Abstract
Initially dismissed as artifacts during sample preparation or as cell debris in bacterial cultures, outer membrane vesicles (OMVs) are now widely accepted as facsimiles of the outer membrane naturally secreted by Gram-negative bacteria. Within the last decades, several studies focused on OMV biogenesis resulting in different, in part complementary models to explain OMV release. Notably, vesicle formation seems to be an essential process as neither a bacterial species nor a mutant lacking vesicle release has been reported so far. Based on the complex physiological roles discussed for OMVs, it is likely that parallel strategies for their production have evolved to ensure their release in diverse conditions. Although, we are still far from a comprehensive mechanistic understanding of OMV release, several studies shed some insights in the processes driving the liberation of OMVs from the bacterial surface. Within this chapter, we will discuss the observations resulting in the current models for OMV formation including their regulation and relevance for bacterial physiology.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
2.1 Introduction
Over the last years, the biogenesis of Gram-negative outer membrane vesicles (OMVs) has been investigated by several groups. This has resulted in a range of models for OMV release from bacteria (Fig. 2.1). Indeed, it is likely that bacteria encompass several complementary mechanisms for vesicle biogenesis.
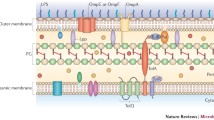
Schematic overview of OMV biogenesis routes discussed in this chapter. Shown are models of OMV formation based on: modulation of outer membrane–peptidoglycan linkages (a), bacterial stress responses (b), filamentous and tubular surface structures (c), and modulation of outer membrane components or outer membrane composition (d). In detail, (a) shows the mechanisms of OMV biogenesis due to loss of linkages between the outer membrane [outer leaflet composed of LPS (green) and inner leaflet composed of phospholipids (black)] and the peptidoglycan layer (gray) either by porins (orange) or proteins (light blue). (b) shows OMV formation due to stressors (red arrows) ranging from membrane attacking substances, high temperature, DNA damage or bacteriophages and accumulation of osmolytes (red). (c) shows OMV formation associated to surface structures like the sheathed flagellum (black) as well as tube-like outer membrane extensions such as nanopods, nanotubes, or nanowires. (d) indicates the mechanism of OMV formation via outer membrane modulation ranging from LPS modulation (green rectangles to triangles), intercalation of curvature-inducing molecules (purple circles), and modulation of phospholipid composition (black rectangles)
First reports indicated that mutants harboring a loss or reduction of linkages between the outer membrane and the periplasmic peptidoglycan layer exhibit increased vesiculation (Deatherage et al. 2009; Suzuki et al. 1978; Schwechheimer et al. 2013; Iwami et al. 2007; Moon et al. 2012; Turner et al. 2015; Llamas et al. 2000; Yeh et al. 2010; Mitra et al. 2016; Bernadac et al. 1998). Although these observations were predominantly achieved by loss-of-function mutants, current data suggest that bacteria can rearrange outer membrane–peptidoglycan linkages or regulate their abundance thereby affecting OMV production (Song et al. 2008; Choi et al. 2017; Schwechheimer et al. 2013).
Envelope stress is a general trigger for OMV release, highlighted by induction of vesiculation upon the presence of physical, chemical, and biological membrane stressors (Kadurugamuwa and Beveridge 1997). Moreover, accumulation of misfolded proteins and metabolites (e.g., peptidoglycan breakdown products) in the periplasm can result in increased OMV secretion, most likely via increased turgor pressure and bulging of the outer membrane (Schwechheimer et al. 2014; Hayashi et al. 2002). This could represent a response to harmful environments to increase the bacterial fitness and allow effective removal of undesired factors.
More recent studies revealed OMV secretion processes independent of mutations and presence of stressors, but allow refined regulation by the bacteria (Mashburn-Warren et al. 2008; Tashiro et al. 2011; Roier et al. 2016; Bonnington and Kuehn 2016; Elhenawy et al. 2016). A first report in this direction investigated small bacterial molecules produced during defined stages of the bacterial life cycle, which induce surface curvature via intercalation into the outer membrane and consequently promote vesiculation (Mashburn-Warren et al. 2008). So far, this strategy seems restricted to certain species as curvature-inducing molecules have only been identified in Pseudomonas aeruginosa (Mashburn and Whiteley 2005). However, alternative OMV biogenesis models relying on modulation of the outer membrane composition that are applicable to a wide range of Gram-negative bacteria have been described. This includes several studies demonstrating that OMV release is affected by alterations in the lipopolysaccharide (LPS) composition, LPS modifications or phospholipid accumulation in the outer membrane via silencing of a retrograde lipid trafficking system (Tashiro et al. 2011; Roier et al. 2016; Bonnington and Kuehn 2016; Elhenawy et al. 2016; Sabra et al. 2003; Haurat et al. 2011). As LPS as well as the retrograde lipid trafficking system are quite conserved among Gram-negative species, these models probably reflect general mechanisms broadly applicable to a diverse set of bacteria.
OMV isolation and quantification protocols have improved within the last years. As OMV research spans over decades, different approaches were used to identify and characterize mechanisms of OMV biogenesis such as phenotypical analyses of OMV formation under differential cultivation conditions or random and site-directed mutants. Furthermore, standard operating procedures for isolation and quantification of bacterial membrane vesicles are currently lacking. Methods for OMV analyses range from protein measurement (e.g., Bradford assay, SDS, BCA assay), dry weight measurement, immune-based assays (e.g., ELISA, immunoblot, dotblot), lipid measurements (e.g., FM4-46, phospholipid measurement with ammonium ferrothiocyanate), LPS quantification (e.g., purpald assay, mass spectrometry analyses), or microscopical analyses (e.g., fluorescence microscopy, electron microscopy, nanosight). Thus, comparison of the results from different OMV biogenesis studies may require careful evaluation of the diverse methodologies used to isolate and quantify OMVs. We kindly refer the interested readers to the original articles cited throughout the chapter.
2.2 Modulation of Outer Membrane–Peptidoglycan Linkages
The cell envelope of Gram-negative bacteria is composed of two bilayered membranes. The so-called inner and the outer membranes are separated by the periplasm, which contains a thin layer of peptidoglycan. While the inner membrane has phospholipids in both leaflets, the outer membrane is composed of phospholipids in the inner leaflet and predominantly LPS in the outer leaflet. This asymmetric structure functions as a selective barrier against the extracellular environment (Ruiz et al. 2006). The peptidoglycan is covalently linked to the outer membrane and determines the shape of the cell’s envelope. Moreover, it prevents Gram-negative cells from lysis due to osmotic changes or mechanical stress (Vollmer and Bertsche 2008). The peptidoglycan is a highly dynamic polymer, which consists of glycan chains that are crosslinked by short peptides (Vollmer and Bertsche 2008). Outer membrane proteins can interact with the peptidoglycan layer, thereby stabilizing the cell envelope by crosslinking the inner and outer membranes with the peptidoglycan. Among these, the main players regarding envelope cross linkage are the Braun’s lipoprotein (Lpp), outer membrane porin A (OmpA) and the Tol-Pal complex. Several reports revealed that modulation of these covalent cross-linkages reduces outer membrane integrity and induces vesiculation, which will be discussed within this chapter.
2.2.1 Braun’s Lipoprotein (Lpp)
Lpp represents the Braun’s lipoprotein in the outer membrane of Escherichia coli, with homologs found in a variety of Gram-negative species including all Enterobacteriaceae (Chang et al. 2012; Deatherage et al. 2009). It is the most abundant lipoprotein in E. coli and was shown to exist in a free or bound form, whereby the latter is covalently linked to the peptidoglycan (Braun 1975). About one third of bacterial Lpp is connected to peptidoglycan via the C-terminal lysine residue, thereby the outer membrane is anchored to the peptidoglycan via Lpp (Braun and Rehn 1969; Braun 1975). While Lpp is quite abundant in the outer membrane of E. coli, its OMVs contain relatively low levels of Lpp (Wensink and Witholt 1981; Hoekstra et al. 1976). In fact, OMVs contained almost none of the peptidoglycan-bound Lpp and only 35% of the free form compared to the corresponding bacterial outer membrane (Wensink and Witholt 1981). These results led to early models of OMV formation, wherein membrane blebbing occurs when the outer membrane expands faster than the underlying peptidoglycan or in surface areas with reduced prevalence of covalent linkages between Lpp in the outer membrane and peptidoglycan (Wensink and Witholt 1981; Hoekstra et al. 1976). Consequently, deletion of lpp leads to increased OMV formation in E. coli and in Salmonella typhimurium (Deatherage et al. 2009; Suzuki et al. 1978). Notably, loss of Lpp is associated with an impairment of the structural integrity of the outer membrane and cellular leakage (Suzuki et al. 1978; Deatherage et al. 2009). Thus, elevated OMV production might simply result from elevated cellular disintegration of lpp mutants. However, Lpp-dependent OMV formation might be more precisely regulated in wild-type cells. This regulation could be based on the reduction of Lpp crosslinks via the modulation of peptidoglycan or a change in the overall amount of Lpp via the small RNA Reg26 (Schwechheimer et al. 2013). Noteworthy, an uneven distribution of (bound) Lpp resulting in outer membrane areas with reduced Lpp amounts, as suggested for cell division sites (Hoekstra et al. 1976), might be more relevant for OMV formation than the overall Lpp cross-linkages as almost no Lpp is found in E. coli OMVs (Schwechheimer et al. 2014; Wensink and Witholt 1981).
2.2.2 Outer Membrane Protein A (OmpA)
OmpA is a major porin in the outer membrane of many Gram-negative species (Palva 1983; Wang 2002). Through its ability to non-covalently bind peptidoglycan via the conserved residues D271 and R286 of the C-terminus, it contributes to the structural integrity and stability of the cell envelope as it connects the peptidoglycan with the outer membrane (Samsudin et al. 2016; Iwami et al. 2007). Indeed, loss of OmpA results in an increase of OMV formation in S. typhimurium, Porphyromonas gingivalis, and Acinetobacter baumannii (Iwami et al. 2007; Moon et al. 2012; Deatherage et al. 2009). Both, Lpp and OmpA interactions, support membrane integrity. Consequently, OMV release by the lpp/ompA double mutant is significantly increased compared to each single mutant in S. typhimurium (Deatherage et al. 2009). In a similar manner, Sonntag and coworkers could demonstrate that an lpp/ompA double mutant in E. coli produced more OMVs than a lpp single mutant (Sonntag et al. 1978). Notably, Lpp-peptidoglycan linkages seem to be a dominant factor for increased OMV production in most bacteria probably due to the higher abundance of Lpp in the outer membrane (Deatherage et al. 2009). Concordantly, the impact of an ompA deletion in S. typhimurium is less pronounced as for lpp mutants and the loss of OmpA–peptidoglycan linkages was not necessarily associated with reduced membrane integrity (Deatherage et al. 2009). Noteworthy, bacteria can regulate ompA expression. For example, the small RNA VrrA of Vibrio cholerae interferes with ompA translation and its overexpression leads to increased vesiculation (Song et al. 2008). Similar results were obtained in E. coli and S. typhimurium by MicA, a homologue of VrrA, which downregulates ompA expression, thereby inducing OMV production in both organisms (Choi et al. 2017). As highlighted above, a similar regulation has also been observed for Lpp via the small RNA Reg26 (Schwechheimer et al. 2013). Interestingly, these small RNAs (VrrA, MicA, and Reg26) are under positive control of σE, an alternative sigma factor induced upon membrane stress, linking OMV formation to the envelope stress response (See Sect. 2.3.1) (Schwechheimer et al. 2013; Udekwu and Wagner 2007; Song et al. 2008).
2.2.3 Tol-Pal Complex
The Tol-Pal complex is widely conserved among Gram-negative bacteria and is crucial for multiple physiological roles including the maintenance of membrane integrity. In E. coli the apparatus consists of five proteins (TolQ-R-A-B and Pal) (Lazzaroni et al. 1999). While TolQ, TolR, and TolA form a complex in the cytoplasm, the periplasmic protein TolB interacts with Pal, which is non-covalently linked to the peptidoglycan (Lazzaroni et al. 1999; Parsons et al. 2006). Moreover, TolB and Pal interact with Lpp and OmpA at a protein level (Clavel et al. 1998; Cascales and Lloubes 2004), indicating that the Tol-Pal complex belongs to a protein network, which connects the periplasmic peptidoglycan layer with the outer membrane. Accordingly, tol-pal mutants exhibit elevated OMV formation associated with loss of membrane integrity and leakage of periplasmic proteins into the extracellular milieu in E. coli (Bernadac et al. 1998). Concordantly, tolQ-, tolR-, and tolA-mutants in Pseudomonas putida produce more OMVs, loss of tolA resulted in more vesicle protein amount in Shigella boydii, deletion of tolB in Helicobacter pylori resulted in increased vesiculation and in Caulobacter crescentus extensive blebbing of OMVs were observed upon mutation of pal, tolA, or tolB (Turner et al. 2015; Llamas et al. 2000; Yeh et al. 2010; Mitra et al. 2016). In tol-pal mutants of E. coli and C. crescentus, a substantial fraction of OMVs originate from the cell poles and division sites where Tol-Pal usually accumulates in wild-type cells. Notably, tol-pal mutants fail to connect the outer membrane and peptidoglycan during cell division, which could explain the increased OMV release (Gerding et al. 2007; Yeh et al. 2010).
2.3 Bacterial Stress Responses Affecting Vesiculation
Facing the outside world, the cell envelope of bacterial cells is exposed to varying environmental conditions, which can be detrimental for microbial viability. Variations in temperature, pH, or the exposure to antimicrobial or toxic compounds can cause impaired protein folding and may result in cell death. Thus, the ability to rapidly sense and respond to these adverse conditions is essential for the survival of bacterial cells. Therefore, bacteria have developed regulatory networks to control gene expression according to their specific requirements. Some of them have been linked to increased vesiculation, which can enhance fitness under stressful conditions.
2.3.1 Envelope Stress Response
In E. coli, adaptation to harsh environments is mediated by the activation of the signal transduction pathways of the bacterial envelope stress responses (Raivio 2005). One of them is the σE stress response activated via DegS, a periplasmic sensor protease, which recognizes misfolded outer membrane proteins. Upon its activation, DegS cuts the anti-sigma factor RseA, which is again cleaved by RseP and ClpXP and finally releases the alternative sigma factor σE. Liberated σE subsequently induces the transcription of genes involved in outer membrane protein and cell wall biogenesis, including DegP, a temperature-dependent periplasmic chaperone and protease (Hayden and Ades 2008). Recently, studies in E. coli and S. typhimurium linked the σE-mediated stress response to OMV biogenesis demonstrating increased vesiculation for mutants of the σE pathway such as degP-, degS-, and rseA-mutants for E. coli and the degP mutant in case of S. typhimurium (McBroom and Kuehn 2007; McBroom et al. 2006). The authors concluded that the impairment of the functionality of the σE pathway leads to an accumulation of misfolded products in the periplasm causing increased turgor pressure, which can be resolved by vesiculation.
The importance of OMV formation as a stress relief mechanism in E. coli was confirmed by mutation of nlpA (resulting in reduced vesiculation) in combination with degP (resulting in accumulation of periplasmic misfolded proteins) (Schwechheimer and Kuehn 2013). The reduced vesiculation of the nlpA/degP-mutant compared to a degP mutant caused a severe growth defect upon higher temperatures as misfolded proteins accumulated in the periplasm and could not be removed via vesiculation (Schwechheimer and Kuehn 2013). By the construction of a fusion protein, which mimicked an unfolded outer membrane protein, McBroom and coworkers could reinforce the model that vesiculation might be an alternative stress relief mechanism as this protein was significantly enriched in OMVs compared to other periplasmic proteins (McBroom and Kuehn 2007). Thus, they introduced OMV formation as a stress response of Gram-negative bacteria, which supports the cell’s efforts to reduce the consequences of unfolded proteins.
Furthermore, the model of OMV formation as a stress relief mechanism could be extended from misfolded proteins to other compounds accumulating in the periplasm. For example, vesiculation was also increased in E. coli upon deletion of ampG, which is an inner membrane permease and amiD, an amidase breaking down large peptidoglycan fragments as well as upon deletion of the autolysin homologue ami in P. gingivalis. These mutations resulted in the accumulation of peptidoglycan fragments in the periplasm and a subsequent increase of turgor pressure (Schwechheimer et al. 2014; Hayashi et al. 2002). In a similar manner, OMV formation was increased in rfaC or rfaG deletion strains in E. coli. Both mutations led to an accumulation of LPS in the periplasm, due to an impaired LPS maturation in the outer membrane (Schwechheimer et al. 2014). Consequently, OMVs of rfaC or rfaG deletion mutants in E. coli exhibited an increase of lipid-to-outer membrane protein ratio (Schwechheimer et al. 2014). The detrimental effect of excessive membrane products was demonstrated by deletion of yciM in E. coli, which increases LPS production and its periplasmic accumulation. Interestingly, E. coli strains lacking yciM required specific suppressor mutations for survival, either involved in LPS synthesis (lpxA, lpxC, or lpxD) or enhance vesiculation (lpp, tolA, pal, galU) (Mahalakshmi et al. 2014; Kulp et al. 2015).
The Kuehn lab extended their OMV biogenesis model by showing that the σE-homologue AlgU of P. aeruginosa also plays a major role in OMV formation (Macdonald and Kuehn 2013). In concordance with earlier observations in E. coli, impairment of the envelope stress response in P. aeruginosa also resulted in enhanced OMV formation (Tashiro et al. 2009; Macdonald and Kuehn 2013). Interestingly, hypervesiculating mutants often showed no altered σE activity in E. coli and deletion of algU in P. aeruginosa did not alter high vesiculation phenotypes of stressed cells, indicating that various mechanisms exist for OMV formation (Macdonald and Kuehn 2013; McBroom and Kuehn 2007). In the case of P. aeruginosa, these additional mechanisms might be modulations of surface structures or composition (see Sects. 2.5.1 and 2.5.2.3).
2.3.2 Cell Wall-Directed Agents
Various chemical and biological substances targeting the outer membrane have been shown to stimulate OMV formation. Increased OMV formation upon presence of sublethal concentrations of membrane-attacking substances (e.g., polymyxin B, LL-37 and colistin) was shown to be beneficial for bacterial survival as OMVs can act as sink for these antimicrobial compounds (Manning and Kuehn 2011; Duperthuy et al. 2013). The increased vesiculation induced by polymyxin B and colistin is based on the displacement of Mg2+ and Ca2+ cations in the outer membrane, which destabilize the electrostatic interactions between negatively charged LPS molecules (Moore and Hancock 1986; Storm et al. 1977). Upon treatment with polymyxin B, increased OMV levels were observed for P. aeruginosa (Macdonald and Kuehn 2013), E. coli and enterotoxigenic Escherichia coli (ETEC) (Manning and Kuehn 2011) as well as enterohemorrhagic Escherichia coli (EHEC) (Bauwens et al. 2017a; Bauwens et al. 2017b). Exposure to colistin resulted in elevated OMV formation in E. coli (Manning and Kuehn 2011). Similarly, the human cathelicidin LL-37, a cationic antimicrobial peptide was shown to induce pore formation in the outer membrane (Brogden 2005) and increases OMV formation in EHEC (Urashima et al. 2017). Presence of bile salts, acting as an emulsifier of biological membranes, also promotes vesicle formation in Campylobacter jejuni (Taheri et al. 2018). Moreover, gentamicin was shown to increase OMV formation in P. aeruginosa via destabilization of the outer membrane by binding to the LPS (Kadurugamuwa and Beveridge 1997).
Furthermore, highly hydrophobic carbon sources such as hexadecane or phenanthrene were reported to interact with the outer membrane of Gram-negative bacteria and are thought to induce vesiculation. Hexadecane stimulates OMV release in Acinetobacter calcoaceticus and phenanthrene induces vesiculation in Delftia acidovorans (Shetty and Hickey 2014; Borneleit et al. 1988). Although this is not a comprehensive list of cell wall-directed agents, it still indicates that substances that disturb or disrupt the outer membrane of Gram-negative bacteria are connected to OMV formation.
It is likely that several other cell wall-directed agents destabilize the outer membrane by similar modes of action as described above and thereby increase bacterial vesiculation. Elevated OMV levels caused by cell wall-directed agents could rather be a consequence of physical or chemical membrane damage, than a controlled OMV formation process actively regulated by the bacterium. Indeed, vesicles can reassemble from membrane material originating from lysed cells (Sych et al. 2018; Turnbull et al. 2016). However, cell wall-directed agents might activate the σE pathway. As a matter of fact, polymyxin B was shown to induce micA in a σE-dependent manner in S. typhimurium (Papenfort et al. 2006) and σE in V. cholerae (Mathur et al. 2007). Moreover, bile induces the SOS response in E. coli and S. typhimurium (Prieto et al. 2006). Thus, OMV formation by sublethal concentrations of cell wall-directed agents could be linked to regulatory pathways implicated in OMV biogenesis (see Sects. 2.3.1 and 2.3.3).
2.3.3 SOS Response and Bacteriophages
In E. coli, one regulatory network facilitating bacterial survival upon DNA damage is the so-called SOS response, representing an inducible DNA repair system (Simmons et al. 2008). Along the SOS response two regulatory proteins, the LexA repressor and the RecA protein, modulate the expression of more than 50 genes comprising, for example, sulA, an inhibitor of cell division; uvr genes, which are involved in DNA repair or sbmC, a DNA gyrase inhibitor (Simmons et al. 2008). While LexA inhibits the expression of the SOS genes during normal cell growth, RecA binds to single-stranded DNA caused by DNA damaging agents (e.g., antibiotics or UV radiation). Binding of RecA to single-stranded DNA activates RecA and stimulates the autocatalytic cleavage of LexA, thereby relieving the repressor from the SOS genes and leading to the expression of LexA repressed genes (Simmons et al. 2008). Recently, some studies have shown that the SOS response is closely linked to OMV biogenesis inducing OMV formation. Such an upregulation of OMV formation has been observed for EHEC and Shigella dysenteriae upon treatment with SOS response stimulating substances like ciprofloxacin or mitomycin C (Bauwens et al. 2017a; Dutta et al. 2004). In P. aeruginosa, Maredia and coworkers demonstrated that the OMV amount was higher under ciprofloxacin treatment compared to non-treated wild-type cells (Maredia et al. 2012). Since ciprofloxacin is an efficient SOS response stimulator, a noncleavable LexA strain (no induction of the SOS response) was used to test if the enhanced vesiculation can be solely attributed to the SOS response. Under ciprofloxacin treatment, OMV levels were higher in wild-type cells than in the noncleavable LexA strain. However, an induction of vesiculation upon ciprofloxacin treatment was also detected in the noncleavable LexA strain, indicating that vesiculation is linked to the SOS response, but also to alternative pathways (Maredia et al. 2012). Moreover, the study of Toyofuku and coworkers confirmed the association of the SOS response with enhanced OMV formation in P. aerugionsa as an increase of the OMV amount was also observed under denitrifying conditions, which induces the SOS response (Toyofuku et al. 2014). Although a concise mechanism is lacking, Maredia and coworkers speculated that the enhanced OMV formation is a consequence of the delay in cell division and cell surface alterations during the SOS response (Maredia et al. 2012).
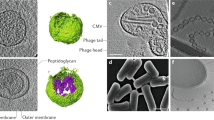
DNA damage also activates lysogenic bacteriophages, which infect, parasitize, and lyse their host cell (Weinbauer 2004). These phages integrate their genome into the bacterial chromosome and replicate through the cell cycle of their host. Upon stressors such as UV radiation or antibiotics, resulting in bacterial DNA damage and induction of the SOS response, lysogenic phages become activated causing excision from the chromosome, proliferation of new phages, and lysis of the bacterial cell (Weinbauer 2004; Feiner et al. 2015). Recently, a novel OMV formation mechanism based on a phage effector was proposed for P. aeruginosa in biofilms (Turnbull et al. 2016). Turnbull and coworkers demonstrated that the SOS-dependent activation of the cryptic phages endolysin, which degrades the bacterial peptidoglycan, results in explosive cell lysis and high amounts of liberated OMVs. These OMVs are formed by shattered membrane fragments of the exploding bacteria, which self-assemble to vesicle structures (Turnbull et al. 2016). Notably, activation of the SOS response in Stenotrophomonas maltophilia triggers formation of vesicles containing outer and inner membranes (Devos et al. 2017). Such cell lysis-induced outer–inner membrane vesicles were also reported for P. aeruginosa, Pseudoalteromonas marina, and Shewanella vesiculosa (Perez-Cruz et al. 2013; Hagemann et al. 2014; Kadurugamuwa and Beveridge 1995). A similar mechanism based on cell lysis was shown to induce vesicle formation in the Gram-positive bacterium Bacillus subtilis mediated by the SOS-dependent activation of an endolysin encoded by a defective prophage (Toyofuku et al. 2017).
In contrast, lytic phages immediately replicate their genome upon infection until the bacterial cell bursts, thereby transferring the phages to new cells. During this process, increased vesicle formation was observed for Yersinia pseudotuberculosis upon infection with a pseudotuberculous diagnostic bacteriophage and for E. coli upon infection with the T4 phage, which is thought to occur due to increased osmotic stress and subsequent self-assembly of outer membrane fragments (Byvalov et al. 2018; Tarahovsky et al. 1994). Prophage(-like) elements are frequently found in bacterial genomes and lytic phages are constantly attacking bacterial cells (Casjens 2003; Weinbauer 2004). Thus, the induction of vesicle formation by phages might be a ubiquitous mechanism for the production of membrane vesicles ranging from Gram-negative to Gram-positive bacteria. Similar to cell wall-directed agents (see Sect. 2.3.2), phage-induced vesicle formation likely represents less of an active mechanism than a physical or chemical consequence of the self-assembly properties of biological membranes, which re-organize themselves to vesicles and vesicle-like structures (Sych et al. 2018).
2.4 Filamentous or Tubular Surface Structures
Recently, new mechanisms of OMV biogenesis have been addressed by several studies dealing with the outer membrane sheathed flagellum or nanotube-like structures of Gram-negative bacteria. Several bacterial species like Brucella melitensis, Vibrio ssp., or H. pylori assemble flagella covered with an outer membrane-derived sheath (Fuerst and Perry 1988; Geis et al. 1993; Ferooz and Letesson 2010). Interestingly, for Vibrio fischeri, B. melitensis, and H. pylori vesicle-like structures were found along the sheath, frequently localized at the distal tip of the flagella (Ferooz and Letesson 2010; Millikan and Ruby 2004; Qin et al. 2016). In the case of V. fischeri, rotation of the flagellum results in release of more LPS molecules, which were later shown to be shed via OMVs (Brennan et al. 2014; Aschtgen et al. 2016). Interestingly, Aschtgen and coworkers also observed different OMV sizes depending on the presence and functionality of the bacterium’s sheathed flagellum. A hyperflagellated mutant produced a higher proportion of smaller OMVs whereas a nonmotile mutant produced fewer OMVs that tend to be slightly larger in electron microscopy than those generated by wild-type cells (Aschtgen et al. 2016). Hampton and coworkers reported similar observations for Vibrio vulnificus comparing the amount and size of OMVs via electron microscopy derived from motile and nonmotile strains (Hampton et al. 2017). Notably, recent studies show that the bacterial growth phase determines OMV size and protein composition in H. pylori, a pathogen with growth phase-dependent expression of flagellar genes (Niehus et al. 2002; Zavan et al. 2019). Taken together, these results not only demonstrate a novel mechanism of OMV formation by bacteria with sheathed flagella, but also raise the question if at least two OMV populations exits, being either released from the cellular body or from the flagellar sheath. Due to their different site of origin, these vesicles may differ in their size and composition (Aschtgen et al. 2016).
Recently, studies showed that bacteria are also able to produce other outer membrane structures such as nanotubes, nanopods, or nanowires, which stay connected to their originating cell and enable them to interact directly with distant surfaces. Among them, the presence of nanopods and nanowires has just recently been reported (Shetty et al. 2011). Nanopods are described as outer membrane tubes filled with OMVs and may represent a new bacterial organelle, still their biogenesis and regulation are currently not well defined (Shetty et al. 2011). In D. acidovorans nanopods were shown to play an important role in the biodegradation of the highly hydrophobic phenanthrene as cells producing almost no nanopods were massively attenuated upon growth on this carbon source (Shetty and Hickey 2014).
Shewanella oneidensis, a metal-reducing bacterium was found to produce extensions of the outer membrane and periplasm called nanowires, which were shown to enable the cells to link the respiratory chain to external (metal) electron acceptors (Pirbadian et al. 2014).
Nanotubes have been described as tube-shaped membranous structures and are considered to be a specialized form of OMV production (McCaig et al. 2013). It has been suggested that these structures connect neighboring cells thereby playing important roles in cell–cell communication or the exchange of molecules in Myxococcus xanthus, enteropathogenic Escherichia coli (EPEC), V. vulnificus, and Francisella novicida (Remis et al. 2014; Hampton et al. 2017; McCaig et al. 2013; Pal et al. 2019). Though their nature is not completely understood, in EPEC nanotubes are thought to be evolutionary related to the injectosome and the flagellum (Pal et al. 2019). A first step toward a regulatory mechanism is a study showing that nanotube production is induced upon amino acid starvation in F. novicida indicating their importance for nutrient acquisition (Sampath et al. 2018).
Taken together, these findings suggest that nanotubes, nanopods and nanowires represent a specialized type of OMVs that stay connected to the cell, thereby facilitating nutrient degradation and acquisition of highly specialized bacteria, aid in energy production cycles or enable cell to cell communication.
2.5 Modulation of Outer Membrane Components or Composition
All domains of life share a bilayered membrane mainly composed of phospholipids in both leaflets or at least in the inner leaflet (i.e., outer membrane of Gram-negative bacteria). For eukaryotic cells, the curvature alteration of membranes plays an important role in defining the cell morphology, organelles, and local subdomains and is therefore crucial for compartmentalizing proteins and enzymatic reactions (McMahon and Boucrot 2015). Besides changes in the phospholipid composition, curvature induction in eukaryotes is predominately mediated via proteins, which may be integral components of the membrane or are able to induce curvature via binding from the inside or outside of the membrane (Zimmerberg and Kozlov 2006; McMahon and Boucrot 2015). Notably, mechanisms for the alteration of membrane curvatures have also been identified in bacteria, which range from the insertion of curvature-inducing small molecules (e.g., Pseudomonas quinolone signal PQS), modification of membrane components (e.g., LPS), or alteration of the membrane composition (e.g., phospholipid accumulation) (Mashburn-Warren et al. 2008; Tashiro et al. 2011; Roier et al. 2016; Bonnington and Kuehn 2016; Elhenawy et al. 2016). All of these processes have also been demonstrated to affect OMV formation, which will be discussed in this chapter.
2.5.1 Pseudomonas Quinolone Signal (PQS)
The quorum sensing signal molecule 2-heptyl-3-hydroxy-4-quinolone (Pseudomonas quinolone signal; PQS) is involved in communication and growth phase coordination of P. aeruginosa. The inter-bacterial delivery of this hydrophobic molecule in aqueous solution was not understood until Mashburn and coworkers demonstrated that PQS is transferred via OMVs (Mashburn and Whiteley 2005). Aside from its role in quorum sensing signaling, PQS also induces OMV formation. In P. aeruginosa it seems to be a dominant biogenesis mechanism as a pqsA mutant, unable to produce PQS, exhibits massively reduced OMV levels (Mashburn and Whiteley 2005). Due to its chemical properties, it is able to interact with the acyl chains of lipid A, a component of LPS, bringing them into a more ordered gel-like state. Thereby PQS reduces membrane fluidity, which facilitates bulging of the membrane and OMV release. Importantly, the high fluidity of the P. aeruginosa outer membrane likely prevents OMV blebbing without PQS (Schertzer and Whiteley 2012; Mashburn and Whiteley 2005; Mashburn-Warren et al. 2008). Consistent with the observation that PQS is strictly produced under aerobic growth, OMV formation under anaerobic conditions is massively reduced (Schertzer and Whiteley 2013). Interestingly, vesicle formation can also be induced upon the addition of PQS to cultures of other Gram-negative or Gram-positive bacteria including E. coli, Burkholderia cepacia, and B. subtilis and is thought to interact with their membranes (Mashburn-Warren et al. 2008; Tashiro et al. 2010). This fascinating observation implicates a unique strategy allowing interspecies controlled induction of vesicle formation in bacteria.
2.5.2 LPS Modifications
LPS is a major component of the outer membrane of Gram-negative bacteria and constitutes its outer leaflet. This molecule consists of three components and is highly variable between species. The O-antigen is the outermost part of the LPS with the highest diversity and is mainly recognized by the host immune system. The core region is in parts highly conserved and plays an important role in outer membrane stability. The lipid A, or so-called endotoxin, represents the membrane anchor of the LPS. In many species, it is a target molecule for defined, inducible modifications to increase resistance against antimicrobial factors, which need to penetrate through the outer membrane or directly attack the outer membrane integrity (Hankins et al. 2012; Gunn 2001). Due to its central role in the bacterial surface it is not surprising that modifications in all three parts of the LPS molecule have been reported to affect OMV formation in various species, which will be discussed below.
2.5.2.1 Lipid A Modifications
The biosynthesis of lipid A is a multistep process wherein many modifications can be introduced such as glycinations, changes in acylation patterns or the attachment of phosphoethanolamine or aminoarabinose (Hankins et al. 2012; Raetz et al. 2007; Brozek and Raetz 1990; Gunn et al. 1998). These lipid A modifications have been shown to promote resistance to antimicrobial peptides, reduce host immune system activation, change in membrane stability, or change in vesicle formation (Raetz et al. 2007; Kawasaki et al. 2004b; Gwozdzinski 2018).
A recent study showed that the addition of a phosphoethanolamine group onto the N-acetylglucosamine disaccharide of lipid A via mcr-1 in E. coli increases OMV formation (Gwozdzinski 2018). The current model suggests this modification causes a charge repulse inducing vesiculation (Gwozdzinski 2018). In S. typhimurium PagL can deacetylate lipid A to downregulate the host immune response. PagL is activated via the two-component system PhoPQ upon entering the macrophages (Kawasaki et al. 2004a; Trent et al. 2001). The deacylation in the 3-position of the lipid A also increases OMV formation inside of macrophages, where OMVs are thought to interfere with host cellular pathways. The loss of the acyl chain is thought to generate a positive curvature of the outer membrane by changing the shape of the lipid A and thereby induce vesiculation (Kawasaki et al. 2004a; Elhenawy et al. 2016).
2.5.2.2 LPS Core Modifications
Mutations in the core region of the LPS can have a massive impact on the bacterial surface structure, colony morphology, and outer membrane integrity. Several LPS core mutants showed elevated OMV levels in P. aeruginosa (Ruhal et al. 2015; Salkinoja-Salonen and Nurmiaho 1978). Similar results were observed in E. coli (Nakao et al. 2012; Schwechheimer et al. 2014). These studies often relied on mutants lacking the majority of the LPS core and, as a consequence, the entire O-antigen. As full length LPS has a pivotal impact on outer membrane stability, such core mutants have an impaired outer membrane integrity, which could explain the increased liberation of outer membrane material. Yet, a study by Schwechheimer and coworkers described a mechanism for the increased vesiculation observed in LPS core mutants (Schwechheimer et al. 2014). According to their model, enhanced vesiculation in E. coli rfaC or rfaB mutants lacking full-length LPS results from accumulation of periplasmic LPS and subsequent increase in turgor pressure (Schwechheimer et al. 2014).
2.5.2.3 O-Antigen Modifications
P. aeruginosa was shown to express two distinct types of LPS varying in the O-antigen composition. One is a homopolymer of d-rhamnose and is named common polysaccharide antigen (CPA, formerly termed A-band), the other is a heteropolymer of three to five distinct sugars in its repeat units known as O-specific antigen (OSA, formerly termed B-band). P. aeruginosa mutants lacking OSA or CPA were shown to produce OMVs with altered size and protein composition (Murphy et al. 2014). While OSA is more immunogenic, CPA plays an important role in binding to human cells (Lam et al. 2011). Interestingly, P. aeruginosa OMVs were shown to be enriched with the negatively charged OSA (Kadurugamuwa and Beveridge 1995). Enrichment of OMVs with OSA is believed to be the consequence of the charge repulsive forces between adjacent OSA and other LPS molecules in the outer membrane. Interestingly, exposure to oxidative stress increased the abundance of negatively charged OSA and OMV formation (Sabra et al. 2003). Thus, increased vesiculation during stress conditions might be induced by the AlgU pathway (see Sect. 2.3.1) and alterations in O-antigen composition to facilitate survival of P. aeruginosa.
P. gingivalis also produces two classes of LPS, carrying either neutral (O-LPS) or negatively charged (A-LPS) O-antigen chains (Paramonov et al. 2001; Rangarajan et al. 2008; Paramonov et al. 2005). OMVs of this human oral pathogen are enriched in A-LPS, which may implicate a role for this O-antigen species in OMV release (Haurat et al. 2011). Indeed, for species with neutral and negatively charged O-antigens, the latter has been found to be enriched in OMVs. Thus, it could be suggested that an interaction between negatively charged O-antigen species contributes to OMV formation comparable to the destabilization of the OM of P. aeruginosa by negatively charged gentamicin (see Sect. 2.3.2; Kadurugamuwa and Beveridge 1997).
2.5.3 Modulation of Phospholipid Composition
The outer membrane of Gram-negative bacteria is composed of an asymmetric bilayer with phospholipids in the inner leaflet and LPS on the outside. This asymmetry is maintained by a putative retrograde phospholipid trafficking system transporting phospholipids from the outer to the inner membrane (Malinverni and Silhavy 2009). This system was first described in E. coli, but is highly conserved among Gram-negative bacteria (Roier et al. 2016; Malinverni and Silhavy 2009). In E. coli, the system is composed of the outer membrane lipoprotein VacJ (MlaA), the periplasmic binding/transport protein YrbC (MlaC), and a permease complex consisting of YrbB (MlaB), YrbD (MlaD), YrbE (MlaE), and YrbF (MlaF) (Malinverni and Silhavy 2009).
Deletion or downregulation of this retrograde phospholipid transport system has been shown to increase OMV formation in multiple species including E. coli, Haemophilus influenzae, V. cholerae, and Neisseria gonorrhoeae (Roier et al. 2016; Baarda et al. 2019). In H. influenzae, comprehensive lipid analyses revealed a higher phospholipid abundance in the OMVs derived from yrbE and vacJ deletion mutants compared to wild type (Roier et al. 2016). This is accompanied by distinct changes of the lipid species composition, such as the enrichment of short-chain fatty acids, in the outer membrane and OMVs of yrbE and vacJ mutants compared to wild type (Roier et al. 2016). Together with the observation that the fatty acid composition of P. aeruginosa OMVs differs from the outer membrane, it can be hypothesized that arrangement of certain phospholipid might promote OMV biogenesis (Tashiro et al. 2011). Generalized models based on accumulation of defined lipid species are hampered by the heterogeneous phospholipid compositions found in the outer membrane of Gram-negative bacteria.
Interestingly, the phospholipid transport system was shown to be transcriptionally silenced upon iron limitation in a Fur (ferric uptake regulator)-dependent manner. As iron limitation is a signal for host environment for many bacteria, it is very likely that vesiculation plays a major role in colonization strategies of pathogenic bacteria (Roier et al. 2016). Along this line, an increased in vivo fitness upon deletion of this system has been reported for P. aeruginosa and N. gonorrhoeae (Shen et al. 2012; Baarda et al. 2019). In contrast, other studies report that mutants in the retrograde lipid transporter of Shigella flexneri and Haemophilus parasuis are attenuated in vivo (Carpenter et al. 2014; Zhao et al. 2017; Suzuki et al. 1994), indicating a current lack of understanding of the complex physiological processes involved. Another example of increased vesiculation due to phospholipid accumulation was observed in Neisseria meningitidis, where higher production of phospholipids triggered by sulfate depletion led to increased OMV production. Moreover, sulfate depletion is associated with oxidative stress, which has already been shown to be a signal for increased vesiculation in P. aeruginosa (see Sect. 2.5.1; Gerritzen et al. 2019; Macdonald and Kuehn 2013).
2.6 Conclusion
Long neglected within the scientific community, the vital role of membrane vesicles is nowadays increasingly studied and has been proven for all branches of life ranging from Gram-negative and Gram-positive bacteria to archaea and even eukaryotic cells (Deatherage and Cookson 2012). Among them, OMVs of Gram-negative bacteria have been most extensively studied. Pioneer work of the Beveridge and Kuehn labs on OMV physiology and biogenesis provided basic knowledge on the existence and multiple physiological roles of these spherical surface derivatives, which ultimately opened this field to become a striving research topic (Horstman and Kuehn 2000; Kadurugamuwa et al. 1993). Noteworthy, a lack of vesiculation has neither been observed for the investigated bacterial species nor any mutants, implicating that outer membrane vesiculation might be an essential process. This chapter provided a concise overview of the most prominent models of OMV formation of Gram-negative bacteria, which have been reported so far (Fig. 2.1). Importantly, all of these mechanisms could act in concert without excluding each other.
While vesiculation due to the loss of outer membrane and peptidoglycan interaction has mostly been observed in mutant strains lacking important structural proteins, recent studies showed that bacteria may actively regulate the abundance of such integral molecules via small RNAs (Song et al. 2008; Choi et al. 2017; Schwechheimer et al. 2013). In a similar manner, bacteria can modulate their surface composition to alter vesiculation. For example, vesiculation models based on modulation of phospholipid abundance or LPS act on structures ubiquitously present in Gram-negative bacteria (Gerritzen et al. 2019; Roier et al. 2016; Kawasaki et al. 2004a; Elhenawy et al. 2016). In contrast, the only curvature-inducing molecule known to date (PQS) is exclusively produced by P. aeruginosa although it induces vesicle formation in other strains including Gram-positive bacteria (Mashburn-Warren et al. 2008; Tashiro et al. 2010). Vesiculation due to external stressors like cell wall-directed agents, UV radiation or high temperature is not only a consequence of bursting cells or increased turgor pressure, but can also be induced by modulation of the σE pathway or the SOS response (McBroom and Kuehn 2007; McBroom et al. 2006; Maredia et al. 2012). Indeed, the increased release of vesicles upon exposure to hostile conditions can be beneficial for bacterial cells due to their abilities to act as decoys or relief of turgor pressure.
The multitude of OMV biogenesis mechanisms as well as the lack of strains showing no vesiculation emphasizes an important role of these surface facsimiles for bacterial fitness. Interestingly, OMVs play an important role in host–pathogen interactions including toxin delivery, immunomodulation, binding of antimicrobial factors as well as the acquisition of nutrients. Thus, it could be speculated that pathogenic or human-associated bacteria are generally prone to high vesiculation in order to aid in pathogenesis.
Recent work has already revealed quite comprehensive and complementary OMV biogenesis models. It is likely that in the same species, several OMV biogenesis mechanisms are simultaneously active or conditionally induced. Thus, one species might produce a variety of OMV types of different sizes or composition, reflecting diverse OMV biogenesis routes. The fast progression of techniques to study OMVs ranging from their isolation, composition analyses, and microscopic visualization will drive future investigation of OMV biogenesis, morphology, and content.
References
Aschtgen MS, Lynch JB, Koch E, Schwartzman J, McFall-Ngai M, Ruby E (2016) Rotation of Vibrio fischeri flagella produces outer membrane vesicles that induce host development. J Bacteriol 198(16):2156–2165. https://doi.org/10.1128/JB.00101-16
Baarda BI, Zielke RA, Le Van A, Jerse AE, Sikora AE (2019) Neisseria gonorrhoeae MlaA influences gonococcal virulence and membrane vesicle production. PLoS Pathog 15(3):e1007385. https://doi.org/10.1371/journal.ppat.1007385
Bauwens A, Kunsmann L, Karch H, Mellmann A, Bielaszewska M (2017a) Antibiotic-mediated modulations of outer membrane vesicles in Enterohemorrhagic Escherichia coli O104:H4 and O157:H7. Antimicrob Agents Chemother 61(9):e00937–e00917. https://doi.org/10.1128/AAC.00937-17
Bauwens A, Kunsmann L, Marejkova M, Zhang W, Karch H, Bielaszewska M, Mellmann A (2017b) Intrahost milieu modulates production of outer membrane vesicles, vesicle-associated Shiga toxin 2a and cytotoxicity in Escherichia coli O157:H7 and O104:H4. Environ Microbiol Rep 9(5):626–634. https://doi.org/10.1111/1758-2229.12562
Bernadac A, Gavioli M, Lazzaroni JC, Raina S, Lloubes R (1998) Escherichia coli tol-pal mutants form outer membrane vesicles. J Bacteriol 180(18):4872–4878
Bonnington KE, Kuehn MJ (2016) Outer membrane vesicle production facilitates LPS remodeling and outer membrane maintenance in Salmonella during environmental transitions. mBio 7(5):e01532-16. https://doi.org/10.1128/mBio.01532-16
Borneleit P, Hermsdorf T, Claus R, Walther P, Kleber HP (1988) Effect of hexadecane-induced vesiculation on the outer membrane of Acinetobacter calcoaceticus. J Gen Microbiol 134(7):1983–1992
Braun V (1975) Covalent lipoprotein from the outer membrane of Escherichia coli. Biochim Biophys Acta 415(3):335–377
Braun V, Rehn K (1969) Chemical characterization, spatial distribution and function of a lipoprotein (murein-lipoprotein) of the E. coli cell wall. The specific effect of trypsin on the membrane structure. Eur J Biochem 10(3):426–438
Brennan CA, Hunt JR, Kremer N, Krasity BC, Apicella MA, McFall-Ngai MJ, Ruby EG (2014) A model symbiosis reveals a role for sheathed-flagellum rotation in the release of immunogenic lipopolysaccharide. elife 3:e01579. https://doi.org/10.7554/eLife.01579
Brogden KA (2005) Antimicrobial peptides: pore formers or metabolic inhibitors in bacteria? Nat Rev Microbiol 3(3):238–250. https://doi.org/10.1038/nrmicro1098
Brozek KA, Raetz CR (1990) Biosynthesis of lipid A in Escherichia coli. Acyl carrier protein-dependent incorporation of laurate and myristate. J Biol Chem 265(26):15410–15417
Byvalov AA, Malkova MA, Chernyad’ev AV, Dudina LG, Litvinets SG, Martinson EA (2018) Influence of specific bacteriophage on the level of vesicle formation and morphology of cells of Yersinia pseudotuberculosis. Bull Exp Biol Med 165(3):403–407. https://doi.org/10.1007/s10517-018-4180-0
Carpenter CD, Cooley BJ, Needham BD, Fisher CR, Trent MS, Gordon V, Payne SM (2014) The Vps/VacJ ABC transporter is required for intercellular spread of Shigella flexneri. Infect Immun 82(2):660–669. https://doi.org/10.1128/IAI.01057-13
Cascales E, Lloubes R (2004) Deletion analyses of the peptidoglycan-associated lipoprotein Pal reveals three independent binding sequences including a TolA box. Mol Microbiol 51(3):873–885
Casjens S (2003) Prophages and bacterial genomics: what have we learned so far? Mol Microbiol 49(2):277–300
Chang TW, Lin YM, Wang CF, Liao YD (2012) Outer membrane lipoprotein Lpp is Gram-negative bacterial cell surface receptor for cationic antimicrobial peptides. J Biol Chem 287(1):418–428. https://doi.org/10.1074/jbc.M111.290361
Choi HI, Kim M, Jeon J, Han JK, Kim KS (2017) Overexpression of MicA induces production of OmpC-enriched outer membrane vesicles that protect against Salmonella challenge. Biochem Biophys Res Commun 490(3):991–996. https://doi.org/10.1016/j.bbrc.2017.06.152
Clavel T, Germon P, Vianney A, Portalier R, Lazzaroni JC (1998) TolB protein of Escherichia coli K-12 interacts with the outer membrane peptidoglycan-associated proteins Pal, Lpp and OmpA. Mol Microbiol 29(1):359–367
Deatherage BL, Cookson BT (2012) Membrane vesicle release in bacteria, eukaryotes, and archaea: a conserved yet underappreciated aspect of microbial life. Infect Immun 80(6):1948–1957. https://doi.org/10.1128/IAI.06014-11
Deatherage BL, Lara JC, Bergsbaken T, Rassoulian Barrett SL, Lara S, Cookson BT (2009) Biogenesis of bacterial membrane vesicles. Mol Microbiol 72(6):1395–1407
Devos S, Van Putte W, Vitse J, Van Driessche G, Stremersch S, Van Den Broek W, Raemdonck K, Braeckmans K, Stahlberg H, Kudryashev M, Savvides SN, Devreese B (2017) Membrane vesicle secretion and prophage induction in multidrug-resistant Stenotrophomonas maltophilia in response to ciprofloxacin stress. Environ Microbiol 19(10):3930–3937. https://doi.org/10.1111/1462-2920.13793
Duperthuy M, Sjostrom AE, Sabharwal D, Damghani F, Uhlin BE, Wai SN (2013) Role of the Vibrio cholerae matrix protein Bap1 in cross-resistance to antimicrobial peptides. PLoS Pathog 9(10):e1003620. https://doi.org/10.1371/journal.ppat.1003620
Dutta S, Iida K, Takade A, Meno Y, Nair GB, Yoshida S (2004) Release of Shiga toxin by membrane vesicles in Shigella dysenteriae serotype 1 strains and in vitro effects of antimicrobials on toxin production and release. Microbiol Immunol 48(12):965–969
Elhenawy W, Bording-Jorgensen M, Valguarnera E, Haurat MF, Wine E, Feldman MF (2016) LPS remodeling triggers formation of outer membrane vesicles in Salmonella. mBio 7(4):e00940-16. https://doi.org/10.1128/mBio.00940-16
Feiner R, Argov T, Rabinovich L, Sigal N, Borovok I, Herskovits AA (2015) A new perspective on lysogeny: prophages as active regulatory switches of bacteria. Nat Rev Microbiol 13(10):641–650. https://doi.org/10.1038/nrmicro3527
Ferooz J, Letesson JJ (2010) Morphological analysis of the sheathed flagellum of Brucella melitensis. BMC Res Notes 3:333. https://doi.org/10.1186/1756-0500-3-333
Fuerst JA, Perry JW (1988) Demonstration of lipopolysaccharide on sheathed flagella of Vibrio cholerae O:1 by protein A-gold immunoelectron microscopy. J Bacteriol 170(4):1488–1494
Geis G, Suerbaum S, Forsthoff B, Leying H, Opferkuch W (1993) Ultrastructure and biochemical studies of the flagellar sheath of Helicobacter pylori. J Med Microbiol 38(5):371–377. https://doi.org/10.1099/00222615-38-5-371
Gerding MA, Ogata Y, Pecora ND, Niki H, de Boer PA (2007) The trans-envelope Tol-Pal complex is part of the cell division machinery and required for proper outer-membrane invagination during cell constriction in E. coli. Mol Microbiol 63(4):1008–1025. https://doi.org/10.1111/j.1365-2958.2006.05571.x
Gerritzen MJH, Martens DE, Uittenbogaard JP, Wijffels RH, Stork M (2019) Sulfate depletion triggers overproduction of phospholipids and the release of outer membrane vesicles by Neisseria meningitidis. Sci Rep 9(1):4716. https://doi.org/10.1038/s41598-019-41233-x
Gunn JS (2001) Bacterial modification of LPS and resistance to antimicrobial peptides. J Endotoxin Res 7(1):57–62
Gunn JS, Lim KB, Krueger J, Kim K, Guo L, Hackett M, Miller SI (1998) PmrA-PmrB-regulated genes necessary for 4-aminoarabinose lipid A modification and polymyxin resistance. Mol Microbiol 27:1171–1182
Gwozdzinski K (2018) Macromolecular modification of the cell wall of Gram-negative bacteria leading to antibiotic resistance and formation of outer membrane vesicles. Justus-Liebig-Universität Gießen, Gießen
Hagemann S, Stoger L, Kappelmann M, Hassl I, Ellinger A, Velimirov B (2014) DNA-bearing membrane vesicles produced by Ahrensia kielensis and Pseudoalteromonas marina. J Basic Microbiol 54(10):1062–1072. https://doi.org/10.1002/jobm.201300376
Hampton CM, Guerrero-Ferreira RC, Storms RE, Taylor JV, Yi H, Gulig PA, Wright ER (2017) The opportunistic pathogen Vibrio vulnificus produces outer membrane vesicles in a spatially distinct manner related to capsular polysaccharide. Front Microbiol 8:2177. https://doi.org/10.3389/fmicb.2017.02177
Hankins JV, Madsen JA, Giles DK, Brodbelt JS, Trent MS (2012) Amino acid addition to Vibrio cholerae LPS establishes a link between surface remodeling in gram-positive and gram-negative bacteria. Proc Natl Acad Sci U S A 109(22):8722–8727
Haurat MF, Aduse-Opoku J, Rangarajan M, Dorobantu L, Gray MR, Curtis MA, Feldman MF (2011) Selective sorting of cargo proteins into bacterial membrane vesicles. J Biol Chem 286(2):1269–1276. https://doi.org/10.1074/jbc.M110.185744
Hayashi J, Hamada N, Kuramitsu HK (2002) The autolysin of Porphyromonas gingivalis is involved in outer membrane vesicle release. FEMS Microbiol Lett 216(2):217–222
Hayden JD, Ades SE (2008) The extracytoplasmic stress factor, sigmaE, is required to maintain cell envelope integrity in Escherichia coli. PLoS One 3(2):e1573. https://doi.org/10.1371/journal.pone.0001573
Hoekstra D, van der Laan JW, de Leij L, Witholt B (1976) Release of outer membrane fragments from normally growing Escherichia coli. Biochim Biophys Acta 455(3):889–899
Horstman AL, Kuehn MJ (2000) Enterotoxigenic Escherichia coli secretes active heat-labile enterotoxin via outer membrane vesicles. J Biol Chem 275(17):12489–12496
Iwami J, Murakami Y, Nagano K, Nakamura H, Yoshimura F (2007) Further evidence that major outer membrane proteins homologous to OmpA in Porphyromonas gingivalis stabilize bacterial cells. Oral Microbiol Immunol 22(5):356–360
Kadurugamuwa JL, Beveridge TJ (1995) Virulence factors are released from Pseudomonas aeruginosa in association with membrane vesicles during normal growth and exposure to gentamicin: a novel mechanism of enzyme secretion. J Bacteriol 177(14):3998–4008
Kadurugamuwa JL, Beveridge TJ (1997) Natural release of virulence factors in membrane vesicles by Pseudomonas aeruginosa and the effect of aminoglycoside antibiotics on their release. J Antimicrob Chemother 40(5):615–621
Kadurugamuwa JL, Clarke AJ, Beveridge TJ (1993) Surface action of gentamicin on Pseudomonas aeruginosa. J Bacteriol 175(18):5798–5805
Kawasaki K, Ernst RK, Miller SI (2004a) 3-O-deacylation of lipid A by PagL, a PhoP/PhoQ-regulated deacylase of Salmonella typhimurium, modulates signaling through Toll-like receptor 4. J Biol Chem 279(19):20044–20048. https://doi.org/10.1074/jbc.M401275200
Kawasaki K, Ernst RK, Miller SI (2004b) Deacylation and palmitoylation of lipid A by Salmonellae outer membrane enzymes modulate host signaling through Toll-like receptor 4. J Endotoxin Res 10(6):439–444. https://doi.org/10.1179/096805104225006264
Kulp AJ, Sun B, Ai T, Manning AJ, Orench-Rivera N, Schmid AK, Kuehn MJ (2015) Genome-wide assessment of outer membrane vesicle production in Escherichia coli. PLoS One 10(9):e0139200. https://doi.org/10.1371/journal.pone.0139200
Lam JS, Taylor VL, Islam ST, Hao Y, Kocincova D (2011) Genetic and functional diversity of Pseudomonas aeruginosa lipopolysaccharide. Front Microbiol 2:118. https://doi.org/10.3389/fmicb.2011.00118
Lazzaroni JC, Germon P, Ray MC, Vianney A (1999) The Tol proteins of Escherichia coli and their involvement in the uptake of biomolecules and outer membrane stability. FEMS Microbiol Lett 177(2):191–197
Llamas MA, Ramos JL, Rodriguez-Herva JJ (2000) Mutations in each of the tol genes of Pseudomonas putida reveal that they are critical for maintenance of outer membrane stability. J Bacteriol 182(17):4764–4772
Macdonald IA, Kuehn MJ (2013) Stress-induced outer membrane vesicle production by Pseudomonas aeruginosa. J Bacteriol 195(13):2971–2981. https://doi.org/10.1128/JB.02267-12
Mahalakshmi S, Sunayana MR, SaiSree L, Reddy M (2014) yciM is an essential gene required for regulation of lipopolysaccharide synthesis in Escherichia coli. Mol Microbiol 91(1):145–157. https://doi.org/10.1111/mmi.12452
Malinverni JC, Silhavy TJ (2009) An ABC transport system that maintains lipid asymmetry in the gram-negative outer membrane. Proc Natl Acad Sci U S A 106(19):8009–8014. https://doi.org/10.1073/pnas.0903229106
Manning AJ, Kuehn MJ (2011) Contribution of bacterial outer membrane vesicles to innate bacterial defense. BMC Microbiol 11:258. https://doi.org/10.1186/1471-2180-11-258
Maredia R, Devineni N, Lentz P, Dallo SF, Yu J, Guentzel N, Chambers J, Arulanandam B, Haskins WE, Weitao T (2012) Vesiculation from Pseudomonas aeruginosa under SOS. TheScientificWorldJournal 2012:402919. https://doi.org/10.1100/2012/402919
Mashburn LM, Whiteley M (2005) Membrane vesicles traffic signals and facilitate group activities in a prokaryote. Nature 437(7057):422–425. https://doi.org/10.1038/nature03925
Mashburn-Warren L, Howe J, Garidel P, Richter W, Steiniger F, Roessle M, Brandenburg K, Whiteley M (2008) Interaction of quorum signals with outer membrane lipids: insights into prokaryotic membrane vesicle formation. Mol Microbiol 69(2):491–502
Mathur J, Davis BM, Waldor MK (2007) Antimicrobial peptides activate the Vibrio cholerae sigmaE regulon through an OmpU-dependent signalling pathway. Mol Microbiol 63(3):848–858
McBroom AJ, Kuehn MJ (2007) Release of outer membrane vesicles by Gram-negative bacteria is a novel envelope stress response. Mol Microbiol 63(2):545–558
McBroom AJ, Johnson AP, Vemulapalli S, Kuehn MJ (2006) Outer membrane vesicle production by Escherichia coli is independent of membrane instability. J Bacteriol 188(15):5385–5392
McCaig WD, Koller A, Thanassi DG (2013) Production of outer membrane vesicles and outer membrane tubes by Francisella novicida. J Bacteriol 195(6):1120–1132. https://doi.org/10.1128/JB.02007-12
McMahon HT, Boucrot E (2015) Membrane curvature at a glance. J Cell Sci 128(6):1065–1070. https://doi.org/10.1242/jcs.114454
Millikan DS, Ruby EG (2004) Vibrio fischeri flagellin A is essential for normal motility and for symbiotic competence during initial squid light organ colonization. J Bacteriol 186(13):4315–4325. https://doi.org/10.1128/JB.186.13.4315-4325.2004
Mitra S, Sinha R, Mitobe J, Koley H (2016) Development of a cost-effective vaccine candidate with outer membrane vesicles of a tolA-disrupted Shigella boydii strain. Vaccine 34(15):1839–1846. https://doi.org/10.1016/j.vaccine.2016.02.018
Moon DC, Choi CH, Lee JH, Choi CW, Kim HY, Park JS, Kim SI, Lee JC (2012) Acinetobacter baumannii outer membrane protein A modulates the biogenesis of outer membrane vesicles. J Microbiol 50(1):155–160. https://doi.org/10.1007/s12275-012-1589-4
Moore RA, Hancock RE (1986) Involvement of outer membrane of Pseudomonas cepacia in aminoglycoside and polymyxin resistance. Antimicrob Agents Chemother 30(6):923–926. https://doi.org/10.1128/aac.30.6.923
Murphy K, Park AJ, Hao Y, Brewer D, Lam JS, Khursigara CM (2014) Influence of O polysaccharides on biofilm development and outer membrane vesicle biogenesis in Pseudomonas aeruginosa PAO1. J Bacteriol 196(7):1306–1317. https://doi.org/10.1128/JB.01463-13
Nakao R, Ramstedt M, Wai SN, Uhlin BE (2012) Enhanced biofilm formation by Escherichia coli LPS mutants defective in Hep biosynthesis. PLoS One 7(12):e51241. https://doi.org/10.1371/journal.pone.0051241
Niehus E, Ye F, Suerbaum S, Josenhans C (2002) Growth phase-dependent and differential transcriptional control of flagellar genes in Helicobacter pylori. Microbiology 148(Pt 12):3827–3837. https://doi.org/10.1099/00221287-148-12-3827
Pal RR, Baidya AK, Mamou G, Bhattacharya S, Socol Y, Kobi S, Katsowich N, Ben-Yehuda S, Rosenshine I (2019) Pathogenic E. coli extracts nutrients from infected host cells utilizing injectisome components. Cell 177(3):683–696 e618. https://doi.org/10.1016/j.cell.2019.02.022
Palva AM (1983) ompA gene in the detection of Escherichia coli and other Enterobacteriaceae by nucleic acid sandwich hybridization. J Clin Microbiol 18(1):92–100
Papenfort K, Pfeiffer V, Mika F, Lucchini S, Hinton JC, Vogel J (2006) SigmaE-dependent small RNAs of Salmonella respond to membrane stress by accelerating global omp mRNA decay. Mol Microbiol 62(6):1674–1688
Paramonov N, Bailey D, Rangarajan M, Hashim A, Kelly G, Curtis MA, Hounsell EF (2001) Structural analysis of the polysaccharide from the lipopolysaccharide of Porphyromonas gingivalis strain W50. Eur J Biochem 268(17):4698–4707
Paramonov N, Rangarajan M, Hashim A, Gallagher A, Aduse-Opoku J, Slaney JM, Hounsell E, Curtis MA (2005) Structural analysis of a novel anionic polysaccharide from Porphyromonas gingivalis strain W50 related to Arg-gingipain glycans. Mol Microbiol 58(3):847–863. https://doi.org/10.1111/j.1365-2958.2005.04871.x
Parsons LM, Lin F, Orban J (2006) Peptidoglycan recognition by Pal, an outer membrane lipoprotein. Biochemistry 45(7):2122–2128. https://doi.org/10.1021/bi052227i
Perez-Cruz C, Carrion O, Delgado L, Martinez G, Lopez-Iglesias C, Mercade E (2013) New type of outer membrane vesicle produced by the Gram-negative bacterium Shewanella vesiculosa M7T: implications for DNA content. Appl Environ Microbiol 79(6):1874–1881. https://doi.org/10.1128/AEM.03657-12
Pirbadian S, Barchinger SE, Leung KM, Byun HS, Jangir Y, Bouhenni RA, Reed SB, Romine MF, Saffarini DA, Shi L, Gorby YA, Golbeck JH, El-Naggar MY (2014) Shewanella oneidensis MR-1 nanowires are outer membrane and periplasmic extensions of the extracellular electron transport components. Proc Natl Acad Sci U S A 111(35):12883–12888. https://doi.org/10.1073/pnas.1410551111
Prieto AI, Ramos-Morales F, Casadesus J (2006) Repair of DNA damage induced by bile salts in Salmonella enterica. Genetics 174(2):575–584. https://doi.org/10.1534/genetics.106.060889
Qin Z, Lin WT, Zhu S, Franco AT, Liu J (2016) Imaging the motility and chemotaxis machineries in Helicobacter pylori by cryo-electron tomography. J Bacteriol 199:e00695–e00616. https://doi.org/10.1128/JB.00695-16
Raetz CR, Reynolds CM, Trent MS, Bishop RE (2007) Lipid A modification systems in Gram-negative bacteria. Annu Rev Biochem 76:295–329. https://doi.org/10.1146/annurev.biochem.76.010307.145803
Raivio TL (2005) Envelope stress responses and Gram-negative bacterial pathogenesis. Mol Microbiol 56(5):1119–1128. https://doi.org/10.1111/j.1365-2958.2005.04625.x
Rangarajan M, Aduse-Opoku J, Paramonov N, Hashim A, Bostanci N, Fraser OP, Tarelli E, Curtis MA (2008) Identification of a second lipopolysaccharide in Porphyromonas gingivalis W50. J Bacteriol 190(8):2920–2932. https://doi.org/10.1128/JB.01868-07
Remis JP, Wei D, Gorur A, Zemla M, Haraga J, Allen S, Witkowska HE, Costerton JW, Berleman JE, Auer M (2014) Bacterial social networks: structure and composition of Myxococcus xanthus outer membrane vesicle chains. Environ Microbiol 16(2):598–610. https://doi.org/10.1111/1462-2920.12187
Roier S, Zingl FG, Cakar F, Durakovic S, Kohl P, Eichmann TO, Klug L, Gadermaier B, Weinzerl K, Prassl R, Lass A, Daum G, Reidl J, Feldman MF, Schild S (2016) A novel mechanism for the biogenesis of outer membrane vesicles in Gram-negative bacteria. Nat Commun 7:10515. https://doi.org/10.1038/ncomms10515
Ruhal R, Antti H, Rzhepishevska O, Boulanger N, Barbero DR, Wai SN, Uhlin BE, Ramstedt M (2015) A multivariate approach to correlate bacterial surface properties to biofilm formation by lipopolysaccharide mutants of Pseudomonas aeruginosa. Colloids Surf B Biointerfaces 127:182–191. https://doi.org/10.1016/j.colsurfb.2015.01.030
Ruiz N, Kahne D, Silhavy TJ (2006) Advances in understanding bacterial outer-membrane biogenesis. Nat Rev Microbiol 4(1):57–66. https://doi.org/10.1038/nrmicro1322
Sabra W, Lunsdorf H, Zeng AP (2003) Alterations in the formation of lipopolysaccharide and membrane vesicles on the surface of Pseudomonas aeruginosa PAO1 under oxygen stress conditions. Microbiology 149(Pt 10):2789–2795. https://doi.org/10.1099/mic.0.26443-0
Salkinoja-Salonen M, Nurmiaho EL (1978) The effect of lipopolysaccharide composition on the ultrastructure of Pseudomonas aeruginosa. J Gen Microbiol 105(1):23–28. https://doi.org/10.1099/00221287-105-1-23
Sampath V, McCaig WD, Thanassi DG (2018) Amino acid deprivation and central carbon metabolism regulate the production of outer membrane vesicles and tubes by Francisella. Mol Microbiol 107(4):523–541. https://doi.org/10.1111/mmi.13897
Samsudin F, Ortiz-Suarez ML, Piggot TJ, Bond PJ, Khalid S (2016) OmpA: a flexible clamp for bacterial cell wall attachment. Structure 24(12):2227–2235. https://doi.org/10.1016/j.str.2016.10.009
Schertzer JW, Whiteley M (2012) A bilayer-couple model of bacterial outer membrane vesicle biogenesis. mBio 3(2):e00297-11. https://doi.org/10.1128/mBio.00297-11
Schertzer JW, Whiteley M (2013) Bacterial outer membrane vesicles in trafficking, communication and the host-pathogen interaction. J Mol Microbiol Biotechnol 23(1–2):118–130. https://doi.org/10.1159/000346770
Schwechheimer C, Kuehn MJ (2013) Synthetic effect between envelope stress and lack of outer membrane vesicle production in Escherichia coli. J Bacteriol 195(18):4161–4173. https://doi.org/10.1128/JB.02192-12
Schwechheimer C, Sullivan CJ, Kuehn MJ (2013) Envelope control of outer membrane vesicle production in Gram-negative bacteria. Biochemistry 52(18):3031–3040. https://doi.org/10.1021/bi400164t
Schwechheimer C, Kulp A, Kuehn MJ (2014) Modulation of bacterial outer membrane vesicle production by envelope structure and content. BMC Microbiol 14:324. https://doi.org/10.1186/s12866-014-0324-1
Shen L, Gao X, Wei J, Chen L, Zhao X, Li B, Duan K (2012) PA2800 plays an important role in both antibiotic susceptibility and virulence in Pseudomonas aeruginosa. Curr Microbiol 65(5):601–609. https://doi.org/10.1007/s00284-012-0196-2
Shetty A, Hickey WJ (2014) Effects of outer membrane vesicle formation, surface-layer production and nanopod development on the metabolism of phenanthrene by Delftia acidovorans Cs1-4. PLoS One 9(3):e92143. https://doi.org/10.1371/journal.pone.0092143
Shetty A, Chen S, Tocheva EI, Jensen GJ, Hickey WJ (2011) Nanopods: a new bacterial structure and mechanism for deployment of outer membrane vesicles. PLoS One 6(6):e20725. https://doi.org/10.1371/journal.pone.0020725
Simmons LA, Foti JJ, Cohen SE, Walker GC (2008) The SOS Regulatory Network. EcoSal Plus 3(1). https://doi.org/10.1128/ecosalplus.5.4.3
Song T, Mika F, Lindmark B, Liu Z, Schild S, Bishop A, Zhu J, Camilli A, Johansson J, Vogel J, Wai SN (2008) A new Vibrio cholerae sRNA modulates colonization and affects release of outer membrane vesicles. Mol Microbiol 70(1):100–111
Sonntag I, Schwarz H, Hirota Y, Henning U (1978) Cell envelope and shape of Escherichia coli: multiple mutants missing the outer membrane lipoprotein and other major outer membrane proteins. J Bacteriol 136(1):280–285
Storm DR, Rosenthal KS, Swanson PE (1977) Polymyxin and related peptide antibiotics. Annu Rev Biochem 46:723–763. https://doi.org/10.1146/annurev.bi.46.070177.003451
Suzuki H, Nishimura Y, Yasuda S, Nishimura A, Yamada M, Hirota Y (1978) Murein-lipoprotein of Escherichia coli: a protein involved in the stabilization of bacterial cell envelope. Mol Gen Genet 167(1):1–9
Suzuki T, Murai T, Fukuda I, Tobe T, Yoshikawa M, Sasakawa C (1994) Identification and characterization of a chromosomal virulence gene, vacJ, required for intercellular spreading of Shigella flexneri. Mol Microbiol 11(1):31–41
Sych T, Mely Y, Romer W (2018) Lipid self-assembly and lectin-induced reorganization of the plasma membrane. Philos Trans R Soc Lond Ser B Biol Sci 373(1747):20170117. https://doi.org/10.1098/rstb.2017.0117
Taheri N, Mahmud A, Sandblad L, Fallman M, Wai SN, Fahlgren A (2018) Campylobacter jejuni bile exposure influences outer membrane vesicles protein content and bacterial interaction with epithelial cells. Sci Rep 8(1):16996. https://doi.org/10.1038/s41598-018-35409-0
Tarahovsky YS, Ivanitsky GR, Khusainov AA (1994) Lysis of Escherichia coli cells induced by bacteriophage T4. FEMS Microbiol Lett 122(1–2):195–199. https://doi.org/10.1111/j.1574-6968.1994.tb07164.x
Tashiro Y, Sakai R, Toyofuku M, Sawada I, Nakajima-Kambe T, Uchiyama H, Nomura N (2009) Outer membrane machinery and alginate synthesis regulators control membrane vesicle production in Pseudomonas aeruginosa. J Bacteriol 191(24):7509–7519
Tashiro Y, Ichikawa S, Nakajima-Kambe T, Uchiyama H, Nomura N (2010) Pseudomonas quinolone signal affects membrane vesicle production in not only gram-negative but also gram-positive bacteria. Microbes Environ 25(2):120–125. https://doi.org/10.1264/jsme2.me09182
Tashiro Y, Inagaki A, Shimizu M, Ichikawa S, Takaya N, Nakajima-Kambe T, Uchiyama H, Nomura N (2011) Characterization of phospholipids in membrane vesicles derived from Pseudomonas aeruginosa. Biosci Biotechnol Biochem 75(3):605–607
Toyofuku M, Zhou S, Sawada I, Takaya N, Uchiyama H, Nomura N (2014) Membrane vesicle formation is associated with pyocin production under denitrifying conditions in Pseudomonas aeruginosa PAO1. Environ Microbiol 16(9):2927–2938. https://doi.org/10.1111/1462-2920.12260
Toyofuku M, Carcamo-Oyarce G, Yamamoto T, Eisenstein F, Hsiao CC, Kurosawa M, Gademann K, Pilhofer M, Nomura N, Eberl L (2017) Prophage-triggered membrane vesicle formation through peptidoglycan damage in Bacillus subtilis. Nat Commun 8(1):481. https://doi.org/10.1038/s41467-017-00492-w
Trent MS, Pabich W, Raetz CR, Miller SI (2001) A PhoP/PhoQ-induced lipase (PagL) that catalyzes 3-O-deacylation of lipid A precursors in membranes of Salmonella typhimurium. J Biol Chem 276(12):9083–9092. https://doi.org/10.1074/jbc.M010730200
Turnbull L, Toyofuku M, Hynen AL, Kurosawa M, Pessi G, Petty NK, Osvath SR, Carcamo-Oyarce G, Gloag ES, Shimoni R, Omasits U, Ito S, Yap X, Monahan LG, Cavaliere R, Ahrens CH, Charles IG, Nomura N, Eberl L, Whitchurch CB (2016) Explosive cell lysis as a mechanism for the biogenesis of bacterial membrane vesicles and biofilms. Nat Commun 7:11220. https://doi.org/10.1038/ncomms11220
Turner L, Praszkier J, Hutton ML, Steer D, Ramm G, Kaparakis-Liaskos M, Ferrero RL (2015) Increased outer membrane vesicle formation in a Helicobacter pylori tolB mutant. Helicobacter 20(4):269–283. https://doi.org/10.1111/hel.12196
Udekwu KI, Wagner EG (2007) Sigma E controls biogenesis of the antisense RNA MicA. Nucleic Acids Res 35(4):1279–1288. https://doi.org/10.1093/nar/gkl1154
Urashima A, Sanou A, Yen H, Tobe T (2017) Enterohaemorrhagic Escherichia coli produces outer membrane vesicles as an active defence system against antimicrobial peptide LL-37. Cell Microbiol 19(11). https://doi.org/10.1111/cmi.12758
Vollmer W, Bertsche U (2008) Murein (peptidoglycan) structure, architecture and biosynthesis in Escherichia coli. Biochim Biophys Acta 1778(9):1714–1734. https://doi.org/10.1016/j.bbamem.2007.06.007
Wang Y (2002) The function of OmpA in Escherichia coli. Biochem Biophys Res Commun 292(2):396–401. https://doi.org/10.1006/bbrc.2002.6657
Weinbauer MG (2004) Ecology of prokaryotic viruses. FEMS Microbiol Rev 28(2):127–181. https://doi.org/10.1016/j.femsre.2003.08.001
Wensink J, Witholt B (1981) Outer-membrane vesicles released by normally growing Escherichia coli contain very little lipoprotein. Eur J Biochem 116(2):331–335
Yeh YC, Comolli LR, Downing KH, Shapiro L, McAdams HH (2010) The Caulobacter Tol-Pal complex is essential for outer membrane integrity and the positioning of a polar localization factor. J Bacteriol 192(19):4847–4858. https://doi.org/10.1128/JB.00607-10
Zavan L, Bitto NJ, Johnston EL, Greening DW, Kaparakis-Liaskos M (2019) Helicobacter pylori growth stage determines the size, protein composition, and preferential cargo packaging of outer membrane vesicles. Proteomics 19(1–2):e1800209. https://doi.org/10.1002/pmic.201800209
Zhao L, Gao X, Liu C, Lv X, Jiang N, Zheng S (2017) Deletion of the vacJ gene affects the biology and virulence in Haemophilus parasuis serovar 5. Gene 603:42–53. https://doi.org/10.1016/j.gene.2016.12.009
Zimmerberg J, Kozlov MM (2006) How proteins produce cellular membrane curvature. Nat Rev Mol Cell Biol 7(1):9–19. https://doi.org/10.1038/nrm1784
Acknowledgements
This work was supported by the BioTechMed-Graz Flagship Project SECRETOME to S.S, the Austrian FWF grants P25691 to S.S., P27654 to S.S., and W901-B12 (DK Molecular Enzymology) to F.G.Z. and S.S.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Zingl, F.G., Leitner, D.R., Schild, S. (2020). Biogenesis of Gram-Negative OMVs. In: Kaparakis-Liaskos, M., Kufer, T. (eds) Bacterial Membrane Vesicles. Springer, Cham. https://doi.org/10.1007/978-3-030-36331-4_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-36331-4_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-36330-7
Online ISBN: 978-3-030-36331-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)