Abstract
The production of extracellular vesicles is a conserved process that is common to all living cells. Both Gram-negative and Gram-positive bacteria produce extracellular vesicles, known as outer membrane vesicles (OMVs) and membrane vesicles (MVs), respectively. Once disregarded as artifacts of bacterial growth, research over the last 50 years has shown that OMVs contribute to numerous bacterial functions. It is now understood that OMVs are purposely secreted by Gram-negative bacteria to aid in bacterial communication and pathogenesis. The OMV field has focused on understanding the mechanisms of OMV biogenesis, the content of OMVs and how OMVs interact with the host immune system and their environment. While there is a wealth of knowledge regarding OMVs, it was only in the last decade that Gram-positive bacteria were found to release MVs. Due to the late discovery of MVs there is little known about MVs in comparison to our knowledge regarding OMVs. However, there is emerging evidence that MVs contain bacterial cargo and may aid in bacterial functions. Research in the field of bacterial vesicles has expanded rapidly within the past decade and continues to be a growing field of interest. Future work aims to manipulate bacterial membrane vesicles as novel therapeutics and nanoparticle technology.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1.1 Introduction to Gram-Negative OMVs
All forms of life, prokaryotic and eukaryotic, naturally release extracellular vesicles as part of their normal growth (Brown et al. 2015; Deatherage and Cookson 2012). Vesicles produced by Gram-negative bacteria are called outer membrane vesicles (OMVs) as they are derived from the outer membrane of the Gram-negative bacterial cell (Hoekstra et al. 1976). First dismissed as bacterial artifacts, early studies visualized OMVs being released from the outer membrane of a range of Gram-negative pathogens by electron microscopy (Knox et al. 1966; Chatterjee and Das 1967). However, it was not until OMVs were identified in the spinal fluid of meningococcal patients (DeVoe and Gilchrist 1975), that interest developed in understanding OMV production, their functions in the host and how they benefit bacteria.
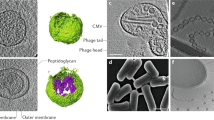
It is now accepted that OMVs are purposely secreted by Gram-negative bacteria to aid in an array of bacterial functions. OMVs range from approximately 20–400 nm in size and contain materials derived from their parent bacterium, including nucleic acids, proteins and enzymes (Fig. 1.1) (Kadurugamuwa and Beveridge 1995; Dorward et al. 1989; Dorward and Garon 1989; Haurat et al. 2011; reviewed in Schwechheimer and Kuehn 2015). It was originally thought that OMV cargo was derived from the bacterial outer membrane and periplasm only, as cytoplasmic components were thought to be unable to cross the inner membrane (Hoekstra et al. 1976; Gankema et al. 1980). However, it is now known that OMVs can also contain components derived from the bacterial cytoplasm, such as nucleic acids and proteins (Lee et al. 2007; Perez-Cruz et al. 2015; Bitto et al. 2017; Renelli et al. 2004; Sjostrom et al. 2015).
OMVs are involved in a range of bacterial functions. Research over the last two decades has highlighted the importance of OMVs in cell-to-cell communication (Mashburn and Whiteley 2005), the transfer of genetic material (Yaron et al. 2000; Dorward et al. 1989), biofilm formation (Yonezawa et al. 2009), inflammation and disease progression (Ismail et al. 2003; Kaparakis et al. 2010; reviewed in Bitto and Kaparakis-Liaskos 2017).
1.1.1 The First Observations of OMVs
OMVs produced by Escherichia coli were the first OMVs to be observed using electron microscopy, appearing as small, spherical “particles” that surrounded the bacterial cell (Knox et al. 1966). These particles were thought to be responsible for the secretion of lipopolysaccharide (LPS) and lipoproteins from bacteria (Knox et al. 1966). Subsequently, OMVs were isolated from the oral bacterium Veillonella parvula by phenol–water extraction (Mergenhagen et al. 1966). These isolated OMVs were heterogeneous in size and the outer leaflet was similar in morphology to the outer membrane of V. parvula cells (Mergenhagen et al. 1966). The release of OMVs from Vibrio cholerae was subsequently observed and it was noted that OMV production occurred only during the log phase of bacterial growth (Chatterjee and Das 1967). Researchers postulated that V. cholerae regulated the release of OMVs from bacterial cells during active growth as a mechanism to secrete bacterial toxins into the extracellular environment (Chatterjee and Das 1967). Despite these first observations, the wider field did not consider OMVs as important or necessary products released by bacteria but merely viewed OMVs as artifacts of bacterial growth.
As research progressed, the release of OMVs from the outer membrane of bacterial cells was proposed to be a continuous and essential process, and not the result of cell lysis (Rothfield and Pearlman-Kothencz 1969; Loeb 1974; Gankema et al. 1980). Importantly, for the first time it was discovered that E. coli and Salmonella enterica serovar Typhimurium (S. Typhimurium) OMVs contained bacterial proteins, lipids and LPS derived from their parent bacteria (Rothfield and Pearlman-Kothencz 1969). Additionally, OMV production was seen to increase when bacterial protein synthesis was inhibited, which was speculated to have been due to stress of the outer membrane (Rothfield and Pearlman-Kothencz 1969). OMVs were next observed to be released by Neisseria meningitidis and were thought to be associated with the release of N. meningitidis toxins into the environment (Devoe and Gilchrist 1973). Similar to V. cholera, N. meningitidis OMVs were not detected as bacteria progressed to stationary phase of growth (Devoe and Gilchrist 1973) supporting the theory that OMVs were only produced during the log phase of bacterial growth. In addition to being released during the exponential phase of bacterial growth, OMVs were also observed to be released in response to treatment with detergents (Leive et al. 1968) and exposure to stress from bacteriophages (Loeb 1974) suggesting they are produced in response to bacterial stress.
1.1.2 Advances in OMV Research
Until 1975, OMV production had only been observed in vitro. OMVs were identified in primary cultures of spinal fluid taken from patients with meningococcal disease (DeVoe and Gilchrist 1975) indicating that the release of OMVs was a normal part of bacterial infection. This was the first study to identify OMVs released from pathogenic bacteria in a physiologically relevant setting.
A decade after their discovery, OMVs were given the name outer membrane vesicles as they closely resembled the outer membrane of their parent bacterium in composition, and were thought to be lacking in cytoplasmic material (Hoekstra et al. 1976). Subsequently, the release of OMVs from E. coli was shown to preferentially occur in locations of the outer membrane that contained newly synthesized proteins (Mug-Opstelten and Witholt 1978). One of the first hypotheses of OMV biogenesis suggested that the incorporation of new proteins into the outer membrane enabled a portion of the outer membrane to bulge from the cell and once large enough, to be released from the bacterial cell (Mug-Opstelten and Witholt 1978). Numerous subsequent studies further elucidated the composition of OMVs from Gram-negative bacterial species including E. coli (Gankema et al. 1980; Wensink and Witholt 1981), Aeromonas spp. (MacIntyre et al. 1980), Brucella melitensis (Gamazo and Moriyon 1987), and Haemophilus influenzae (Deich and Hoyer 1982).
As OMVs were known to contain LPS, they were suggested to be able to interact with host cells (MacIntyre et al. 1980). Functional studies of OMVs produced by Porphyromonas gingivalis suggested that OMVs may contribute to the progression of periodontal disease, as P. gingivalis OMVs contained bacterial toxins and enzymes and promoted bacterial adhesion (Grenier and Mayrand 1987). Importantly, it was found that immunization of mice with OMVs from H. influenzae type B resulted in an increase in the permeability of the blood–brain barrier, a similar response to that observed when mice were treated with H. influenzae LPS (Wispelwey et al. 1989). These studies highlighted the importance of OMVs in an infection setting and demonstrated the role of OMVs as vehicles for bacterial cargo. Furthermore, these studies were some of the first to describe how OMVs may contribute to bacterial pathogenesis.
Due to their pathogenic cargo, OMVs were also thought to be ideal vaccine candidates. The first OMV vaccine was trialed in 1991 against the pathogen N. meningitidis, which causes group B meningococcal disease (Bjune et al. 1991). Subsequent studies determined that three doses of the OMV-based vaccine increased the vaccine efficiency and therefore conferred protection, leading to the production of the MenB vaccine (Rosenqvist et al. 1995) that is now licensed for human use (Arnold et al. 2011; Vernikos and Medini 2014). Currently, there are ongoing efforts to develop new vaccines for other diseases caused by Gram-negative pathogens (Chen et al. 2010; Nieves et al. 2011).
1.1.3 Outer Membrane Vesicles Research in the Last Decade
OMVs from numerous pathogenic bacteria have been investigated for their ability to induce an immune response in host cells. OMVs isolated from Helicobacter pylori and Pseudomonas aeruginosa can induce an interleukin-8 (IL-8) response in host epithelial cells, resulting in inflammation (Ismail et al. 2003; Bauman and Kuehn 2006). Additionally, OMVs isolated from Treponema denticola can disrupt the epithelial cell layer and cross to the basolateral side of epithelial cells (Chi et al. 2003). Furthermore H. pylori OMVs can enter host epithelial cells via lipid rafts and interact with intracellular nucleotide-binding oligomerization domain-containing protein 1 (NOD1) causing an inflammatory response in host cells (Kaparakis et al. 2010; Allison et al. 2009). We now have a greater understanding of how OMVs can interact with and sometimes cross the host epithelial cell layer to elicit a pro-inflammatory immune response in the host. These studies highlight the important role of OMVs in contributing to the immunogenicity of bacteria.
While there was increasing interest in understanding the inflammatory nature of OMVs, there was still relatively little known about their production. Although early studies hypothesized the mechanisms of OMV biogenesis, the last decade of research has emphasized that OMV biogenesis is a complex and varied process that is still not well understood. It is now known that OMV release can be mediated by membrane proteins, LPS, O-polysaccharides, and phospholipids (Murphy et al. 2014; Roier et al. 2016; Elhenawy et al. 2016). Future studies should aim to identify other novel mechanisms of OMV biogenesis that may be either conserved to specific bacterial species or common to all Gram-negative bacteria.
Due to the nature of their biogenesis, there has been debate as to whether cargo is selectively packaged into OMVs. Selective packaging has since been identified in P. gingivalis, where OMVs were enriched in virulence proteins such as gingipains, while excluding numerous outer membrane proteins (Haurat et al. 2011). There is now greater interest in providing in-depth proteomic analyses of OMVs to further elucidate OMV content. For example, the proteomes of E. coli, P. aeruginosa, and H. pylori have been examined to provide insights into how bacterial growth stage, biofilms, and infection settings can determine the protein content of OMVs (Zavan et al. 2019; Ayalew et al. 2013; Pierson et al. 2011; Turner et al. 2018; Park et al. 2015) highlighting that there are a number of conditions that can determine OMV composition.
These works highlight only some areas of OMV research that has been the focus in recent years. Interest in OMVs has vastly increased over the last decade and continued research shows there is still much that remains unknown. Future efforts may focus on understanding what regulates OMV production and composition, how OMV content can be used by recipient bacteria, and further elucidating the role of OMVs in inflammation and disease. Collectively, these studies will broaden our knowledge regarding OMVs and will facilitate their development as novel therapeutics.
1.1.4 Biogenesis of OMVs
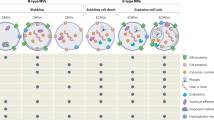
The Gram-negative cell membrane is composed of the outer membrane, inner membrane, and the periplasmic space, which contains a thick peptidoglycan layer (reviewed in Costerton et al. 1974). Embedded within the membranes and connecting them together are proteins which allow the bacterial cell to maintain its shape (Schnaitman 1970). Additionally, the bacterial outer membrane contains lipids, lipoproteins, and LPS that dictate membrane fluidity, curvature, and integrity (reviewed in Schwechheimer and Kuehn 2015). Disruption to these fundamental building blocks of the bacterial outer membrane can result in changes to OMV biogenesis. Here we summarize some of the mechanisms of OMV biogenesis, and a detailed discussion of this topic can be found in Chap. 2.
One mechanism of OMV biogenesis observed in numerous Gram-negative species is a process known as budding or blebbing. Blebbing of OMVs occurs when a portion of the outer membrane bulges at the cell surface and is liberated from the membrane to create a vesicle (reviewed in Schwechheimer and Kuehn 2015). Blebbing of OMVs from the outer membrane has been studied in numerous bacterial species and can be the result of a disruption at the cell membrane during protein modification or lipid remodeling (Bernadac et al. 1998; Elhenawy et al. 2016).
Protein modifications that occur in the outer membrane and surrounding regions are known to affect the production of OMVs in differing ways. For example, mutations in tolB or tolC of the Tol-Pal complex spanning the inner and outer membrane of E. coli cause a decrease in the release of OMVs, while in H. pylori Tol-Pal mutants cause an increase in OMV production (Turner et al. 2015; Bernadac et al. 1998). Additionally, the overexpression or misfolding of membrane proteins can cause vesiculation to increase up to 100-fold, which may be a response to the increasing pressure at the outer membrane (McBroom and Kuehn 2007; reviewed in Vasilyeva et al. 2009).
Furthermore, other key components of the bacterial outer membrane such as phospholipids and LPS have been implicated in OMV biogenesis. For example, the accumulation of phospholipids in the outer membrane of H. influenzae and V. cholerae can regulate OMV biogenesis (Roier et al. 2016). Additionally, the remodeling of Lipid A in phospholipids of S. Typhimurium was found to be required for the formation of OMVs (Elhenawy et al. 2016).
Moreover, research has shown that explosive cell lysis events in P. aeruginosa, caused by the production of prophage endolysins, can also result in OMV production (Turnbull et al. 2016). This mechanism has not been observed for other bacterial species; however, research continues to investigate the numerous mechanisms of OMV biogenesis.
While there have been numerous studies detailing specific changes in the outer membrane that result in OMV release, there are other circumstances that lead to variation in OMV production. Cellular stresses such as growth conditions (Park et al. 2015), bacterial growth stage (Zavan et al. 2019), changes in temperature (McMahon et al. 2012), or the presence of antibiotics (Kadurugamuwa and Beveridge 1997, 1995) can all alter OMV biogenesis.
The mechanisms described are widely varied but highlight the numerous factors which influence the release of OMVs. However, future research is needed to understand what other factors may impact or regulate OMV production. Expanding our knowledge regarding the mechanisms of Gram-negative OMV biogenesis will provide further understanding of the regulation of OMV composition and subsequent functions by bacteria.
1.1.5 Outer Membrane Vesicles in Bacterial Communication
OMVs contain parental proteins, enzymes, and nucleic acids that can be delivered to surrounding bacterial cells (Kadurugamuwa and Beveridge 1995; Mashburn and Whiteley 2005; Dorward et al. 1989; Bomberger et al. 2009). Importantly, it has been shown that P. aeruginosa OMVs carrying proteins and toxins from their parent bacteria could kill neighboring bacterial cells (Kadurugamuwa and Beveridge 1996). This research has since sparked interest into examining the ability of OMVs to confer a selective advantage to their parent bacterium.
OMVs from Neisseria gonorrhoeae, Acinetobacter baylyi and P. aeruginosa can carry both chromosomal and plasmid DNA derived from their parent bacteria (Dorward et al. 1989; Fulsundar et al. 2014; Renelli et al. 2004). DNA contained in some OMVs can be transferred to neighboring cells, including DNA that encodes for antibiotic resistance, highlighting the potential role of OMVs in horizontal gene transfer. Furthermore, it has recently been discovered that OMVs can contain RNA including messenger RNA (mRNA), ribosomal RNA (rRNA), and small RNA (sRNA) (Blenkiron et al. 2016; Koeppen et al. 2016; Sjostrom et al. 2015; Choi et al. 2017). Additionally, it was recently shown that OMVs can deliver sRNA to host cells, where the sRNA modulates the innate immune molecules of the host (Koeppen et al. 2016; Choi et al. 2017). However, it is still not well understood how or why RNA is packaged into OMVs.
OMVs can package other bacterial molecules such as the Pseudomonas quinolone signal (PQS) molecule, which contributes to cell-to-cell communication and coordination of the formation of biofilms (Mashburn and Whiteley 2005; Pesci et al. 1999). Removal of OMVs from P. aeruginosa cultures stops bacterial communication and group behaviors that are mediated by PQS, highlighting OMVs as a key contributor to cell-to-cell communication of P. aeruginosa (Mashburn and Whiteley 2005). Collectively, these works highlight the variety of materials that can be packaged into OMVs and how bacteria can use OMVs to communicate and interact with neighboring cells. A more detailed discussion about the ability of OMVs to function in inter-bacterial communication can be found in Chap. 5.
1.1.6 Outer Membrane Vesicles in Host–Pathogen Interactions
Due to the nature of their contents, OMVs are immunostimulatory to eukaryotic hosts. OMVs contain a number of microbe-associated molecular patterns (MAMPs) such as LPS, DNA, and peptidoglycan, all of which are capable of inducing a host pro-inflammatory response (reviewed in Ellis and Kuehn 2010). Toll-like receptors (TLRs) are pattern recognition receptors (PRRs) located on the membranes of host cells that can detect bacterial MAMPs (reviewed in Kawai and Akira 2010). TLR4 has been shown to detect LPS contained in E. coli and N. meningitidis OMVs, with the detection of E. coli OMVs leading to the production of pro-inflammatory cytokines (Mirlashari and Lyberg 2003; Soderblom et al. 2005). Alternatively, when OMVs interact with epithelial cells they can be internalized by lipid rafts on the cell surface and their peptidoglycan cargo can be detected by cytosolic NOD1 (Kaparakis et al. 2010). The detection of OMVs by PRRs activates a signaling cascade, resulting in the initiation of an innate immune response, the production of pro-inflammatory cytokines and the recruitment of immune cells (Kaparakis et al. 2010; Ismail et al. 2003; Bielig et al. 2011). These reports represent a small portion of the research that has been undertaken to determine how bacterial OMVs interact with innate immune receptors of the host to mediate inflammation. They also highlight the ability of OMVs to function as an important secretory system for immunostimulatory molecules and demonstrates their role in bacterial infection and inflammation. A more detailed discussion of the pathogenic and immunostimulatory functions of OMVs can be found in Chaps. 7 and 8.
Commensal bacteria have adapted mechanisms to enable their persistence in the host (reviewed in Hooper and Gordon 2001; Hooper 2004). Commensal bacteria that reside in the gut cannot cross the mucus layer to interact directly with epithelial cells (Johansson et al. 2008). However, it was recently identified that commensal and probiotic bacteria such as Bacteroides fragilis (Shen et al. 2012) and E. coli (Cañas et al. 2016) are capable of producing OMVs. These OMVs can be used as a bacterial delivery system, as they can cross the mucus layer and enter host epithelial cells via endocytic pathways (Cañas et al. 2016). It is now known that OMVs from commensal bacteria are able to modulate the host immune system to prevent inflammation and protect against diseases such as colitis (Shen et al. 2012; Kang et al. 2013). Additionally, OMVs isolated from commensal and probiotic E. coli prime the host immune system via NOD1 activation which may aid in the elimination of pathogenic bacteria (Cañas et al. 2018). These recent works highlight that OMVs from commensal bacteria may interact with the host immune system to maintain host–microbe homeostasis and may aid in the prevention of infections, and these topics are discussed in further detail in Chap. 9.
1.1.7 Distribution of OMVs in the Environment
Although OMVs have been thoroughly studied in the context of human pathogenic bacteria, more recently we have begun to understand their presence and functions in the environment. Here we give a brief overview of the distribution of bacterial OMVs in the environment, and this topic is discussed in further detail in Chap. 4. Two marine bacterial species, Prochlorococcus sp. and Synechoccocus sp., were found to be able to produce OMVs in their ecosystem (Biller et al. 2014). OMVs produced by Prochlorococcus sp. contained carbon and were able to aid in the growth of other marine bacterial species as the sole carbon source provided (Biller et al. 2014). Along with carbon, Prochlorococcus sp. OMVs contain DNA, RNA, and a range of proteins. In addition to the presence of OMVs in marine ecosystems, early research had identified OMVs were produced by freshwater bacterial biofilms (Beveridge 1999). However, it was not until recently that freshwater OMVs were further explored. Electron microscopy images of autotrophic freshwater bacterial species showed the release of OMVs from the outer membrane of bacterial cells into the environment (Silva et al. 2014). However, the importance of OMVs in freshwater aquatic environments is currently unknown and requires further investigation.
Environmental bacteria predominantly reside in the form of biofilms (Costerton et al. 1978). Biofilms are composed of a mucus layer containing bacteria in a scaffold-like structure known as the extracellular matrix. The extracellular matrix of biofilms contains exopolysaccharides, proteins, and extracellular DNA (eDNA) (Danese et al. 2000; Allesen-Holm et al. 2006; Jurcisek and Bakaletz 2007; Whitchurch et al. 2002). It was shown that P. aeruginosa OMVs make up an important and necessary component of the biofilm matrix (Schooling and Beveridge 2006; Whitchurch et al. 2002). It is now known that OMV size and content can differ between biofilm and planktonic cultures, as demonstrated for P. aeruginosa and H. pylori, suggesting that the role of OMVs in bacterial biofilms determines their composition (Park et al. 2015; Grande et al. 2015). These works highlight biofilm OMVs as an important component of the extracellular matrix and suggests that changes in OMV composition are in response to the role and necessity of OMVs in biofilms.
Finally, OMVs have recently been found within household environments. Gram-negative OMVs have been identified in household dust in the air and in mattresses (Kim et al. 2013). It was speculated that dust OMVs may be inhaled by residents and internalized by epithelial cells of the airway to cause disease. Mouse models have shown that internalization of dust OMVs leads to an inflammatory response that can be blocked by Polymyxin B (Kim et al. 2013). This suggests that like pathogenic OMVs, LPS from dust OMVs is detected by PRRs and can cause an inflammatory response (Kim et al. 2013).
Collectively, these studies indicate that Gram-negative OMVs can be identified in a number of environments suggesting that they are an essential part of bacterial growth and survival. As research continues, the extent to which OMVs can be found in the environment and the roles that OMVs play in these settings will become apparent.
1.2 Introduction to Gram-Positive MVs
The last decade of research has uncovered that Gram-positive bacteria can also produce vesicles, known as membrane vesicles (MVs). The discovery of Gram-positive MVs occurred much later than the discovery of Gram-negative OMVs, as researchers thought that the thick cell wall that surrounds Gram-positive bacteria would prevent the release of MVs. Despite this, MVs were reported to be produced by Gram-positive bacteria as early as 1976 (Bisschop and Konings 1976), as well as in a number of other early reports, however, these findings were dismissed by the wider bacterial vesicle field (Dorward and Garon 1990; Ruhr and Sahl 1985). Here we provide a brief discussion of the discovery, biogenesis, and functions of Gram-positive MVs, and an extensive review of this topic can be found in Chap. 3.
In 2009, electron microscopy showed for the first time the release of MVs from the surface of the Gram-positive organism, Staphylococcus aureus (Lee et al. 2009). This renewed interest in the existence of Gram-positive MVs, and soon reports emerged of MVs being produced by other Gram-positive species, including Bacillus anthracis (Rivera et al. 2010), Listeria monocytogenes (Lee et al. 2013b), Clostridium perfringens (Jiang et al. 2014), and Streptococcus sp. (Liao et al. 2014; Resch et al. 2016).
Due to the years between the discovery of Gram-negative OMVs and Gram-positive MVs, MVs remain poorly understood in comparison to their Gram-negative counterparts. While interest in MVs is increasing, there is still much to be uncovered surrounding their roles in inter-bacterial communication and host–pathogen interactions.
1.2.1 Production and Biogenesis of Gram-Positive MVs
Gram-positive MVs are similar in size to Gram-negative OMVs, ranging from 20 to 400 nm (Jiang et al. 2014; Brown et al. 2014; Haas and Grenier 2015; Tartaglia et al. 2018). However, the mechanism of Gram-positive MV biogenesis and release through their thick peptidoglycan layer is unclear. Studies have suggested that surfactant-like enzymes may be involved in disrupting the cytoplasmic membrane (Wang et al. 2018; Schlatterer et al. 2018), as well as endolysins that may alter the permeability of the peptidoglycan-rich cell wall thereby enabling the release of MVs (Toyofuku et al. 2017). It has also been suggested that cytoskeletal changes may contribute to the formation of MVs (Mayer and Gottschalk 2003). Furthermore, MV production is increased during stress conditions such as antibiotic exposure (He et al. 2017; Andreoni et al. 2019) suggesting that environmental factors may regulate their biogenesis. However, since MV biogenesis is still in the early stages of exploration, more studies are needed to reach a consensus on their mechanisms of biogenesis and the factors that influence it.
1.2.2 Contents of Gram-Positive MVs
The first characterization of Gram-positive MVs was a proteomic study of S. aureus MVs (Lee et al. 2009). These findings revealed that S. aureus MVs contain a variety of proteins that may serve biological roles in inter-bacterial communication, antibiotic resistance, virulence, and regulation of MV biogenesis (Lee et al. 2009). Moreover, this study suggested an enrichment of specific proteins in MVs compared to their parent bacteria indicating a selective packaging of protein cargo (Lee et al. 2009). Further studies confirmed that S. aureus MVs carry a range of pathogenic proteins including beta-lactamase (Lee et al. 2013a), alpha-toxin (Thay et al. 2013), and other virulence-related proteins (Lee et al. 2013b; Tartaglia et al. 2018). Similar findings have since been reported for MVs isolated from other Gram-positive species, including B. anthracis (Rivera et al. 2010), Enterococcus faecium (Wagner et al. 2018), C. perfringens (Jiang et al. 2014), L. monocytogenes (Coelho et al. 2019), and Streptococcus sp. (Haas and Grenier 2015; Resch et al. 2016). These studies suggest that MVs may serve as a Gram-positive secretion system for the delivery of biologically active proteins.
Early reports suggested that Gram-positive MVs do not carry nucleic acids (Dorward and Garon 1990). However, more recent findings have demonstrated that MVs from a variety of Gram-positive species contain DNA and RNA, including C. perfringens (Jiang et al. 2014), Streptococcus sp. (Liao et al. 2014; Resch et al. 2016), and Lactobacillus reuteri (Grande et al. 2017). While it is unclear how this DNA is packaged, the amount of DNA contained in Streptococcus MVs changes at different growth stages, suggesting that this process may be regulated by their parent bacterium during bacterial growth (Liao et al. 2014). Moreover, there are currently only a few reports describing the detection of RNA associated with Gram-positive MVs. RNA species detected in MVs include ribosomal RNA (rRNA) (Resch et al. 2016), transfer RNA (tRNA) (Resch et al. 2016), and small RNA (sRNA) (Choi et al. 2018). Differences in the abundance of RNA species carried by MVs when compared to their parent bacteria suggests that RNA may be selectively packaged into MVs (Resch et al. 2016).
These studies highlight that Gram-positive MVs can contain a range of molecules from their parent bacterium including proteins and nucleic acids. While it is still not well understood as to how and why these molecules are packaged into MVs, researchers are beginning to understand how the contents of MVs may aid in bacterial functions.
1.2.3 Role of MVs in Inter-Bacterial Communication
While there are limited studies describing the role of MVs in aiding bacterial functions compared to OMVs, their contents suggest that MVs are involved in inter-bacterial communication and the delivery of molecules between bacteria (reviewed in Brown et al. 2015). Proteins with bacteriolytic function and proteins that may facilitate transfer of molecules in an inter-bacterial manner have been identified in S. aureus MVs (Lee et al. 2009). Additionally, MVs have been implicated in biofilm production and formation. For example, DNA contained in Streptococcus mutans MVs are thought to be a component of S. mutans biofilms (Liao et al. 2014), while S. aureus MV production is upregulated during biofilm formation (He et al. 2017). Additionally, recent work determined that E. faecium MVs carry proteins that facilitate the production of bacterial biofilms (Wagner et al. 2018). While horizontal gene transfer via MVs is yet to be demonstrated, transfer of functional beta-lactamase protein via MVs has been shown, whereby MVs from an ampicillin resistant S. aureus strain transferred resistance to ampicillin-sensitive strains of E. coli, Salmonella enterica ser. Enteritidis, and Staphylococcus sp. via the transfer of the BlaZ protein (Lee et al. 2013a). These studies indicate that MVs may play an important role in communicating with other bacterial cells in the environment to promote bacterial survival.
1.2.4 Role of MVs in Host–Pathogen Interactions
Like OMVs, MVs are able to interact with eukaryotic host cells, including both epithelial cells (Gurung et al. 2011; Kim et al. 2012) and immune cells (Haas and Grenier 2015; Rivera et al. 2010; Jiang et al. 2014). Although there are few studies investigating the mechanisms of MV entry into target cells, there is evidence that S. aureus MVs enter host cells via cholesterol-dependent fusion (Thay et al. 2013), and that they are likely to enter host cells via a number of other mechanisms. Entry into host cells enables MVs to deliver their immunogenic cargo to mediate pathogenesis, similar to Gram-negative OMVs.
MVs carry a range of MAMPs, however, there is limited knowledge surrounding the innate and adaptive immune responses that they induce. Reports have shown that S. aureus MVs induce inflammation and cell death in host cells (Gurung et al. 2011; Jeon et al. 2016; Hong et al. 2011; Jun et al. 2017). The first reports of MVs activating innate immune pathways showed that S. aureus MVs activate TLR2 and NOD2, leading to the production of pro-inflammatory cytokines (Hong et al. 2011; Jun et al. 2017; Kim et al. 2012). Furthermore, MVs isolated from feces were shown to cause sepsis through the activation of TLR2 (Park et al. 2018).
The ability of Gram-positive MVs to induce adaptive immune responses has also been reported. C. perfringens MVs were shown to produce high-titer immunoglobulin G1 (IgG1) responses in mice (Jiang et al. 2014), while B. anthracis MVs produce a robust IgM response in mice when they encounter toxins carried by the MVs (Rivera et al. 2010). Due to the ability of MVs to activate the adaptive immune response, researchers have investigated their efficacy as a vaccine platform. Studies have shown that MVs from Streptococcus pneumoniae induce a protective response in mice when exposed to bacterial challenge (Olaya-Abril et al. 2014). Similarly, administration of S. aureus MVs to mice has been shown to be protective against S. aureus lung infection (Choi et al. 2015). These studies indicate that MVs warrant further investigation into their potential as alternative vaccine candidates.
Compared to OMV research, these few studies highlight that there is still little knowledge regarding how MVs interact with the innate and adaptive immune system of their host. Future work focused on how MVs from a variety of Gram-positive bacteria can enter host cells and modulate the host immune system will provide better understanding of the role of MVs in the context of Gram-positive bacterial infections.
1.3 Conclusions
It has become apparent that OMVs are important biological products that contribute to numerous bacterial functions including cell-to-cell communication and bacterial pathogenesis (Mashburn and Whiteley 2005; reviewed in Kaparakis-Liaskos and Ferrero 2015). OMVs contain a range of materials such as proteins and nucleic acids that aid in bacterial functions (Haurat et al. 2011; Dorward et al. 1989; Ciofu et al. 2000); however, the mechanisms of selective packaging of materials into OMVs remains elusive. Research is now focusing on understanding the mechanisms of OMV biogenesis and how OMVs modulate the innate and adaptive immune system of their host in order to develop their use as novel therapeutics. In the last decade it was shown that Gram-positive bacteria can also produce vesicles as part of their natural growth (Lee et al. 2009). It has become apparent that like OMVs, MVs can carry a range of cargo from their parent bacterium that may be able to aid in bacterial communication and pathogenesis (Resch et al. 2016; Lee et al. 2009). Nevertheless, the roles of MVs in bacterial functions are still not well understood. Research is continuously progressing in both OMV and MV fields to further understand the fundamental production of vesicles and the packaging of materials into them. Importantly, elucidating how OMVs and MVs interact with host epithelial and immune cells is necessary to determine the role of membrane vesicles in contributing to bacterial survival and disease progression. Understanding the production of bacterial membrane vesicles and manipulating vesicles for therapeutic use will have broad implications in how we consider host–pathogen interactions and bacterial diseases. Overall, these works demonstrate the multifaceted, but not exhaustive, roles bacterial membrane vesicles play in contributing to bacterial survival, communication, and pathogenesis.
References
Allesen-Holm M, Barken KB, Yang L, Klausen M, Webb JS, Kjelleberg S, Molin S, Givskov M, Tolker-Nielsen T (2006) A characterization of DNA release in Pseudomonas aeruginosa cultures and biofilms. Mol Microbiol 59(4):1114–1128. https://doi.org/10.1111/j.1365-2958.2005.05008.x
Allison CC, Kufer TA, Kremmer E, Kaparakis M, Ferrero RL (2009) Helicobacter pylori induces MAPK phosphorylation and AP-1 activation via a NOD1-dependent mechanism. J Immunol 183(12):8099–8109. https://doi.org/10.4049/jimmunol.0900664
Andreoni F, Toyofuku M, Menzi C, Kalawong R, Mairpady Shambat S, Francois P, Zinkernagel AS, Eberl L (2019) Antibiotics stimulate formation of vesicles in Staphylococcus aureus in both phage-dependent and -independent fashions and via different routes. Antimicrob Agents Chemother 63(2):e01439. https://doi.org/10.1128/AAC.01439-18
Arnold R, Galloway Y, McNicholas A, O’Hallahan J (2011) Effectiveness of a vaccination programme for an epidemic of meningococcal B in New Zealand. Vaccine 29(40):7100–7106. https://doi.org/10.1016/j.vaccine.2011.06.120
Ayalew S, Confer AW, Shrestha B, Wilson AE, Montelongo M (2013) Proteomic analysis and immunogenicity of Mannheimia haemolytica vesicles. Clin Vaccine Immunol 20(2):191–196. https://doi.org/10.1128/CVI.00622-12
Bauman SJ, Kuehn MJ (2006) Purification of outer membrane vesicles from Pseudomonas aeruginosa and their activation of an IL-8 response. Microbes Infect 8(9–10):2400–2408. https://doi.org/10.1016/j.micinf.2006.05.001
Bernadac A, Gavioli M, Lazzaroni JC, Raina S, Lloubes R (1998) Escherichia coli tol-pal mutants form outer membrane vesicles. J Bacteriol 180(18):4872–4878
Beveridge TJ (1999) Structures of Gram-negative cell walls and their derived membrane vesicles. J Bacteriol 181(16):4725–4733
Bielig H, Rompikuntal PK, Dongre M, Zurek B, Lindmark B, Ramstedt M, Wai SN, Kufer TA (2011) NOD-like receptor activation by outer membrane vesicles from Vibrio cholerae non-O1 non-O139 strains is modulated by the quorum-sensing regulator HapR. Infect Immun 79(4):1418–1427. https://doi.org/10.1128/IAI.00754-10
Biller SJ, Schubotz F, Roggensack SE, Thompson AW, Summons RE, Chisholm SW (2014) Bacterial vesicles in marine ecosystems. Science 343(6167):183–186. https://doi.org/10.1126/science.1243457
Bisschop A, Konings WN (1976) Reconstitution of reduced nicotinamide adenine dinucleotide oxidase activity with menadione in membrane vesicles from the menaquinone-deficient Bacillus subtilis aro D. relation between electron transfer and active transport. Eur J Biochem 67(2):357–365
Bitto NJ, Kaparakis-Liaskos M (2017) The therapeutic benefit of bacterial membrane vesicles. Int J Mol Sci 18(6):E1287. https://doi.org/10.3390/ijms18061287
Bitto NJ, Chapman R, Pidot S, Costin A, Lo C, Choi J, D’Cruze T, Reynolds EC, Dashper SG, Turnbull L, Whitchurch CB, Stinear TP, Stacey KJ, Ferrero RL (2017) Bacterial membrane vesicles transport their DNA cargo into host cells. Sci Rep 7(1):7072. https://doi.org/10.1038/s41598-017-07288-4
Bjune G, Hoiby EA, Gronnesby JK, Arnesen O, Fredriksen JH, Halstensen A, Holten E, Lindbak AK, Nokleby H, Rosenqvist E et al (1991) Effect of outer membrane vesicle vaccine against group B meningococcal disease in Norway. Lancet 338(8775):1093–1096
Blenkiron C, Simonov D, Muthukaruppan A, Tsai P, Dauros P, Green S, Hong J, Print CG, Swift S, Phillips AR (2016) Uropathogenic Escherichia coli releases extracellular vesicles that are associated with RNA. PLoS One 11(8):e0160440. https://doi.org/10.1371/journal.pone.0160440
Bomberger JM, Maceachran DP, Coutermarsh BA, Ye S, O’Toole GA, Stanton BA (2009) Long-distance delivery of bacterial virulence factors by Pseudomonas aeruginosa outer membrane vesicles. PLoS Pathog 5(4):e1000382. https://doi.org/10.1371/journal.ppat.1000382
Brown L, Kessler A, Cabezas-Sanchez P, Luque-Garcia JL, Casadevall A (2014) Extracellular vesicles produced by the Gram-positive bacterium Bacillus subtilis are disrupted by the lipopeptide surfactin. Mol Microbiol 93(1):183–198. https://doi.org/10.1111/mmi.12650
Brown L, Wolf JM, Prados-Rosales R, Casadevall A (2015) Through the wall: extracellular vesicles in Gram-positive bacteria, mycobacteria and fungi. Nat Rev Microbiol 13(10):620–630. https://doi.org/10.1038/nrmicro3480
Cañas MA, Gimenez R, Fabrega MJ, Toloza L, Baldoma L, Badia J (2016) Outer membrane vesicles from the probiotic Escherichia coli Nissle 1917 and the commensal ECOR12 enter intestinal epithelial cells via clathrin-dependent endocytosis and elicit differential effects on DNA damage. PLoS One 11(8):e0160374. https://doi.org/10.1371/journal.pone.0160374
Cañas M-A, Fábrega M-J, Giménez R, Badia J, Baldomà L (2018) Outer membrane vesicles from probiotic and commensal Escherichia coli activate NOD1-mediated immune responses in intestinal epithelial cells. Front Microbiol 9:498. https://doi.org/10.3389/fmicb.2018.00498
Chatterjee SN, Das J (1967) Electron microscopic observations on the excretion of cell-wall material by Vibrio cholerae. J Gen Microbiol 49(1):1–11. https://doi.org/10.1099/00221287-49-1-1
Chen DJ, Osterrieder N, Metzger SM, Buckles E, Doody AM, DeLisa MP, Putnam D (2010) Delivery of foreign antigens by engineered outer membrane vesicle vaccines. Proc Natl Acad Sci U S A 107(7):3099–3104. https://doi.org/10.1073/pnas.0805532107
Chi B, Qi M, Kuramitsu HK (2003) Role of dentilisin in Treponema denticola epithelial cell layer penetration. Res Microbiol 154(9):637–643. https://doi.org/10.1016/j.resmic.2003.08.001
Choi SJ, Kim MH, Jeon J, Kim OY, Choi Y, Seo J, Hong SW, Lee WH, Jeon SG, Gho YS, Jee YK, Kim YK (2015) Active immunization with extracellular vesicles derived from Staphylococcus aureus effectively protects against staphylococcal lung infections, mainly via Th1 cell-mediated immunity. PLoS One 10(9):e0136021. https://doi.org/10.1371/journal.pone.0136021
Choi JW, Kim SC, Hong SH, Lee HJ (2017) Secretable small RNAs via outer membrane vesicles in periodontal pathogens. J Dent Res 96(4):458–466. https://doi.org/10.1177/0022034516685071
Choi J-W, Kwon T-Y, Hong S-H, Lee H-J (2018) Isolation and characterization of a microRNA-size secretable small RNA in Streptococcus sanguinis. Cell Biochem Biophys 76(1):293–301. https://doi.org/10.1007/s12013-016-0770-5
Ciofu O, Beveridge TJ, Kadurugamuwa J, Walther-Rasmussen J, Hoiby N (2000) Chromosomal beta-lactamase is packaged into membrane vesicles and secreted from Pseudomonas aeruginosa. J Antimicrob Chemother 45(1):9–13
Coelho C, Brown L, Maryam M, Vij R, Smith DFQ, Burnet MC, Kyle JE, Heyman HM, Ramirez J, Prados-Rosales R, Lauvau G, Nakayasu ES, Brady NR, Hamacher-Brady A, Coppens I, Casadevall A (2019) Listeria monocytogenes virulence factors, including listeriolysin O, are secreted in biologically active extracellular vesicles. J Biol Chem 294(4):1202–1217. https://doi.org/10.1074/jbc.RA118.006472
Costerton JW, Ingram JM, Cheng KJ (1974) Structure and function of the cell envelope of Gram-negative bacteria. Bacteriol Rev 38(1):87–110
Costerton JW, Geesey GG, Cheng KJ (1978) How bacterial stick. Sci Am 238(1):86–95
Danese PN, Pratt LA, Kolter R (2000) Exopolysaccharide production is required for development of Escherichia coli K-12 biofilm architecture. J Bacteriol 180(12):3593–3596
Deatherage BL, Cookson BT (2012) Membrane vesicle release in bacteria, eukaryotes, and archaea: a conserved yet underappreciated aspect of microbial life. Infect Immun 80(6):1948–1957. https://doi.org/10.1128/IAI.06014-11
Deich RA, Hoyer LC (1982) Generation and release of DNA-binding vesicles by Haemophilus influenzae during induction and loss of competence. J Bacteriol 152(2):855–864
Devoe IW, Gilchrist JE (1973) Release of endotoxin in the form of cell wall blebs during in vitro growth of Neisseria meningitidis. J Exp Med 138(5):1156–1167
DeVoe IW, Gilchrist JE (1975) Pili on meningococci from primary cultures of nasopharyngeal carriers and cerebrospinal fluid of patients with acute disease. J Exp Med 141:297. https://doi.org/10.1084/jem.141.2.297
Dorward DW, Garon CF (1989) DNA-binding proteins in cells and membrane blebs of Neisseria gonorrhoeae. J Bacteriol 171(8):4196–4201
Dorward DW, Garon CF (1990) DNA is packaged within membrane-derived vesicles of Gram-negative but not Gram-positive bacteria. Appl Environ Microbiol 56(6):1960–1962
Dorward DW, Garon CF, Judd RC (1989) Export and intercellular transfer of DNA via membrane blebs of Neisseria gonorrhoeae. J Bacteriol 171(5):2499–2505
Elhenawy W, Bording-Jorgensen M, Valguarnera E, Haurat MF, Wine E, Feldman MF (2016) LPS remodeling triggers formation of outer membrane vesicles in Salmonella. MBio 7(4):e00940–e00916. https://doi.org/10.1128/mBio.00940-16
Ellis TN, Kuehn MJ (2010) Virulence and immunomodulatory roles of bacterial outer membrane vesicles. Microbiol Mol Biol Rev 74(1):81–94. https://doi.org/10.1128/MMBR.00031-09
Fulsundar S, Harms K, Flaten GE, Johnsen PJ, Chopade BA, Nielsen KM (2014) Gene transfer potential of outer membrane vesicles of Acinetobacter baylyi and effects of stress on vesiculation. Appl Environ Microbiol 80(11):3469–3483. https://doi.org/10.1128/AEM.04248-13
Gamazo C, Moriyon I (1987) Release of outer membrane fragments by exponentially growing Brucella melitensis cells. Infect Immun 55(3):609–615
Gankema H, Wensink J, Guinee PA, Jansen WH, Witholt B (1980) Some characteristics of the outer membrane material released by growing enterotoxigenic Escherichia coli. Infect Immun 29(2):704–713
Grande R, Marcantonio MC, Robuffo I, Pompilio A, Celia C, Marzio L, Paolino D, Codagnone M, Muraro R, Stoodley P, Hall-Stoodley L, Mincione G (2015) Helicobacter pylori ATCC 43629/NCTC 11639 outer membrane vesicles (OMVs) from biofilm and planktonic phase associated with extracellular DNA (eDNA). Front Microbiol 6:1369. https://doi.org/10.3389/fmicb.2015.01369
Grande R, Celia C, Mincione G, Stringaro A, Di Marzio L, Colone M, Di Marcantonio MC, Savino L, Puca V, Santoliquido R, Locatelli M, Muraro R, Hall-Stoodley L, Stoodley P (2017) Detection and physicochemical characterization of membrane vesicles (MVs) of Lactobacillus reuteri DSM 17938. Front Microbiol 8:1040. https://doi.org/10.3389/fmicb.2017.01040
Grenier D, Mayrand D (1987) Functional characterization of extracellular vesicles produced by Bacteroides gingivalis. Infect Immun 55(1):111–117
Gurung M, Moon DC, Choi CW, Lee JH (2011) Staphylococcus aureus produces membrane-derived vesicles that induce host cell death. PLoS One 6:e27958. https://doi.org/10.1371/journal.pone.0027958
Haas B, Grenier D (2015) Isolation, characterization and biological properties of membrane vesicles produced by the swine pathogen Streptococcus suis. PLoS One 10(6):e0130528. https://doi.org/10.1371/journal.pone.0130528
Haurat MF, Aduse-Opoku J, Rangarajan M, Dorobantu L, Gray MR, Curtis MA, Feldman MF (2011) Selective sorting of cargo proteins into bacterial membrane vesicles. J Biol Chem 286(2):1269–1276. https://doi.org/10.1074/jbc.M110.185744
He X, Yuan F, Lu F, Yin Y, Cao J (2017) Vancomycin-induced biofilm formation by methicillin-resistant Staphylococcus aureus is associated with the secretion of membrane vesicles. Microb Pathog 110:225–231. https://doi.org/10.1016/j.micpath.2017.07.004
Hoekstra D, van der Laan J, de Leij L, Witholt B (1976) Release of outer membrane fragments from normally growing Escherichia coli. Biochim Biophys Acta Biomembr 455(3):889–899. https://doi.org/10.1016/0005-2736(76)90058-4
Hong SW, Kim MR, Lee EY, Kim JH, Kim YS, Jeon SG, Yang JM, Lee BJ, Pyun BY, Gho YS, Kim YK (2011) Extracellular vesicles derived from Staphylococcus aureus induce atopic dermatitis-like skin inflammation. Allergy 66(3):351–359. https://doi.org/10.1111/j.1398-9995.2010.02483.x
Hooper LV (2004) Bacterial contributions to mammalian gut development. Trends Microbiol 12(3):129–134. https://doi.org/10.1016/j.tim.2004.01.001
Hooper LV, Gordon JI (2001) Commensal host-bacterial relationships in the gut. Science 292(5519):1115–1118
Ismail S, Hampton MB, Keenan JI (2003) Helicobacter pylori outer membrane vesicles modulate proliferation and interleukin-8 production by gastric epithelial cells. Infect Immun 71(10):5670–5675
Jeon H, Oh MH, Jun SH, Kim SI, Choi CW, Kwon HI, Na SH, Kim YJ, Nicholas A, Selasi GN, Lee JC (2016) Variation among Staphylococcus aureus membrane vesicle proteomes affects cytotoxicity of host cells. Microb Pathog 93:185–193. https://doi.org/10.1016/j.micpath.2016.02.014
Jiang Y, Kong Q, Roland KL, Curtiss R (2014) Membrane vesicles of Clostridium perfringens type a strains induce innate and adaptive immunity. Int J Med Microbiol 304(3–4):431–443. https://doi.org/10.1016/j.ijmm.2014.02.006
Johansson ME, Phillipson M, Petersson J, Velcich A, Holm L, Hansson GC (2008) The inner of the two Muc2 mucin-dependent mucus layers in colon is devoid of bacteria. Proc Natl Acad Sci U S A 105(39):15064–15069. https://doi.org/10.1073/pnas.0803124105
Jun SH, Lee JH, Kim SI, Choi CW, Park TI, Jung HR, Cho JW, Kim SH, Lee JC (2017) Staphylococcus aureus-derived membrane vesicles exacerbate skin inflammation in atopic dermatitis. Clin Exp Allergy 47(1):85–96. https://doi.org/10.1111/cea.12851
Jurcisek JA, Bakaletz LO (2007) Biofilms formed by nontypeable Haemophilus influenzae in vivo contain both double-stranded DNA and type IV pilin protein. J Bacteriol 189(10):3868–3875. https://doi.org/10.1128/JB.01935-06
Kadurugamuwa JL, Beveridge TJ (1995) Virulence factors are released from Pseudomonas aeruginosa in association with membrane vesicles during normal growth and exposure to gentamicin: a novel mechanism of enzyme secretion. J Bacteriol 177(14):3998–4008
Kadurugamuwa JL, Beveridge TJ (1996) Bacteriolytic effect of membrane vesicles from Pseudomonas aeruginosa on other bacteria including pathogens: conceptually new antibiotics. J Bacteriol 178(10):2767–2774
Kadurugamuwa JL, Beveridge TJ (1997) Natural release of virulence factors in membrane vesicles by Pseudomonas aeruginosa and the effect of aminoglycoside antibiotics on their release. J Antimicrob Chemother 40:615. https://doi.org/10.1093/jac/40.5.615
Kang CS, Ban M, Choi EJ, Moon HG, Jeon JS, Kim DK, Park SK, Jeon SG, Roh TY, Myung SJ, Gho YS, Kim JG, Kim YK (2013) Extracellular vesicles derived from gut microbiota, especially Akkermansia muciniphila, protect the progression of dextran sulfate sodium-induced colitis. PLoS One 8(10):e76520. https://doi.org/10.1371/journal.pone.0076520
Kaparakis M, Turnbull L, Carneiro L, Firth S, Coleman HA, Parkington HC, Le Bourhis L, Karrar A, Viala J, Mak J, Hutton ML, Davies JK, Crack PJ, Hertzog PJ, Philpott DJ, Girardin SE, Whitchurch CB, Ferrero RL (2010) Bacterial membrane vesicles deliver peptidoglycan to NOD1 in epithelial cells. Cell Microbiol 12(3):372–385. https://doi.org/10.1111/j.1462-5822.2009.01404.x
Kaparakis-Liaskos M, Ferrero RL (2015) Immune modulation by bacterial outer membrane vesicles. Nat Rev Immunol 15(6):375–387. https://doi.org/10.1038/nri3837
Kawai T, Akira S (2010) The role of pattern-recognition receptors in innate immunity: update on toll-like receptors. Nat Immunol 11(5):373–384. https://doi.org/10.1038/ni.1863
Kim MR, Hong SW, Choi EB, Lee WH, Kim YS, Jeon SG, Jang MH, Gho YS, Kim YK (2012) Staphylococcus aureus-derived extracellular vesicles induce neutrophilic pulmonary inflammation via both Th1 and Th17 cell responses. Allergy 67(10):1271–1281. https://doi.org/10.1111/all.12001
Kim YS, Choi EJ, Lee WH, Choi SJ (2013) Extracellular vesicles, especially derived from Gram-negative bacteria, in indoor dust induce neutrophilic pulmonary inflammation associated with both Th1 and Th17 cell responses. Clin Exp Allergy 43:443–454
Knox KW, Vesk M, Work E (1966) Relation between excreted lipopolysaccharide complexes and surface structures of a lysine-limited culture of Escherichia coli. J Bacteriol 92(4):1206–1217
Koeppen K, Hampton TH, Jarek M, Scharfe M, Gerber SA, Mielcarz DW, Demers EG, Dolben EL, Hammond JH, Hogan DA, Stanton BA (2016) A novel mechanism of host-pathogen interaction through sRNA in bacterial outer membrane vesicles. PLoS Pathog 12(6):e1005672. https://doi.org/10.1371/journal.ppat.1005672
Lee EY, Bang JY, Park GW, Choi DS, Kang JS, Kim HJ, Park KS, Lee JO, Kim YK, Kwon KH, Kim KP, Gho YS (2007) Global proteomic profiling of native outer membrane vesicles derived from Escherichia coli. Proteomics 7(17):3143–3153. https://doi.org/10.1002/pmic.200700196
Lee EY, Choi DY, Kim DK, Kim JW, Park JO, Kim S, Kim SH, Desiderio DM, Kim YK, Kim KP, Gho YS (2009) Gram-positive bacteria produce membrane vesicles: proteomics-based characterization of Staphylococcus aureus-derived membrane vesicles. Proteomics 9(24):5425–5436. https://doi.org/10.1002/pmic.200900338
Lee J, Lee E-Y, Kim S-H, Kim D-K, Park K-S, Kim KP, Kim Y-K, Roh T-Y, Gho YS (2013a) Staphylococcus aureus extracellular vesicles carry biologically active β-lactamase. Antimicrob Agents Chemother 57(6):2589–2595. https://doi.org/10.1128/AAC.00522-12
Lee JH, Choi C-W, Lee T, Kim S (2013b) Transcription factor σB plays an important role in the production of extracellular membrane-derived vesicles in Listeria monocytogenes. PLoS One 8(8):e73196. https://doi.org/10.1371/journal.pone.0073196
Leive L, Shovlin VK, Mergenhagen SE (1968) Physical chemical and immunological properties of lipopolysaccharide released from Escherichia coli by Ethylenediaminetetraacetate. J Biol Chem 243(24):6384+
Liao S, Klein MI, Heim KP, Fan Y, Bitoun JP, Ahn SJ, Burne RA, Koo H, Brady LJ, Wen ZT (2014) Streptococcus mutans extracellular DNA is upregulated during growth in biofilms, actively released via membrane vesicles, and influenced by components of the protein secretion machinery. J Bacteriol 196(13):2355–2366. https://doi.org/10.1128/jb.01493-14
Loeb MR (1974) Bacteriophage T4-mediated release of envelope components from Escherichia coli. J Virol 13:631–641
MacIntyre S, Trust TJ, Buckley JT (1980) Identification and characterization of outer membrane fragments released by Aeromonas sp. Can J Biochem 58(10):1018–1025
Mashburn LM, Whiteley M (2005) Membrane vesicles traffic signals and facilitate group activities in a prokaryote. Nature 437(7057):422–425. https://doi.org/10.1038/nature03925
Mayer F, Gottschalk G (2003) The bacterial cytoskeleton and its putative role in membrane vesicle formation observed in a Gram-positive bacterium producing starch-degrading enzymes. J Mol Microbiol Biotechnol 6(3–4):127–132. https://doi.org/10.1159/000077243
McBroom AJ, Kuehn MJ (2007) Release of outer membrane vesicles by Gram-negative bacteria is a novel envelope stress response. Mol Microbiol 63(2):545–558. https://doi.org/10.1111/j.1365-2958.2006.05522.x
McMahon KJ, Castelli ME, Garcia Vescovi E, Feldman MF (2012) Biogenesis of outer membrane vesicles in Serratia marcescens is thermoregulated and can be induced by activation of the Rcs phosphorelay system. J Bacteriol 194(12):3241–3249. https://doi.org/10.1128/JB.00016-12
Mergenhagen SE, Bladen HA, Hsu KC (1966) Electron microscopic localization of endotoxic lipopolysaccharide in Gram-negative organisms. Ann N Y Acad Sci 133(2):279–291
Mirlashari MR, Lyberg T (2003) Expression and involvement of toll-like receptors (TLR)2, TLR4, and CD14 in monocyte TNF-alpha production induced by lipopolysaccharides from Neisseria meningitidis. Med Sci Monit 9(8):BR316–BR324
Mug-Opstelten D, Witholt B (1978) Preferential release of new outer membrane fragments by exponentially growing Escherichia coli. Biochim Biophys Acta 508(2):287–295
Murphy K, Park AJ, Hao Y, Brewer D, Lam JS, Khursigara CM (2014) Influence of O polysaccharides on biofilm development and outer membrane vesicle biogenesis in Pseudomonas aeruginosa PAO1. J Bacteriol 196(7):1306–1317
Nieves W, Asakrah S, Qazi O, Brown KA, Kurtz J, Aucoin DP, McLachlan JB, Roy CJ, Morici LA (2011) A naturally derived outer-membrane vesicle vaccine protects against lethal pulmonary Burkholderia pseudomallei infection. Vaccine 29(46):8381–8389. https://doi.org/10.1016/j.vaccine.2011.08.058
Olaya-Abril A, Prados-Rosales R, McConnell MJ, Martín-Peña R, González-Reyes JA, Jiménez-Munguía I, Gómez-Gascón L, Fernández J, Luque-García JL, García-Lidón C, Estévez H, Pachón J, Obando I, Casadevall A, L-a P, Rodríguez-Ortega MJ (2014) Characterization of protective extracellular membrane-derived vesicles produced by Streptococcus pneumoniae. J Proteome 106:46–60
Park AJ, Murphy K, Surette MD, Bandoro C, Krieger JR, Taylor P, Khursigara CM (2015) Tracking the dynamic relationship between cellular systems and extracellular subproteomes in Pseudomonas aeruginosa biofilms. J Proteome Res 14(11):4524–4537. https://doi.org/10.1021/acs.jproteome.5b00262
Park K-S, Lee J, Lee C, Park HT, Kim J-W, Kim OY, Kim SR, Rådinger M, Jung H-Y, Park J, Lötvall J, Gho YS (2018) Sepsis-like systemic inflammation induced by nano-sized extracellular vesicles from feces. Front Microbiol 9:1735. https://doi.org/10.3389/fmicb.2018.01735
Perez-Cruz C, Delgado L, Lopez-Iglesias C, Mercade E (2015) Outer-inner membrane vesicles naturally secreted by Gram-negative pathogenic bacteria. PLoS One 10(1):e0116896. https://doi.org/10.1371/journal.pone.0116896
Pesci EC, Milbank JB, Pearson JP, McKnight S, Kende AS, Greenberg EP, Iglewski BH (1999) Quinolone signaling in the cell-to-cell communication system of Pseudomonas aeruginosa. Proc Natl Acad Sci U S A 96(20):11229–11234
Pierson T, Matrakas D, Taylor YU, Manyam G, Morozov VN, Zhou W, van Hoek ML (2011) Proteomic characterization and functional analysis of outer membrane vesicles of Francisella novicida suggests possible role in virulence and use as a vaccine. J Proteome Res 10(3):954–967. https://doi.org/10.1021/pr1009756
Renelli M, Matias V, Lo RY, Beveridge TJ (2004) DNA-containing membrane vesicles of Pseudomonas aeruginosa PAO1 and their genetic transformation potential. Microbiology 150(Pt 7):2161–2169. https://doi.org/10.1099/mic.0.26841-0
Resch U, Tsatsaronis JA, Le Rhun A, Stubiger G, Rohde M, Kasvandik S, Holzmeister S, Tinnefeld P, Wai SN, Charpentier E (2016) A two-component regulatory system impacts extracellular membrane-derived vesicle production in group a Streptococcus. MBio 7(6):e00207–e00216. https://doi.org/10.1128/mBio.00207-16
Rivera J, Cordero RJB, Nakouzi AS, Frases S (2010) Bacillus anthracis produces membrane-derived vesicles containing biologically active toxins. Proc Natl Acad Sci U S A 107(44):19002–19007. https://doi.org/10.1073/pnas.1008843107
Roier S, Zingl FG, Cakar F, Durakovic S, Kohl P, Eichmann TO, Klug L, Gadermaier B, Weinzerl K, Prassl R, Lass A, Daum G, Reidl J, Feldman MF, Schild S (2016) A novel mechanism for the biogenesis of outer membrane vesicles in Gram-negative bacteria. Nat Commun 7:10515. https://doi.org/10.1038/ncomms10515
Rosenqvist E, Hoiby EA, Wedege E, Bryn K, Kolberg J, Klem A, Ronnild E, Bjune G, Nokleby H (1995) Human antibody responses to meningococcal outer membrane antigens after three doses of the Norwegian group B meningococcal vaccine. Infect Immun 63(12):4642–4652
Rothfield L, Pearlman-Kothencz M (1969) Synthesis and assembly of bacterial membrane components: a lipopolysaccharide-phospholipid-protein complex excreted by living bacteria. J Mol Biol 44:477
Ruhr E, Sahl HG (1985) Mode of action of the peptide antibiotic nisin and influence on the membrane potential of whole cells and on cytoplasmic and artificial membrane vesicles. Antimicrob Agents Chemother 27(5):841–845
Schlatterer K, Beck C, Hanzelmann D, Lebtig M, Fehrenbacher B, Schaller M, Ebner P, Nega M, Otto M, Kretschmer D, Peschel A (2018) The mechanism behind bacterial lipoprotein release: phenol-soluble modulins mediate toll-like receptor 2 activation via extracellular vesicle release from Staphylococcus aureus. MBio 9(6):e01851–e01818. https://doi.org/10.1128/mBio.01851-18
Schnaitman CA (1970) Protein composition of the cell wall and cytoplasmic membrane of Escherichia coli. J Bacteriol 104(2):890–901
Schooling SR, Beveridge TJ (2006) Membrane vesicles: an overlooked component of the matrices of biofilms. J Bacteriol 188(16):5945–5957. https://doi.org/10.1128/JB.00257-06
Schwechheimer C, Kuehn MJ (2015) Outer-membrane vesicles from Gram-negative bacteria: biogenesis and functions. Nat Rev Microbiol 13(10):605–619. https://doi.org/10.1038/nrmicro3525
Shen Y, Giardino Torchia ML, Lawson GW, Karp CL, Ashwell JD, Mazmanian SK (2012) Outer membrane vesicles of a human commensal mediate immune regulation and disease protection. Cell Host Microbe 12(4):509–520. https://doi.org/10.1016/j.chom.2012.08.004
Silva TP, Noyma NP, Duque TLA, Gamalier JP, Vidal LO, Lobao LM, Chiarini-Garcia H, Roland F, Melo RCN (2014) Visualizing aquatic bacteria by light and transmission electron microscopy. Antonie Van Leeuwenhoek 105:1–14
Sjostrom AE, Sandblad L, Uhlin BE, Wai SN (2015) Membrane vesicle-mediated release of bacterial RNA. Sci Rep 5:15329. https://doi.org/10.1038/srep15329
Soderblom T, Oxhamre C, Wai SN, Uhlen P, Aperia A, Uhlin BE, Richter-Dahlfors A (2005) Effects of the Escherichia coli toxin cytolysin a on mucosal immunostimulation via epithelial Ca2+ signalling and toll-like receptor 4. Cell Microbiol 7(6):779–788. https://doi.org/10.1111/j.1462-5822.2005.00510.x
Tartaglia NR, Breyne K, Meyer E, Cauty C, Jardin J, Chrétien D, Dupont A, Demeyere K, Berkova N, Azevedo V, Guédon E, Le Loir Y (2018) Staphylococcus aureus extracellular vesicles elicit an immunostimulatory response in vivo on the murine mammary gland. Front Cell Infect Microbiol 8:277. https://doi.org/10.3389/fcimb.2018.00277
Thay B, Wai SN, Oscarsson J (2013) Staphylococcus aureus alpha-toxin-dependent induction of host cell death by membrane-derived vesicles. PLoS One 8(1):e54661. https://doi.org/10.1371/journal.pone.0054661
Toyofuku M, Cárcamo-Oyarce G, Yamamoto T, Eisenstein F, Hsiao C-C, Kurosawa M, Gademann K, Pilhofer M, Nomura N, Eberl L (2017) Prophage-triggered membrane vesicle formation through peptidoglycan damage in Bacillus subtilis. Nat Commun 8(1):481. https://doi.org/10.1038/s41467-017-00492-w
Turnbull L, Toyofuku M, Hynen AL, Kurosawa M, Pessi G, Petty NK, Osvath SR, Carcamo-Oyarce G, Gloag ES, Shimoni R, Omasits U, Ito S, Yap X, Monahan LG, Cavaliere R, Ahrens CH, Charles IG, Nomura N, Eberl L, Whitchurch CB (2016) Explosive cell lysis as a mechanism for the biogenesis of bacterial membrane vesicles and biofilms. Nat Commun 7:11220. https://doi.org/10.1038/ncomms11220
Turner L, Praszkier J, Hutton ML, Steer D, Ramm G, Kaparakis-Liaskos M, Ferrero RL (2015) Increased outer membrane vesicle formation in a Helicobacter pylori tolB mutant. Helicobacter 20(4):269–283. https://doi.org/10.1111/hel.12196
Turner L, Bitto NJ, Steer DL, Lo C, D’Costa K, Ramm G, Shambrook M, Hill AF, Ferrero RL, Kaparakis-Liaskos M (2018) Helicobacter pylori outer membrane vesicle size determines their mechanisms of host cell entry and protein content. Front Immunol 9:1466
Vasilyeva NV, Tsfasman IM, Suzina NE, Stepnaya OA, Kulaev IS (2009) Outer membrane vesicles of Lysobacter sp. Dokl Biochem Biophys 426:139–142
Vernikos G, Medini D (2014) Bexsero(R) chronicle. Pathog Glob Health 108(7):305–316. https://doi.org/10.1179/2047773214Y.0000000162
Wagner T, Joshi B, Janice J, Askarian F, Škalko-Basnet N, Hagestad OC, Mekhlif A, Wai SN, Hegstad K, Johannessen M (2018) Enterococcus faecium produces membrane vesicles containing virulence factors and antimicrobial resistance related proteins. J Proteome 187:28–38. https://doi.org/10.1016/j.jprot.2018.05.017
Wang X, Thompson CD, Weidenmaier C, Lee JC (2018) Release of Staphylococcus aureus extracellular vesicles and their application as a vaccine platform. Nat Commun 9(1):1379. https://doi.org/10.1038/s41467-018-03847-z
Wensink J, Witholt B (1981) Outer-membrane vesicles released by normally growing Escherichia coli contain very little lipoprotein. Eur J Biochem 116(2):331–335
Whitchurch CB, Tolker-Nielsen T, Ragas PC, Mattick JS (2002) Extracellular DNA required for bacterial biofilm formation. Science 295(5559):1487. https://doi.org/10.1126/science.295.5559.1487
Wispelwey B, Hansen EJ, Scheld WM (1989) Haemophilus influenzae outer membrane vesicle-induced blood-brain barrier permeability during experimental meningitis. Infect Immun 57(8):2559–2562
Yaron S, Kolling GL, Simon L, Matthews KR (2000) Vesicle-mediated transfer of virulence genes from Escherichia coli O157:H7 to other enteric bacteria. Appl Environ Microbiol 66(10):4414–4420
Yonezawa H, Osaki T, Kurata S, Fukuda M, Kawakami H, Ochiai K, Hanawa T, Kamiya S (2009) Outer membrane vesicles of Helicobacter pylori TK1402 are involved in biofilm formation. BMC Microbiol 9:197. https://doi.org/10.1186/1471-2180-9-197
Zavan L, Bitto NJ, Johnston EL, Greening DW, Kaparakis-Liaskos M (2019) Helicobacter pylori growth stage determines the size, protein composition, and preferential cargo packaging of outer membrane vesicles. Proteomics 19:e1800209. https://doi.org/10.1002/pmic.201800209
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Zavan, L., Bitto, N.J., Kaparakis-Liaskos, M. (2020). Introduction, History, and Discovery of Bacterial Membrane Vesicles. In: Kaparakis-Liaskos, M., Kufer, T. (eds) Bacterial Membrane Vesicles. Springer, Cham. https://doi.org/10.1007/978-3-030-36331-4_1
Download citation
DOI: https://doi.org/10.1007/978-3-030-36331-4_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-36330-7
Online ISBN: 978-3-030-36331-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)