN can be performed by several different inhibitors, since FASN is a multienzyme protein complex with two identical polypeptides. The enzyme complex includes several catalytic domains with ACP, malonyl/acetyltransferase (MAT), β-ketoacyl-ACP synthase, β-ketoacyl-ACP reductase, 3-hydroxyacyl-ACP dehydrase, enoyl-CoA reductase, and palmitoyl-ACP thioesterase. Several inhibitors block β-ketoacyl-ACP synthase of FASN, namely, cerulenin, C75 (4-methylene-2-octyl-5-oxotetrahydrofuran-3-carboxylic acid, cerulenin-derived semisynthetic FASN inhibitor with improved stability), and epigallocatechin-3 gallate (EGCG) [44]. Cerulenin and C75 have been tested in several cancer xenograft models like for ovary, prostate, mesothelioma, breast, and colon cancer. Intraperitoneally administered cerulenin also suppresses liver metastasis of colon cancer cells in mice [55]. Blockage of FASN with EGCG has been considered for a broad range of cancer types such as prostate, lung, breast, and colorectal cancer [56, 57]. EGCG inhibits pancreatic cancer cell proliferation, and antiproliferative effects are also observed with catechin gallate (CG) and epicatechin gallate (ECG) [58]. EGCG inhibits growth of pancreatic tumor cells orthotopically implanted in mice [59]. For inhibiting β-ketoacyl-ACP reductase, several compounds like TVB-2640, TVB-3166, and GSK2194069 have been proposed. TVB-2640 has entered clinical trials, e.g., for colon cancer, breast cancer, and astrocytoma. Treatment with TVB-3166 leads to inhibition of proliferation and reduction in tumor growth of multiple cancer cell lines and pancreatic cancer xenografts [57, 60]. The β-lactone orlistat blocks palmitoyl-ACP thioesterase, and enoyl-CoA reductase can be blocked by triclosan [44, 61]. Orlistat is a US food and Drug Administration (FDA)-approved anti-obesity drug, and it has been shown that orlistat reduces human pancreatic cancer cell growth [47, 62]. Inhibition of FASN with orlistat suppresses growth of EGFR tyrosine kinase inhibitor-resistant cancer cells and also tumors in EGFR mutant transgenic mice [63]. One main limitation of orlistat is its low oral bioavailability, and improved formulation of orlistat-like inhibitors may be required in the future. Alternatively, other inhibitors of palmitoyl-ACP thioesterase can be identified via in silico screening of FDA-approved drugs. Lansoprazole, rabeprazole, omeprazole, and pantoprazole are proton pump inhibitors, but also function as inhibitors of thioesterase activity, which can induce pancreatic cancer cell death [64]. In conclusion, a number of inhibitors of ACLY, ACC, and FASN have been proposed and show significant effects in cancer therapy.
Fatty Acid Desaturases: Not Just a Modifier
The main product of FA synthesis in the cytoplasm is 16-carbon saturated palmitic acid. Longer FAs are formed by reactions catalyzed by several enzymes on the cytosolic side of the endoplasmic reticulum (ER). The desaturation of fatty acids occurs also in ER membranes. These modifications support the production of a wide variety of FAs and lipids. In mammalian cells, three types of fatty acid desaturases introduce carbon double bonds at Δ5 (Δ5-eicosatrienoyl-CoA desaturase), Δ6 (Δ6-oleoyl(linolenoyl)-CoA desaturase), or Δ9 (Δ9-stearoyl-CoA desaturase) (SCD). SCD is the rate-limiting enzyme catalyzing the synthesis of monounsaturated 16- or 18-carbon-like palmitoleate and oleate from palmitoyl-CoA and stearoyl-CoA [65]. Enhanced FA synthesis in cancer cells also increases the requirement of enzymes for modifying FAs and lipids. SCD1 (the main isoform) has been associated with insulin resistance and diabetes. Expression of SCD1 is associated with tumor promotion, shorter survival of lung cancer patients [66], and with sorafenib resistance in liver cancer patients [67]. SCD1 expression is upregulated in human colorectal cancer tissues, and patients with high SCD1 expression levels have a shorter overall survival [68]. It has also been suggested that increased SCD1 expression is associated with shorter survival of pancreatic cancer patients [69]. SCD1 contributes to the maintenance of cancer cell stemness, and knockdown of SCD1 reduces the expression of stemness markers like SOX2 and NANOG [70]. Cancer stemness may be responsible not only for tumor initiation but also for metastasis [71]. Taken together, targeting SCD1 could be a promising option.
Targeting Fatty Acid Desaturases
However, the role of SCD1 remains controversial and requires further investigation. In a murine intestinal cancer model with a mutant allele Min (multiple intestinal neoplasia) of the Apc (adenomatous polyposis coli) locus (called ApcMin/+ mice), conditional deletion of Scd1 in the intestinal epithelium promotes inflammation and tumorigenesis [72]. On the other hand, the inhibitor A939572 has been applied for renal cell carcinoma treatment. Oral administration of A939572 inhibits the development of tumor xenografts in mice [73]. Intraperitoneal injection with another SCD1 inhibitor (BZ36) reduces prostate cancer xenografts in mice [74]. Furthermore, pretreatment with the SCD1 inhibitor CAY10566 suppresses ovarian tumor growth after inoculation of cancer stem cells, where inhibition of SCD1 impairs cancer cell stemness [70]. The effects these inhibitors have on pancreatic cancer cells are currently not known.
Sterol Regulatory Element-Binding Proteins: Master Regulators of Lipid Biogenesis and Cholesterol Metabolism
Expression of genes involved in FA synthesis and modification such as ACLY, ACACA/B (coding ACCs), FASN, and SCD is regulated by the transcription factor sterol regulatory element-binding protein 1c (SREBP-1c) that is itself regulated transcriptionally and/or posttranslationally by several signaling pathways and factors such as PI3K/Akt and MEK/ERK [75]. EGFR signaling is required for oncogenic KRASG12D-induced pancreatic tumorigenesis [76, 77], and EGFR activation also induces upregulation of FASN in pancreatic cancer cells in an ERK-dependent manner [78]. Along this line, PDAC patients with high SREBP1 expression have a shorter overall survival than patients with low SREBP1 expression, and knockdown of SREBF1 (for SREBP1 expression) decreases pancreatic cancer cell viability and proliferation [79]. Taken together, oncogenic signaling pathways activate expression of lipogenic enzymes leading to aberrant activation of FA synthesis, which supports cancer cell development.
There are three SREBP isoforms, SREBP-1a, SREBP-1c, and SREBP-2. Both SREBP-1a and SREBP-1c are derived from a single gene but through alternative transcription start sites. Whereas SREBP-1c preferentially regulates genes of FA metabolism, SREBP-1a is a potent activator of all SREBP-responsive genes, and SREBP-2 regulates cholesterol biosynthesis [80]. Cholesterol is an essential structural component of cell membranes together with various phospholipids, sphingomyelin, and glycolipids. Cholesterol is de novo synthesized from cytoplasmic acetyl-CoA through the mevalonate pathway. The rate-limiting step of the pathway is the conversion of 3-hydroxy-3-methylglutaryl-CoA (HMG-CoA, also known as β-hydroxy-β-methylglutaryl-CoA) to mevalonate by HMG-CoA reductase [81]. In addition to the mevalonate pathway, cells can increase their cholesterol contents thought receptor-mediated endocytosis of low-density lipoproteins (LDLs) [82]. The LDL receptor (LDLR) and HMG-CoA reductase are both transcriptional targets of SREBP-2 [80]. Expression of HMG-CoA reductase and LDLR is elevated in an oncogenic KRASG12D pancreatic cancer mouse model [83]. It has been suggested that cholesterol intake is associated with increased risk of pancreatic cancer [84]. Increased expression of Ldlr has no significant effect on overall survival of pancreatic cancer patients, but high Ldlr expression is associated with an increased risk of tumor recurrence. Since LDLR silencing reduces ERK signaling as well as proliferation of PDAC cells, silencing also enhances response to gemcitabine chemotherapy [83].
Targeting Cholesterol Synthesis and SREBP
The development of LDLR-inactivating agents is currently an ongoing issue. Alternatively, SREBP-1c and SREBP-2 can be potential targets for cancer therapy, since these are key regulators of FASN expression and other enzymes in fatty acid synthesis like ACLY and ACC, and it also regulates expression of SCD, LDLR, and HMG-CoA reductase. SREBPs interact with the SREBP cleavage-activating protein (SCAP), and the complex stays with the ER membrane proteins INSIG1 and INSIG2. Under physiological conditions, reduction of cellular lipid levels results in conformational change of SCAP that abrogates its interaction with INSIGs. Dissociation of the SREBP/SCAP complex from INSIGs leads to transport of the complex from the ER to the Golgi where SREBP is cleaved and activated [85]. Glucose can enhance SCAP stability and reduce its association with INSIGs allowing transport of the SREBP/SCAP complex to the Golgi [86]. Betulin and fatostatin have been proposed as SREBP inhibitors through inhibition of ER-Golgi translocation. Betulin has initially been shown to improve hyperlipidemia and insulin resistance and to reduce atherosclerotic plaques [87]. Intraperitoneal injection of betulinic acid combined with mithramycin A (DNA and RNA polymerase inhibitor) blocks the development of pancreatic cancer xenografts in mice [88]. Fatostatin injection reduces expression of FASN, ACC, SCD1, ACLY, and also Hmgcr (HMG-CoA reductase) and Ldlr transcription to a lesser extent in obese mice [85]. The inhibitor has been tested in glioblastoma and prostate cancer cell xenografts. There, intraperitoneal treatment with fatostatin reduced xenograft growth in mice [89, 90]. Inhibiting de novo cholesterol synthesis by blockage of the rate-limiting enzyme HMG-CoA reductase has also been considered for cancer therapy. Several statin derivatives such as atorvastatin, lovastatin, pravastatin, rosuvastatin, and simvastatin have entered clinical trials. Among the derivatives, atorvastatin and simvastatin have been considered for pancreatic cancer treatment [57]. Taken together, there are several therapeutic options targeting SREBP and the mevalonate pathway, and a number of cancer studies are currently ongoing.
Glutamine and Acetate Metabolism in Pancreatic Cancer
Glutamine Metabolism : It Works Also Without Mitochondria
By modulating the activity of several metabolic pathways including glutamine metabolism, cancer cells aim for continuous generation of FAs necessary for cell growth. Glutamine is the most abundant and nonessential amino acid that can be synthesized from glucose. In the canonical route of mitochondrial glutamine catabolism (glutaminolysis), glutaminase (GLS) catalyzes glutamine to glutamate (Fig. 13.3). Glutamate is further converted by glutamate dehydrogenase (GLUD1) to α-ketoglutarate (α-KG), and α-KG can then be integrated into the tricarboxylic acid cycle (TCA cycle). Glutamine is an essential nutrient for the proliferation of human cancer cells [91], and several oncogenes which activate glutaminolysis have been identified. Oncogenic c-Myc enhances expression of mitochondrial GLS supporting canonical glutaminolysis [92]. Pancreatic cancer cells rely on a cytoplasmic noncanonical glutaminolysis pathway producing pyruvate via aspartate transaminase (GOT1, catalyzes aspartate/oxaloacetate), malate dehydrogenase (MDH1, catalyzes malate/oxaloacetate), and malate enzyme (ME1, catalyzes malate/pyruvate). Oncogenic KRAS induces a shift from canonical to noncanonical glutaminolysis by inhibiting mitochondrial GLUD1 and activating cytoplasmic GOT1 [93]. By reprogramming of glutamine metabolism from the mitochondrial to the cytoplasmic system , pancreatic cancer can keep synthesis of FAs intact, because cytoplasmic isocitrate dehydrogenase (IDH1) can catalyze α-KG/isocitrate under hypoxic conditions or even with defective mitochondria [94,95,96].
Targeting Glutamine Metabolism
Several drugs such as 968, BPTES, and CB-839 have been developed to inhibit GLS glutamate synthesis. Treatment with 968 or with BPTES reduces pancreatic cancer cell viability [93]. Intravenous injection of BPTES nanoparticles reduces pancreatic cancer xenograft growth in mice, and combination with intraperitoneal injection of metformin enhances therapeutic effects [97]. CB-839 has already been tested in several clinical studies including a broad range of cancer types, such as clear cell renal carcinoma, breast cancer, and colorectal cancer. However , oral gavage of CB-839 has no antitumor activity in mice with oncogenic KRASG12D combined with Trp53 deficiency. In addition, mice treated with CB-839 show marginally shorter survival than the group without CB-839 treatment [98]. Further investigations are therefore required to judge whether GLS inhibition is a potential therapeutic option for pancreatic cancer patients. EGCG and R162 have been considered to inhibit GLUD1 [99]. EGCG has been described as a FASN β-ketoacyl-ACP synthase inhibitor and shown to inhibit pancreatic cancer cell proliferation (see Targeting Fatty Acid Synthesis in Cancer), and it is also recognized as a GLUD1 inhibitor. Treatment with R162 inhibits proliferation of several cancer cells including primary leukemia cells. Furthermore, intraperitoneal injection of R162 inhibits the development of lung cancer xenografts in mice [100]. Oncogenic KRASG12D has been suggested to inhibit GLUD1 and preferentially activate the noncanonical glutaminolysis pathway (see Glutamine Metabolism: It Works Also Without Mitochondria); thus, GLUD1 inhibition might be ineffective in pancreatic cancer. Methyl 3-(3-(4-(2,4,4-trimethylpentan-2-yl)phenoxy)-propanamido)benzoate (named compound 16c) has been synthesized to inhibit the noncanonical glutaminolysis pathway as a MDH inhibitor. This inhibitor blocks both cytoplasmic MDH1 and mitochondrial MDH2 enzymes. It has been shown that intraperitoneal administration of this inhibitor attenuates the development of colon cancer xenografts [101]. Since inhibition of MDH1 activity leads to suppression of glutamine metabolism and reduction of pancreatic cancer cell growth [102], inhibitors for the noncanonical glutaminolysis pathway could be potential candidates for pancreatic cancer therapy.
Acetate Metabolism : Cancer Cells Are Experts in Bridging the Gap
Acetyl-CoA represents a central metabolite not only for lipid synthesis but also for regulating gene expression as a key determinant of protein/histone acetylation [103, 104]. Cancer cells preferentially convert pyruvate into lactate rather than to transport it into the mitochondria for PDH reaction and the TCA cycle. Although the IDH1-mediated non-canonical glutaminolysis pathway (see Glutamine Metabolism: It Works Also Without Mitochondria) may compensate to provide acetyl-CoA in the cytoplasm, alternative sources of acetyl-CoA could still be necessary for sufficient supporting lipid synthesis and cancer cell growth. Cells with ACLY deficiency remain viable and proliferate, where acetate supports acetyl-CoA generation and de novo lipid synthesis is supported by the enzyme called ACSS2 [105]. There have been 26 acyl-CoA synthetases (ACS) identified in the human genome. Among those, three enzymes, the short-chain ACS (ACSS) family (acetyl-CoA synthetase), are capable of catalyzing synthesis of acetyl-CoA from acetate in an ATP-dependent manner [106]. ACSS1 and ACSS3 are mitochondrial enzymes, and ACSS2 localizes to both the cytoplasmic and nuclear compartments. Silencing of ACSS2 in cancer cells reduces incorporation of acetyl units from acetate into either lipids or histones. ACSS2 is highly expressed in several human tumors, and loss of ACSS2 suppresses tumor development in certian mouse liver cancer models including c-Myc combined with PTEN knockout [107]. Under metabolic stress such as hypoxia and/or low-nutrition conditions, expression of ACSS2 is elevated, and it promotes acetate uptake for lipid synthesis and membrane phospholipids in several cancers including pancreatic cancer cells [108, 109].
Inhibitors specifically targeting ACSS2 remain largely unexplored. So far a compound 1-(2,3-di(thiophen-2-yl)quinoxalin-6-yl)-3-(2-methoxyethyl)urea (PubChem CID: 2300455; here referred to as 508186-14-9) has been proposed as a ACSS2-specific inhibitor [107]. The inhibitor has been tested and showed decreased lipid contents in bladder cancer cells, but not in non-cancer cells [110]. Targeting ACSS2 and acetate metabolism would be a highly interesting concept for treating pancreatic cancer.
Conclusion
Extensive research on cancer metabolism has revealed that a number of enzymes and metabolites are involved in reprogramming strategies of many cancer types including pancreatic cancer. Furthermore, it is evident that overexpression of specific enzymes is not only related with metabolic reprogramming but also with cellular stemness. Several studies with inhibitors targeting specific catalyzing steps in selected metabolic pathways have shown convincing effects in inhibiting cancer development and progression. Cancers may however still find other ways to generate necessary metabolic intermediates and cellular components. Therefore, it is important to further understand not only the cross talk between oncogenic signaling pathways and metabolism but also between metabolic pathways for offering stratified and more effective therapies in the future.
References
Kleeff J, Korc M, Apte M, La Vecchia C, Johnson CD, Biankin AV, et al. Pancreatic cancer. Nat Rev Dis Primers. 2016;2:16022.
Rahib L, Smith BD, Aizenberg R, Rosenzweig AB, Fleshman JM, Matrisian LM. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 2014;74:2913–21.
Hingorani SR, Petricoin EF, Maitra A, Rajapakse V, King C, Jacobetz MA, et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell. 2003;4(6):437–50.
Perera RM, Bardeesy N. Pancreatic cancer metabolism: breaking it down to build it Back up. Cancer Discov. 2015;5(12):1247–61.
White E. Exploiting the bad eating habits of Ras-driven cancers. Genes Dev. 2013;27(19):2065–71.
Hanahan D, Weinberg RA. Hallmark of cancer: the next generation. Cell. 2011;144:646–74.
Warburg O, Minami S. Versuche an Überlebendem Carcinomgewebe. Klin Wochenschr. 1923;2(17):776–7.
Warburg O, Posener K, Negelein E. Über den Stoffwechsel der Carcinomzelle. Biochem Z. 1924;152:309–44.
Warburg O, Wind F, Negelein E. The metabolism of tumors in the body. J Gen Physiol. 1927;8(6):519–30.
Warburg O. On respiratory impairment in cancer cells. Science. 1956;124(3215):269–70.
Ying H, Kimmelman AC, Lyssiotis CA, Hua S, Chu GC, Fletcher-Sananikone E, et al. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell. 2012;149(3):656–70.
Muller PA, Vousden KH. p53 mutations in cancer. Nat Cell Biol. 2013;15(1):2–8.
Zhang C, Liu J, Liang Y, Wu R, Zhao Y, Hong X, et al. Tumour-associated mutant p53 drives the Warburg effect. Nat Commun. 2013;4:2935.
Papageorgis P, Cheng K, Ozturk S, Gong Y, Lambert AW, Abdolmaleky HM, et al. Smad4 inactivation promotes malignancy and drug resistance of colon cancer. Cancer Res. 2011;71(3):998–1008.
Zhang W, Liu Y, Chen X, Bergmeier SC. Novel inhibitors of basal glucose transport as potential anticancer agents. Bioorg Med Chem Lett. 2010;20(7):2191–4.
Liu Y, Cao Y, Zhang W, Bergmeier S, Qian Y, Akbar H, et al. A small-molecule inhibitor of glucose transporter 1 downregulates glycolysis, induces cell-cycle arrest, and inhibits cancer cell growth in vitro and in vivo. Mol Cancer Ther. 2012;11(8):1672–82.
Shibuya K, Okada M, Suzuki S, Seino M, Seino S, Takeda H, et al. Targeting the facilitative glucose transporter GLUT1 inhibits the self-renewal and tumor-initiating capacity of cancer stem cells. Oncotarget. 2015;6(2):651–61.
Wang J, Ye C, Chen C, Xiong H, Xie B, Zhou J, et al. Glucose transporter GLUT1 expression and clinical outcome in solid tumors: a systematic review and meta-analysis. Oncotarget. 2017;8(10):16875–86.
Sharen G, Peng Y, Cheng H, Liu Y, Shi Y, Zhao J. Prognostic value of GLUT-1 expression in pancreatic cancer: results from 538 patients. Oncotarget. 2017;8(12):19760–7.
Atsumi T, Chesney J, Metz C, Leng L, Donnelly S, Makita Z, et al. High expression of inducible 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase (iPFK-2; PFKFB3) in human cancers. Cancer Res. 2002;62(20):5881–7.
Minchenko A, Leshchinsky I, Opentanova I, Sang N, Srinivas V, Armstead V, et al. Hypoxia-inducible factor-1-mediated expression of the 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase-3 (PFKFB3) gene. Its possible role in the Warburg effect. J Biol Chem. 2002;277(8):6183–7.
De Bock K, Georgiadou M, Schoors S, Kuchnio A, Wong BW, Cantelmo AR, et al. Role of PFKFB3-driven glycolysis in vessel sprouting. Cell. 2013;154(3):651–63.
Gustafsson NMS, Färnegårdh K, Bonagas N, Ninou AH, Groth P, Wiita E, et al. Targeting PFKFB3 radiosensitizes cancer cells and suppresses homologous recombination. Nat Commun. 2018;9(1):3872.
Conradi LC, Brajic A, Cantelmo AR, Bouché A, Kalucka J, Pircher A, et al. Tumor vessel disintegration by maximum tolerable PFKFB3 blockade. Angiogenesis. 2017;20(4):599–613.
Dayton TL, Jacks T, Vander Heiden MG. PKM2, cancer metabolism, and the road ahead. EMBO Rep. 2016;17(12):1721–30.
Christofk HR, Vander Heiden MG, Harris MH, Ramanathan A, Gerszten RE, Wei R, et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature. 2008;452(7184):230–3.
Anastasiou D, Yu Y, Israelsen WJ, Jiang JK, Boxer MB, Hong BS, et al. Pyruvate kinase M2 activators promote tetramer formation and suppress tumorigenesis. Nat Chem Biol. 2012;8(10):839–47.
Luengo A, Gui DY, Vander Heiden MG. Targeting metabolism for cancer therapy. Cell Chem Biol. 2017;24(9):1161–80.
Hillis AL, Lau AN, Devoe CX, Dayton TL, Danai LV, Di Vizio D, et al. PKM2 is not required for pancreatic ductal adenocarcinoma. Cancer Metab. 2018;6:17.
Doherty JR, Cleveland JL. Targeting lactate metabolism for cancer therapeutics. J Clin Invest. 2013;123(9):3685–92.
Le A, Cooper CR, Gouw AM, Dinavahi R, Maitra A, Deck LM, et al. Inhibition of lactate dehydrogenase A induces oxidative stress and inhibits tumor progression. Proc Natl Acad Sci U S A. 2010;107(5):2037–42.
Rajeshkumar NV, Dutta P, Yabuuchi S, de Wilde RF, Martinez GV, Le A, et al. Therapeutic targeting of the Warburg effect in pancreatic cancer relies on an absence of p53 function. Cancer Res. 2015;75(16):3355–64.
Patra KC, Hay N. The pentose phosphate pathway and cancer. Trends Biochem Sci. 2014;39(8):347–54.
Santana-Codina N, Roeth AA, Zhang Y, Yang A, Mashadova O, Asara JM, et al. Oncogenic KRAS supports pancreatic cancer through regulation of nucleotide synthesis. Nat Commun. 2018;9(1):4945.
Tong X, Zhao F, Thompson CB. The molecular determinants of de novo nucleotide biosynthesis in cancer cells. Curr Opin Genet Dev. 2009;19(1):32–7.
Liu H, Huang D, McArthur DL, Boros LG, Nissen N, Heaney AP. Fructose induces transketolase flux to promote pancreatic cancer growth. Cancer Res. 2010;70(15):6368–76.
Aune D, Chan DS, Vieira AR, Navarro Rosenblatt DA, Vieira R, Greenwood DC, et al. Dietary fructose, carbohydrates, glycemic indices and pancreatic cancer risk: a systematic review and meta-analysis of cohort studies. Ann Oncol. 2012;23(10):2536–46.
Shukla SK, Purohit V, Mehla K, Gunda V, Chaika NV, Vernucci E, et al. MUC1 and HIF-1alpha signaling crosstalk induces anabolic glucose metabolism to impart gemcitabine resistance to pancreatic cancer. Cancer Cell. 2017;32(1):71–87.e7.
Cunningham JT, Moreno MV, Lodi A, Ronen SM, Ruggero D. Protein and nucleotide biosynthesis are coupled by a single rate-limiting enzyme, PRPS2, to drive cancer. Cell. 2014;157(5):1088–103.
Becker MA, Heidler SA, Bell GI, Seino S, Le Beau MM, Westbrook CA, et al. Cloning of cDNAs for human phosphoribosylpyrophosphate synthetases 1 and 2 and X chromosome localization of PRPS1 and PRPS2 genes. Genomics. 1990;8(3):555–61.
Qian X, Li X, Tan L, Lee JH, Xia Y, Cai Q, et al. Conversion of PRPS hexamer to monomer by AMPK-mediated phosphorylation inhibits nucleotide synthesis in response to energy stress. Cancer Discov. 2018;8(1):94–107.
Zhang H, Qian DZ, Tan YS, Lee K, Gao P, Ren YR, et al. Digoxin and other cardiac glycosides inhibit HIF-1alpha synthesis and block tumor growth. Proc Natl Acad Sci U S A. 2008;105(50):19579–86.
Clerc P, Bensaadi N, Pradel P, Estival A, Clemente F, Vaysse N. Lipid-dependent proliferation of pancreatic cancer cell lines. Cancer Res. 1991;51:3633–8.
Menendez JA, Lupu R. Fatty acid synthase and the lipogenic phenotype in cancer pathogenesis. Nat Rev Cancer. 2007;7:763–77.
Baenke F, Peck B, Miess H, Schulze A. Hooked on fat: the role of lipid synthesis in cancer metabolism and tumour development. Dis Model Mech. 2013;6:1353–63.
Walter K, Hong SM, Nyhan S, Canto M, Fedarko N, Klein A, et al. Serum fatty acid synthase as a marker of pancreatic neoplasia. Cancer Epidemiol Biomark Prev. 2009;18:2380–5.
Tadros S, Shukla SK, King RJ, Gunda V, Vernucci E, Abrego J, et al. De novo lipid synthesis facilitates gemcitabine resistance through endoplasmic reticulum stress in pancreatic cancer. Cancer Res. 2017;77:5503–17.
Yang Y, Liu H, Li Z, Zhao Z, Yip-Schneider M, Fan Q, et al. Role of fatty acid synthase in gemcitabine and radiation resistance of pancreatic cancers. Int J Biochem Mol Biol. 2011;2:89–98.
Hatzivassiliou G, Zhao F, Bauer DE, Andreadis C, Shaw AN, Dhanak D, et al. ATP citrate lyase inhibition can suppress tumor cell growth. Cancer Cell. 2005;8:311–21.
Currie E, Schulze A, Zechner R, Walther TC, Farese RV Jr. Cellular fatty acid metabolism and cancer. Cell Metab. 2013;18:153–61.
Nishi K, Suzuki K, Sawamoto J, Tokizawa Y, Iwase Y, Yumita N, et al. Inhibition of fatty acid synthesis induces apoptosis of human pancreatic cancer cells. Anticancer Res. 2016;36(9):4655–60.
Li S, Qiu L, Wu B, Shen H, Zhu J, Zhou L, et al. TOFA suppresses ovarian cancer cell growth in vitro and in vivo. Mol Med Rep. 2013;8:373–8.
Svensson RU, Parker SJ, Eichner LJ, Kolar MJ, Wallace M, Brun SN, et al. Inhibition of acetyl-CoA carboxylase suppresses fatty acid synthesis and tumor growth of non-small-cell lung cancer in preclinical models. Nat Med. 2016;22:1108–19.
Petrova E, Scholz A, Paul J, Sturz A, Haike K, Siegel F, et al. Acetyl-CoA carboxylase inhibitors attenuate WNT and Hedgehog signaling and suppress pancreatic tumor growth. Oncotarget. 2017;8:48660–70.
Murata S, Yanagisawa K, Fukunaga K, Oda T, Kobayashi A, Sasaki R, et al. Fatty acid synthase inhibitor cerulenin suppresses liver metastasis of colon cancer in mice. Cancer Sci. 2010;101:1861–5.
Niedzwiecki A, Roomi MW, Kalinovsky T, Rath M. Anticancer efficacy of polyphenols and their combinations. Nutrients. 2016;8(9):pii: E552.
Sunami Y, Rebelo A, Kleeff J. Lipid metabolism and lipid droplets in pancreatic cancer and stellate cells. Cancers (Basel). 2017;10(1):pii: E3.
Kürbitz C, Heise D, Redmer T, Goumas F, Arlt A, Lemke J, et al. Epicatechin gallate and catechin gallate are superior to epigallocatechin gallate in growth suppression and anti-inflammatory activities in pancreatic tumor cells. Cancer Sci. 2011;102(4):728–34.
Shankar S, Marsh L, Srivastava RK. EGCG inhibits growth of human pancreatic tumors orthotopically implanted in Balb C nude mice through modulation of FKHRL1/FOXO3a and neuropilin. Mol Cell Biochem. 2013;372:83–94.
Röhrig F, Schulze A. The multifaceted roles of fatty acid synthesis in cancer. Nat Rev Cancer. 2016;16(11):732–49.
Jones SF, Infante JR. Molecular pathways: fatty acid synthase. Clin Cancer Res. 2015;21:5434–8.
Sokolowska E, Presler M, Goyke E, Milczarek R, Swierczynski J, Sledzinski T. Orlistat reduces proliferation and enhances apoptosis in human pancreatic cancer cells (PANC-1). Anticancer Res. 2017;37:6321–7.
Ali A, Levantini E, Teo JT, Goggi J, Clohessy JG, Wu CS, et al. Fatty acid synthase mediates EGFR palmitoylation in EGFR mutated non-small cell lung cancer. EMBO Mol Med. 2018;10(3):pii: e8313.
Fako VE, Wu X, Pflug B, Liu JY, Zhang JT. Repositioning proton pump inhibitors as anticancer drugs by targeting the thioesterase domain of human fatty acid synthase. J Med Chem. 2015;58:778–84.
Peck B, Schulze A. Lipid desaturation – the next step in targeting lipogenesis in cancer? FEBS J. 2016;283(15):2767–78.
Huang J, Fan XX, He J, Pan H, Li RZ, Huang L, et al. SCD1 is associated with tumor promotion, late stage and poor survival in lung adenocarcinoma. Oncotarget. 2016;7:39970–9.
Ma MKF, Lau EYT, Leung DHW, Lo J, Ho NPY, Cheng LKW, et al. Stearoyl-CoA desaturase regulates sorafenib resistance via modulation of ER stress-induced differentiation. J Hepatol. 2017;67:979–90.
Ran H, Zhu Y, Deng R, Zhang Q, Liu X, Feng M, et al. Stearoyl-CoA desaturase-1 promotes colorectal cancer metastasis in response to glucose by suppressing PTEN. J Exp Clin Cancer Res. 2018;37(1):54.
Macášek J, Vecka M, Žák A, Urbánek M, Krechler T, Petruželka L, et al. Plasma fatty acid composition in patients with pancreatic cancer: correlations to clinical parameters. Nutr Cancer. 2012;64:946–55.
Li J, Condello S, Thomes-Pepin J, Ma X, Xia Y, Hurley TD, et al. Lipid desaturation is a metabolic marker and therapeutic target of ovarian cancer stem cells. Cell Stem Cell. 2017;20:303–314.e5.
Marquardt JU, Thorgeirsson SS. Stem cells in hepatocarcinogenesis: evidence from genomic data. Semin Liver Dis. 2010;30(1):26–34.
Ducheix S, Peres C, Härdfeldt J, Frau C, Mocciaro G, Piccinin E, et al. Deletion of stearoyl-CoA desaturase-1 from the intestinal epithelium promotes inflammation and tumorigenesis, reversed by dietary oleate. Gastroenterology. 2018;155(5):1524–1538.e9.
Von Roemeling CA, Marlow LA, Wei JJ, Cooper SJ, Caulfield TR, Wu K, et al. Stearoyl-CoA desaturase 1 is a novel molecular therapeutic target for clear cell renal cell carcinoma. Clin Cancer Res. 2013;19:2368–80.
Fritz V, Benfodda Z, Rodier G, Henriquet C, Iborra F, Avancès C, et al. Abrogation of de novo lipogenesis by stearoyl-CoA desaturase 1 inhibition interferes with oncogenic signaling and blocks prostate cancer progression in mice. Mol Cancer Ther. 2010;9:1740–54.
Wang Y, Viscarra J, Kim SJ, Sul HS. Transcriptional regulation of hepatic lipogenesis. Nat Rev Mol Cell Biol. 2015;16(11):678–89.
Ardito CM, Grüner BM, Takeuchi KK, Lubeseder-Martellato C, Teichmann N, Mazur PK, et al. EGF receptor is required for KRAS-induced pancreatic tumorigenesis. Cancer Cell. 2012;22:304–17.
Navas C, Hernández-Porras I, Schuhmacher AJ, Sibilia M, Guerra C, Barbacid M. EGF receptor signaling is essential for k-Ras oncogene-driven pancreatic ductal adenocarcinoma. Cancer Cell. 2012;22:318–30.
Bian Y, Yu Y, Wang S, Li L. Up-regulation of fatty acid synthase induced by EGFR/ERK activation promotes tumor growth in pancreatic cancer. Biochem Biophys Res Commun. 2015;463:612–7.
Sun Y, He W, Luo M, Zhou Y, Chang G, Ren W, et al. SREBP1 regulates tumorigenesis and prognosis of pancreatic cancer through targeting lipid metabolism. Tumor Biol. 2015;36:4133–41.
Horton JD, Goldstein JL, Brown MS. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest. 2002;109:1125–31.
Ikonen E. Cellular cholesterol trafficking and compartmentalization. Nat Rev Mol Cell Biol. 2008;9:125–38.
Goldstein JL, Brown MS, Anderson RG, Russell DW, Schneider WJ. Receptor-mediated endocytosis: concepts emerging from the LDL receptor system. Annu Rev Cell Biol. 1985;1:1–39.
Guillaumond F, Bidaut G, Ouaissi M, Servais S, Gouirand V, Olivares O, et al. Cholesterol uptake disruption, in association with chemotherapy, is a promising combined metabolic therapy for pancreatic adenocarcinoma. Proc Natl Acad Sci U S A. 2015;112:2473–8.
Chen H, Qin S, Wang M, Zhang T, Zhang S. Association between cholesterol intake and pancreatic cancer risk: evidence from a meta-analysis. Sci Rep. 2015;5:8243.
Soyal SM, Nofziger C, Dossena S, Paulmichl M, Patsch W. Targeting SREBPs for treatment of the metabolic syndrome. Trends Pharmacol Sci. 2015;36(6):406–16.
Cheng C, Ru P, Geng F, Liu J, Yoo JY, Wu X, et al. Glucose-mediated N-glycosylation of SCAP is essential for SREBP-1 activation and tumor growth. Cancer Cell. 2015;28(5):569–81.
Tang JJ, Li JG, Qi W, Qiu WW, Li PS, Li BL, et al. Inhibition of SREBP by a small molecule, betulin, improves hyperlipidemia and insulin resistance and reduces atherosclerotic plaques. Cell Metab. 2011;13:44–56.
Gao Y, Jia Z, Kong X, Li Q, Chang DZ, Wie D, et al. Combining betulinic acid and mithramycin a effectively suppresses pancreatic cancer by inhibiting proliferation, invasion, and angiogenesis. Cancer Res. 2011;71:5182–93.
Williams KJ, Argus JP, Zhu Y, Wilks MQ, Marbois BN, York AG, et al. An essential requirement for the SCAP/SREBP signaling axis to protect cancer cells from lipotoxicity. Cancer Res. 2013;73:2850–62.
Li X, Chen YT, Hu P, Huang WC. Fatostatin displays high antitumor activity in prostate cancer by blocking SREBP-regulated metabolic pathways and androgen receptor signaling. Mol Cancer Ther. 2014;13(4):855–66.
Eagle H. Nutrition needs of mammalian cells in tissue culture. Science. 1955;122:501–14.
Gao P, Tchernyshyov I, Chang TC, Lee YS, Kita K, Ochi T, et al. c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature. 2009;458:762–5.
Son J, Lyssiotis CA, Ying H, Wang X, Hua S, Ligorio M, et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature. 2013;496:101–5.
Metallo CM, Gameiro PA, Bell EL, Mattaini KR, Yang J, Hiller K, et al. Reductive glutamine metabolism by IDH1 mediates lipogenesis under hypoxia. Nature. 2011;481:380–4.
Mullen AR, Wheaton WW, Jin ES, Chen PH, Sullivan LB, Cheng T, et al. Reductive carboxylation supports growth in tumour cells with defective mitochondria. Nature. 2011;481:385–8.
Anastasiou D, Cantley LC. Breathless cancer cells get fat on glutamine. Cell Res. 2012;22:443–6.
Elgogary A, Xu Q, Poore B, Alt J, Zimmermann SC, Zhao L, et al. Combination therapy with BPTES nanoparticles and metformin targets the metabolic heterogeneity of pancreatic cancer. Proc Natl Acad Sci U S A. 2016;113:E5328–36.
Biancur DE, Paulo JA, Małachowska B, Quiles Del Rey M, Sousa CM, Wang X, et al. Compensatory metabolic networks in pancreatic cancers upon perturbation of glutamine metabolism. Nat Commun. 2017;8:15965.
Altman BJ, Stine ZE, Dang CV. From Krebs to clinic: glutamine metabolism to cancer therapy. Nat Rev Cancer. 2016;16:619–34.
Jin L, Li D, Alesi GN, Fan J, Kang HB, Lu Z, et al. Glutamate dehydrogenase 1 signals through antioxidant glutathione peroxidase 1 to regulate redox homeostasis and tumor growth. Cancer Cell. 2015;27:257–70.
Naik R, Ban HS, Jang K, Kim I, Xu X, Harmalkar D, et al. Methyl 3-(3-(4-(2,4,4-Trimethylpentan-2-yl)phenoxy)-propanamido)benzoate as a novel and dual malate dehydrogenase (MDH) 1/2 inhibitor targeting cancer metabolism. J Med Chem. 2017;60:8631–46.
Wang YP, Zhou W, Wang J, Huang X, Zuo Y, Wang TS, et al. Arginine methylation of MDH1 by CARM1 inhibits glutamine metabolism and suppresses pancreatic cancer. Mol Cell. 2016;64:673–87.
Choudhary C, Weinert BT, Nishida Y, Verdin E, Mann M. The growing landscape of lysine acetylation links metabolism and cell signaling. Nat Rev Mol Cell Biol. 2014;15:536–50.
Pietrocola F, Galluzzi L, Bravo-San Pedro JM, Madeo F, Kroemer G. Acetyl coenzyme A: a central metabolite and second messenger. Cell Metab. 2015;21:805–21.
Zhao S, Torres A, Henry RA, Trefely S, Wallace M, Lee JV, et al. ATP-citrate lyase controls a glucose-to-acetate metabolic switch. Cell Rep. 2016;17(4):1037–52.
Watkins PA, Maiguel D, Jia Z, Pevsner J. Evidence for 26 distinct acyl-coenzyme A synthetase genes in the human genome. J Lipid Res. 2007;48:2736–50.
Comerford SA, Huang Z, Du X, Wang Y, Cai L, Witkiewicz AK, et al. Acetate dependence of tumors. Cell. 2014;159:1591–602.
Schug ZT, Peck B, Jones DT, Zhang Q, Grosskurth S, Alam IS, et al. Acetyl-CoA synthetase 2 promotes acetate utilization and maintains cancer cell growth under metabolic stress. Cancer Cell. 2015;27:57–71.
Bulusu V, Tumanov S, Michalopoulou E, van den Broek NJ, MacKay G, Nixon C, et al. Acetate recapturing by nuclear acetyl-CoA synthetase 2 prevents loss of histone acetylation during oxygen and serum limitation. Cell Rep. 2017;18:647–58.
Lee MY, Yeon A, Shahid M, Cho E, Sairam V, Figlin R, et al. Reprogrammed lipid metabolism in bladder cancer with cisplatin resistance. Oncotarget. 2018;9(17):13231–43.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Sunami, Y. (2020). Targeting Metabolism. In: Michalski, C., Rosendahl, J., Michl, P., Kleeff, J. (eds) Translational Pancreatic Cancer Research. Molecular and Translational Medicine. Humana, Cham. https://doi.org/10.1007/978-3-030-49476-6_13
Download citation
DOI: https://doi.org/10.1007/978-3-030-49476-6_13
Published:
Publisher Name: Humana, Cham
Print ISBN: 978-3-030-49475-9
Online ISBN: 978-3-030-49476-6
eBook Packages: MedicineMedicine (R0)