Abstract
Estimation of structural biomarkers and covariance networks from MRI have provided valuable insight into the morphological processes and organisation of the human brain. State-of-the-art analyses such as linear mixed effects (LME) models and pairwise descriptive correlation networks are usually performed independently, providing an incomplete picture of the relationships between the biomarkers and network organisation. Furthermore, descriptive network analyses do not generalise to the population level. In this work, we develop a Bayesian generative model based on wombling that allows joint statistical inference on biomarkers and connectivity covariance structure. The parameters of the wombling model were estimated via Markov chain Monte Carlo methods, which allow for simultaneous inference of the brain connectivity matrix and the association of participants’ biomarker covariates. To demonstrate the utility of wombling on real data, the method was used to characterise intrahemispheric cortical thickness and networks in a study cohort of subjects with Alzheimer’s disease (AD), mild-cognitive impairment and healthy ageing. The method was also compared with state-of-the-art alternatives. Our Bayesian modelling approach provided posterior probabilities for the connectivity matrix of the wombling model, accounting for the uncertainty for each connection. This provided superior inference in comparison with descriptive networks. On the study cohort, there was a loss of connectivity across diagnosis levels from healthy to Alzheimer’s disease for all network connections (posterior probability ≥ 0.7). In addition, we found that wombling and LME model approaches estimated that cortical thickness progressively decreased along the dementia pathway. The major advantage of the wombling approach was that spatial covariance among the regions and global cortical thickness estimates could be estimated. Joint modelling of biomarkers and covariance networks using our novel wombling approach allowed accurate identification of probabilistic networks and estimated biomarker changes that took into account spatial covariance. The wombling model provides a novel tool to address multiple brain features, such as morphological and connectivity changes facilitating a better understanding of disease pathology.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Conditional autoregressive model
- Markov chain Monte Carlo
- Spatial statistics
- Wombling
- Cortical thickness
- Alzheimer’s disease
- Structural MRI
1 Introduction
Alzheimer’s disease (AD) is the most common form of dementia [13, 67]. While clinical diagnosis of AD is often derived from psychological assessments, neuroimaging studies have found that the structural and functional changes in the brain that align with AD pathology can be identified prior to the detection of cognitive symptoms [2, 65].
Structural neuroimaging studies typically use two approaches: region of interest (ROI) analyses to estimate morphological biomarkers for each region, such as thickness, volume and the rate of tissue loss; and cortical networks to investigate associations between multiple ROIs. This two pronged approach is important as biomarkers in one region are likely to influence the morphological properties of connected regions. For example, highly correlated ROIs (often quantified through cortical networks) are often a part of a system that is known to be associated with particular behavioural or cognitive functions [3, 47]. Nonetheless these approaches are often performed independently, providing valuable insight into the differences in brain organisation and degeneration patterns for multiple regions between healthy and pathological groups [9, 15, 34, 55, 58, 62, 72]. For example, Bernal-Rusiel et al. [9] found that their models for ROIs were able to characterise changes in individuals’ measurements at multiple time points while handling up to 45.5% patient drop out. Furthermore, analyses on cortical thickness networks have demonstrated a reduction in connectivity efficiency between healthy groups and groups with neurological disorders such as schizophrenia and AD [8, 15, 37, 46, 59, 71].
An advantage of analyses conducted on a single region is the direct biological interpretation on the estimation of tissue features, such as thickness and estimated annual rate of tissue loss [9]. However, it is difficult to ascertain a brain-wide picture of all ROIs under such analyses, as this requires multiple comparison corrections in order to account for the high number of hypothesis tests [21, 34, 60, 72]. Alternatively, cortical networks provide a summary measure on the topological brain network organisation which conveniently encompasses the complex information across all ROIs [15, 37, 46, 58]. However, direct biological interpretations of such networks are difficult as the relationship between the ROI node and corresponding links represent a covariance measure among ROIs, and not physical connections [3]. Furthermore, generalising to a population cortical network is difficult to achieve from descriptive analyses as such methods are not generative models and do not take into account the variability of each connection [46, 59,60,61, 71].
In practice both approaches complement each other with participants who are healthy, in general, tend to have thicker cortical tissue and highly organised networks compared to pathological groups such as AD [9, 17, 18, 37, 55, 71]. The aforementioned shortcomings of these methods could be resolved by combining both approaches into a unified framework. Such a framework could avoid multiple comparisons and provide a cortical network whose links reflect the uncertainty of the data.
In this work, we propose a Bayesian hierarchical (generative) model that jointly performs network-based inference in conjunction with neuroimaging biomarker estimates. This approach enforces consistency between any spatial interactions and biomarker estimates (for network and cortical thickness) at the population and participant level, while handling correlated measures from within and between individuals in conjunction with covariates in a statistically principled manner.
1.1 Technical Survey of Previous Work of Bayesian Hierarchical Models
Bayesian hierarchical models have been extensively applied to unify independent analyses, for example, combining the joint estimation of voxel and ROI analyses [12, 19, 69], and combining diffusion and functional MRI into a single model [70]. Accommodating both within and between participant variation from longitudinal observations, as well as high patient drop out (unbalanced design) has been previously achieved through a related method called the mass univariate Bayesian hierarchical analysis [72]. Previously, Bayesian linear mixed effect (LME) models, which are a type of hierarchical model, have been applied independently to key ROIs associated with AD progression [17]. An advantage of Bayesian inference is that it can detect significant differences among groups of interest through the direct comparison of the marginal posterior distributions, without the need for hypothesis tests or multiple comparisons corrections. However, LME models (Bayesian and non-Bayesian) applied to neuroimaging data usually analyse each ROI independently and do not account for the covarying measurements between several brain regions.
Recent probabilistic brain networks in the Bayesian framework have shown great potential to estimate a population network for clinical groups. Bayesian brain networks are probabilistic rather than binary, and so are easy to interpret [38, 41, 61]. However, these probabilistic networks are not easily extended to include additional neuroimaging biomarker estimation, such as cortical thickness, volumes, or fluorodeoxyglucose uptake as measured by positron emission topography. To account for the correlation between measurements on ROIs, several neuroimaging studies [33, 36, 48, 53] have used spatial dependence modelling via a Gaussian Markov random field (GMRF, Gössl et al. [32] and Woolrich et al. [68]). However, an underlying and potentially invalid assumption is that the adjacency structure of the correlations are known and fixed, and most are constrained to nearest neighbour configurations. This was highlighted in the Bayesian hierarchical spatial models by Bowman [11] and Bowman et al. [12] suggesting that the underlying physical and biological processes may not always be contiguous, and relationships among ROIs are not restricted to regions which are immediate anatomical neighbours.
1.1.1 Previous Work on Wombling
Wombling refers to the estimation of a neighbourhood matrix through the covariance structure of a GMRF that is estimated under a Bayesian framework [42, 49, 50]. This neighbourhood structure can be incorporated as an additional parameter in the Bayesian hierarchical model, and can be estimated in addition to participant specific covariates such as gender and other biomarkers associated with AD factors [22, 24, 31].
1.2 Overview of Our Work
In this work, we propose Bayesian hierarchical wombling models that jointly performs network based inference in conjunction with regional biomarker estimates. This approach estimates the complex covariance associations among several regions without assuming contiguous relationships via estimation of a connectivity structure. Furthermore, biomarker estimates at the population and participant level handle correlated measures from within and between individuals in conjunction with covariates. This enables full statistical inference of biomarker estimates and produces a probabilistic network.
To this end, this chapter is organised as follows: Sect. 7.2.2 outlines the proposed Bayesian hierarchical wombling model. The wombling model is validated via a simulation study described in Sect. 7.2.3 and results are reported in Sect. 7.3.1. Sections 7.2.1 and 7.3.2 present the application of brain wombling on the Australian Imaging, Biomarkers and Lifestyle (AIBL) study of ageing data on healthy controls (HC), mild cognitive impaired (MCI) and AD diagnosed groups as well as in age ranges discretised into three groups. A comparison of the results from the wombling approach with comparable independent analyses are presented in Sects. 7.3.2.2 and 7.3.2.4. A discussion of our work appears in Sect. 7.4.
2 Materials and Methods
The overarching objective of this work is to develop and validate a joint analysis of biomarker and covariance networks facilitated by the proposed wombling approach. The flowchart in Fig. 7.1 provides an overview of the experiments presented in this work, showing the inputs and outputs for each analysis. Case study data will be based on cortical thickness estimates into a study of into a study of healthy ageing, MCI and AD participants. The wombling method will be compared to Pearson pairwise correlation networks and Bayesian LME models. In addition, as this work was the first to investigate the wombling approach for joint analysis of cortical networks and biomarker estimates, a simulation study was used to evaluate the performance of the wombling algorithm to recover the true connectivity structure and simulated biomarker values.
Overview of the analysis workflow. Arrows show the relationships between the rectangle methods sections. The results from both methods are denoted by the circular plots on the far right and are compared to each other to assess the performance of joint analyses facilitated by the wombling approach (Sect. 7.2.2) in comparison with the state-of-the-art independent analyses (Sect. 7.2.4)
2.1 AIBL Study of Ageing
In this work, we applied our proposed method to data from the Australian Imaging, Biomarkers and Lifestyle (AIBL) longitudinal study of ageing. AIBL is an ongoing study which aims to discover which biomarkers such as cognitive assessment, neuroimaging, lifestyle and demographic factors potentially influence the development of AD. The AIBL study was approved by the institutional ethics committees of Austin Health, St Vincent’s Health, Hollywood Private Hospital and Edith Cowan University. All study volunteers gave written informed consent prior to participating in the study. MRI data were collected at baseline and at several ∼18 month follow-up intervals (replicates) from a subset of 167 participants. This resulted in a total of 597 sets of ROI observations. Only those observations from participants with two or more replicates were retained; these included 120 HC, 21 and 26 clinically diagnosed MCI individuals and AD participants respectively. Of the 167 participants, 77 were male (46%) and 90 were female (54%). Mean baseline ages was numerically higher in those diagnosed with MCI (HC: 73.1 ± 6.7, MCI: 77.0 ± 6.4 and AD: 73.8 ± 7.5, p = 0.055).
The structural T1W MRI images were first segmented into grey/white matter and cerebral spinal fluid using an in-house implementation of the expectation maximisation algorithm applied to a Gaussian mixture model [64]. Cortical thickness was computed along the grey matter based on a combined Lagrangian-Eulerian partial differential equation approach [1]. The grey matter was parcellated following the Automated Anatomical Labelling (AAL) atlas [63] using a multi-atlas registration approach [10]. For this work, we used 35 ROI cortical thickness regions from the left hemisphere of the brain, as listed in Table 7.1.
2.2 Wombling Model Formulation and Parameter Estimation
In this section, we present the wombling generative model used to jointly estimate cortical brain connectivity and thickness in a Bayesian hierarchical framework. Wombling is a type of a LME model that accounts for correlations between regions, after accounting for fixed effects. In this work, wombling does not provide age related estimates as it is not a longitudinal model, such an extension is beyond the scope of this work and motivates future work. The wombling model comprises of two parts; a mixed effect model and connectivity estimation with their respective set of assumptions. LME model assumptions include: a linear relationship exists between the response and the exploratory variables; the response is normally distributed about a mean, although for non-normal responses we may extend this assumption to the exponential family and apply generalised linear mixed models [51]; the variances across fixed and random effects are unknown but constant, and observations for a region can be correlated with its neighbours, but observations between non-neighbouring regions are assumed to be conditionally independent. Connectivity matrix assumptions are twofold. Firstly, the underlying connectivity structure quantified by matrix W is the same across all individuals in a specified group. Secondly, relationships between regions are equally weighted, as our framework estimates the probability of each pairwise connection and not the connection strength. This implies that if region j is a neighbour of region k, then region k is also neighbour of region j, and regions are not neighbours with themselves, w ii = 0 ∀i.
The hierarchical structure of the model separates the variation of the data into two levels; fixed effects (A) and random effects (B) shown in Fig. 7.2. At level A, the linear predictor for person i, at repeated measure r on region k comprises of participant i′s covariate vector x i (covariate matrix for all participants is denoted by X), parameter vector β, spatial random effects b i and residual variance σ 2. Level B consists of the spatial random effects b i which follow a multivariate normal distribution with a mean of 0 and a covariance matrix \(\sigma ^2_sQ\). The product, \(\sigma ^2_sQ\), comprises of the spatial scale variance term, \(\sigma ^2_s\), which controls the variation of the random effects and a function of the connectivity structure matrix Q.
Visualisation of brain wombling model (7.1) via a directed acyclic graph. Nodes in circles and rectangle denote parameters and observed variables respectively. Direction of arrows indicates direction of influence or dependence. Rectangle B denotes the second layer of model, which accounts for spatial dependence among ROI conditional and nested in rectangle A, which is the upper-most layer with fixed effects β parameter. The spatial random effects vector for participant i, b i, is modelled as a multivariate normal, whose covariance structure is a function of the binary symmetric adjacency matrix W, of dimension K × K and w jk = 1 or w jk = 0 implies that region j and k are connected or disconnected respectively
The cortical thickness of region k = 1, 2, …, K within participant i = 1, 2, …, I who has r = 1, …, R i replicates is y irk measured in millimetres. The brain wombling model is of the following form:
Details of the formulation for the connectivity structure are as follows: matrix D w is a diagonal matrix with elements given by the row sums (or number of neighbours) \(\sum ^K_{j = 1}w_{jk}\) for k = 1, 2, …, K. The matrix W is a zero-diagonal, binary symmetric matrix, with elements w jk = 1 if regions j and k are neighbours or zero otherwise, and identity matrix \(\mathbb {I}\) has dimension K × K. The value of ρ determines the global level of the spatial correlation [43] where values of ρ close to zero correspond to (near) independence in the spatial random effect, and ρ close to one denotes high spatial correlation. While ρ can be an additional parameter in our wombling model, in this application we fix ρ = 0.9, to enforce high spatial correlation and avoid the difficult and computationally intensive task of estimating ρ, as described in Lu et al. [50] and Lee [42]. For completeness we investigated the effect of ρ at various values to assess the recovery of W; refer to Sect. 7.2.3 for further details.
The parametrisation of Q −1 defined in (7.1) was chosen due to its superior ability to handle a range of spatial strengths [42, 45]. This parametrisation has also been favoured in other wombling and spatial clustering applications [4, 43]. Visualisation of model parameters conditional on the observed regional biomarker response, such as cortical thickness, and participant specific covariates are shown in Fig. 7.2.
In a Bayesian framework the likelihood corresponding to the model in (7.1) is p(y|b, σ 2, β, X), which is conditional on the spatial random effects and the model parameters. Note the data is conditionally independent of the network structure W and spatial scale variance, \(\sigma ^2_s\). The resultant joint posterior distribution for the model parameters and the random effects given the data is
In the Bayesian paradigm the population parameters are random variables, and priors \(p(\boldsymbol {\beta }), p(\sigma ^2), p(\sigma ^2_s)\) and p(W) are assigned to each parameter. Details on prior specification are described in Sects. 7.2.2.1 and 7.2.2.3. Markov chain Monte Carlo (MCMC) methods were used to sample from the joint posterior probability distribution of the parameters [57], which samples from the marginal posterior distributions as a by-product [29]. At each MCMC step, samples are iteratively drawn from the full conditionals of the parameters with a Metropolis-Hastings (M-H, Chib and Greenberg [20], Metropolis et al. [52]) update for W. Following a burn-in period, samples will eventually be drawn from the joint posterior distribution of the parameters [12].
Full conditional distributions in closed form were derived for parameters \(\boldsymbol {\beta }, \sigma ^2_s\) and σ 2 which were sampled via a Gibbs sampler as described in Sect. 7.2.2.1. As the matrix W is symmetric, the off-diagonal, upper triangular elements were updated one at a time via a M-H sampler as described in Sect. 7.2.2.3.
2.2.1 Prior and Conditional Distributions for \(\sigma ^2_s\), σ 2 and β
A semi-conjugate prior in the form of an inverse gamma distribution, IG(c, d), was chosen for the spatial scale variance \(\sigma ^2_s\), with shape and rate values c and d, respectively. Likewise, the prior for the residual variance σ 2 was an IG(e, f) distribution. Hyperparameters were chosen to provide support over a wide range of possible values for \(\sigma ^2_s\) and σ 2. The full conditional distributions for \(\sigma ^2_s\) and σ 2 are as follows
and
where N is the total number of observations.
The prior for the fixed effect parameter β is a multivariate normal distribution MV N(μ 0, Σ 0), and in keeping with wombling literature [50], μ 0 and Σ 0 were chosen so that the prior on β is vague. It can be shown that the full conditional distribution for β is
The response in long vector form is \(\mathbf {y} = [y_{111}, y_{112}, \ldots , y_{11K}, y_{121}, \ldots , y_{IR_{I}K}]\) and the covariate matrix X with superscript L is \(\overset {L}X = [{\mathbf {x}}_1,{\mathbf {x}}_1, \ldots , {\mathbf {x}}_2, {\mathbf {x}}_2, \ldots , {\mathbf {x}}_I]\), hence the individual specific covariate vector x i is repeated R i times, where \(\overset {L}X\) is an N by p matrix; where p is the total number of covariates for the model, including the intercept. Similarly, the long vector form for the spatial random effects is \(\overset {L}{\mathbf {b}} = [b_{111}, b_{121}, \ldots , b_{1KR_{1}}, b_{211}, \ldots , b_{IR_{I}K}]\).
2.2.2 Full Conditional Distribution for Spatial Random Effects b i
From Model (7.1) we can derive meaningful participant specific estimates of cortical thickness for each of the ROIs analysed, and investigate how this deviates from the population average (β 0). The individual-specific estimates of cortical thickness for each ROI in our analysis are derived from the full conditional distribution of b i given by
where the unit vector e = [1, 1, …, 1] is of length K.
2.2.3 Prior and Posterior Sampling for Brain Connectivity Matrix W
According to the posterior distribution in (7.2), the full conditional for matrix W is of the form
Elements of the matrix W are updated one at a time. As W is symmetric, we only require estimation of the off-diagonal, upper triangular elements. To facilitate a data driven method to estimate the brain connectivity matrix, our prior knowledge of the probability of a link between any pair of ROIs is 0.5, that is, p(w ij = 1) = p(w ij = 0) = 0.5 for all values of i and j.
We use the M-H algorithm within a Gibbs sampler to draw posterior simulations for W. We update W element-wise by drawing independent proposals, \(w_{kj}^*\), from the prior of W and accepting a proposal with probability
where the covariance precision evaluated at the proposed value is \(Q^{-1}_{w_{kj}^*}\).
2.3 Simulation Studies
The proposed Bayesian brain wombling approach accommodates for both network based inference and biomarker estimates. For this reason the aims of the simulation study are twofold. Firstly, we aim to evaluate the performance of this model at ‘recovering’ two underlying connectivity matrices W (structured and contiguous configurations). In the context of this manuscript, by recovery we refer to whether the credible intervals of the estimator contain the true solution. The assumed true matrices for W are shown in Fig. 7.3A and F respectively. Our second aim is to illustrate that our model recovers the simulated biomarker estimates via fixed effect vector β in addition to simulated participant specific estimates through their spatial random effects (b i).
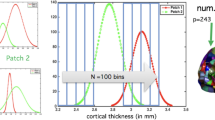
Data generated from binary W matrices for structured (a) and contiguous (f) configurations. A single wombling simulation run results in a posterior distribution for W, whose mean represents an average connectivity matrix. The mean of the posterior means over all 50 simulations in each scenario are the connectivity matrices shown in b, d, g and i, which show the average posterior probability of region j being a neighbour of region k. Top row: Mean of posterior means for structured balanced (b) and unbalanced (d) simulation designs, with corresponding binary matrices (c and e respectively) whose elements are equated to one if their value is greater than threshold τ = 0.6 and zero otherwise. Bottom row: Similarly for the contiguous simulation study, mean of posterior means for balanced (g) and unbalanced (i) matrices with corresponding binarised matrices (h and i) at probability threshold of 0.6 and above
In order to relate the simulation study to the real data application, the values used to generate the simulated study data were chosen to reflect features of the AIBL study, such as the number of simulated ROIs, number of repeated measures (replicates) in the unbalanced design, the number of participants and range of biomarker values.
2.3.1 Wombling Simulated Analyses
For both configurations of W, the vector β = [β 0, β 1] = [3, 0.5] was assumed as the intercept and gender effect, and x sim was specified as a binary vector with male participant as baseline (i.e. x i,sim = 1 to simulate a female participant and x i,sim = 0 a male participant). The average global human cortical thickness can range from approximately 1 to 4.5 mm [27], hence the prior for β was chosen to remain physiologically feasible around this value. For this reason the hyperparameters for the precision matrix Σ 0 had zero off-diagonals and diagonal elements of value 1/10, and the hyperparameter for μ 0 was chosen to be 0. We note that these are the same priors used for the real data application described in Sect. 7.2.4. Variance terms for both W configurations were set to \(\sigma ^2_s = 1\) and σ 2 = 0.5, with relatively uninformative inverse gamma priors specified as IG(1, 1) and IG(1, 0.5) respectively. Priors for both configurations of W matrices are described in Sect. 7.2.2.3.
Our simulation studies were undertaken by generating 50 independent data sets from Model (7.1). We fitted our model to each data set to obtain (50) posterior distributions for our parameters. Here, we considered a balanced design whereby each simulated participant had the same number of repeated measures, and the more realistic unbalanced alternative, where the number of replicates for each participant varied.
Data for I = 100 participants were simulated from Model (7.1), where each participant had K = 35 simulated ROI as listed on Table 7.1, and each participant had R i = 7 replicates as a balanced design. The unbalanced simulation design comprised of participants with 4–7 replicates (mean 5.8). Parameter values and prior information as described above were set for balanced and unbalanced designs, whereby each design was explored as structured and contiguous W configurations, for a total of four scenarios.
Each scenario resulted in a mean of posterior means for W, representing the probabilistic network. These scenarios were binarised for ease of comparison to assess the recovery of W. Values w jk = 1 if the average posterior probability of a connection between regions j and k was equal to or greater than τ = 0.6, and w jk = 0 otherwise. We note that binary W is dependent on the choice of τ, and that τ = 0.6 is sufficiently far away from the prior (p(w ij = 1) = p(w ij = 0) = 0.5).
Further details of the simulation analyses including percentage of recovery of the assumed true values and MCMC convergence checks are provided in section “Simulation Study” of Appendix.
2.4 Application to Study Cohort
We hypothesised that each population group has an underlying cortical brain network, denoted as matrix W, while expecting differences in W between groups, as each group represented progressive levels of neurodegeneration in both cortical thickness estimates and structural brain networks.
The Bayesian brain wombling Model (7.1) was applied independently to data from three diagnosis groups (HC, MCI and AD) as well as three age groups (A: 59–69y; B: 69–79y; C: 79–93y). In order to compare the wombling model with current state-of-the-art methods that provide cortical networks, population and participant specific estimates, we derived Pearson pairwise correlation networks and Bayesian LME models to the aforementioned case study groups. The subsections below describe how the marginal posterior draws were processed after the wombling model was applied to the AIBL case study, as well as details of the independent analyses methods applied to produce comparable biomarker estimates as described in literature [9, 16, 17, 34, 39].
2.4.1 Probabilistic Connectivity Matrices via Wombling
Inference on the brain wombling models were estimated by the MCMC scheme described in Sect. 7.2.2, which was applied to each group and was run using four chains. Each chain ran for M = 500,000 iterations. The first 50,000 runs (burn-in) were discarded and every 50th iteration retained (thinning).
The resultant elements of the posterior mean of W matrices are \(\bar {w}_{kj}\), and represent the probability that region k is connected to region j in a cortical structural network. These networks represent the underlying average network of a group estimated from our sample. Binary matrices were derived for a given probability threshold (0 < τ < 1) for each element of W. This threshold determines the level of confidence in our brain network, and allows for straightforward comparisons across groups. However as noted in He et al. [37] and Yao et al. [71], a high threshold on brain networks may lead to disconnected networks and this may make topological network metrics difficult to analyse. In this work, we set τ = 0.7 as this value is substantially higher than the prior value of 0.5, and is greater than our 0.6 value from our simulated study in Sect. 7.2.3, thus providing a more stringent level on the certainty of the resultant networks, resulting in a high level of confidence regarding the potential connections between nodes.
2.4.2 Descriptive Pearson Cortical Networks
Following the methods of Bassett et al. [8], we applied Pearson pairwise correlation networks at both baseline (which consisted of all observations being independent and identically distributed (IID)) as well as on the whole data with repeated measures treated as IID.
2.4.3 Wombled Population and Participant ROI Biomarker Estimates
In the Bayesian paradigm, the posterior distributions of parameters can be compared directly to make probabilistic statements about each other, or in regards to other biologically relevant quantities. The probability that parameter β 0,A from group A is within the lower 2.5% and upper 97.5% quantiles of the posteriori β 0,B from group B (denoted by Y L and Y H), is estimated by

where the indicator function  is equal to one if \(Y_L < \beta _{0, A}^m < Y_H\) and zero otherwise. The length of the MCMC chain for β
0,A is M. Comparison of all groups are computed in a similar manner, whose results are listed in Table 7.2 of Sect. 7.3.2.3.
is equal to one if \(Y_L < \beta _{0, A}^m < Y_H\) and zero otherwise. The length of the MCMC chain for β
0,A is M. Comparison of all groups are computed in a similar manner, whose results are listed in Table 7.2 of Sect. 7.3.2.3.
While our algorithm provides cortical thickness estimates on all participants in the analysis for each ROI, we focused on the nine key regions often used to describe the cortical signature of AD [21, 23]: the inferior, medial and superior temporal lobes; supramarginal, angular, posterior cingulate and the precuneus gyrus. Results for all 35 ROI can be found in Figs. 7.17 and 7.18.
Low cortical thickness estimates are often indicative of neurodegeneration. For this reason, at the participant level analyses in Sect. 7.3.2.3, we expected an increasing atrophy pattern to be associated with diagnosis, from AD to MCI to HC, as well as among age groups, from old to young. Participants which differ from this pattern may be showing early signs of AD pathology, thus this analysis could be also be used to flag sub-groups of participants to follow up.
2.4.4 Bayesian LME ROI Analyses
Bayesian LME models were applied independently on each ROI in a similar manner as Bernal-Rusiel et al. [9], Guillaume et al. [34], Caselli et al. [16], Holland et al. [39] and Cespedes et al. [17] who applied LME models at the ROI level. Similarly, others who applied these models at the voxel scale in AD and in other neurological applications [35, 72]. Refer to section “Bayesian Linear Mixed Effect Models on Each ROI” in Appendix for model specifications. The wombling and combined Bayesian LME models were compared by the Watanabe-Akaike information criterion (WAIC, Watanabe [66]). The survey by Gelman et al. [30] describes how the WAIC has been shown to be the preferred approach for model comparison in the Bayesian community. For this reason, it is applied to the models this work.
2.5 Statistical Analysis
All analyses were undertaken using the open-source software R [56]. Source code and data used in the simulation study are available at https://github.com/MarcelaCespedes/Brain_wombling. Simulation experiments were performed using a high performance computer cluster. We note that a single MCMC instance of the Bayesian brain wombling model ran on a single central processing unit (CPU) and took approximately 24 h to run on a standard computer (four core 3.40 GHz Intel i7-4770 processor).
3 Results
3.1 Simulation Studies
3.1.1 Wombling Simulated Analyses
Figure 7.3A and F show the comparison between the W we should recover, and the average estimated W for the structured configuration (Fig. 7.3B and D), and contiguous configuration (Fig. 7.3G and I). Section 7.2.3 describes how the mean of the posterior mean matrices in Fig. 7.3B, D, G and I were binarised. The resultant binarised matrices for the structured balanced and unbalanced designs recovered 83% and 82% of the networks’ solution (Fig. 7.3C, E). The binarised matrices for the contiguous balanced and unbalanced designs recovered 70% and 65% of the contiguous configuration (Fig. 7.3H and J). The parameter dimension in the 35 ROI simulation study consisted of the off-diagonals of W, (K(K − 1)∕2 = 595) in addition to β, σ 2 and \(\sigma ^2_s\), which was a total of 599 parameters. As can be seen by Fig. 7.3, the wombling model showed the desired recovery of the connectivity matrices in both configurations and in the balanced and unbalanced designs, despite the high parameter dimension.
To assess whether the random effects were recovered appropriately, we evaluated their 95% credible intervals. These results showed that the true values of the random effects were recovered on average 95% of the time indicating that the variation of the posterior distribution is appropriate. See Table 7.3 and Fig. 7.9 for details of these results. Likewise, the recovery of the solution vector β was within the 95% of the credible intervals approximately 95% of the time in all simulation configurations and scenarios, demonstrating that in addition to recovery of connectivity networks, the wombling model was able to recover the biomarker and participant level estimates.
A sensitivity analysis with respect to the prior information on \(\sigma ^2, \sigma ^2_s,\) and β, was conducted on the structured W configuration. This entailed re-running the analysis using various specifications of the prior information. The subsequent posterior summaries did not vary considerably based on different prior information. Hence we postulate that estimation of \(\sigma ^2, \sigma ^2_s,\) and β are relatively robust to the priors specified in this work.
The results described above relate to two fixed W configurations with the same values on β, σ 2, and \(\sigma ^2_s\) for each scenario. We investigated the effect of different values for variance terms (\(\sigma ^2_s\) and σ 2) on the recovery of W and fixed and random effects. We found the results to be very similar to those reported here (model results for different variance terms not shown). Furthermore, we investigated the effect of the value of ρ on the recovery of W with ρ ∈{0.85, 0.9, 0.95, 0.99} using the balanced structured simulation scenario. There is some wombling literature which suggests that the choice of ρ can affect the recovery of \(\sigma ^2_s\) and W [42, 44, 50]; we found a choice of ρ = 0.9 provided appropriate recovery of parameters of interest. Refer to Table 7.4 for results on a range of ρ values.
In summary, our simulation study showed the recovery of W proved to be appropriate, which implies that the estimation of Q −1 is reliable. However the spatial scale variance \((\sigma ^2_s)\) was typically overestimated, a finding that is not uncommon in wombling literature [50]. Despite this, the simulation study also showed adequate recovery on biomarker and participant estimates, as such our estimates for β, σ 2 and b i are reliable.
3.2 Application to Real Data
In this section, we present the results of the joint analysis derived by the wombling model, and compared them with the results from the independent analyses (overview in Fig. 7.1).
The MCMC algorithm was utilised to draw posterior samples from the wombling model applied on diagnosis and age groups of the AIBL case study. As described in Sect. 7.2.5, informal diagnostic measures were assessed, such as trace, density and autocorrelation plots, as well as formal measures to investigate between and within chain variation with the Gelman-Rubin convergence measure [14]. All plots suggested convergence to a stationarity distribution according to the Gelman-Rubin convergence checks. Furthermore, posterior predictive checks on all models in these analyses showed the models fit the data well; there were no systematic departures from the model predictions and 95–99% of all response values were within the 95% credible intervals of the posterior predictive distributions; refer to Table 7.5 and Figs. 7.10, 7.11 for results.
3.2.1 Probabilistic Connectivity Matrices via Wombling
The networks corresponding to the probabilistic matrices in Fig. 7.4 show the results for diagnosis levels HC (top: A and B), MCI (middle: C and D) and AD (bottom: E and F). The varying level of uncertainty between matrices is indicated by elements with probability values close to 0.5, in contrast with connections which have high or low probabilities. This is partly due to a sample size effect, as there were 120 participants who were diagnosed as HC compared to MCI (21) and AD (26) participants.
Left: Posterior mean of W for each diagnosis, top to bottom; HC (a and b), MCI (c and d) and AD (e and f). Right: Cortical networks from binarised posterior matrix W with threshold τ = 0.7 for the respective diagnosis groups. Node size reflects the number of edges on each vertice. Total number of edges for each network (top to bottom) are 156, 124 and 112 for HC, MCI and AD networks respectively
The networks on the right of Fig. 7.4 show those connections with a probability equal to or greater than 0.7. The network configurations reflect the underlying estimated population networks. The total number of edges in these networks show HC participants have a more complex cortical network structure with 156 connections, in comparison with the MCI network which had 124 connections. Furthermore, the AD network has a lower degree (112 connections) in contrast with the MCI and HC networks, suggesting a higher loss of network communication among the ROIs. The middle temporal lobe is one of the earliest regions known to be affected by the onset of AD [40]; with a probability greater than 0.7, our results indicate the number of connections of the HC, MCI and AD networks for this region are 7, 6 and 4 respectively, suggesting a loss of connections along the AD pathway. A similar reduction in node degree, in general, can be observed on the entire cortical mantle, across the frontal, temporal, parietal and occipital lobes.
Baseline age differences are observed in cortical networks in Fig. 7.5. The networks on the right of Fig. 7.5 show a re-organisation of connections, rather than a direct loss of total network degree with an increase of age. The older age Group C (79–93y) consists of 62 participants of which 41 were diagnosed as HC at baseline and 8 were diagnosed as AD. Hence the analysis in this group is dominated by HC, and the resultant network better aligns to healthy ageing rather than onset of AD. The diagnosis ratio of participants in the younger and middle age Groups A and B (59–69y and 69–79y respectively) have higher ratio of AD and MCI participants in contrast to HC. Hence the averaged networks across these groups include participants with a broader spectrum across healthy ageing, and progression to AD or other dementias, in contrast with age Group C.
Left: Posterior means of W for age groups (top to bottom) A (59–69y), B (69–79y) and C (79–93y) shown in plots A, C and E respectively. Right: Cortical network from binarised posterior matrix W with threshold τ = 0.7 for the respective age groups for the respective age groups A, B and C shown in plots B, D and F. Node size reflects the number of edges on each vertice
3.2.2 Descriptive Pearson Cortical Networks
The Pearson pairwise network analyses on diagnosis and age groups were sensitive to data with repeated measures, as connections varied across both groups between networks derived from IID and data with replicates. This finding is interesting as the studies by Li et al. [46] and Fan et al. [25] used repeated measures in their pairwise correlation network analyses. However, in our analyses, only the IID Pearson cortical networks were used for comparison with the wombling model. Once the IID correlation networks were binarised by placing a link between ROIs whose absolute correlation values greater than τ = 0.7, the diagnosis group did not support biologically meaningful networks: the Pearson pairwise correlations were considerably higher in the MCI group, followed by AD and HC with fewer connections. The age correlation matrices were binarised in the same manner, and the resulting sparse networks had a loss of connections from young to older age groups, of A (47) to B (38) to C (19). Refer to Figs. 7.26, 7.27, 7.28, 7.29, 7.30, and 7.31 for full Pearson pairwise correlation network results.
3.2.3 Wombled Population and Participant ROI Biomarker Estimates
In our application of brain wombling with Model (7.1), the vector β = [β 0, β 1] contains fixed effect parameters, where the intercept β 0 represents the mean thickness of the left cortex hemisphere of the brain, for a particular group and β 1 is the gender effect, with females as baseline and covariate x i = 1 for male. In all groups analysed, the gender effect was not substantive (95% credible intervals for β 1 included zero), thus we conclude there are no significant gender differences in global cortical thickness between the groups analysed.
The median cortical thickness mantle in HC groups (β 0,HC) is significantly higher than AD clinical diagnosis, as the 95% credible interval of β 0,HC lies outside of the 95% credible interval of the AD distribution (β 0,AD). While the posterior median of the MCI group was not significantly different from the medians of the HC or AD groups, from Fig. 7.6, the cascading order of degeneration on the cortex can be seen in the disease progression from HC to MCI to AD.
Posterior marginal distributions of total cortical thickness (β 0) across groups. Median of each distribution shown in each box plot, whiskers indicate 95% credible interval for each parameter. Diagnosis groups; healthy control (HC), mild cognitive impaired (MCI), Alzheimer’s disease (AD) and age Groups A, B and C correspond to age ranges 59–69y, 69–79y and 79–93y respectively
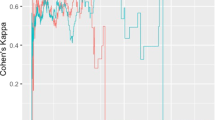
Ordered ROI cortical thickness posterior means and 95% credible intervals over participants random effects b i, for nine key regions associated with AD cortical signature. Colour coded as follows; AD (red), MCI (orange) and HC (green) diagnosis groups. Refer to Table 7.1 for full name of cortex regions
Posterior mean and 95% credible interval on participants estimated cortical thickness (random effects b i), on regions of interest, namely those that showed the highest differences. Age Groups colour coded as follows; (dark green) A, younger age group; (orange) B, middle age group and (brown) C, oldest age group
Each scenario (structured balanced and unbalanced, contiguous balanced and unbalanced) had 100 simulated participants and each participant had 35 ROI (3500 random effects in total for each simulation). As each scenario comprised of 50 simulations, there are 50 × 3500 random effects to assess. Each histogram denoted the percentage of the number of random effects recovered (that is random effects whose solution within the 95% credible interval)
As described in Sect. 7.2.4.3, we can make probabilistic comparisons among the median cortical thickness between groups. The probability that, a posteriori β 0,HC is within the AD L and AD H quantiles of the posterior distribution of the AD is 0.02, which implies there is a significant difference between the cortex of HC and AD groups. The probability that β 0,HC is within the MCI L and MCI H quantiles of the MCI diagnosis (β 0,MCI) is 0.49. This high probability is reflected in the third quartile of the MCI box plot overlapping the HC box plot in Fig. 7.6. The comparison of the MCI and AD box plots in Fig. 7.6 reflect the overlapping of the upper and lower distribution tail ends, which is reflected in the distribution for β 0,MCI, whose posteriori probability of being within AD L and AD H is 0.9.
There were subtle differences in the posterior cortical thickness estimates among age Groups A, B and C shown in Fig. 7.6. Unlike the large differences between diagnosis groups shown in Fig. 7.6 and probability comparisons in Table 7.2, the posteriori of a younger age group lies inside the credible interval of an older age group with a probability ≥0.79. These results suggest there were no significant differences between the median cortex of the age groups. However, as expected, there is a cascading order of cortical degeneration from thicker to thinner estimates from age Groups (A, B) to C, that is age ranges 59–79y and 79–93y respectively.
In addition to brain network and global cortical thickness estimates, the hierarchical structure of the wombling approach allowed for participant level estimates for all ROIs. The caterpillar plots in Fig. 7.7 show distinct patterns of participant clusters of AD, MCI and HC groups, particularly in the nine key regions, as they are the most likely to be influenced in the early stages of AD. AD participants had the lowest cortical thickness estimates as a result of higher cortical atrophy. MCI are midway in the degeneration scope with slightly higher cortical thickness estimates than AD, but lower than HC. Regions in which diagnosis groups differed particularly included the temporal poles of the middle and superior temporal gyrus and posterior cingulate gyrus. Regions which showed AD participants were not clustered exclusively at the lowest range of the cortical thickness estimates include the temporal poles (middle and superior) as well as the angular gyrus. Excluding these regions, for the remainder of the diagnosis clusters among participants were consistent in all other ROI plots (see Figs. 7.17 and 7.18). Note that these differences in diagnosis levels are consistent with the loss of network connectivity in Fig. 7.4 and total average cortical thickness estimates in Fig. 7.6.
The results of ranked participants were analysed with respect to age groups and selected regions are shown in Fig. 7.8; refer to Figs. 7.17 and 7.18 for the remaining ROI plots. The regions in Fig. 7.8 showed pronounced age group specific clusters. Key regions which age Group A had consistently higher cortical thickness estimates in contrast with age Groups B and C were the calcarine fissure, fusiform, heschl, middle temporal and precentral gyrus.
Left: Posterior mean of W heat map for APOE ε4 carriers (top) and non-carriers (bottom). Right: Cortical network from binarised heat map with threshold τ = 0.7 for the respective APOE ε4 carrier groups. Node size reflects the number of edges on each vertice. Total number of edges for each network (top and bottom) are 152 and 150 for APOE ε4 non-carriers and carriers groups
3.2.4 Bayesian LME ROI Analyses
Participant specific estimates via the Bayesian LME models were assessed and the results align with those from the wombling model in both key ROI which support strong distinctions among groups (particularly in the diagnosis groups) as well as in instances which all ROI showed little difference among groups, such as as those in the age groups; refer to Figs. 7.20, 7.21, 7.22, 7.23 for all Bayesian LME results. For example, the superior middle and inferior temporal regions had distinct HC, MCI and AD participant clusters, as well as the supramarginal and the posterior cingulate. The WAIC values of the wombled model in all groups were found to be substantially lower than the WAIC values of the Bayesian LME models. These results show that the wombling model is a more parsimonious approach to model biomarker estimates compared to independent LME models on each ROI, and hence a desirable model for this data. Refer to Table 7.6 for WAIC results.
4 Discussion
This work demonstrated and validated the Bayesian wombling approach using intrahemispherical cortical thickness observations of the brain in both a simulation study, and applied to an Alzheimer’s disease cohort study. Each analysis was applied across HC, MCI and AD diagnosis categories as well as three age groups. Wombling provides a novel way to combine both regression and network analyses into a single unified model. This takes into account the uncertainty of all possible links to estimate a network, but also allows group comparisons from independent measurements (for example, participants’ cortical volumes for many ROIs) without the need for multiple comparison correction.
4.1 Simulation Study
The ability of the wombling algorithm to successfully recover the underlying connectivity solution while appropriately accounting for the variance was assessed in Sect. 7.3.1. Figure 7.3 shows the overall average performance of the wombling algorithm as probability and thresholded networks. The wombling algorithm consistently and correctly detected the absence of connections in the structured configuration, and recovered 82% and above of the true values of W. On the more difficult contiguous scenario, Fig. 7.3H and I show that 65% and above the contiguous solution was recovered, at a probability threshold greater than 0.6, which as expected was less certain than the structured configuration.
Approximately 95% of the cortical thickness estimates were recovered at the population and participant level. Recovery in the statistical sense refers to whether the intervals of the estimator contain the true solution. Thus, an algorithm that recovers the known solution 100% of the time, could potentially do so by simply overestimating the variance. In our simulation studies, based on 95% credible intervals the wombling parameters were recovered approximately 95% of the time (see Table 7.3). This indicated that the wombled model appropriately estimated the variability in the parameters.
While the simulation study was designed to mimic features typical of longitudinal study data (in this case we matched some of the characteristics of the AIBL study, such as the number of participants, replicates and connectivity configuration), the practical performance of the wombling algorithm is better assessed when it is applied to the real data and directly compared with the alternative state-of-the-art methods.
4.2 Application to Study Cohort
4.2.1 Cortical Networks
The results from the brain wombling model were compared with those of alternative independent analyses on the AIBL data. Figure 7.4A, C and E shows a decrease in connections from HC (156) to MCI (124) to AD (112), which reflect the biological order of neurodegeneration [15, 18, 58]. As expected from previous work [6], the loss of connections on the wombled networks reflect the strong differences in the diagnosis groups which is also reflected in the wombled cortical thickness estimates shown in Figs. 7.6, 7.7 and Table 7.2, as well as on the Bayesian LME analyses in Figs. 7.20 and 7.22.
At the same threshold as the wombling networks (τ = 0.7), the Pearson pairwise correlation networks on baseline observations did not show a biological decrease of connections. Specifically, both MCI and AD had 34 connections, 12 more than the HC network with 22 connections. These results suggest that in this work, the wombled networks provided superior connectivity information in comparison to the Pearson pairwise correlation method.
Pearson pairwise correlation networks showed a decrease in overall connectivity across baseline age Groups A to B to C with 47, 38 and 19 total connections respectively, suggesting age dependent loss of connections. However, further investigation into these results is required as the Bayesian LME and wombling models did not support participant age clusters; suggesting there were no age differences in the data (see Figs. 7.22 and 7.23). Furthermore, age Group C comprises of predominately HC and MCI participants, as 18 of the 26 AD participants are in age Groups A and B, which suggests age Group C should not reflect high neurodegeneration estimates.
In addition to these improvements, unlike the descriptive networks from the Pearson pairwise correlation approach, the wombling model provided full posterior distributions which quantified the uncertainty in all possible links. As the Pearson pairwise correlation networks do not take into account the uncertainty of each connection, they cannot correctly estimate the group population networks.
One potential question about the modelling approach proposed in this work is whether the inclusion of additional terms in the mean of the model would significantly change the inference about W. Such terms could include fixed effects to estimate ROI means. If we consider the covariance between data for two ROIs, then, in principle, the correlation structure should be unaffected if, for example, the data were standardised such that data for each ROI had a mean of zero and a variance of one. However, such a simplistic scenario may not be directly applicable to the complex model fitted in this work. Thus we investigated this by extending the wombling model to allow for the estimation of ROI means through the inclusion of fixed effect parameters in the mean of the model. The results showed that the inference about W for the HC group was similar to that presented in Sect. 7.3.2.1 (see section “Wombling Cortical Thickness Estimates at the ROI Level” in Appendix). However, with the MCI and AD groups, this model provided large amounts of uncertainty in the posterior distribution for W, limiting our ability to determine whether inference is impacted by the estimation of ROI means. We believe this is due to the additional (35 fixed effect) parameters included into the model, and this appears to have a major impact in the MCI and AD groups as they have much smaller sample sizes (21 and 26, respectively) compared the HC group (120). These smaller sample sizes appear to have led to a loss of information about the network connection for these groups.
The choice of which wombling model to apply, whether it be the model presented in this work or the extended version which includes ROI means depends on the research questions which one wishes to address and the data which are available. If there are only approximately 20 to 30 individuals in a group of interest and intrahemispheric data are available, then the wombling model presented here can provide meaningful inferences about W but not on ROI means. However if there are over 120 individuals in the groups of interest, then the more complex model with additional ROI parameters would provide joint estimates on the ROI means as well as on the participant and covariance networks. We note that in our analyses it was reassuring to find that the estimates for W were similar in both models.
Further, there is potential for the inference about W to change if important covariates are included into the model. That is, perceived covariance may be due to the influence of unobserved covariate information. Our model can easily incorporate covariates, and indeed it also does this in demonstration through the inclusion of sex, and other covariates could be similarly included (and tested for importance).
4.2.2 Biomarker Estimates
In all groups analysed, the WAIC values for the wombling model were substantially lower compared to the independent Bayesian LME models combined across all ROIs. In this work, this result shows that the wombling model was the preferred parsimonious approach for modelling biomaker estimates compared to the independent analyses. Refer to Table 7.6 for WAIC results. At the participant level estimates of cortical thickness, both approaches demonstrated comparable differences in diagnosed participant clusters as shown in Figs. 7.7 and 7.8 and the Bayesian LME model estimates in Figs. 7.21 and 7.23. These results further demonstrate the flexibility of the wombling approach to jointly analyse cortical networks in addition to biomarker estimates. Above the third quartile of the MCI posterior distribution had a large degree of overlap with the HC posterior distribution (Fig. 7.6). With a probability of 0.49, the posterior distribution of the HC total cortical thickness average is within the upper and lower 95% quantiles of the MCI distribution (Table 7.2). Such a high probability suggests that this overlap could be due a subset of MCI participants in the study who are not on the AD pathway [26, 54]. Hence further investigation into MCI participants further divided into subgroups, such as participants with documented memory complaints, amnestic and non-amnestic is suggested to identify potential non-AD converters.
4.3 Sensitivity Analyses
Two sensitivity analyses were conducted on the application of the Bayesian wombling approach to real data. The first analyses were with respect to the chosen value of ρ, as described in Sect. 7.3.1.1. A number of authors [42, 44, 50] have discussed the limitations of including ρ as an additional parameter to be estimated. Following these recommendations, we fixed ρ = 0.9 throughout all our simulations and application studies, and conducted a sensitivity assessment to evaluate the impact of this choice. Table 7.4 showed that at ρ = 0.9 and the parameters W, β, σ 2 and b i were recovered well. Our results support those of Lu et al. [50], Lee [42] and Lee and Mitchell [44], and we recommend fixing ρ at 0.9 for future wombling model extensions.
The second sensitivity analysis was with respect to the prior specification described in Sect. 7.3.1.1. Since the resulting posterior summaries did not vary considerably based on different prior information, we conclude that our results are relatively robust to the priors specified in this work. The rationale for using vague priors is to ensure that the information in the data primarily governs the results. Alternatively, informative priors may be employed when relevant information is available [12, 70]. In particular, investigating the best use of diffusion or functional network priors (or patient specific networks) would be an interesting future research avenue.
4.4 Limitations and Future Work
The intended application of the wombling model in this work is to demonstrate its utility. Due to the limited sample sizes in this study cohort, the biological interpretation and comparison of each group, in this work, is limited to the total number of connections for each network. Additional cortical network metrics which assess the organisational structure, such as small world topology and characteristic path length [7, 15, 58], is beyond the scope of this work. Future work and clinical application of the wombling model will greatly benefit from matched sampled groups which have similar age ranges, number of replicates, gender and other features known to be associated with the pathology of interest.
A primary drawback of wombling models is the computation time. As mentioned by Bowman et al. [12], limitations of a Bayesian hierarchical framework in spatial analysis include extensive and long computational times, often restricting attention to small ROI analysis or localised voxel-wise analysis. For example, the study by Bowman [11] considered only ROI in the cerebellum to limit the computational extent. Although computationally intensive, our brain wombling approach is not prohibitively so: a single MCMC run of the algorithm can also be computed in approximately a day on a standard desktop computer (see Sect. 7.2.5).
As the dimension of W increases, the parameter space increases dramatically, and this is considered a drawback of the wombling model. For example, our 35 ROI model resulted in a 599-dimensional parameter space, which ran for 500, 000 MCMC iterations. This issue motivates future work to investigate inducing sparsity on W based on prior information as suggested in Babacan et al. [5], as this could potentially reduce the computational burden of the wombling model. Nevertheless, in the present study, the added insight and corroboration between networks and cortical thickness estimates were deemed to be worth the additional computational time.
A second limitation of the present study is that our analysis was restricted to participants with four or more repeated measures, as this affected the ability of the wombling model to converge (results not shown). Such repeated measures can be prohibitive in smaller neuroimaging studies, as patient drop out is a common occurrence. For use of this method in neuroimaging studies with a limited number (< 4) of time points, future work detailing the performance of the wombling model is needed. Nonetheless, our algorithm performed remarkably well for small sample sizes (N AD and N MCI < 21 < 35 ROI) on data where all participants had repeated measures. We conjecture that the probabilistic networks from the wombling model will better distinguish between a link and the absence of a link (i.e. network probabilities will be closer to zero or one), and result in narrower credible intervals on biomarker estimates as the sample size increases.
The final limitation of the present study was the relative simplicity of the two layered linear random effects model, as shown in Fig. 7.2. This is not a fixed limitation of the approach presented here; the hierarchical Bayesian framework is capable of handling complex models, such as models with two or more nested layers to account for complex data structures [28, 29]. Extensions of this nature will allow the modelling of cerebral morphological features across multiple ROIs over participants’ age, and expand our spatial approach into a spatio-temporal domain.
4.5 Conclusion
In this work, we have demonstrated the advantages of the Bayesian brain wombling approach applied in the neuroimaging field over state-of-the-art independent analyses. The ability of the wombling model to recover the connectivity and biomarker effect estimates give confidence on our results from the cohort study. Taking into account of the uncertainty of each network, the population wombled networks across diagnosis levels from healthy to Alzheimer’s disease showed a loss of connectivity (posterior probability \(\geqslant 0.7\)). Compared to independent LME models, we found that both approaches estimated cortical thickness progressively along the dementia pathway. Although applied here to cortical thickness, this method can be applied to other types of neuroimaging data, unifying existing previously independent analyses that are aimed at exploring the same underlying biological system. This powerful analysis tool provides the potential to extend our understanding of the human brain functions and effects of brain disorders on both local and network scale.
References
O. Acosta, P. Bourgeat, M.A. Zuluaga, J. Fripp, O. Salvado, S. Ourselin, A.D.N. Initiative, et al., Automated voxel-based 3D cortical thickness measurement in a combined Lagrangian–Eulerian PDE approach using partial volume maps. Med. Image Anal. 13(5), 730–743 (2009)
A. Adaszewski, J. Dukart, F. Kherif, R. Frackowiak, B. Draganski, How early can we predict Alzheimer’s disease using computational anatomy. Neurobiol. Aging 34(12), 2815–2826 (2013)
A. Alexander-Bloch, J. N. Giedd, et al., Imaging structural co-variance between human brain regions. Nat. Rev. Neurosci. 14(5), 322 (2013)
C. Anderson, D. Lee, N. Dean, Identifying clusters in Bayesian disease mapping. Biostatistics 15(3), 457–469 (2014)
S.D. Babacan, M. Luessi, R. Molina, A.K. Katsaggelos, Sparse Bayesian methods for low-rank matrix estimation. IEEE Trans. Signal Process. 60(8), 3964–3977 (2012)
A. Bakkour, J.C. Morris, D.A. Wolk, B.C. Dickerson, The effects of aging and Alzheimer’s disease on cerebral cortical anatomy: specificity and differential relationships with cognition. NeuroImage 76, 332–344 (2013)
D.S. Bassett, E.T. Bullmore, Small-world brain networks revisited. Neuroscientist (2016). https://doi.org/10.1177/1073858416667720
D.S. Bassett, E. Bullmore, B.A. Verchinski, V.S. Mattay, D.R. Weinberger, A. Meyer-Lindenberg, Hierarchical organization of human cortical networks in health and schizophrenia. J. Neurosci. 28(37), 9239–9248 (2008)
J. Bernal-Rusiel, D.N. Greve, M. Reuter, B. Fischl, M.R. Sabuncu, Statistical analysis of longitudinal neuroimage data with Linear Mixed Effects models. NeuroImage 66, 249–60 (2013)
P. Bourgeat, G. Chetelat, V. Villemagne, J. Fripp, P. Raniga, K. Pike, O. Acosta, C. Szoeke, S. Ourselin, D. Ames, et al., β-Amyloid burden in the temporal neocortex is related to hippocampal atrophy in elderly subjects without dementia. Neurology 74(2), 121–127 (2010)
F.D. Bowman, Spatiotemporal models for region of interest analyses of functional neuroimaging data. J. Am. Stat. Assoc. 102(478), 442–453 (2007)
F.D. Bowman, B. Caffo, S.S. Bassett, C. Kilts, A Bayesian hierarchical framework for spatial modeling of fMRI data. NeuroImage 39(1), 146–156 (2008)
M.R. Brier, J.B. Thomas, A.M. Fagan, J. Hassenstab, D.M. Holtzman, T.L. Benzinger, J.C. Morris, B.M. Ances, Functional connectivity and graph theory in preclinical Alzheimer’s disease. Neurobiol. Aging 35(4), 757–768 (2014)
S.P. Brooks, A. Gelman, General methods for monitoring convergence of iterative simulations. J. Comput. Graph. Stat. 7(4), 434–455 (1998)
E. Bullmore, O. Sporns, Complex brain networks: graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10(3), 186–198 (2009)
R.J. Caselli, A.C. Dueck, D. Osborne, M.N. Sabbagh, D.J. Connor, G.L. Ahern, L.C. Baxter, S.Z. Rapcsak, J. Shi, B.K. Woodruff, et al., Longitudinal modeling of age-related memory decline and the APOE ε4 effect. New Engl. J. Med. 361(3), 255–263 (2009)
M.I. Cespedes, J. Fripp, J.M. McGree, C.C. Drovandi, K. Mengersen, J.D. Doecke, Comparisons of neurodegeneration over time between healthy ageing and Alzheimer’s disease cohorts via Bayesian inference. BMJ Open, 7(2), e012174 (2017)
Z.J. Chen, Y. He, P. Rosa-Neto, G. Gong, A.C. Evans, Age-related alterations in the modular organization of structural cortical network by using cortical thickness from MRI. NeuroImage 56(1), 235–245 (2011)
S. Chen, F.D. Bowman, H.S. Mayberg, A Bayesian hierarchical framework for modeling brain connectivity for neuroimaging data. Biometrics 72(2), 596–605 (2016). https://doi.org/10.1111/biom.12433
S. Chib, E. Greenberg, Understanding the Metropolis-Hastings algorithm. Am. Stat. 49(4), 327–335 (1995)
B.C. Dickerson, A. Bakkour, D.H. Salat, E. Feczko, J. Pacheco, D.N. Greve, F. Grodstein, C.I. Wright, D. Blacker, H.D. Rosas, et al., The cortical signature of Alzheimer’s disease: regionally specific cortical thinning relates to symptom severity in very mild to mild AD dementia and is detectable in asymptomatic amyloid-positive individuals. Cereb. Cortex 19(3), 497–510 (2009)
V. Doré, V.L. Villemagne, P. Bourgeat, J. Fripp, O. Acosta, G. Chetélat, L. Zhou, R. Martins, K.A. Ellis, C.L. Masters, et al., Cross-sectional and longitudinal analysis of the relationship between Aβ deposition, cortical thickness, and memory in cognitively unimpaired individuals and in Alzheimer disease. JAMA Neurol. 70(7), 903–911 (2013)
A.-T. Du, N. Schuff, J.H. Kramer, H.J. Rosen, M.L. Gorno-Tempini, K. Rankin, B.L. Miller, M.W. Weiner, Different regional patterns of cortical thinning in Alzheimer’s disease and frontotemporal dementia. Brain 130(4), 1159–1166 (2007)
K.A. Ellis, A.I. Bush, D. Darby, D. De Fazio, J. Foster, P. Hudson, N.T. Lautenschlager, N. Lenzo, R.N. Martins, P. Maruff, et al., The Australian Imaging, Biomarkers and Lifestyle (AIBL) study of aging: methodology and baseline characteristics of 1112 individuals recruited for a longitudinal study of Alzheimer’s disease. Int. Psychogeriatr. 21(04), 672–687 (2009)
Y. Fan, F. Shi, J.K. Smith, W. Lin, J.H. Gilmore, D. Shen, Brain anatomical networks in early human brain development. NeuroImage 54(3), 1862–1871 (2011)
F.L. Ferreira, S. Cardoso, D. Silva, M. Guerreiro, A. de Mendonça, S.C. Madeira, Improving prognostic prediction from mild cognitive impairment to Alzheimer’s disease using genetic algorithms, in Alzheimer’s Disease: Advances in Etiology, Pathogenesis and Therapeutics, chapter 14, ed. by K. Iqbal, S.S. Sisodia, B. Winbald (Springer, New York, 2017)
B. Fischl, A.M. Dale, Measuring the thickness of the human cerebral cortex from magnetic resonance images. Proc. Natl. Acad. Sci. 97(20), 11050–11055 (2000)
A. Gelman, J. Hill, Data Analysis Using Regression and Multilevel/Hierarchical Models (Cambridge University Press, Cambridge, 2006))
A. Gelman, J.B. Carlin, H.S. Stern, D.B. Dunson, A. Vehtari, D.B. Rubin, Bayesian Data Analysis, 2nd edn. (CRC Press, Boca Raton, 2013)
A. Gelman, J. Hwang, A. Vehtari, Understanding predictive information criteria for Bayesian models. Stat. Comput. 24(6), 997–1016 (2014)
A. Goldstone, S.D. Mayhew, I. Przezdzik, R.S. Wilson, J.R. Hale, A.P. Bagshaw, Gender specific re-organization of resting-state networks in older age. Front. Aging Neurosci. 8, 285 (2016)
C. Gössl, D.P. Auer, L. Fahrmeir, Bayesian spatiotemporal inference in functional magnetic resonance imaging. Biometrics 57(2), 554–562 (2001)
A.R. Groves, M.A. Chappell, M.W. Woolrich, Combined spatial and non-spatial prior for inference on MRI time-series. NeuroImage 45(3), 795–809 (2009)
B. Guillaume, X. Hua, P.M. Thompson, L. Waldorp, T.E. Nichols, Fast and accurate modelling of longitudinal and repeated measures neuroimaging data. NeuroImage 94, 287–302 (2014)
Y. Guo, F. DuBois Bowman, C. Kilts, Predicting the brain response to treatment using a Bayesian hierarchical model with application to a study of schizophrenia. Hum. Brain Mapp. 29(9), 1092–1109 (2008)
L.M. Harrison, G.G. Green, A Bayesian spatiotemporal model for very large data sets. NeuroImage 50(3), 1126–1141 (2010)
Y. He, Z. Chen, A. Evans, Structural insights into aberrant topological patterns of large-scale cortical networks in Alzheimer’s disease. J. Neurosci. 28(18), 4756–4766 (2008)
M. Hinne, T. Heskes, M.A.J. van Gerven, Bayesian inference of whole-brain networks. 1–10 (2012). arXiv:1202.1696
D. Holland, R.S. Desikan, A.M. Dale, L.K. McEvoy, A.D.N. Initiative, et al., Rates of decline in Alzheimer disease decrease with age. PloS One 7(8), e42325 (2012)
C.R. Jack, H.J. Wiste, S.D. Weigand, D.S. Knopman, M.M. Mielke, P. Vemuri, V. Lowe, M.L. Senjem, J.L. Gunter, D. Reyes, et al., Different definitions of neurodegeneration produce similar amyloid/neurodegeneration biomarker group findings. Brain 138(12), 3747–3759 (2015)
R.J. Janssen, M. Hinne, T. Heskes, M.A.J. van Gerven, Quantifying uncertainty in brain network measures using Bayesian connectomics. Front. Comput. Neurosci. 8, 126 (2014)
D. Lee, A comparison of conditional autoregressive models used in Bayesian disease mapping. Spatial Spatio-temporal Epidemiol. 2(2), 79–89 (2011)
D. Lee, R. Mitchell, Boundary detection in disease mapping studies. Biostatistics 13(3), 415–426 (2012)
D. Lee, R. Mitchell, Locally adaptive spatial smoothing using conditional auto-regressive models. J. R. Stat. Soc. Ser. C: Appl. Stat. 62(4), 593–608 (2013)
B.G. Leroux, X. Lei, N. Breslow, Estimation of disease rates in small areas: a new mixed model for spatial dependence, in Statistical Models in Epidemiology, the Environment, and Clinical Trials, ed. by H.M. Elizabeth, D. Berry (Springer, New York, 2000), pp. 179–191
Y. Li, Y. Wang, G. Wu, F. Shi, L. Zhou, W. Lin, D. Shen, A.D.N. Initiative, et al., Discriminant analysis of longitudinal cortical thickness changes in Alzheimer’s disease using dynamic and network features. Neurobiol. Aging 33(2), 427-e15 (2012)
X. Li, F. Pu, Y. Fan, H. Niu, S. Li, D. Li, Age-related changes in brain structural covariance networks. Front. Hum. Neurosci. 7, 98 (2013)
K. Liu, Z.L. Yu, W. Wu, Z. Gu, Y. Li, S. Nagarajan, Bayesian electromagnetic spatio-temporal imaging of extended sources with Markov Random Field and temporal basis expansion. NeuroImage 139, 385–404 (2016)
H. Lu, B.P. Carlin, Bayesian areal wombling for geographical boundary analysis. Geograph. Anal. 37(3), 265–285 (2005)
H. Lu, C.S. Reilly, S. Banerjee, B.P. Carlin, Bayesian areal wombling via adjacency modeling. Environ. Ecol. Stat. 14(4), 433–452 (2007)
P. McCullagh, J.A. Nelder, Generalized Linear Models, vol. 37 (CRC Press, Boca Raton, 1989)
N. Metropolis, A.W. Rosenbluth, M.N. Rosenbluth, A.H. Teller, E. Teller, Equation of state calculations by fast computing machines. J. Chem. Phys. 21(6), 1087–1092 (1953)
M.F. Miranda, H. Zhu, J.G. Ibrahim, Bayesian spatial transformation models with applications in neuroimaging data. Biometrics 69(4), 1074–1083 (2013)
R.C. Petersen, Mild cognitive impairment as a diagnostic entity. J. Intern. Med. 256(3), 183–194 (2004)
A. Pfefferbaum, T. Rohlfing, M.J. Rosenbloom, W. Chu, I.M. Colrain, E.V. Sullivan, Variation in longitudinal trajectories of regional brain volumes of healthy men and women (ages 10 to 85 years) measured with atlas-based parcellation of MRI. NeuroImage 65, 176–193 (2013)
R Core Team, R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2015)
C. Robert, G. Casella, Monte Carlo Statistical Methods. Springer Texts in Statistics (Springer, New York, 2010)
M. Rubinov, O. Sporns, Complex network measures of brain connectivity: uses and interpretations. NeuroImage 52(3), 1059–1069 (2010)
W.W. Seeley, R.K. Crawford, J. Zhou, B.L. Miller, M.D. Greicius, Neurodegenerative diseases target large-scale human brain networks. Neuron 62(1), 42–52 (2009)
S.L. Simpson, F. Bowman, P.J. Laurienti, Analyzing complex functional brain networks: fusing statistics and network science to understand the brain. Stat. Surv. 7, 1 (2013)
M.R. Sinke, R.M. Dijkhuizen, A. Caimo, C.J. Stam, W.M. Otte, Bayesian exponential random graph modeling of whole-brain structural networks across lifespan. NeuroImage 135, 79–91 (2016)
A.B. Storsve, A.M. Fjell, C.K. Tammes, L.T. Westlye, K. Overbye, H.W. Aasland, K.B. Walhovd, Differential longitudinal changes in cortical thickness, surface area and volume across the adult life span: regions of accelerating and decelerating change. J. Neurosci. 34, 8488–8498 (2014)
N. Tzourio-Mazoyer, B. Landeau, D. Papathanassiou, F. Crivello, O. Etard, N. Delcroix, B. Mazoyer, M. Joliot, Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. NeuroImage 15(1), 273–289 (2002)
K. Van Leemput, F. Maes, D. Vandermeulen, P. Suetens, Automated model-based tissue classification of MR images of the brain. IEEE Trans. Med. Imaging 18(10), 897–908 (1999)
V. Villemagne, S. Burnham, P. Bourgeat, B. Brown, K. Ellis, O. Salvado, C. Szoeke, S. Macaulay, R. Martins, P. Maruff, D. Ames, C. Rowe, C. Masters, Amyloid β deposition, neurodegeneration and cognitive decline in sporadic Alzheimer’s disease. Lancet Neurol. 12, 357–367 (2013)
S. Watanabe, A widely applicable Bayesian information criterion. J. Mach. Learn. Res. 14, 867–897 (2013)
M.W. Weiner, D.P. Veitch, P.S. Aisen, L.A. Beckett, N.J. Cairns, R.C. Green, D. Harvey, C.R. Jack, W. Jagust, E. Liu, et al., The Alzheimer’s Disease Neuroimaging Initiative: a review of papers published since its inception. Alzheimer’s Dementia 9(5), e111–e194 (2013)
M.W. Woolrich, T.E. Behrens, C.F. Beckmann, M. Jenkinson, S.M. Smith, Multilevel linear modelling for FMRI group analysis using Bayesian inference. NeuroImage 21(4), 1732–1747 (2004)
L. Xu, T.D. Johnson, T.E. Nichols, D.E. Nee, Modeling inter-subject variability in fMRI activation location: a Bayesian hierarchical spatial model. Biometrics 65(4), 1041–1051 (2009)
W. Xue, F.D. Bowman, A.V. Pileggi, A.R. Mayer, A multimodal approach for determining brain networks by jointly modeling functional and structural connectivity. Front. Comput. Neurosci. 9, 22 (2015)
Z. Yao, Y. Zhang, L. Lin, Y. Zhou, C. Xu, T. Jiang, A.D.N. Initiative, et al., Abnormal cortical networks in mild cognitive impairment and Alzheimer’s disease. PLoS Comput. Biol. 6(11), e1001006 (2010)
G. Ziegler, W.D. Penny, G.R. Ridgway, S. Ourselin, K.J. Friston, A.D.N. Initiative, et al., Estimating anatomical trajectories with Bayesian mixed-effects modeling. NeuroImage 121, 51–68 (2015)
Acknowledgements
We wish to thank the Australian Imaging, Biomarkers and Lifestyle longitudinal study of ageing (www.aibl.csiro.au), including all the clinicians, scientists, participants and their families. MIC was jointly funded by the Research Training Program (RTP), the Commonwealth Scientific and Industrial Research Organisation (CSIRO) Health and Biosecurity division, and supported by the ARC Centre of Excellence for Mathematical & Statistical Frontiers (ACEMS). CCD was supported by an Australian Research Council’s Discovery Early Career Researcher Award funding scheme DE160100741. Computational resources and services used in this work were provided by the High Performance Computing (HPC) and Research Support Group, Queensland University of Technology, Brisbane, Australia.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Appendices
Appendix: Methods and Applications
Additional material to supplement simulation study results, posterior diagnostic checks, wombling ROI cortical thickness estimates at the population and participant levels, independent Bayesian mixed effect model results, WAIC values and Pearson correlation networks can be found in this Appendix. R code to implement the wombling model can be found at the following GitHub repository https://github.com/MarcelaCespedes/Brain_wombling.
Simulation Study
The simulation study described in Sect. 7.2.3 provided a thorough assessment of the Bayesian brain wombling algorithm. The four scenarios in the simulation study are; contiguous balanced (each person had an equal number of replicates) and unbalanced (the number of replicates varied per person), and a structured balanced and unbalanced designs. The results for fixed effect parameters β and residual variance σ 2 are shown in Table 7.3. While the results for the structured configuration show a slightly lower recovery of fixed effect parameters, they do not represent a potential biological configuration. Hence performance of the wombling algorithm is better assessed on the contiguous configuration, whose performance of the recovery of the parameters is approximately 95%.
As discussed in Sect. 7.2.3.1, spatial scale variance \(\sigma ^2_s\) is a biased estimate and was not recovered in our simulation study.
Figure 7.9 shows the histograms on the percentage of the recovered random effects for each scenario. The simulation study comprised of 50 independently simulated data sets for each scenario, each data set consisted of I = 100 simulated participants, each with K = 35 ROI resulting in 3500 random effects per simulated data set to estimate. Overall we can see that there is approximately 95% recovery of the random effects for each scenario.
As ρ in Model (7.1) is a fixed value, we investigated the effect recovering the parameters in the structured scenario for ρ values [0.85, 0.9, 0.95, 0.99]. Table 7.4 summarises the results.
Contact the author for additional simulation study results such as MCMC convergence checks, estimation of credible intervals, and posterior predictive plots.
Posterior Diagnostic Checks for AIBL Data Set
Posterior predictive plots for each AIBL group analysed were used to assess goodness-of-fit for each wombled model. The plots in Figs. 7.10, 7.11 and 7.12 show the expected mean of the data was recovered well, however there is a slight overestimation of the variance, as the proportion of predicted values inside the 95% credible intervals is slightly over 0.95. However these results show our models adequately captured the uncertainty in the data.
Table 7.5 shows the Gelman-Rubin diagnostic, upper 95% credible interval for convergence checks of the four chains for β 0, β 1, σ 2 and \(\sigma ^2_s\).
Wombling Cortical Thickness Estimates at the ROI Level
As discussed in Sect. 7.4.2.1, we investigated an adaptation to the wombling model to account for ROI means via fixed effect parameters. The extended model is of the form
Where the response (y irk), spatial random effects (b ik), residual (σ 2) and spatial scale variance \((\sigma ^2_s)\) terms are the same as those presented in Sect. 7.2.2. The precentral gyrus is the baseline ROI whose cortical thickness (in mm) is estimated by β 0. The fixed effect parameter β k−1 estimates the deviation of ROI k away from β 0 when the binary indicator variable R k is equal to one. Estimation of β is attained by the same conditional distribution described in Sect. 7.2.2, with minor modifications to account for the design matrix R rather than X. Figures 7.17 and 7.18 show participant specific cortical thickness estimates as caterpillar plots (β k + b ik) colour coded for diagnosis and age groups respectively. Figure 7.13 shows the posterior means of W for HC (top), MCI (middle) and AD (bottom) groups. While the posterior mean for the HC group is similar that in Fig. 7.4, with the same 36 links present in both networks and 468 absent connections in common, the matrices for MCI and AD group show the probability of each link is close to 0.5. We believe that the reason for this is because the HC group has a substantially larger sample size (120 individuals) compared to the MCI and AD groups (with 21 and 26 individuals respectively). Hence, the more complex model in Eq. (7.9) requires data with larger sample sizes, compared to the original wombling model, in order to derive meaningful W estimates.
Figure 7.14 shows the marginal posterior densities for the ROI means for 35 regions. These results resemble the independent Bayesian LME ROI estimates in Fig. 7.20, particularly for ROIs associated with early onset of AD such as the inferior, middle and superior temporal gyrus, posterior cingulate gyrus.
Wombling Cortical Thickness Estimates at the Participant Level
As described in Sect. 7.2.4.4 and discussed in Sect. 7.3.2.4, the wombling model derived participant specific estimates on all ROIs. Figures 7.15 and 7.16 shows the posterior means and 95% credible intervals (as error bars) for each participant.
APOE Wombling Results
Carriers of the Apolipoprotein (APOE) ε4 gene have known to be at higher risk of developing AD compared to non-carriers, hence in neuroimaging studies, it is a key biomarker to investigate. For exploration purposes, we applied the wombling model on AIBL data divided into APOE ε4 carrier and non-carrier groups. Figures 7.17, 7.18 and 7.19 show the cortical networks, global estimates across all ROI and participant specific rankings for key AD regions as described in Sect. 7.2.4.3.
There were no strong differences APOE ε4 carrier and non-carrier groups in any of the ROI. We believe the reason for this is due to APOE ε carrier and non-carrier groups comprising of participants across the entire spectrum (HC, MCI and AD), large variety of ages and many other AD biomarkers, making it difficult to assess the deterioration differences associated with the APOE ε gene. Unfortunately due to our low sample size, we did not have sufficient data to investigate more meaningful biomarker groups such as APOE ε4 carrier and non-carrier groups that were clinically diagnosed as HC or AD.
Bayesian Linear Mixed Effect Models on Each ROI
As described in Sect. 7.2.4.4 and discussed in Sect. 7.3.2.4, Bayesian linear mixed effect models were independently applied to each ROI on groups; diagnosis levels HC, MCI and AD and age groups A, B and C. For exploration purposes we also investigated APOE ε4 allele carriers and non-carriers. All models were of the form
In order to make the models comparable with the wombling approach, covariate x i is gender as described in Sect. 7.2.3.1, with x i = 1 for male and 0 otherwise. The residual variance prior for σ 2 and the random effects prior, \(\sigma ^2_0\), is the same as discussed in Sect. 7.2.3.1. Similarly, the prior for the intercept effect μ 0 is also relatively vague with a N(0, 10) distribution.
Figures 7.20, 7.21, 7.22, 7.23, 7.24 and 7.25 show the marginal posterior mean population distributions and participants ranked according to posterior means with 95% credible interval.
WAIC Results
As described in Sect. 7.2.4.4, we applied the WAIC criterion on the wombled and independent Bayesian LME models to assess model choice. Table 7.6 shows the results of the WAIC for the wombling model applied to each group, and the combined WAIC criterion for the independent Bayesian LME analyses for each region.
Pearson Correlation Networks for Each Group
Cortical networks derived by Pearson’s pairwise correlation networks for each group are shown in Figs. 7.26, 7.27, 7.28, 7.29, 7.30, and 7.31. As Pearson’s pairwise networks do not accommodate the repeated measure structure of the data, we derived networks at both baseline (independent and identically distributed (IID) observations) as well as on the whole data, with repeated measures treated as IID to investigate any potential differences.
Rights and permissions
Copyright information
© 2020 The Editor(s) (if applicable) and The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Cespedes, M.I. et al. (2020). A Bayesian Hierarchical Approach to Jointly Model Cortical Thickness and Covariance Networks. In: Mengersen, K., Pudlo, P., Robert, C. (eds) Case Studies in Applied Bayesian Data Science. Lecture Notes in Mathematics, vol 2259. Springer, Cham. https://doi.org/10.1007/978-3-030-42553-1_7
Download citation
DOI: https://doi.org/10.1007/978-3-030-42553-1_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-42552-4
Online ISBN: 978-3-030-42553-1
eBook Packages: Mathematics and StatisticsMathematics and Statistics (R0)