Abstract
Zika virus is a member of the Flaviviridae virus family, similar to other viruses that affect humans, such as hepatitis C and dengue virus. After its first appearance in 1947, Zika virus reappeared in 2016 causing an international public health emergency. Zika virus was considered a non dangerous human pathogen; however, it is currently considered a pathogen with serious consequences for human health, showing association with neurological complications such as Guillain-Barre syndrome and microcephaly. Then, it is necessary to get antivirals able to inhibit the replication of the Zika virus since vaccines for this virus are not yet available. Zika virus structure is similar to hepatitis C virus structure. This characteristic suggests that anti-hepatitis C virus agents can be used as alternative in treatments against the Zika virus. This work aims to determine a non-nucleoside analogue antivirals that can be considered possible antivirals against Zika virus. In this study, we used computational methods to analyze the Docking and the modeling of the NS5 polymerase of Zika virus and antivirals.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
The Zika virus (ZIKV) is a pathogen, which is part of the Flaviviridae family and is transmitted by mosquitoes, similar to other viruses, such as yellow fever virus (YFV), dengue virus (DENV), West Nile virus (WNV), and Japanese encephalitis (JEV) [2, 19]. This virus comes from Uganda, specifically from the Zika forest and was initially identified in 1947, when the Rhesus Macaque monkeys with suspected yellow fever were studied at the Forest Research Institute-Uganda. The presence of the virus in humans, considered initially an occasional host, was confirmed through serological studies in 1952 and it was not until 1968 when the virus was isolated from a patient in Nigeria [5, 7].
In America, Zika epidemics occurred from 2015 to 2016, when the World Health Organization (WHO) declared it a public health emergency of international interest. The great epidemic outbreak occurred in Brazil and then quickly spread to other countries in the South American region [2].
Among the emerging diseases of the 21st century, ZIKV disease is among the biggest concerns for public health worldwide. Share the same vector, mosquitoes of the genus Aedes, with other arboviruses of particular importance to Public Health in the Americas, such as Dengue and Chikungunya, in addition to the Yellow Fever Urban [18].
Currently, there are no vaccines or drugs with an effective license for the treatment of ZIKV infections. Considering the need to mitigate the morbidities associated with ZIKV, it is necessary to provide antiviral against this virus. Consequently, the aim of this work is to determine in silico possible drugs with an effective license for the hepatitis C virus as possible drugs for the treatment of Zika infection.
The structure of the paper is as follows: Sect. 2 provides a background related to Zika virus, Sect. 3 presents the related work, Sect. 4 presents the materials and methods used in this research, Sect. 5 reports our results, and Sect. 6 presents the conclusions.
2 Background
2.1 History
No cases of ZIKV were detected for almost 70 years. From October 2013 to March 2014 a major epidemic occurred in French Polynesia, then, during 2014 it was introduced in Brazil from the Pacific Islands [8]. In 2015, Brazil reported an epidemic of ZIKV in the northeast. From this moment, the epidemic was spread very fast to other areas and also it was spread to almost all countries in South America. More than 300,000 cases were confirmed based on laboratory results; however, it is estimated that much more cases occurred because only 15% of cases have symptoms [22].
Its main reservoirs are monkeys and humans, however, serological studies have shown that it could infect other mammals such as rodents, elephants and cats [16]. Its main vector are female mosquitoes of the genus Aedes (Ae.). The most common being Ae. Aegypti and Ae. Albopictus, the latter of special interest due to its anthropomorphic habits and high adaptability to them (humans); the vector competence that is associated to the capacity of the virus to multiply to high viral titers within the vector, and the effectiveness of the vector to transfer during the phage process, which makes them a very efficient vector in the transmission of febrile diseases [16].
2.2 Zika Virus
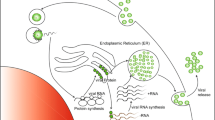
The ZIKV is a particular about 50 nm, which contains (a) an internal nucleocapsid that presents an icosahedral symmetry composed of a positive chain genomic RNA and several copies of the viral capsid protein (C) and (b) the lipid bilayer derived from the host cell external that contains 180 copies of each of two proteins: the viral membrane (M, a cleavage product of the prM protein) and the envelope protein (E) [14].
The genome has a length of 10,794 kb, consisting of positive-sense single-stranded RNA. It is composed by two non-coding regions (5 and 3 NCR) and a single open reading frame (ORF), which together encodes a polyprotein. Furthermore, it is divided into three structural proteins such as capsid (C), envelope (E), membrane precursor (prM) and seven non-structural proteins (NS) [24].
2.3 Pathology
In most cases, infections are asymptomatic or lightly symptomatic. Most of the described symptoms include fever, arthralgia, rash, fatigue, myalgia, conjunctivitis and headache. In humans, the evolution since the bite of the mosquito occurs until the outbreak is between 3 to 12 days [23]. More than 90% of patients present an outbreak of maculopapular eruptions; thus, this symptom allows describing infection by ZIKV [11]. Only about 18 % of the reported cases of ZIKV infections were recorded as symptomatic [26].
2.4 Transmission
ZIKV is mainly transmitted to humans through the bite of infected mosquitoes such as: Aedes and Aedes aegypti. Aedes mosquitoes usually bite during the day, and they also are able to transmit dengue, chikungunya fever and yellow fever [30]. Zika has been detected for prolonged periods in semen and documented cases of sexual transmission are increasing. Intrauterine/perinatal transmission also occurs. In theory, the Zika virus could also be contracted from infected breast milk or organ/tissue transplants [13].
2.5 Treatment
Usually, Zika virus is relatively light and thus, it does not required a specific treatment. Infected patients just need to rest, drink a lot of liquids and take medications against fever. If the patients continues with the symptoms, they must consult the doctor [18], although microcephaly associated with ZIKV and GBS emphasize that antiviral interventions are urgent [25].
An effective vaccine for ZIKV is not yet available, although about 40 candidate vaccines against ZIKV are in the process of being developed. Five of them are close to entering, in the phase I of clinical trials, which evaluates the safety of the vaccine and its effectiveness to produce an immune response [18].
Currently, drug reuse studies have been implemented, where they evaluate the efficacy of medications approved by the Food and Drug Administration (FDA) for antiviral activity against ZIKV infection. Therefore, the systematic screening of drugs approved by the FDA may reveal new agents to treat the infection by ZIKV [1].
The ZIKV structure is very similar to other flaviviruses. Specially, it is similar to the hepatitis C virus (HCV) [25]. Then, it represents an advance in the possible development of treatments against the ZIKV, considered by the WHO as a health emergency. The structure of the ZIKV provides potential regions for a therapeutic treatment, which could be used to evaluate an antiviral [25].
It has been hypothesized that ZIKV uses the NS5 non-structural protein or enzyme RNA-dependent RNA polymerase (RdRp), together with cofactors, to replicate, maintain and express its RNA genome [3].
The predicted three-dimensional structure and the amino acids of the NS5 protein of the ZIKV indicate that the NS5B protein (RdRp enzyme) of HCV shares many residues in common, particularly at the active site, with the NS5 protein of the ZIKV. Therefore, it is possible that certain nucleoside analogues (NA) and non-nucleoside analogues (NNA) that inhibit HCV NS5B protein may have inhibitory potency against ZIKV.
2.6 Antivirals
An antiviral is a type of drug used to treat infections caused by viruses. Viruses, unlike bacteria and other microorganisms, use the biosynthetic machinery of the cell they infect to replicate. Due to this intimate relationship with the host cell they have fewer targets of their own that allow the selective action of a drug. That is why just from the exhaustive study of the mechanisms of viral replication and the recognition of exclusive steps of these agents, antiviral drugs have acquired great interest [32]. The development of new, more effective drugs has been achieved, with fewer side effects, and specifically directed against virus proteins, called direct action antivirals (DAAS) [12].
2.7 Mechanism of Action of Antivirals
To understand the mechanism of action of antivirals, it is necessary to know the complete life cycle of a virus, which comprises 5 steps or stages (adhesion, penetration-loss of the coating, duplication of the genome and viral proteins, assembly and release). Knowledge of these stages has provided scientists with a potential target for antiviral drugs. Each step of the life cycle of a virus is a potential target for inhibitory molecules It should be noted that between these target proteins there are two types, depending on whether they are structural proteins (involved in adhesion) or non-structural proteins (enzymes involved in the multiplication) [21].
Non-nucleoside analogue antivirals act by blocking the enzyme from initiation, through the inhibition of a conformational change necessary to proceed with the elongation of nascent RNA, that is, they bind to less conserved sites outside the active center (allosteric sites), inhibiting the catalytic efficiency of the active center [15].
3 Related Work
Currently, there are some studies that aim to find antiviral molecules. Those studies have been focused on essential enzymes in the infection process. It is done by two processes: (a) the indirect or direct inhibition of their biological functions or (b) the block of the viral replication [17].
One of the most used targets that is employed for this purpose is the NS2B/NS3 protease and NS5 RNA-dependent RNA polymerase [33]. In addition, flavivirus proteins such as NS2B/NS3 and NS5 have been found essential for viral infectivity and replication.
Some studies are based on a structure-based approach that targets the ZIKV RNA-dependent RNA polymerase (RdRp), which is conducted in silico screening of a library of 100,000 small molecules and tested the top ten lead compounds for their ability to inhibit the virus replication in cell-based in vitro assays. [20].
Others studies are focused to find the ability of antiviral drugs in preventing vertical transmission of ZIKV in the mouse model by using a well-established flaviviral inhibitor NITD008, an adenosine analog, that has been shown to inhibit the replication by directly inhibiting the RNA-dependent RNA polymerase activity through chain-termination [34], and also shown to reduce viremia in mice infected with ZIKV.
There are many emerging and re-emerging globally prevalent viruses for which there are no licensed vaccines or antiviral medicines. Arbidol (ARB, umifenovir), used clinically for decades in several countries as an anti-infuenza virus drug, inhibits many other viruses. Thus, ARB, was demonstrated by some researchers, which is a broadly acting anti-viral agent with a well-established safety profile, inhibits ZIKV, likely by blocking viral entry. [9]
Therefore, it is currently being studied, not only in the Zika virus, but also in other viruses of the Flavivirus genus such as dengue virus. The purpose of discovering small molecule inhibitors that are suitable as treatment options for the disease.
4 Materials and Methods
According to the literature, the antivirals used for the treatment of HCV that present in their attributes, ideal characteristics for treatment of ZIKV, are those that inhibit the NS5B polymerase of HCV [10]. This enzyme it is essential for HCV replication, since it catalyzes the synthesis of RNA complementary negative chain and subsequent genomic RNA strand positive. The main applicants for ZIKV can probably be established using HCV inhibitors, since NS5B polymerase of HCV [27,28,29], particularly at the active site, it shares a lot of waste in common with the ZIKV NS5 polymerase, and the structures presented can provide opportunities for intervention strategies to treat ZIKV. For this work, we used the following two antivirals: Dasabuvir and Nesbuvir. These antivirals have possible activity in Zika. In addition, they are used for the treatment of hepatitis C virus. Furthermore, they present in their attributes, ideal characteristics for the treatment of Zika, low toxicity, and potential action against the virus protein. The three-dimensional structures of the possible antivirals proposed were obtained from the DrugBankFootnote 1 database.
Figure 1 presents the Dasabuvir molecular structure that has chemical notation \(C_{26}H_{27}N_{3}O_{5}S\) with molecular wheight 493.58 [g/mol]. Dasabuvir is a competitive inhibitor of HCV polymerase and inhibits the extension of the RNA chain after incorporation into the RNA of the nascent HCV.
Figure 2 presents the Nesbubir molecular structure that has chemical notation \(C_{22}H_{23}FN_{2O5}S\) with molecular weight 446.492 [g/mol]. Nesbuvir joins the pocket of HCV NS5B polymerase palm II site and inhibits the replication of viral genome.
Moreover, we used a docking analysis of ZIKV protein with three-dimensional structure resolved with antiviral. The antivirals proposed were evaluated in vitro, by bioinformatics prediction of anti-ZIKV activity. Molecular Docking (computational method) AutoDock Tools Version 1.5.6 [31] were use to analyze drug-target interactions (antiviral in interaction with Zika NS5 protein).
Antivirals were represented with the three-dimensional structure of the NS5 protein of the ZIKV using the software PyMOL Molecular Graphics SystemFootnote 2, which is an appropriate molecular viewer to produce 3D images of biological molecules, such as proteins, which allows the visualization of multiple conformations.
The NS5 protein of the ZIKV, extracted from the Protein DataBank (PDB) database from the corresponding identity code (5U04)Footnote 3 was used as the working structure.
5 Results
5.1 Molecular Docking Analysis
The NS5 protein of ZIKV was determined in silico with the corresponding ligands, i.e., the antivirals: Dasabuvir and Nesbuvir.
A 3D model of the NS5 protein of the ZIKV was created. It was represented by tapes and according to the subdomains were colored as follows:
-
blue: the domain corresponding to the thumb
-
green: the domain of the fingers
-
yellow: the domain corresponding to the palm
In the molecular coupling analysis of the NS5 polymerase of ZIKV and Dasabuvir, the antiviral was located towards the interior of the active site of the NS5 polymerase of the ZIKV. Figure 3 presents the three dimensional structure of the Protein NS5 of the ZIKV with the antiviral Dasabuvir, while Fig. 4 presents the molecular docking of the three dimensional structure of the Protein NS5 of the ZIKV with the antiviral Dasabuvir.
Finally, the antiviral Nesbuvir was located in the region furthest from the active site (palm) of the NS5 polymerase of ZIKV Fig. 5 presents the three dimensional structure of the Protein NS5 of the ZIKV with the antiviral Nesbuvir, while Fig. 6 the molecular docking of the three dimensional structure of the Protein NS5 of the ZIKV with the antiviral Nesbuvir.
Consequently, the gene coding for RNA polymerase in the Flaviviridae family shows the highest degree of conservation. Then, new therapeutic alternatives against hepatitis C virus, especially those that targets viral RNA polymerase, might provide a broader spectrum than other members of the Flaviviridae family. With this in mind, the antivirals presented in this work were clinically approved in recent years for the therapeutic intervention against HCV infection. Overall, these results enables studying whether the chemical structures of antivirals have anti-ZIKV activity.
In addition, broad-spectrum antivirals, such as ribavirin, interferon, and Favipiravir are harmful to women and pregnant animal models. Nucleoside analogue anti-HCV drugs, Sofosbuvir, and the non-nucleoside analogues, Dasabuvir and Nesbuvir, have not been associated with teratogenicity [25]. Some previous studies have shown that the antimalarial drug chloroquine and the new nucleoside analogs inhibit the replication of ZIKV [4].
Zmurko et al. [35] discovered that the 7DMA viral polymerase inhibitor (7-deaza-2’-C-methyladenosine) inhibits the in vitro replication of ZIKV. Therefore, 7DMA can be used as a reference comparator compound in future studies.
Eyer et al. [6] demonstrated that nucleoside analogues exert activity against ZIKV under in vitro conditions. These compounds provide a basis for optimization based on the structure and rational design of effective prodrugs, which will be further tested in rodent models for the treatment of ZIKV infection. In this work, we show in silico that anti-HCV drugs, non-nucleoside analogues, as possible therapeutic agents for ZIKV infection.
In our molecular coupling analysis of the NS5 structure of ZIKV, Dasabuvir was located towards the interior of the active site; whereas Nesbuvir was located in a region further away from the active site. The antivirals proposed in this study were evaluated in silico, in order to be evaluated in vitro in the future.
Consequently, the results presented in this work will help to provide alternatives of antiviral agents for ZIKV infection, could be used to prevent and contain the emerging epidemic and combat the public health crisis.
6 Conclusions
Based on the computational methods used in this studey, we could determined in silico a correct chemical interaction of Dasabuvir and Nesbuvir (non-nucleoside analogue), with the active site of the Zika virus NS5 protein, which allows us to conclude that these drugs have an effective license for the treatment of hepatitis C virus, they can have effects on the inhibition of Zika virus replication. Of the two antivirals analyzed, Dasabuvir, could be more effective for the treatment of Zika virus infection because of its location of binding to the active site of the NS5 protein, while Nesbuvir would be less effective, due to its location in close union to the active site of the NS5 protein that could affect its efficacy.
These results would indicate that an antiviral effect against this flavivirus could be achieved, avoiding the side effects of these viral infections such as microcephaly or congenital malformations associated with ZIKV, when they are not treated in time.
Finally, the absence of specific and efficient treatment protocols (there are no vaccines or drugs with an effective license for the treatment of infections) would allow these drugs to be a potential solution to improve treatments for orphaned or neglected diseases, and to mitigate the morbidities associated with ZIKV. This would imply a progress in the potential treatment against this virus, considered by WHO as a Health emergency and an urgent need.
In this way, antivirals tested in this study are promising to move on to in vitro studies and could be use in the future as treatment for ZIKV.
References
Alam, A., Imam, N., Ali, S., Malik, M.Z., Ishrat, R., et al.: Recent trends in ZIKV research: a step away from cure. Biomed. Pharmacother. 91, 1152–1159 (2017)
Calvet, G., et al.: Detection and sequencing of Zika virus from amniotic fluid of fetuses with microcephaly in Brazil: a case study. Lancet Infect. Dis. 16(6), 653–660 (2016)
Cox, B.D., Stanton, R.A., Schinazi, R.F.: Predicting Zika virus structural biology: challenges and opportunities for intervention. Antivir. Chem. Chemother. 24(3–4), 118–126 (2015)
Delvecchio, R., et al.: Chloroquine, an endocytosis blocking agent, inhibits Zika virus infection in different cell models. Viruses 8(12), 322 (2016)
Dick, G., Kitchen, S., Haddow, A., et al.: Zika virus (II). Pathogenicity and physical properties. Trans. R. Soc. Trop. Med. Hyg. 46(5), 521–534 (1952)
Eyer, L., et al.: Nucleoside inhibitors of Zika virus. J. Infect. Dis. 214(5), 707–711 (2016)
Fagbami, A.: Zika virus infections in Nigeria: virological and seroepidemiological investigations in Oyo state. Epidemiol. Infect. 83(2), 213–219 (1979)
Fauci, A.S., Morens, D.M.: Zika virus in the Americas–yet another arbovirus threat. N. Engl. J. Med. 374(7), 601–604 (2016)
Fink, S.L., et al.: The antiviral drug arbidol inhibits Zika virus. Sci. Rep. 8(1), 8989 (2018)
Florez, H., Salvatierra, K.: Bioinformatics study of mutations of resistance to antivirals in the NS5A gen of HCV. Int. Inf. Inst. (Tokyo) Inf. 20(9), 6665–6672 (2017)
Hamel, R., et al.: Zika virus: epidemiology, clinical features and host-virus interactions. Microbes Infect. 18(7–8), 441–449 (2016)
Kieffer, T.L., Kwong, A.D., Picchio, G.R.: Viral resistance to specifically targeted antiviral therapies for hepatitis C (STAT-Cs). J. Antimicrob. Chemother. 65(2), 202–212 (2009)
Koenig, K.L., Almadhyan, A., Burns, M.J.: Identify-isolate-inform: a tool for initial detection and management of Zika virus patients in the emergency department. West. J. Emerg. Med. 17(3), 238 (2016)
Kostyuchenko, V.A., et al.: Structure of the thermally stable Zika virus. Nature 533(7603), 425 (2016)
Kwong, A.D., McNair, L., Jacobson, I., George, S.: Recent progress in the development of selected hepatitis C virus NS3.4A protease and NS5B polymerase inhibitors. Curr. Opin. Pharmacol. 8(5), 522–531 (2008)
Li, C., et al.: Zika virus disrupts neural progenitor development and leads to microcephaly in mice. Cell Stem Cell 19(1), 120–126 (2016)
Lou, Z., Sun, Y., Rao, Z.: Current progress in antiviral strategies. Trends Pharmacol. Sci. 35(2), 86–102 (2014)
Organization, W.H., et al.: Zika virus research agenda (2016)
Pang, T., Mak, T.K., Gubler, D.J.: Prevention and control of dengue–the light at the end of the tunnel. Lancet Infect. Dis. 17(3), e79–e87 (2017)
Pattnaik, A., et al.: Discovery of a non-nucleoside rna polymerase inhibitor for blocking Zika virus replication through in silico screening. Antivir. Res. 151, 78–86 (2018)
Penié, J.B., González-Piñera, J.G., Rodríguez, M.A.R., Alfonso, P.P.P.: Medicamentos antivirales. Acta Médica 8(1), 86–100 (1998)
Petersen, L.R., Jamieson, D.J., Powers, A.M., Honein, M.A.: Zika virus. N. Engl. J. Med. 374(16), 1552–1563 (2016)
Plourde, A.R., Bloch, E.M.: A literature review of Zika virus. Emerg. Infect. Dis. 22(7), 1185 (2016)
Rather, I.A., Lone, J.B., Bajpai, V.K., Paek, W.K., Lim, J.: Zika virus: an emerging worldwide threat. Front. Microbiol 8, 1417 (2017)
Sacramento, C.Q., et al.: The clinically approved antiviral drug sofosbuvir impairs Brazilian Zika virus replication. BioRxiv, p. 061671 (2016)
Saiz, J.C., Vázquez-Calvo, Á., Blázquez, A.B., Merino-Ramos, T., Escribano-Romero, E., Martín-Acebes, M.A.: Zika virus: the latest newcomer. Front. Microbiol. 7, 496 (2016)
Salvatierra, K., Florez, H.: Analysis of hepatitis C virus in hemodialysis patients. Infectio 20(3), 130–137 (2016)
Salvatierra, K., Florez, H.: Biomedical mutation analysis (BMA): a software tool for analyzing mutations associated with antiviral resistance. F1000Research 5, 1141 (2016)
Salvatierra, K., Florez, H.: Prevalence of hepatitis B and C infections in hemodialysis patients. F1000Research 5, 1–6 (2016)
Song, B.H., Yun, S.I., Woolley, M., Lee, Y.M.: Zika virus: history, epidemiology, transmission, and clinical presentation. J. Neuroimmunol. 308, 50–64 (2017)
Trott, O., Olson, A.J.: Autodock vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31(2), 455–461 (2010)
Tuset, M., José, M., Del Cacho, E., Alberdi, A., Codina, C., Ribas, J., et al.: Características de los fármacos antivirales. Enfermedades infecciosas y microbiologia clinica 21(8), 433–458 (2003)
Yang, C.C., et al.: A novel dengue virus inhibitor, BP13944, discovered by high-throughput screening with dengue virus replicon cells selects for resistance in the viral NS2B/NS3 protease. Antimicrob. Agents Chemother. 58(1), 110–119 (2014)
Yin, Z., et al.: An adenosine nucleoside inhibitor of dengue virus. Proc. Natl. Acad. Sci. 106(48), 20435–20439 (2009)
Zmurko, J., Marques, R.E., Schols, D., Verbeken, E., Kaptein, S.J., Neyts, J.: The viral polymerase inhibitor 7-deaza-2’-c-methyladenosine is a potent inhibitor of in vitro Zika virus replication and delays disease progression in a robust mouse infection model. PLoS Negl. Trop. Dis. 10(5), e0004695 (2016)
Acknowledgment
Authors are grateful for the support received from Universidad Nacional de Misiones, Posadas (Argentina), and the Information Technologies Innovation (ITI) Research Group, Universidad Distrital Francisco Jose de Caldas, Bogota (Colombia).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Salvatierra, K., Vera, M., Florez, H. (2019). Bioinformatics Methods to Discover Antivirals Against Zika Virus. In: Florez, H., Leon, M., Diaz-Nafria, J., Belli, S. (eds) Applied Informatics. ICAI 2019. Communications in Computer and Information Science, vol 1051. Springer, Cham. https://doi.org/10.1007/978-3-030-32475-9_1
Download citation
DOI: https://doi.org/10.1007/978-3-030-32475-9_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-32474-2
Online ISBN: 978-3-030-32475-9
eBook Packages: Computer ScienceComputer Science (R0)