Abstract
Retinal imaging is a valuable tool in diagnosing many eye diseases but offers opportunities to have a direct view to central nervous system and its blood vessels. The accurate measurement of the characteristics of retinal vessels allows not only analysis of retinal diseases but also many systemic diseases like diabetes and other cardiovascular or cerebrovascular diseases. This analysis benefits from precise blood vessel characterization. Automatic machine learning methods are typically trained in the supervised manner where a training set with ground truth data is available. Due to difficulties in precise pixelwise labeling, the question of the reliability of a trained model arises. This paper addresses this question using Bayesian deep learning and extends recent research on the uncertainty quantification of retinal vasculature and artery-vein classification. It is shown that state-of-the-art results can be achieved by using the trained model. An analysis of the predictions for cases where the class labels are unavailable is given.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
A number of eye and systemic diseases influence the vasculature of the retina in different ways. The blood vessel characteristics in retinal images may provide visible evidence about numerous diseases such as hypertensive retinopathy, diabetic retinopathy, as well as other cardio- and cerebrovascular diseases [12]. The related characteristics include the shape and size of retinal vessels, arteriovenous ratio and arteriovenous crossing [14]. These characteristics may be obtained by using blood vessel segmentation masks produced by automatic machine learning techniques [5].
The topic of blood vessels segmentation is well studied by the community [1]. However, the artery-vein (AV) classification task remains challenging not only for machines, but also for humans. Despite the fact that discriminative features based on color and geometry are described, it is still difficult to distinguish arteries from veins [14] due to imperfect imaging conditions and limited visibility of the retinal blood vessels.
Recently, deep convolutional neural networks have become a common trend for retinal vasculature segmentation and AV classification because of the ability to automatically learn meaningful features. Welikala et al. [16] proposed a method based on a convolutional neural network (CNN) classifying arteries and veins in a patch-wise manner. The authors considered the problem as a multi-class classification task placing a softmax layer at the end of the network. The UK Biobank database was used from which 100 images were labeled and classification accuracy of 82.26% for arteries and veins was reported. Girard et al. [5] proposed to use a modified U-Net [15] with likelihood score propagation in the minimum spanning tree effectively utilizing information about the global vessel topology. The approach was tested on the DRIVE data set [8] and it achieved 94.93% accuracy for the AV classification. Badawi et al. [2] proposed to train a CNN with multiloss function consisting of pixelwise cross entropy loss and segment-level loss to overcome training issues appearing because of inconsistent thickness of blood vessels. The authors also created a new data set consisting of labeled subsets of EPIC and MESSIDOR [3] data sets and classification accuracy of 96.5% was reported. Hemelings et al. [7] applied the U-Net architecture for the task of AV classification stating the problem as a multi-class classification problem predicting labels for four classes (background, vein, artery, and unknown) with classification accuracy of 94.42% and 94.11% for arteries and veins, respectively. Zhang et al. [18] proposed cascade refined U-net which modifies the original model with multi-scale loss training and includes sub-networks for simultaneous AV and blood vessel segmentation. The authors achieved 97.27% arteriovenous classification accuracy evaluated on the automatically detected vessels.
In this work, a multi-label classification approach is considered with the uncertainty quantification experiments presented. Our approach is most similar to the method proposed by Zhang et al. [18] in a way how three-component loss is used. The main difference is that in this work, classification of arteries and veins are not conditioned on blood vessel predictions, but vessel labels are conditioned on arteries and veins. Using the multi-label classification approach, there is no need to separately model the AV crossings and background. To the best of authors’ knowledge, this work is the first presenting uncertainty quantification experiments for the of AV classification. For the experiments, the RITE data set is utilized.
2 Data and Methods
2.1 DRIVE and RITE Data Sets
The DRIVE database is a common benchmark for the retinal blood vessel segmentation task [8]. It contains 20 train and 20 test images with two sets of manual blood vessel segmentations. The RITE data set [9] extends DRIVE with an AV reference standard containing four types of labels: arteries (red), veins (blue), overlapping (green), and uncertain vessels (white). An example test image is shown in Fig. 1.
2.2 AV Classification
Let \(f\) be a model with parameters \(\pmb {\theta }\) that maps an input image \(\mathbf {x}\) to a map of logits with the same spatial dimensionality as the original image:
Given predicted logits \(\mathbf {\hat{y}} = \left[ \hat{y}_{\text {artery}} \ \hat{y}_{\text {vein}} \right] \), probabilities of assigning labels to arteries and veins can be calculated as follows:
In the multi-label setup, the same pixel can be classified with both artery and vein labels, which is meaningful in the case of AV crossings. A vessel probability label can then be naturally inferred by a simple formula:
Since the data set contains the masks for both the AV classification and blood vessel segmentation, it is possible to state the following optimization problem

where \(\mathcal {L}\) denotes the binary cross entropy loss for the corresponding labels. This way even if the labels for arteries and veins are not given for uncertain vessel labels, it is possible to enforce a model to predict correct labels for the blood vessels.
2.3 Aleatoric and Epistemic Uncertainties
The approach described in the previous section gives only point estimates for the label probabilities and the model parameters are considered to be deterministic. In order to better capture imperfect data labeling and image noise, one can consider the model outputs and the parameters to be random variables. The first approach captures the heteroscedastic aleatoric uncertainty that depends on the input data, whereas the second represents the epistemic uncertainty that models a distribution of the learned parameters. More detailed explanations for the uncertainties can be found in [13] and [4]. In this work, a brief explanation for the AV classification task is given below.
Aleatoric uncertainty can be captured by modifying the original model to predict the mean and standard deviations of logits:
In order to predict standard deviations, a second layer similar and parallel to the one used for logits is added to the output of the network. In order to ensure that the predicted standard deviations are positive, an additional absolute value activation is added to the output of the layer. The probabilities of the labels can then be calculated as follows:
where \(\odot \) stands for the Hadamard product and \(\pmb {\epsilon }\) are sampled during inference.
The main inference scheme for AV remains the same with the exception that instead of a point estimate, the model now yields \(N_A\) samples that are then used to calculate the loss (5). The final minimized loss is just an average over the predicted losses for each sample.
Epistemic uncertainty can be captured by considering the model parameters to be a random variable and considering the following posterior predictive:
where \(\mathcal {D}\) denotes a data set of input-output pairs. Typically, the parameter’s posterior \(p \left( \pmb {\theta } \,\vert \,\mathcal {D} \right) \) for complex models such as deep neural networks is intractable and variational approximations are used. The posterior in (8) can be replaced by a simpler distribution \(q\left( \pmb {\theta }\right) \) and the training procedure can then be formulated as the minimization of the Kullback-Leibler divergence between the true posterior and the approximation.
In this work, the model \(f\) is parameterized as a dense fully-convolutional network (Dense-FCN) and Monte-Carlo dropout [4] is used for the variational approximation. The description of the utilized architecture is given below.
2.4 Architecture
The architecture utilized in this work is a Dense-FCN. It has been shown that Dense-FCNs have less parameters and may outperform other fully-convolutional network (FCN) architectures in a variety of different segmentation tasks [11]. Here we adapt the Dense-FCN architecture for the AV classification tasks.
The main building block of Dense-FCN is a dense convolutional block (DCB) where the input of each layer is a concatenation of the outputs of the previous layers. The block consists of repeating batch normalization (BN), ReLU, convolution and dropout \(p = 0.5\) layers resulting in \(g\) feature maps (growth rate).
The main concept of Dense-FCN is similar to other encoder-decoder architectures in the sense that the input is first compressed to a hidden representation by the downsampling part, and then the segmentation masks are recovered by an upsampling part. The downsampling part consists of DCBs and downsampling transitions with skip connections to the upsampling part. The upsampling part consists of DCBs and upsampling transitions. An example of two blocks in downsampling and upsampling paths of a Dense-FCN is given in Fig. 2. The architectural parameters used are given below:
-
Growth rate for all DCBs: \(g = 16\).
-
Downsampling path consists of five DCBs with depths \(D_{\text {down}} = \left[ 4, 5, 7, 10, 12, 15\right] \).
-
Upsampling also consists of five DCBs with depths \(D_{\text {up}} = \left[ 12, 10, 7, 5, 4\right] \).
-
The first and last convolution layers are the same as in Fig. 2.
Dense-FCN architecture: Dense stands for a DCB; C is a tensor concatenation; H is a block consisting of BN, ReLU and a convolutional layer with growth rate \(g\); Down is a transition down block with \(F\) output feature maps; Up is a transition up with \(F\) output feature maps and \(2 \times 2\) stride. logits std denotes standard deviations of logits.
2.5 Image Preprocessing
It was noticed in the experimental part of the work that simple preprocessing involving contrast enhancement and channel normalization improves the convergence and performance of the trained models. First, contrast-limited adaptive histogram equalization [19] with the clip limit of 2 and the grid size of \(8 \times 8\) is applied and then each image channel is normalized to values between 0 and 255. The preprocessing scheme was used to reduce the effects of uneven illumination fields of the channel images (Fig. 3).
2.6 Training Details
The Dense-FCN was pretrained for 200 epochs with 1000 steps per epoch on random patches \(224 \times 224\) with the batch size equal to 5. Then it was fine-tuned for 50 epochs with 500 steps per epoch on full size images with the batch size equal to 1.
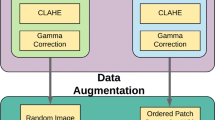
The weights were initialized using HeNormal [6]. In addition to dropout, \(l2\) regularization with the weight decay factor \(10^{-4}\) was used. As the optimizer, Adadelta [17] with the learning rate \(l=1\) and the decay rate \(\rho = 0.95\) was used for both the pretraining and fine-tuning. The learning rate was dropped by a factor of 10 if the training loss was not decreased by 0.005 for 10 epochs. Data augmentation by using flipping, reflecting and rescaling (with scale rates 0.8 and 1.2) was applied in both cases. During the fine-tuning stage, the images were randomly padded to size \(608 \times 608\) so that the size is divisible by 32 and could be properly compressed by the downsampling path. The parameter values were determined empirically based on initial experiments with the RITE database.
3 Experiments and Results
3.1 Training and Evaluation Strategies
Considering the given reference standard, the question arises of how to use the uncertain class labels and its effect on the final training results. Possible ways for utilizing this information are to consider these pixels to be arteries and veins simultaneously including uncertain (IU), or to exclude them from the training completely excluding uncertain (EU). In this work, a comparison of both training strategies is provided. The crossing labels are considered to be veins and arteries simultaneously. Both strategies are evaluated against the reference standard with excluded uncertain labels, and the vessels classification metrics are given by evaluating against the reference standard provided by the second expert.
Since the AV classification problem stated being multilabel, binary classification metrics were calculated for each class separately: area under receiver operating characteristic curve (ROC-AUC), accuracy, sensitivity and specificity.
During the inference stage, the model parameters are sampled 100 times and the number of inferred samples is \(N_A = 50\). The final posterior predictive mean is calculated over all predicted samples, and the outputs aleatoric uncertainty \(U_A\) and epistemic uncertainty \(U_E\) are calculated as in [10]:
where \( \mathbb {E} \) and \( \mathbb {V} \) denote expectation and variance, respectively.
3.2 Experimental Results
The receiver operating characteristic (ROC) curves calculated after training with both strategies are shown in Fig. 4. The corresponding performance metrics are given in Tables 1 and 2. From the tables, it is clear that the AV classification performances are high, not far from the vessel pixel classification performance. Including uncertain labels into the training set leads to reduced classification accuracy for arteries and veins, but it slightly improves the performance of vessel classification. It is also clear that the Including uncertain strategy increases classification sensitivity, since the training procedure now takes all labeled vessels into account during the AV inference stage.
The segmentation results for two example images from the test set are illustrated in Fig. 5. Comparing the results for the training strategies shows that the network trained with the EU strategy tends to be more discriminative for arteries and veins in the areas closer to the optic disc. The common issue for both strategies is the learned bias about the thin vessels being arteries and incapacity to capture connectivity patterns of the predicted segmentation masks inferring vein branches to be arteries.
The aforementioned problems can also be visualized as predicted epistemic and aleatoric uncertainties which are presented in Fig. 6 for the same images shown in Fig. 5. From the figure, it is clear that the epistemic uncertainty is larger near the optic disc where blood vessels cross. Further away from the optic disc it is concentrated mostly on the vessels’ edges with a pattern similar to the one of the aleatoric uncertainty. Similar observations can be made from Fig. 7 where the uncertainties are compared for the two training strategies. The regions of highest uncertainty include vessel crossings and thin vessels even in the case correct classification.
3.3 Comparison with the State of the Art
The Table 3 shows a comparison of the proposed method with recently proposed methods. It is troublesome to directly compare the methods, since the evaluation methods and metrics used by different authors vary. The method proposed by Zhang et al. [18] is clearly superior compared to all the other methods, including the method studied in this work, but the authors use 5-fold cross-validation split, meaning that they have at least 32 images in the training set, whereas in this work the experiments were carried out using standard split with 20 images in the training set. Nevertheless, the performance obtained in this work is comparable with those recently published by Girard et al. [5] and Hemelings et al. [7].
Visualization of estimated uncertainty: from left to right, targets with removed uncertain labels and crossings, posterior predictive mean, epistemic uncertainty, and aleatoric uncertainty. The results are obtained using the excluding uncertain (top row) and including uncertain (bottom row) strategy.
4 Conclusion
In this work, multilabel classification of arteries and veins using a Bayesian fully-convolutional network was studied. It was shown that the misclassified areas on the images can be visualized using uncertainty estimates. The proposed approach is comparable with recent state-of-the-art approaches for blood vessel segmentation and AV classification methods.
The main topics for the future research are how to reduce the epistemic uncertainty and more careful study on the classification of uncertain labels in the RITE database. Retinal vasculature segmentation and AV classification methods typically include preprocessing procedures that affect the data. One of the opened questions, how different preprocessing techniques change the aleatoric uncertainty estimates. Other possible directions include differentiable end-to-end methods for modeling the connectivity and regularizations similar to [5] and [2].
References
Almotiri, J., Elleithy, K., Elleithy, A.: Retinal vessels segmentation techniques and algorithms: a survey. Appl. Sci. 8(2), 155 (2018)
Badawi, S., Fraz, M.: Multiloss function based deep convolutional neural network for segmentation of retinal vasculature into arterioles and venules. BioMed Res. Int. 2019, 1–17 (2019). https://doi.org/10.1155/2019/4747230
Decencière, E., et al.: Feedback on a publicly distributed image database: the messidor database. Image Anal. Stereol. 33(3), 231–234 (2014)
Gal, Y., Ghahramani, Z.: Dropout as a Bayesian approximation: representing model uncertainty in deep learning. In: International Conference on Machine Learning, pp. 1050–1059 (2016)
Girard, F., Kavalec, C., Cheriet, F.: Joint segmentation and classification of retinal arteries/veins from fundus images. Artif. Intell. Med. 94, 96–109 (2019). https://doi.org/10.1016/j.artmed.2019.02.004
He, K., Zhang, X., Ren, S., Sun, J.: Delving deep into rectifiers: surpassing human-level performance on ImageNet classification. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 1026–1034 (2015)
Hemelings, R., Elen, B., Stalmans, I., Van Keer, K., De Boever, P., Blaschko, M.B.: Artery-vein segmentation in fundus images using a fully convolutional network. Comput. Med. Imaging Graph. 76, 101636 (2019)
Hoover, A., Kouznetsova, V., Goldbaum, M.: Locating blood vessels in retinal images by piecewise threshold probing of a matched filter response. IEEE Trans. Med. Imaging 19(3), 203–210 (2000)
Hu, Q., Abràmoff, M.D., Garvin, M.K.: Automated separation of binary overlapping trees in low-contrast color retinal images. In: Mori, K., Sakuma, I., Sato, Y., Barillot, C., Navab, N. (eds.) MICCAI 2013. LNCS, vol. 8150, pp. 436–443. Springer, Heidelberg (2013). https://doi.org/10.1007/978-3-642-40763-5_54
Hu, S., Worrall, D., Knegt, S., Veeling, B., Huisman, H., Welling, M.: Supervised uncertainty quantification for segmentation with multiple annotations. arXiv preprint arXiv:1907.01949 (2019)
Jégou, S., Drozdzal, M., Vazquez, D., Romero, A., Bengio, Y.: The one hundred layers tiramisu: fully convolutional DenseNets for semantic segmentation. In: 2017 IEEE Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), pp. 1175–1183. IEEE (2017)
Jogi, R.: Basic Ophthalmology. Jaypee Brothers Medical Publishers, New Delhi (2008)
Kendall, A., Gal, Y.: What uncertainties do we need in Bayesian deep learning for computer vision? In: Advances in Neural Information Processing Systems, pp. 5574–5584 (2017)
Malek, J., Tourki, R.: Blood vessels extraction and classification into arteries and veins in retinal images. In: 10th International Multi-conferences on Systems, Signals Devices 2013 (SSD 2013), pp. 1–6, March 2013
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Welikala, R., et al.: Automated arteriole and venule classification using deep learning for retinal images from the UK Biobank cohort. Comput. Biol. Med. 90, 23–32 (2017). https://doi.org/10.1016/j.compbiomed.2017.09.005
Zeiler, M.D.: ADADELTA: an adaptive learning rate method. Technical report, December 2012. arXiv: 1212.5701http://arxiv.org/abs/1212.5701
Zhang, S., et al.: Simultaneous arteriole and venule segmentation of dual-modal fundus images using a multi-task cascade network. IEEE Access 7, 57561–57573 (2019). https://doi.org/10.1109/ACCESS.2019.2914319
Zuiderveld, K.: Contrast limited adaptive histogram equalization. In: Heckbert, P.S. (ed.) Graphics Gems IV, pp. 474–485. Academic Press Professional Inc., San Diego (1994). http://dl.acm.org/citation.cfm?id=180895.180940
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Garifullin, A., Lensu, L., Uusitalo, H. (2020). On the Uncertainty of Retinal Artery-Vein Classification with Dense Fully-Convolutional Neural Networks. In: Blanc-Talon, J., Delmas, P., Philips, W., Popescu, D., Scheunders, P. (eds) Advanced Concepts for Intelligent Vision Systems. ACIVS 2020. Lecture Notes in Computer Science(), vol 12002. Springer, Cham. https://doi.org/10.1007/978-3-030-40605-9_8
Download citation
DOI: https://doi.org/10.1007/978-3-030-40605-9_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-40604-2
Online ISBN: 978-3-030-40605-9
eBook Packages: Computer ScienceComputer Science (R0)