Abstract
In scoliosis diagnosis and treatment, estimation of spinal curvature plays an important role. Compared with the traditional method, which is time-consuming and unreliable, automated estimation has been more and more popular. But it remains to be such a great challenge that direct estimation has poor precision due to the lack of information. To meet this challenge, we propose a Multi-Task learning method with pyramidal feature aggregation. Our method is one-stage. It means that we can directly estimate the angles without detecting landmarks. To enhance the feature extraction and collect more information, we make the fusion of the pyramidal features and extend the base model by adding an extra branch for spinal segmentation. We evaluate our method on the validation set from the challenge (Accurate Automated Spinal Curvature Estimation, MICCAI 2019) and obtain a symmetric mean absolute percentage error of 12.97.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
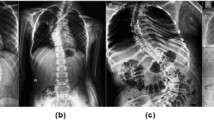
Cobb angle is a measurement of the bending deformity of the vertebral column. It has been widely used for scoliosis treatment, including structural, lateral, rotated curvature of the spine. Large reports show that there is a sustained increment of the prevalence rate of scoliosis. Therefore, it’s likely essential to have a reliable estimation of Cobb angles.
In clinical practice, each X-ray contains seventeen vertebras. For each one of them, doctors find four landmarks and draw the outline accordingly. Then they locate the key vertebras and calculate the angles. The main obstacle is that this method is highly time-consuming and can be easily mistaken.
To address this problem, researchers have designed lots of automated frameworks. Two-stage methods like [9] simulate clinical practice. They firstly segment anatomical structures and then estimate the measurement based on segmentation. Some recent studies adopt the detector, instead of the segmentation model, to observe the key structure [5]. But these studies are limited by the selection of key vertebras and bias of different operators. Direct estimation methods [6,7,8] aim to obtain the functional connection of medical images and clinical measurement. While these studies are limited by the multi-type views (anterior-posterior and lateral X-ray images). In the case of a single view, they perform much worse due to the insufficient use of global information.
For the purpose of full use of global information, we propose a direct estimation method based on pyramidal feature aggregation. In contrast to earlier direct methods, we add an extra branch to predict spinal masks inspired by [1], which can benefit the angles’ estimation. To further enhance the feature extraction, we make a fusion of the decoder’s feature map with multiple scales. Overall, our method achieves the highest symmetric mean absolute percentage error (SMAPE) of 12.8 on the 125 scans from the challenge.
2 Methods
2.1 System Framework
As is shown in Fig. 1, the network is based on the encoder-decoder structure [3]. The last feature map of the encoder is transformed into a vector via global average pooling (GAP), and the biggest feature map of the decoder outputs the mask via activation function (Softmax). In an attempt to make full use of a decoder, we add GAP into each stage of the decoder except the last one. For the different lengths, the feature vectors are simply concatenated and output the three angles through the dense layer.
For network training, our loss function includes mask loss and angle loss which can be defined as:
Dice similarity coefficient (DSC) is a metric function to measure the degree of similarity and always used in the medical image segmentation, so we calculate the mask loss by DSC. As for the Cobb angle, we choose SMAPE, which is usually used to evaluate the angles’ estimation.
Here A and X are ground truth, B and Y are predicted result, N is the number of test data.
2.2 Image Pre-processing
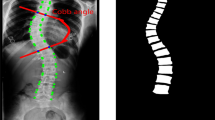
The direct estimation method hasn’t taken it into consideration that different ratio (rate of height and width) will obtain different Cobb angles (see Fig. 2). There is a certain problem while resizing all the images into the same size. For instance, if the original ratio is smaller than the model’s input ratio, Cobb angles will be smaller in the result. Hence we keep the ratio while resizing and use padding to fit the input shape. What’s more, we augment the training data by randomly shifting resized images.
3 Experiments
3.1 Datasets and Implementation Details
Datasets are collected from the challenge (Accurate Automated Spinal Curvature Estimation, MICCAI 2019), and it’s composed of three parts, among which only train and validation sets’ annotations are provided. As a result, we take the experiments only on the two sets. We count the three angles’ value and draw the histogram as Fig. 3. From top to bottom, we number the Cobb angles. It’s apparent that the angle above is more likely to be bigger than the angle below. From the graph, we can also see that most angles are small.
Although samples in the training set may come from the same patient, considered that there is still bias due to interval of operation, we randomly select \(10\%\) samples from the public train set as the validation and take public validation set as our test set. For all experiments, we use the same optimizer (Adam) and the learning rate (\(1e-4\)). By initialized with 100 epochs, the training process is terminated by a strategy called Earlystop.
3.2 Results
Through the same setting described above, we have taken a series of experiments to make the discussion. First of all, we talk about the impact of different encoder: Vgg11 [4], ResNet-50 [2]. Thanks to the single channel of X-ray images, weights pre-trained on ImageNet cannot be transferred into this task. Thus we train both networks from scratch and haven’t adopted deeper encoder. As the second and third rows show, there is a significant difference between the two conditions. Vgg11 outperforms so that we take it as our encoder while adding PFA and a new branch. The last two rows suggest that estimation is more precise by using our method (Table 1).
In detail, correlation coefficients between the three different predicted angles and ground truth are given as Fig. 4. The figure demonstrates that the estimation of the first angle is much better than the rest, and the second one is better than the third one. It has the same variation tendency as angle’s range that the smaller angles have worse performance (see Fig. 3).
4 Conclusion
In this paper, we have proposed a Multi-Task network with pyramid feature aggregation to estimate Cobb angles automatically. Inspired by Multi-Task learning, we add a new branch for segmentation to enhance the feature extraction. Typical aggregation of pyramid features is used to catch features at different levels. The two strategies both provide a more precise estimation of Cobb angles, and our method finally achieves high performance of SMAPE (12.97).
However, imbalanced estimation exists that our method estimates the top angle much better than the rest. There are several possible explanations for this result. Since big Cobb angles have a distinct appearance, the model can easily achieve excellent performance. On contrast, estimation of a straight spine with small Cobb angles has obvious relative fluctuation.
References
He, K., Gkioxari, G., Dollár, P., Girshick, R.: Mask R-CNN. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 2961–2969 (2017)
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition (2015)
Ronneberger, O., Fischer, P., Brox, T.: U-net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556 (2014)
Wu, H., Bailey, C., Rasoulinejad, P., Li, S.: Automatic landmark estimation for adolescent idiopathic scoliosis assessment using BoostNet. In: Descoteaux, M., Maier-Hein, L., Franz, A., Jannin, P., Collins, D.L., Duchesne, S. (eds.) MICCAI 2017. LNCS, vol. 10433, pp. 127–135. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-66182-7_15
Wu, H., Bailey, C., Rasoulinejad, P., Li, S.: Automated comprehensive adolescent idiopathic scoliosis assessment using mvc-net. Med. Image Anal. 48, 1–11 (2018)
Xue, W., Islam, A., Bhaduri, M., Li, S.: Direct multitype cardiac indices estimation via joint representation and regression learning. IEEE Trans. Med. Imaging 36(10), 2057–2067 (2017)
Xue, W., Nachum, I.B., Pandey, S., Warrington, J., Leung, S., Li, S.: Direct estimation of regional wall thicknesses via residual recurrent neural network. In: Niethammer, M., et al. (eds.) IPMI 2017. LNCS, vol. 10265, pp. 505–516. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-59050-9_40
Zhang, K., Xu, N., Yang, G., Wu, J., Fu, X.: An automated cobb angle estimation method using convolutional neural network with area limitation. In: Shen, D., et al. (eds.) MICCAI 2019. LNCS, vol. 11769, pp. 775–783. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-32226-7_86
Acknowledgement
This work was supported by National Natural Science Foundation of China (Grant No. 61671399) and by the Fundamental Research Funds for the Central Universities (Grant No. 20720190012).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Wang, J., Wang, L., Liu, C. (2020). A Multi-task Learning Method for Direct Estimation of Spinal Curvature. In: Cai, Y., Wang, L., Audette, M., Zheng, G., Li, S. (eds) Computational Methods and Clinical Applications for Spine Imaging. CSI 2019. Lecture Notes in Computer Science(), vol 11963. Springer, Cham. https://doi.org/10.1007/978-3-030-39752-4_14
Download citation
DOI: https://doi.org/10.1007/978-3-030-39752-4_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-39751-7
Online ISBN: 978-3-030-39752-4
eBook Packages: Computer ScienceComputer Science (R0)