Abstract
RNA degradation is considered a critical posttranscriptional regulatory checkpoint, maintaining the correct functioning of organisms. When a specific RNA transcript is no longer required in the cell, it is signaled for degradation through a number of highly regulated steps. Ribonucleases (or simply RNases) are key enzymes involved in the control of RNA stability. These enzymes can perform the RNA degradation alone or cooperate with other proteins in RNA degradation complexes. Important findings over the last years have shed light into eukaryotic RNA degradation by members of the RNase II/RNB family of enzymes. DIS3 enzyme belongs to this family and represents one of the catalytic subunits of the multiprotein complex exosome. This RNase has a diverse range of functions, mainly within nuclear RNA metabolism. Humans encode two other DIS3-like enzymes: DIS3L (DIS3L1) and DIS3L2. DIS3L1 also acts in association with the exosome but is strictly cytoplasmic. In contrast, DIS3L2 acts independently of the exosome and shows a distinctive preference for uridylated RNAs. These enzymes have been shown to be involved in important cellular processes, such as mitotic control, and associated with human disorders like cancer. This review shows how the impairment of function of each of these enzymes is implicated in human disease.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Introduction
RNA is a labile molecule, by its chemical nature, and a plethora of events and factors ensure its protection or decay. The termini of cytoplasmic mRNAs are usually protected by a 7-methyl guanosine (m7GpppG) at the 5′ end cap, and by a long terminal poly(A) tail at the 3′ end, which promotes the association of Poly(A) Binding Proteins (PABPs) (Fig. 4.1a). These RNA modifications in the ends of the mRNA molecules warrant their stability. Specific protein factors associate with each of the structures in the mRNA extremities, and its physical interaction holds the mRNA in a circular conformation [1, 2]. This closed-loop mRNA structure is recognized by specific translation initiation complexes [2,3,4,5]. When a specific RNA is no longer required for the cellular metabolism, the molecule is signaled for degradation through a number of highly regulated steps (Fig. 4.1a).
(a) Overview of RNA degradation pathways in humans. RNA transcripts first undergo removal of the 3′ poly(A) tail through a deadenylation step (performed by PAN2/PAN3 complex followed by the CCR4/NOT). Following this step, the RNA becomes vulnerable and can be degraded 3′-5′ by the Exosome-DIS3/DIS3L1 or by DIS3L2 (that prefers uridylated RNA substrates), independently of the exosome. Following deadenylation, RNAs can also be decapped by DCP1/DCP2 (removal of the 5′-cap), a step stimulated by Lsm1–7–Pat1 complex, exposing the transcripts to the 5′-3′ exoribonuclease XRN1. (b) Correlation between DIS3 enzymes and human disease. Scheme of human diseases that have been related with overexpression or dysfunction of DIS3, DIS3L1 and DIS3L2. Dashed arrows correspond to correlations that need further confirmation. Each enzyme-disease association represented in the picture is developed in the main text, with the respective references
4.2 Mechanisms of Cytoplasmic mRNA Degradation in Humans
The stability of mRNAs depends on intrinsic features of its sequence and on the cellular demands for the protein it encodes. For instance, specific features such as AU rich elements (AREs) that consist on stretches of adenine and uracil nucleotides within the 3′UTR of some mRNAs [6,7,8,9] are able to promote or protect RNA from degradation [10,11,12]. The 3′UTR of the mRNAs can also contain other specific sequences that target its decay trough the binding of regulatory microRNAs (miRNAs) [13,14,15].
Under particular circumstances, like in cellular quality control mechanisms, endonucleolytic cleavage can directly disrupt the closed-circle RNA conformation and trigger subsequent decay, from either of the RNA extremities ([16, 17]; see other chapters in this volume for details on quality control mechanisms). Though, in general, mRNAs must be first deadenylated or decapped to enable the access of exoribonucleases and trigger degradation.
For many cytoplasmic mRNAs, decay starts with the shortening or removal of the poly(A) tail. Deadenylation releases the PABPs that protect the (A) tail and leaves the 3′ end exposed to exoribonucleases. This is considered the main event that signals mRNAs for degradation from the 3′ end. The length of the poly(A) tail depends on the organism, but the regulation of its extension is always a dynamic process that involves the concerted action of poly(A) polymerases (PAPs) and poly(A) specific 3′ exonucleases (deadenylases). This allows the fine-tuning control of mRNA stability. Eukaryotic genomes encode a wide variety of deadenylases [18]. In humans, initial deadenylation is performed by the PAN2/PAN3 complex (reviewed in [19,20,21]) followed by the action of the CCR4/NOT complex (Fig. 4.1a) [22,23,24,25,26].
Following deadenylation, degradation can alternatively proceed from the 5′ end requiring the prior removal of the 5′-cap of the mRNA. The deccaping process is regulated by a plethora of activators and inhibitors [27,28,29]. One of these, the Lsm1–7-Pat1 complex, preferentially binds the shortened 3′-terminal adenosine extensions of the deadenylated mRNAs to stimulate decapping and inhibit exosome attachment [30, 31]. The reaction products of decapping enzymes are the 5′ m7GDP cap and an unprotected 5′ monophosphate RNA that is then accessible for XRN1 processive and complete degradation [32, 33]. Decapping process commits RNAs to 5′-3′ degradation since XRN1 directly interacts with the decapping enzymes (DCP1/DCP2) [32] (Fig. 4.1a).
In contrast to the 5′-3′ pathway, where XRN1 is the only known cytoplasmic 5′-3′ exoribonuclease, there are several options in the degradation from the 3′ end. After initial deadenylation, the multisubunit RNA exosome complex may further degrade the shortened oligo(A) tail and proceed with the 3′-5′ degradation into the mRNA body. Exosome activity depends on the presence of specific cofactors, called “superkillers” or Ski proteins that regulate its activity. Human homologs of the Ski family – SKIV2L (Ski2), TTC37 (Ski3) and WDR61 (Ski8), associate in the Ski complex [26, 34]. The remaining mRNA fragment with its 5′-cap (m7GpppG) is hydrolyzed by the scavenger decapping enzyme (DCPS) [35]. DCPS is a m7G-specific pyrophosphatase that shows specificity towards RNA fragments not longer than 10 nts (Fig. 4.1a) [36,37,38].
DIS3 enzyme is the essential catalytic subunit of the exosome. The human genome encodes three members of DIS3 family: DIS3, DIS3L1 and DIS3L2 enzymes and their characteristics and specifications will be described therein in this chapter. While both DIS3 and DIS3L1 interact with the exosome ring, in different cellular locations [39,40,41,42,43], DIS3L2 does not. The enzyme represents a 3′-5′ RNA decay pathway alternative to degradation by XRN1 and the exosome [44, 45]. DIS3L2 shows a distinctive preference towards uridylated substrates, which prompted the discovery of new roles for 3′-uridylation in cytoplasmic mRNA decay. It was proposed that the requirement of deadenylation as an mRNA decay signal can be overcome trough 3′ oligouridylation of transcripts. It can either stimulate decapping and consequent degradation in the 5′-3′ direction, through Lsm1–7/Pat1 binding, or directly activate 3′-5′ DIS3L2 dependent degradation [46] (Fig. 4.1a). The importance of this uridylation-dependent pathway in bulk mRNA degradation was highlighted by the substantial technological progresses in RNA analysis in the recent years. Novel approaches as TAIL-seq revealed that 3′ end mRNA modifications such as urydilation, cytidylation or guanylation are also frequent [47]. The use of oligo(dT)-based priming methods, due to the long poly(A) tails in the 3’ end of eukaryotic mRNAs, had previously underestimated its presence.
In general, regulation of gene expression in eukaryotic cells occurs through multiple parallel, partially redundant, mRNA decay pathways. This is further illustrated by the multiplicity of enzymes, which are able to catalyze the same reaction, and their functional redundancy. RNases’ activity is also important for RNA surveillance and processing. Their high degree of conservation in different organisms and the specific phenotypes following their individual loss suggest defined roles within the cell.
Beyond the advances on the mechanisms whereby these enzymes affect cellular processes, structural information is crucial to explain the mechanism of action and exact function of the protein alone or in the context of a multiprotein complex. Structural changes across species provide insight into the evolution and conservation of the protein architecture. In fact, important findings over the last years have shed new light onto the mechanistic details of RNA degradation by members of the RNase II/RNB family of exoribonucleases, including DIS3 enzymes [45, 48,49,50,51,52,53,54,55,56]. A phylogenetic comparison of different Dis3 homologues in eukaryotes indicates a clear division of three Dis3-like protein families, with the Dis3 group being the most conserved and the Dis3L1 and Dis3L2 groups being more divergent [45].
A growing number of publications associate DIS3-enzymes with several human diseases [57,58,59,60]. In this review we will try to sum up the mechanistic and structural details of these RNase II-like enzymes, and how human disorders can result from the associated defects.
4.2.1 DIS3
DIS3 (defective in sister chromatid joining) gene is located on human chromosome 13 (q22·1), and it encodes for a highly conserved ribonuclease (also known as Rrp44 in yeast) that contains both 3′-5′ exoribonuclease and endoribonuclease activities [40, 42, 61]. This RNase constitutes an essential catalytic subunit of the exosome [62]. This multiprotein complex is composed of a catalytically inert ring-shaped 9-subunit core with a prominent central channel and associated catalytic subunits [61, 63,64,65]. The composition of its catalytic subunits varies accordingly to cellular localization. Exosome-associated DIS3 enzyme exists mainly in the nucleus, where it acts on a vast array of different RNAs, and has a diverse range of functions in RNA metabolism including processing, maturation and quality control [66,67,68,69,70,71].
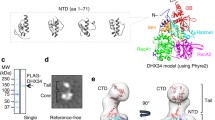
The specific domains of DIS3 dictate its degradation preferences. DIS3 is composed by two cold shock domains (CSD1 and CDS2), followed by an RNB and an S1 domain. Both CSDs and the S1 domain contribute to RNA binding. At the N-terminal region, it contains a PilT N-terminal (PIN) domain [40, 42, 51, 72] and also a CR3 motif involved in the binding of DIS3 to the exosome [73]. Both RNB and PIN domains are responsible for RNA degradation activity. The RNB catalytic domain is a hallmark of the RNase II protein family, and confers to DIS3 the ability to cleave RNA in a highly processive manner [51, 61, 74]. Arraiano’s lab contributed to the resolution of the crystal structure of the family prototype, E. coli RNase II, and to its extensive functional characterization [49,50,51]. This was an important breakthrough in the understanding of the mechanism of action of this ubiquitous family of proteins. Moreover, the determination of the electron microscopy structure of yeast Rrp44 (Dis3) suggested that the RNA recruitment mechanism is conserved [75]. The knowledge acquired by these model organisms was crucial for the construction of the 3D model of human DIS3. Its exoribonuclease activity is dependent on four conserved aspartic acid residues (D488, D487, D485, D489) that coordinate two magnesium ions in the catalytic center [61, 63]. The RNB active site in DIS3 is responsible to hydrolyze single-stranded RNA (ssRNA) in a 3′–5′ direction, releasing one nucleotide at a time and leaving an end product of 4 nts [51, 70, 76,77,78]. Only ssRNAs with a minimum length of 7 nts can be cleaved [76]. Dis3/Rrp44 is also able to unwind and digest structured RNAs as long as there is an unstructured region of ~4 nts at the 3′ terminus [79].
The PIN domain in DIS3 confers the ability to cleave RNA endoribonucleolytically [40, 42, 43]. PIN-like domains constitute a widespread superfamily of nucleases with representatives in all kingdoms of life [80, 81]. Combination of endoribonuclease and exoribonuclease activities is a widespread feature of RNA-degrading machines from bacteria to humans [48]. Both DIS3 activities cooperate with each other in the degradation of RNA molecules [40, 42, 43, 82]. The PIN domain is able to cleave circular and linear ssRNAs, preferentially with a 5′ monophosphate [40, 42]. Its active site is composed of four acidic amino acids essential for endoribonuclease activity (E97, D69, D177 and D146) that coordinate two divalent metal ions, and Mn2+ is the preferred ion for its activity [40, 42]. It was proposed that the role of the PIN domain is to assist in the release of RNA substrates that are stalled at sites with strong secondary structures [83]. Besides its endoribonuclease activity, the PIN domain has also a structural role, being necessary for DIS3 association with the core exosome [39, 42, 43, 84].
Like in other organisms, such as Drosophila and yeast, human DIS3 is essential for survival [85, 86]. This RNase together with the exosome complex play a crucial role in maintaining the fidelity of gene expression. In the nucleus, the exosome-associated DIS3 is involved in the degradation of a vast range of RNAs, including protein-coding RNAs, stable RNA species such as ribosomal RNA (rRNA), transfer RNA (tRNAs) and small nucleolar RNAs (snoRNAs), introns, long non-coding RNAs (ncRNAs), miRNAs and also unstable RNAs products, like Promoter Upstream Transcripts (PROMPTs) [87].
An impaired RNA surveillance system can compromise RNA homeostasis, having detrimental consequences for multiple biological processes, which may result in malignancy [88]. Indeed, an increasing number of publications have associated dysregulation of DIS3 with human disease, namely cancer (reviewed by [60, 89]). Sequencing data have identified DIS3 gene as one of the most frequently mutated genes in multiple myeloma (on average in 11–18.5% of patients) [58, 90,91,92,93]. This constitutes the second most frequent hematologic tumor after lymphomas [94]. In multiple myeloma patients, DIS3 mutations were detected in highly conserved regions along PIN, CSD2, RNB and S1 domains [58, 90, 91, 93]. However, the mutations in the RNB domain seem to be more prevalent for the development of the disease [58, 90, 91], and contain mutational hotspots (D488, E665 and R780) [90, 93].
Tomecki and co-workers have studied Multiple myeloma mutations that abolish or cause dysfunction, without total inactivation, of DIS3 RNB activity in vitro [85]. The results indicated that the point mutations D487N, S477R, G766R and R780K cause significant aberrations on exoribonucleolytic activity. DIS3 amino acid changes with significantly decreased activity in vitro gave rise to a slower cellular proliferation rate in HEK293-derived cells [85]. DIS3-mutations such as R780K, an amino acid involved in RNA binding, also revealed an abnormal RNA metabolism, with accumulation of 5.8S processing intermediates, tRNAs, RNA polymerase III transcripts and PROMPTs [85].
Szczepińska and co-workers also observed a major role of DIS3 in maintaining RNA polimerase II transcriptome homeostasis and in the regulation of PROMPTs [87]. PROMPTs are transcribed in reverse orientation to most active protein coding genes, and cover ∼1% of the human genome [72, 95]. Although the biological role of PROMPTs has yet to be elucidated, there are evidences that these RNAs could serve important functions in human cells [96]. For instance, the PROMPT HIF2PUT was suggested to be a novel regulator of osteosarcoma, the most common primary bone malignancy [96]. The observation that PROMPTs were the most prominent targets of DIS3 (>50-fold increase in DIS3 mutant cells), indicates that there are no alternative pathways for their decay, making their connection with DIS3 and disease necessary to be explored.
Multiple myeloma associated mutations were also mapped on PIN domain, showing only small effects on cell growth [85]. However, when mutations in PIN and RNB domains are combined, a synergistic effect in the proliferation and metabolic activity is observed in human cells [85, 87].
Human cells bear two DIS3 isoforms that differ in the size of the PIN domain. Isoform 1 encodes a full-length PIN domain, whereas the PIN domain of isoform 2 is shorter and misses a segment with conserved amino acids [52]. A study by Robinson and co-workers [52] anticipated that different ratios of the two PIN isoforms could be characteristic of several haematological cancers, namely Multiple Myeloma [52]. Isoform 1 was found in higher levels than isoform 2 in Multiple Myeloma patient samples and all cancer cell lines tested [52]. Contrastingly, healthy donors and Acute Myeloid Leukemia and Chronic Myelomonocytic Leukaemia patients have similar levels of both isoforms. Regarding leukaemias, in Acute Myeloid Leukemia (a cancer of the myeloid line of blood cells) missense mutations in DIS3 account for 4% of patients and were all found in the RNB domain [59]. In patients with Chronic Lymphocytic Leukemia (a monoclonal disorder characterized by a progressive accumulation of abnormal lymphocytes) DIS3 locus, 13q22, is often deleted [97]. This evidence together with DIS3 mutations in several cancers, suggest that DIS3 may function as a tumor suppressor gene.
Increased levels of DIS3 mRNA and protein have also been proposed as one of the causes of other types of cancer. This is the case of epithelial ovarian cancer in which DIS3 was observed to be significantly up-regulated in plasma from patients in the late stage of the disease (FIGO III/IV) [98]. The majority of cancer deaths are due to metastasis of neoplastic cells from the primary tumor to distant organs. Metastasis is thus the most important factor that determines bad prognoses for cancer patients. In 1997, DIS3 was reported to have a 38-fold higher expression in primary tumors and metastatic cells from patients with colorectal cancers and liver metastases (compared to adenomas) [99]. The same study has classified DIS3 as an oncogene, being positively correlated with the incidence of metastasis and consistent with its involvement in the regulation of mitosis in Schizosaccharomyces pombe, Saccharomyces cerevisiae and Drosophila [66, 100,101,102]. Other studies also reported a significant overexpression of DIS3 in colorectal carcinomas (compared to adenomas) [103, 104]. This overexpression could be explained by an amplification of the DIS3 locus, 13q22, frequently observed in colorectal cancer.
DIS3 was also found to be differentially expressed in melanoma cells [105]. Specifically, in superficial spreading melanoma cells, DIS3 has a reduced expression, contrarily to nodular melanoma cells, where DIS3 is overexpressed (compared to normal melanocytes) [105] However, in 2013, a wide-genome analysis resultant from five melanoma microarrays datasets did not recognize DIS3 as a melanoma biomarker [106]. DIS3 has also a role in pancreatic tumorigenesis and breast cancer, however their linkage has to be further explored [107, 108].
All examples presented here strongly suggest that both DIS3 overexpression and lack of function can lead to the manifestation of different cancers. This seems contradictory, but it is in agreement with the fact that DIS3 may function as either oncogene or tumor suppressor. Some genes are known to have both functions, and recently it was reported that most of these genes are transcription factors or kinases that can regulate transcription positively and negatively [109]. Since RNases are responsible to control post-transcriptionally gene expression, it is not surprising that DIS3 would also have a dual function. For instance, DIS3 is known to facilitate the maturation of the tumor suppressor let-7 miRNA. When the levels of the mature let-7 miRNA are reduced, translation of oncogenes (MYC and RAS) increases, enhancing tumorigenesis [110]. Human DIS3 also appear to function in the Ran signaling pathway required for nuclear import of proteins [111]. Ran was associated to cancer progression and has been investigated as a target for cancer therapy. In sum, the precise and dual role of DIS3 in cancer is not fully understood lacking further investigation.
4.2.2 DIS3L1
Human DIS3 and DIS3L1 have a similar domain composition, however only the first has an active endoribonuclease domain. Two important residues (E97 and D146N) are absent in DIS3L1 PIN domain rendering it inactive. The aspartic acid D146 is the most conserved in the PIN domains and its single mutation is reported to abolish its activity in vivo and in vitro [43, 112, 113]. The E97 is not strictly conserved across the PIN-domain family [112].
Both human DIS3 and DIS3L1 associate with the exosome ring. In contrast to the mainly nuclear localization of DIS3, DIS3L1 is strictly cytoplasmic [57, 72, 114]. The stable association of DIS3L1 with the cytoplasmic exosome suggests that it acts in concert with the core of the exosome in the degradation of cytoplasmic RNAs. One of the RNA substrates degraded by the exosome-associated DIS3L1 is the 28S rRNA. The degradation proceeds through polyadenylated intermediates, which accumulate upon DIS3L1 knockdown [115, 116]. DIS3L1 was also implicated in the degradation of intermediary products generated by DNA-based antisense oligonucleotides (ASOs), as part of the RNA surveillance machinery. These agents recruit RNase H1 that after endonucleolytic cleavage of the ASO-targeted mRNAs generates both a 5′ and a 3′ fragments. DIS3L1 appears to be involved, together with the exosome, on the 3′-5′ exoribonucleolytic degradation of the cytoplasmic upstream cleavage products [117].
Less is known about DIS3L1 association with human disease, but there are a few reports showing its implication on diverse pathologies, directly or indirectly related with its function over specific substrates. It was suggested that DIS3L1 could be implicated in the regulation of steady state levels of Y RNAs, an abundant class of small non-coding RNAs with a role in a range of cellular processes, such as RNA quality control, DNA replication, cellular stress responses and histone mRNA processing [118, 119]. Several enzymes are involved in Y RNAs maturation, both on its 3′ end adenylation and subsequent trimming. Poly(A)-specific ribonuclease (PARN) is one of the enzymes involved on its 3′ end processing and, in its absence, oligo(A) tails are degraded by the exoribonuclease DIS3L1. It was seen that PARN mutations cause a severe form of dyskeratosis congenita (DC), a telomere biology disorder characterized by dysplastic nails, lacy reticular pigmentation of the upper chest and/or neck, and oral leukoplakia [120]. The loss of PARN reduces the levels of human Y RNAs. At the same time, low levels of Y RNAs intensify the effect of PARN depletion on telomere maintenance, leading to the same severe DC phenotype. PARN seems to be responsible to stabilize Y RNAs by removing the oligoadenylated tails that recruit DIS3L1 for degradation [121]. Moreover, it was recently demonstrated that PARN is also involved in miRNAs stabilization by removing their oligo(A) tails – the signal for the recruitment of the cytoplasmic exonucleases DIS3L1 or DIS3L2. Therefore, upon PARN knockdown there is decrease in miRNAs’ levels, namely several that target p53 mRNA (a gene that plays a central role in cancer). This was the missing link to explain the p53 accumulation that is observed in PARN-defective patients [122].
DIS3L1 also seems to play a role in cancer, similarly to DIS3 (see above) and DIS3L2 (see bellow) homologues. The Hedgehog (Hh) pathway controls cell proliferation and differentiation in response to a gradient of secreted Hh ligands, and its aberrant activation can promote tumorigenesis. The transcriptional factor Zfx is a common cell-intrinsic regulator of diverse Hh-induced tumors. hDIS3L1, the human gene encoding DIS3L1 was identified as direct transcriptional target of Zfx, in the context of skin basal cell carcinoma (BCC) and cerebellar medulloblastoma (MB) models in vivo and in vitro [123].
Two independent exome sequencing studies (technique that sequences all protein-coding genes in a genome) have reported an association of hDIS3L1 with cardiac risk. First, in a study to identify genetic variants that confer susceptibility to myocardial infarction (MI) in the Asian population (Korean individuals), several single nucleotide polymorphisms (SNPs) on hDIS3L1 gene were associated with MI risk. However, how the gene influences MI pathogenesis would have to be determined and confirmed in other ethnic populations [124]. In another study, novel SNPs were also identified in the hDIS3L1 gene of individuals with Hyperalphalipoproteinemia (HALP). This condition of high-density lipoprotein cholesterol (HDL) levels is inversely correlated with coronary heart disease (CHD), and hDIS3L1 gene was identified as a candidate gene associated with HALP [125].
4.2.3 DIS3L2
Human DIS3L2 is the third member of the RNase II/RNB family of enzymes. This protein is a processive 3′-5′ exoribonuclease (mainly cytoplasmic) able to degrade structured RNA molecules, as long as they possess a 2 nt 3′ overhang as a “landing platform” [44, 45]. Unlike its family counterparts (DIS3 and DIS3L1), DIS3L2 lacks the PIN and the CR3 domains on its structure, both necessary for the interaction with the exosome complex [39, 43, 45, 72, 73].
In mammalian cells, DIS3L2 is involved in miRNA maturation and in the decay of numerous RNA-species, namely bulk mRNA, ARES and ncRNAs [126,127,128,129]. Several studies associated DIS3L2 with a degradation pathway that relies on the addition of untemplated uridines to several classes of RNAs, in a process called uridylation [44, 45, 127, 129,130,131,132]. This process was reported for the first time in S. pombe [133], in which Dis3L2 was shown to degrade uridylated poly(A)-containing mRNAs [45]. Later on, uridylation was found to be widespread and to have a decisive impact on RNA′s fate. There is a negative correlation between the addition of short (1–4) uridine residues with mRNA stability [47, 130]. Oligo(U) tailed mRNAs are recognized by Lsm1–7 complex stimulating 3′-5′ degradation by DIS3L2, however it can also trigger decapping by DCP2 allowing 5′-3′ degradation by XRN1 [44, 45, 129, 130, 133,134,135]. DIS3L2 crystal structure unveiled the DIS3L2 RNA pathway, revealing three uracil-specific zones that explain how DIS3L2 recognizes, binds and processes preferentially oligo(U)-tailed RNAs [132].
The uridylation process is achieved by proteins termed uridyltransferases (TUTases). Humans have seven TUTases that are strictly cytoplasmic, except TUTase-1 (TUT1) that can also be found in mitochondria [133, 136]. Two TUTases were implicated in mRNA uridylation at the 3′ end, TUT4 and TUT7 [130]. In the same study, Lim and colleagues showed that these TUTases were able to sense the length of the poly(A) tail, and preferentially uridylate mRNAs with a tail ranging between 0–25 As. On the contrary, PABPs preferentially bind longer poly(A) tails protecting them from the action of TUT4/7 [130].
This so called TUT-DIS3L2 mRNA decay mechanism was found to prevail in cells under apoptosis. Apoptosis is the most common physiological program of cell death, which plays a vital role in pathogen immune defense, removal of damaged cells, cancer surveillance and cancer therapy effectiveness [137]. Thomas and colleagues [138] observed in human apoptotic cells, that apoptosis triggers global mRNA decay, and the RNA products generated are 3′-uridylated by TUT4/7 and subsequently degraded by DIS3L2. Knockdown of the exoribonuclease inhibits mRNA decay and suppresses cell death; conversely, DIS3L2 overexpression enhances apoptosis, supporting that mRNA decay is a hallmark of cell death [138]. Recently, Liu and colleagues [139], brought another player to this process, a mitochondrial exoribonuclease called PNPT1 that evolved from bacterial PNPase [48, 139, 140]. The work performed in human colon cancer cells, demonstrated that, upon apoptosis triggering, PNPT1 and DIS3L2 act in the same pathway. PNPT1 is released from mitochondria, and starts to degrade RNA from the 3′ end. PNPT1 stops whenever it encounters an obstacle (e.g. ribosome, RNA-binding protein or highly structured sequence), being the RNAs further degraded by the TUT-DIS3L2 pathway [139].
DIS3L2 is also involved in the regulation of let-7 miRNA expression in pluripotent cells, establishing a role of this enzyme in cell differentiation [128, 131, 132, 141]. Indeed, let-7 pre-miRNA biogenesis is one of the best characterized DIS3L2-mediated pathways. miRNAs from the let-7 family function as tumor suppressors and are involved in stem cell renewal [128, 131]. In undifferentiated cells, the expression of let-7 miRNAs is blocked by Lin28, a pluripotency factor that also functions as an oncogene in several cancers [142]. This RNA-binding protein binds to let-7 precursors and promotes their uridylation by TUT4/7. These RNA precursors are thus marked for DIS3L2 degradation, leading to inhibition of let-7 biogenesis.
A recent study has also found a role of DIS3L2 in nonsense-mediated decay (NMD), a quality control pathway that degrades aberrant and physiological mRNAs to maintain cellular homeostasis (as discussed in Chap. 3). In this context, DIS3L2 acts over 3′ ends of NMD decay intermediates that were previously subject to uridylation (L. Romão, personal communication and [143]).
The involvement of DIS3L2 in such cellular important processes, like apoptosis, cell differentiation and RNA quality control (NMD) anticipates its role in human disease. In fact, this RNase has been related with several human disorders. DIS3L2 is associated with Perlman syndrome, which is a rare congenital overgrowth disease [57, 144]. Children affected with Perlman syndrome display macrocephaly, facial abnormalities, neurodevelopmental delay, fetal gigantism, kidney abnormal enlargement and high neonatal mortality. These children also present nephroblastomatosis, an important precursor for Wilms’ tumor, a kidney cancer also known as nephroblastoma. Astuti et al. [57] demonstrated that the affected children have germline mutations consistent with DIS3L2 loss of function. DIS3L2 mutations were also associated with Wilms tumor susceptibility [57, 144]. It has been recently suggested that regulation of the growth-promoting gene, insulin growth factor 2 (IGF2), by DIS3L2, could be the link between this RNase and Wilms tumorigenesis [145]. Interestingly, Gregory RI and colleagues have found that DIS3L2 has no effect on the steady state mRNAs levels in DIS3L2-deficient cell lines and knockout mouse kidneys. Instead, it rather specifically perturbs endoplasmic reticulum (ER)-mediated translation (R.I. Gregory, personal communication).
Besides its well documented role in Perlman syndrome and Wilms’ tumor, DIS3L2 was also found to be mutated in 3–6% of carcinomas [57, 146]. Also, DIS3L2 has been associated with a Marfan-like syndrome with skeletal overgrowth [147]. The Marfan syndrome is a disorder of the connective tissue that causes high mortality for untreated patients, mainly due to aortic complications [148]. Patients in which DIS3L2 gene was affected showed skeletal overgrowth and malformations, including severe scoliosis (abnormal curvature of the spine), arachnodactyly (long, slender fingers, curvature of the hands and feet) and mild syndactyly (interdigital webbing) [147, 149, 150].
4.3 Concluding Remarks
RNA degradation is a set of highly regulated steps that maintain cellular integrity and homeostasis. DIS3-enzymes act over a panoply of RNA substrates in eukaryotic cells and it is clear their role in human disease, namely in cancer development and progression. In this chapter, we explored the consequences of DIS3-enzymes impairment on the physiology of human cells. From this group of proteins, the most well-characterized is DIS3, however its role in cancer is not completely understood. Less is known about the mechanism of action and specific RNA targets of its homologs DIS3L1 and DIS3L2. The molecular mechanisms that link both proteins with disease are still unexplored. Despite the progress that has been made, there is still much work to perform in order to completely understand how DIS3-enzymes regulate cellular pathways, and how they are related with disease progression. Clinical medicine will certainly benefit from this kind of fundamental research.
References
Archer SK et al (2015) Probing the closed-loop model of mRNA translation in living cells. RNA Biol 12(3):248–254
Wells SE et al (1998) Circularization of mRNA by eukaryotic translation initiation factors. Mol Cell 2(1):135–140
Darnell JE Jr (2013) Reflections on the history of pre-mRNA processing and highlights of current knowledge: a unified picture. RNA 19(4):443–460
Kahvejian A et al (2005) Mammalian poly(A)-binding protein is a eukaryotic translation initiation factor, which acts via multiple mechanisms. Genes Dev 19(1):104–113
Mangus DA, Evans MC, Jacobson A (2003) Poly(A)-binding proteins: multifunctional scaffolds for the post-transcriptional control of gene expression. Genome Biol 4(7):223
Barreau C, Paillard L, Osborne HB (2005) AU-rich elements and associated factors: are there unifying principles? Nucleic Acids Res 33(22):7138–7150
Chen CY, Shyu AB (1995) AU-rich elements: characterization and importance in mRNA degradation. Trends Biochem Sci 20(11):465–470
Eulalio A et al (2009) Deadenylation is a widespread effect of miRNA regulation. RNA 15(1):21–32
Wu L, Fan J, Belasco JG (2006) MicroRNAs direct rapid deadenylation of mRNA. Proc Natl Acad Sci U S A 103(11):4034–4039
Abdelmohsen K, Gorospe M (2010) Posttranscriptional regulation of cancer traits by HuR. Wiley Interdiscip Rev RNA 1(2):214–229
Murray EL, Schoenberg DR (2007) A+U-rich instability elements differentially activate 5′-3′ and 3′-5′ mRNA decay. Mol Cell Biol 27(8):2791–2799
Peng SS et al (1998) RNA stabilization by the AU-rich element binding protein, HuR, an ELAV protein. EMBO J 17(12):3461–3470
Huntzinger E, Izaurralde E (2011) Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nat Rev Genet 12(2):99–110
Oliveto S et al (2017) Role of microRNAs in translation regulation and cancer. World J Biol Chem 8(1):45–56
Valinezhad Orang A, Safaralizadeh R, Kazemzadeh-Bavili M (2014) Mechanisms of miRNA-mediated gene regulation from common downregulation to mRNA-specific upregulation. Int J Genomics 2014:970607
Eberle AB et al (2009) SMG6 promotes endonucleolytic cleavage of nonsense mRNA in human cells. Nat Struct Mol Biol 16(1):49–55
Huntzinger E et al (2008) SMG6 is the catalytic endonuclease that cleaves mRNAs containing nonsense codons in metazoan. RNA 14(12):2609–2617
Pavlopoulou A et al (2013) A comprehensive phylogenetic analysis of deadenylases. Evol Bioinformatics Online 9:491–497
Uchida N, Hoshino S, Katada T (2004) Identification of a human cytoplasmic poly(A) nuclease complex stimulated by poly(A)-binding protein. J Biol Chem 279(2):1383–1391
Wahle E, Winkler GS (2013) RNA decay machines: deadenylation by the Ccr4-not and Pan2-Pan3 complexes. Biochim Biophys Acta 1829(6–7):561–570
Wolf J, Passmore LA (2014) mRNA deadenylation by Pan2-Pan3. Biochem Soc Trans 42(1):184–187
Chen CY, Shyu AB (2011) Mechanisms of deadenylation-dependent decay. Wiley Interdiscip Rev RNA 2(2):167–183
Doidge R et al (2012) Deadenylation of cytoplasmic mRNA by the mammalian Ccr4-Not complex. Biochem Soc Trans 40(4):896–901
Lau NC et al (2009) Human Ccr4-Not complexes contain variable deadenylase subunits. Biochem J 422(3):443–453
Temme C, Simonelig M, Wahle E (2014) Deadenylation of mRNA by the CCR4-NOT complex in Drosophila: molecular and developmental aspects. Front Genet 5:143
Yamashita A et al (2005) Concerted action of poly(A) nucleases and decapping enzyme in mammalian mRNA turnover. Nat Struct Mol Biol 12(12):1054–1063
Coller J, Parker R (2004) Eukaryotic mRNA decapping. Annu Rev Biochem 73:861–890
Franks TM, Lykke-Andersen J (2008) The control of mRNA decapping and P-body formation. Mol Cell 32(5):605–615
Li Y, Kiledjian M (2010) Regulation of mRNA decapping. Wiley Interdiscip Rev RNA 1(2):253–265
Sharif H, Conti E (2013) Architecture of the Lsm1-7-Pat1 complex: a conserved assembly in eukaryotic mRNA turnover. Cell Rep 5(2):283–291
Tharun S (2009) Lsm1-7-Pat1 complex: a link between 3′ and 5′-ends in mRNA decay? RNA Biol 6(3):228–232
Braun JE et al (2012) A direct interaction between DCP1 and XRN1 couples mRNA decapping to 5′ exonucleolytic degradation. Nat Struct Mol Biol 19(12):1324–1331
Nissan T et al (2010) Decapping activators in Saccharomyces cerevisiae act by multiple mechanisms. Mol Cell 39(5):773–783
Halbach F et al (2013) The yeast ski complex: crystal structure and RNA channeling to the exosome complex. Cell 154(4):814–826
Chen N et al (2005) Crystal structures of human DcpS in ligand-free and m7GDP-bound forms suggest a dynamic mechanism for scavenger mRNA decapping. J Mol Biol 347(4):707–718
Liu H et al (2002) The scavenger mRNA decapping enzyme DcpS is a member of the HIT family of pyrophosphatases. EMBO J 21(17):4699–4708
Milac AL, Bojarska E, Wypijewska del Nogal A (2014) Decapping Scavenger (DcpS) enzyme: advances in its structure, activity and roles in the cap-dependent mRNA metabolism. Biochim Biophys Acta 1839(6):452–462
Shen V et al (2008) DcpS scavenger decapping enzyme can modulate pre-mRNA splicing. RNA 14(6):1132–1142
Bonneau F et al (2009) The yeast exosome functions as a macromolecular cage to channel RNA substrates for degradation. Cell 139(3):547–559
Lebreton A et al (2008) Endonucleolytic RNA cleavage by a eukaryotic exosome. Nature 456(7224):993–996
Mamolen M, Smith A, Andrulis ED (2010) Drosophila melanogaster Dis3 N-terminal domains are required for ribonuclease activities, nuclear localization and exosome interactions. Nucleic Acids Res 38(16):5507–5517
Schaeffer D et al (2009) The exosome contains domains with specific endoribonuclease, exoribonuclease and cytoplasmic mRNA decay activities. Nat Struct Mol Biol 16(1):56–62
Schneider C et al (2009) The N-terminal PIN domain of the exosome subunit Rrp44 harbors endonuclease activity and tethers Rrp44 to the yeast core exosome. Nucleic Acids Res 37(4):1127–1140
Lubas M et al (2013) Exonuclease hDIS3L2 specifies an exosome-independent 3′-5′ degradation pathway of human cytoplasmic mRNA. EMBO J 32(13):1855–1868
Malecki M et al (2013) The exoribonuclease Dis3L2 defines a novel eukaryotic RNA degradation pathway. EMBO J 32(13):1842–1854
Arribas-Layton M et al (2013) Structural and functional control of the eukaryotic mRNA decapping machinery. Biochim Biophys Acta 1829(6–7):580–589
Chang H et al (2014) TAIL-seq: genome-wide determination of poly(A) tail length and 3′ end modifications. Mol Cell 53(6):1044–1052
Arraiano CM et al (2010) The critical role of RNA processing and degradation in the control of gene expression. FEMS Microbiol Rev 34(5):883–923
Barbas A et al (2008) New insights into the mechanism of RNA degradation by ribonuclease II: identification of the residue responsible for setting the RNase II end product. J Biol Chem 283(19):13070–13076
Barbas A et al (2009) Determination of key residues for catalysis and RNA cleavage specificity: one mutation turns RNase II into a “SUPER-ENZYME”. J Biol Chem 284(31):20486–20498
Frazao C et al (2006) Unravelling the dynamics of RNA degradation by ribonuclease II and its RNA-bound complex. Nature 443(7107):110–114
Robinson SR et al (2018) DIS3 isoforms vary in their endoribonuclease activity and are differentially expressed within haematological cancers. Biochem J 475(12):2091–2105
Viegas SC et al (2015) Surprises in the 3′-end: ‘U’ can decide too! FEBS J 282(18):3489–3499
Matos RG et al (2014) The importance of proteins of the RNase II/RNB-family in pathogenic bacteria. Front Cell Infect Microbiol 4:68
Matos RG et al (2012) The rnb gene of Synechocystis PCC6803 encodes a RNA hydrolase displaying RNase II and not RNase R enzymatic properties. PLoS One 7(3):e32690
Matos RG et al (2011) Swapping the domains of exoribonucleases RNase II and RNase R: conferring upon RNase II the ability to degrade ds RNA. Proteins 79(6):1853–1867
Astuti D et al (2012) Germline mutations in DIS3L2 cause the Perlman syndrome of overgrowth and Wilms tumor susceptibility. Nat Genet 44(3):277–284
Chapman MA et al (2011) Initial genome sequencing and analysis of multiple myeloma. Nature 471(7339):467–472
Ding L et al (2012) Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 481(7382):506–510
Reis FP et al (2013) The RNase II/RNB family of exoribonucleases: putting the ‘Dis’ in disease. Wiley Interdiscip Rev RNA 4(5):607–615
Dziembowski A et al (2007) A single subunit, Dis3, is essentially responsible for yeast exosome core activity. Nat Struct Mol Biol 14(1):15–22
Wasmuth EV, Lima CD (2012) Exo- and endoribonucleolytic activities of yeast cytoplasmic and nuclear RNA exosomes are dependent on the noncatalytic core and central channel. Mol Cell 48(1):133–144
Liu Q, Greimann JC, Lima CD (2006) Reconstitution, activities, and structure of the eukaryotic RNA exosome. Cell 127(6):1223–1237
Makino DL, Baumgartner M, Conti E (2013) Crystal structure of an RNA-bound 11-subunit eukaryotic exosome complex. Nature 495(7439):70–75
Wasmuth EV, Januszyk K, Lima CD (2014) Structure of an Rrp6-RNA exosome complex bound to poly(A) RNA. Nature 511(7510):435–439
Allmang C et al (1999) Functions of the exosome in rRNA, snoRNA and snRNA synthesis. EMBO J 18(19):5399–5410
Bousquet-Antonelli C, Presutti C, Tollervey D (2000) Identification of a regulated pathway for nuclear pre-mRNA turnover. Cell 102(6):765–775
Chen CY et al (2001) AU binding proteins recruit the exosome to degrade ARE-containing mRNAs. Cell 107(4):451–464
Milligan L et al (2005) A nuclear surveillance pathway for mRNAs with defective polyadenylation. Mol Cell Biol 25(22):9996–10004
Mitchell P et al (1997) The exosome: a conserved eukaryotic RNA processing complex containing multiple 3′-->5′ exoribonucleases. Cell 91(4):457–466
Mukherjee D et al (2002) The mammalian exosome mediates the efficient degradation of mRNAs that contain AU-rich elements. EMBO J 21(1–2):165–174
Tomecki R et al (2010) The human core exosome interacts with differentially localized processive RNases: hDIS3 and hDIS3L. EMBO J 29(14):2342–2357
Schaeffer D et al (2012) The CR3 motif of Rrp44p is important for interaction with the core exosome and exosome function. Nucleic Acids Res 40(18):9298–9307
Schneider C, Anderson JT, Tollervey D (2007) The exosome subunit Rrp44 plays a direct role in RNA substrate recognition. Mol Cell 27(2):324–331
Wang HW et al (2007) Architecture of the yeast Rrp44 exosome complex suggests routes of RNA recruitment for 3′ end processing. Proc Natl Acad Sci U S A 104(43):16844–16849
Lorentzen E et al (2008) Structure of the active subunit of the yeast exosome core, Rrp44: diverse modes of substrate recruitment in the RNase II nuclease family. Mol Cell 29(6):717–728
Amblar M et al (2006) Characterization of the functional domains of Escherichia coli RNase II. J Mol Biol 360(5):921–933
Amblar M et al (2007) The role of the S1 domain in exoribonucleolytic activity: substrate specificity and multimerization. RNA 13(3):317–327
Lee G et al (2012) Elastic coupling between RNA degradation and unwinding by an exoribonuclease. Science 336(6089):1726–1729
Matelska D, Steczkiewicz K, Ginalski K (2017) Comprehensive classification of the PIN domain-like superfamily. Nucleic Acids Res 45(12):6995–7020
Senissar M, Manav MC, Brodersen DE (2017) Structural conservation of the PIN domain active site across all domains of life. Protein Sci 26(8):1474–1492
Drazkowska K et al (2013) The RNA exosome complex central channel controls both exonuclease and endonuclease Dis3 activities in vivo and in vitro. Nucleic Acids Res 41(6):3845–3858
Schneider C et al (2012) Transcriptome-wide analysis of exosome targets. Mol Cell 48(3):422–433
Malet H et al (2010) RNA channelling by the eukaryotic exosome. EMBO Rep 11(12):936–942
Tomecki R et al (2014) Multiple myeloma-associated hDIS3 mutations cause perturbations in cellular RNA metabolism and suggest hDIS3 PIN domain as a potential drug target. Nucleic Acids Res 42(2):1270–1290
Hou D, Ruiz M, Andrulis ED (2012) The ribonuclease Dis3 is an essential regulator of the developmental transcriptome. BMC Genomics 13:359
Szczepinska T et al (2015) DIS3 shapes the RNA polymerase II transcriptome in humans by degrading a variety of unwanted transcripts. Genome Res 25(11):1622–1633
Morton DJ et al (2018) The RNA exosome and RNA exosome-linked disease. RNA 24(2):127–142
Robinson SR et al (2015) The 3′ to 5′ exoribonuclease DIS3: from structure and mechanisms to biological functions and role in human disease. Biomol Ther 5(3):1515–1539
Lionetti M et al (2015) A compendium of DIS3 mutations and associated transcriptional signatures in plasma cell dyscrasias. Oncotarget 6(28):26129–26141
Lohr JG et al (2014) Widespread genetic heterogeneity in multiple myeloma: implications for targeted therapy. Cancer Cell 25(1):91–101
Walker BA et al (2012) Intraclonal heterogeneity and distinct molecular mechanisms characterize the development of t(4;14) and t(11;14) myeloma. Blood 120(5):1077–1086
Weissbach S et al (2015) The molecular spectrum and clinical impact of DIS3 mutations in multiple myeloma. Br J Haematol 169(1):57–70
Laubach J, Richardson P, Anderson K (2011) Multiple myeloma. Annu Rev Med 62:249–264
Preker P et al (2008) RNA exosome depletion reveals transcription upstream of active human promoters. Science 322(5909):1851–1854
Wang Y et al (2015) A novel long non-coding RNA, hypoxia-inducible factor-2alpha promoter upstream transcript, functions as an inhibitor of osteosarcoma stem cells in vitro. Mol Med Rep 11(4):2534–2540
Ng D et al (2007) Identification of a novel chromosome region, 13q21.33-q22.2, for susceptibility genes in familial chronic lymphocytic leukemia. Blood 109(3):916–925
Pils D et al (2013) A combined blood based gene expression and plasma protein abundance signature for diagnosis of epithelial ovarian cancer--a study of the OVCAD consortium. BMC Cancer 13:178
Lim J et al (1997) Isolation of murine and human homologues of the fission-yeast dis3+ gene encoding a mitotic-control protein and its overexpression in cancer cells with progressive phenotype. Cancer Res 57(5):921–925
Kinoshita N, Goebl M, Yanagida M (1991) The fission yeast dis3+ gene encodes a 110-kDa essential protein implicated in mitotic control. Mol Cell Biol 11(12):5839–5847
Ohkura H et al (1988) Cold-sensitive and caffeine-supersensitive mutants of the Schizosaccharomyces pombe dis genes implicated in sister chromatid separation during mitosis. EMBO J 7(5):1465–1473
Towler BP et al (2015) The 3′-5′ exoribonuclease Dis3 regulates the expression of specific microRNAs in Drosophila wing imaginal discs. RNA Biol 12(7):728–741
Camps J et al (2013) Genetic amplification of the NOTCH modulator LNX2 upregulates the WNT/beta-catenin pathway in colorectal cancer. Cancer Res 73(6):2003–2013
de Groen FL et al (2014) Gene-dosage dependent overexpression at the 13q amplicon identifies DIS3 as candidate oncogene in colorectal cancer progression. Genes Chromosom Cancer 53(4):339–348
Rose AE et al (2011) Integrative genomics identifies molecular alterations that challenge the linear model of melanoma progression. Cancer Res 71(7):2561–2571
Liu W, Peng Y, Tobin DJ (2013) A new 12-gene diagnostic biomarker signature of melanoma revealed by integrated microarray analysis. PeerJ 1:e49
Rozenblum E et al (2002) A genomic map of a 6-Mb region at 13q21-q22 implicated in cancer development: identification and characterization of candidate genes. Hum Genet 110(2):111–121
Hoskins JW et al (2016) Functional characterization of a chr13q22.1 pancreatic cancer risk locus reveals long-range interaction and allele-specific effects on DIS3 expression. Hum Mol Genet 25(21):4726–4738
Shen L, Shi Q, Wang W (2018) Double agents: genes with both oncogenic and tumor-suppressor functions. Oncogene 7(3):25
Segalla S et al (2015) The ribonuclease DIS3 promotes let-7 miRNA maturation by degrading the pluripotency factor LIN28B mRNA. Nucleic Acids Res 43(10):5182–5193
Noguchi E et al (1996) Dis3, implicated in mitotic control, binds directly to ran and enhances the GEF activity of RCC1. EMBO J 15(20):5595–5605
Glavan F et al (2006) Structures of the PIN domains of SMG6 and SMG5 reveal a nuclease within the mRNA surveillance complex. EMBO J 25(21):5117–5125
Bleichert F et al (2006) The PINc domain protein Utp24, a putative nuclease, is required for the early cleavage steps in 18S rRNA maturation. Proc Natl Acad Sci U S A 103(25):9464–9469
Staals RH et al (2010) Dis3-like 1: a novel exoribonuclease associated with the human exosome. EMBO J 29(14):2358–2367
Slomovic S et al (2010) Addition of poly(A) and poly(A)-rich tails during RNA degradation in the cytoplasm of human cells. Proc Natl Acad Sci U S A 107(16):7407–7412
Slomovic S et al (2006) Polyadenylation of ribosomal RNA in human cells. Nucleic Acids Res 34(10):2966–2975
Lima WF et al (2016) RNA cleavage products generated by antisense oligonucleotides and siRNAs are processed by the RNA surveillance machinery. Nucleic Acids Res 44(7):3351–3363
Kohn M et al (2015) The Y3∗∗ ncRNA promotes the 3′ end processing of histone mRNAs. Genes Dev 29(19):1998–2003
Kowalski MP, Krude T (2015) Functional roles of non-coding Y RNAs. Int J Biochem Cell Biol 66:20–29
Savage SA (1993) Dyskeratosis congenita. In: Adam MP et al (eds) GeneReviews((R)). University of Washington, Seattle
Shukla S, Parker R (2017) PARN modulates Y RNA stability and its 3′-end formation. Mol Cell Biol 37(20):e00264
Shukla S et al (2019) The RNase PARN controls the levels of specific miRNAs that contribute to p53 regulation. Mol Cell 73:1204
Palmer CJ et al (2014) Zfx facilitates tumorigenesis caused by activation of the hedgehog pathway. Cancer Res 74(20):5914–5924
Lee JY et al (2017) Genome-based exome sequencing analysis identifies GYG1, DIS3L and DDRGK1 are associated with myocardial infarction in Koreans. J Genet 96(6):1041–1046
Oates CP et al (2018) Novel polymorphisms associated with hyperalphalipoproteinemia and apparent cardioprotection. J Clin Lipidol 12(1):110–115
Labno A et al (2016) Perlman syndrome nuclease DIS3L2 controls cytoplasmic non-coding RNAs and provides surveillance pathway for maturing snRNAs. Nucleic Acids Res 44(21):10437–10453
Pirouz M et al (2016) Dis3l2-mediated decay is a quality control pathway for noncoding RNAs. Cell Rep 16(7):1861–1873
Ustianenko D et al (2013) Mammalian DIS3L2 exoribonuclease targets the uridylated precursors of let-7 miRNAs. RNA 19(12):1632–1638
Ustianenko D et al (2016) TUT-DIS3L2 is a mammalian surveillance pathway for aberrant structured non-coding RNAs. EMBO J 35(20):2179–2191
Lim J et al (2014) Uridylation by TUT4 and TUT7 marks mRNA for degradation. Cell 159(6):1365–1376
Chang HM et al (2013) A role for the Perlman syndrome exonuclease Dis3l2 in the Lin28-let-7 pathway. Nature 497(7448):244–248
Faehnle CR, Walleshauser J, Joshua-Tor L (2014) Mechanism of Dis3l2 substrate recognition in the Lin28-let-7 pathway. Nature 514(7521):252–256
Rissland OS, Mikulasova A, Norbury CJ (2007) Efficient RNA polyuridylation by noncanonical poly(A) polymerases. Mol Cell Biol 27(10):3612–3624
Mullen TE, Marzluff WF (2008) Degradation of histone mRNA requires oligouridylation followed by decapping and simultaneous degradation of the mRNA both 5′ to 3′ and 3′ to 5′. Genes Dev 22(1):50–65
Song MG, Kiledjian M (2007) 3′ terminal oligo U-tract-mediated stimulation of decapping. RNA 13(12):2356–2365
Tomecki R et al (2004) Identification of a novel human nuclear-encoded mitochondrial poly(A) polymerase. Nucleic Acids Res 32(20):6001–6014
Shimizu S et al (2014) Autophagic cell death and cancer. Int J Mol Sci 15(2):3145–3153
Thomas MP et al (2015) Apoptosis triggers specific, rapid, and global mRNA decay with 3′ uridylated intermediates degraded by DIS3L2. Cell Rep 11(7):1079–1089
Liu X et al (2018) PNPT1 release from mitochondria during apoptosis triggers decay of Poly(A) RNAs. Cell 174(1):187–201.e12
Briani F, Carzaniga T, Deho G (2016) Regulation and functions of bacterial PNPase. Wiley Interdiscip Rev RNA 7(2):241–258
Bussing I, Slack FJ, Grosshans H (2008) let-7 microRNAs in development, stem cells and cancer. Trends Mol Med 14(9):400–409
Thornton JE, Gregory RI (2012) How does Lin28 let-7 control development and disease? Trends Cell Biol 22(9):474–482
Kurosaki T et al (2018) NMD-degradome sequencing reveals ribosome-bound intermediates with 3′-end non-templated nucleotides. Nat Struct Mol Biol 25(10):940–950
Schilke K et al (2000) A case of Perlman syndrome: fetal gigantism, renal dysplasia, and severe neurological deficits. Am J Med Genet 91(1):29–33
Hunter RW et al (2018) Loss of Dis3l2 partially phenocopies Perlman syndrome in mice and results in up-regulation of Igf2 in nephron progenitor cells. Genes Dev 32(13–14):903–908
Morris MR, Astuti D, Maher ER (2013) Perlman syndrome: overgrowth, Wilms tumor predisposition and DIS3L2. Am J Med Genet C: Semin Med Genet 163C(2):106–113
Tassano E et al (2013) Genotype-phenotype correlation of 2q37 deletions including NPPC gene associated with skeletal malformations. PLoS One 8(6):e66048
von Kodolitsch Y et al (2010) Marfan syndrome and the evolving spectrum of heritable thoracic aortic disease: do we need genetics for clinical decisions? Vasa 39(1):17–32
Bocciardi R et al (2007) Overexpression of the C-type natriuretic peptide (CNP) is associated with overgrowth and bone anomalies in an individual with balanced t(2;7) translocation. Hum Mutat 28(7):724–731
Moncla A et al (2007) A cluster of translocation breakpoints in 2q37 is associated with overexpression of NPPC in patients with a similar overgrowth phenotype. Hum Mutat 28(12):1183–1188
Acknowledgements
This work was supported by project LISBOA-01-0145-FEDER-007660 (Microbiologia Molecular, Estrutural e Celular) funded by FEDER funds through COMPETE2020 – Programa Operacional Competitividade e Internacionalização (POCI) and by national funds from FCT (Fundação para a Ciência e a Tecnologia); project PTDC/BIA-MIC/1399/2014 to CMA and project PTDC/BIM-MEC/3749/2014 to SCV. In addition, FCT provides postdoctoral grant ref. SFRH/BPD/109464/2015 to MS. SCV was financed by program IF of FCT (ref IF/00217/2015).
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Saramago, M., da Costa, P.J., Viegas, S.C., Arraiano, C.M. (2019). The Implication of mRNA Degradation Disorders on Human DISease: Focus on DIS3 and DIS3-Like Enzymes. In: Romão, L. (eds) The mRNA Metabolism in Human Disease. Advances in Experimental Medicine and Biology, vol 1157. Springer, Cham. https://doi.org/10.1007/978-3-030-19966-1_4
Download citation
DOI: https://doi.org/10.1007/978-3-030-19966-1_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-19965-4
Online ISBN: 978-3-030-19966-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)