Abstract
Ryanodine receptor calcium release channels (RyRs) play central roles in controlling intracellular calcium concentrations in excitable and non-excitable cells. RyRs are located in the sarcoplasmic or endoplasmic reticulum, intracellular Ca2+ storage compartment, and release Ca2+ during cellular action potentials or in response to other cellular stimuli. Mammalian cells express three structurally related isoforms of RyR. RyR1 and RyR2 are the major RyR isoforms in skeletal and cardiac muscle, respectively, and RyR3 is expressed in various tissues along with the other two isoforms. A prominent feature of RyRs is that the Ca2+ release channel activities of RyRs are regulated by calcium ions; therefore, intracellular Ca2+ release controls positive- and negative-feedback phenomena through the RyRs. RyR channel activities are also regulated by Ca2+ indirectly, i.e. through Ca2+ binding proteins at both cytosolic and sarco/endoplasmic reticulum luminal sides. Here, I summarize Ca2+-dependent feedback regulation of RyRs including recent progress in the structure/function aspects.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Ryanodine receptor
- Excitation-contraction coupling
- Calcium release channel
- Intracellular calcium
- Calmodulin
Transient increase of intracellular Ca2+ concentration plays a pivotal role in numerous cell functions, including muscle contraction, neuronal plasticity, and immune responses. Multiple sources of Ca2+ are involved in this signaling, including Ca2+ influx from the extracellular spaces and Ca2+ release from intracellular Ca2+ stores: the endo/sarcoplasmic reticulum (ER/SR), nuclear envelope, and mitochondria [1,2,3]. Ryanodine receptors (RyRs) are Ca2+ release channels located in the ER/SR and play a primary role in Ca2+ release from SR during skeletal and cardiac muscle contraction [4,5,6]. Additionally, RyRs play an important role in smooth muscle, neurons, and other cell types by co-existing with another family of Ca2+ release channels called inositol-trisphosphate receptors [3]. RyRs release Ca2+ from the ER/SR and are regulated by Ca2+ [4,5,6], suggesting that RyRs have a self-regulatory mechanism controlled by Ca2+, i.e. RyR-released Ca2+ regulates the same or neighboring RyRs. This chapter focuses on the structure-function aspect of RyR regulation by Ca2+ and Ca2+ binding proteins such as calmodulin and calsequestrin (Fig. 13.1).
A schematic of the RyR Ca 2+ release channel regulation by Ca 2+ . RyRs have been suggested to possess at least three Ca2+ regulatory sites; cytoplasmic high affinity activation (A) and low affinity inhibition (I) sites, and luminal regulatory (L) site. Mg2+ inhibits RyRs by binding to A or I sites. Calcium ions passing through the RyR from ER/SR lumen to the cytoplasm are considered to bind to RyR-A and I sites and cytoplasmic accessory proteins, CaM and S100A1. The low affinity luminal Ca2+ binding protein, CSQ, forms a macromolecular complex via its interaction with RyR accessory proteins, triadin (TRD) and junctin (JTN). CSQ may also bind to RyR directly
13.1 Molecular Structure of RyRs
In striated muscle, RyRs are localized in the junctional SR membrane in close proximity to transverse (T)-tubule membranes, invaginations of the plasma membrane into the myofibrils. In skeletal muscle the SR is typically on both sides of the T-tubule (triad), while in cardiomyocytes, the SR is only on one side of the T-tubule (dyad). In both triadic and dyadic junctions, electron microscopy shows foot structures spanning between the SR and T-tubule [4]. Molecular identification of RyRs was first performed with rabbit fast twitch skeletal muscle using ryanodine, a specific ligand of RyRs. Isolated RyRs are homotetramers of a ∼500 kDa polypeptide. Morphological analysis of the reconstituted purified proteins identified RyRs as the foot structures [7,8,9,10]. Molecular cloning of RyRs showed that mammals express three different RyR isoforms [11,12,13,14,15,16]. Skeletal muscle expresses primarily RyR1. The dominant RyR isoform in cardiac muscle is RyR2. RyR3 was initially identified in the brain; however, the brain expresses all three RyR isoforms. Although expression patterns depend on the locations in the brain, in general, RyR2 is widely dispersed over the whole brain [17]. RyR3 is also expressed together with RyR1 in the diaphragm and slow twitch muscle [18, 19], thus functional characterization of RyR1 is mainly performed with fast twitch muscle. In amphibians and avian skeletal muscle, two RyR isoforms, αRyR and βRyR, are recognized and correspond to the mammalian RyR1 and RyR3, respectively [20,21,22].
All three isoforms of RyR have a large cytoplasmic domain, which possesses multiple regulatory sites for channel activity. The carboxyl-terminal end of the RyR spans the SR membrane six times, in which a pore helix and the transmembrane segment form the channel pore [23,24,25,26,27]. RyRs are activated by micromolar Ca2+ and adenine nucleotide, and are inhibited by millimolar Ca2+ and Mg2+ [4,5,6]. A number of proteins have been found to interact with RyRs and regulate their channel activity. These include triadin, junctin, FK506-binding proteins, protein kinases and phosphatases, and Ca2+ binding proteins such as calmodulin and S100A1. Recently, cryo-electron microscopy and 3D image reconstruction of the purified full-length RyRs and crystal structural analysis of truncated recombinant RyRs have detailed the structures of RyR1 and RyR2 at near atomic resolution [24,25,26,27,28,29,30].
13.2 Activation by Cytoplasmic Ca2+
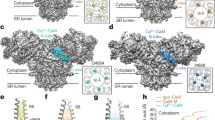
Skeletal and cardiac muscle contractions are triggered by SR Ca2+ release mediated by RyR1 and RyR2. Two different mechanisms are now recognized to open RyRs. In skeletal muscle, direct interaction between RyR1 and the T-tubule voltage sensor, also recognized as the DHP receptor L-type Ca2+ channel (Cav1.1), opens RyR1 during skeletal action potential (voltage-induced Ca2+ release) [31]. Alternatively, in cardiac muscle, small Ca2+ influx through the cardiac L-type Ca2+ channel (Cav1.2) increases intracellular Ca2+, and at ∼micromolar concentrations opens RyR2 by means of Ca2+-induced Ca2+ release (CICR) [32]. The CICR mechanism was initially recognized in skeletal muscle contraction [33, 34]; however, elimination of Ca2+ from the extracellular space or blocking Ca2+ influx through Cav1.1 did not abolish voltage-dependent intracellular Ca2+ transients [35, 36]. Thus, CICR in skeletal muscle (RyR1) is not considered a trigger for muscle contraction. Furthermore, slower kinetics of CICR in contrast to the rapid Ca2+ release in skeletal muscle also supported the idea that CICR is not a physiological trigger for skeletal muscle contraction [37]. However, CICR may play a role in amplifying Ca2+ signaling by activating RyR1s which do not couple with DHP receptors or the small population of RyR3s [38]. Calcium-dependent activation of RyR1 can be altered by RyR1 missense mutations associated with skeletal myopathies such as malignant hyperthermia, thus, CICR may impact these pathologies [37, 39]. Ca2+-dependent activation of RyRs has been well characterized using isolated membrane fractions, intact cells and muscle fibers, purified RyR proteins, and recombinant RyRs by several different methods including muscle tension measurements, Ca2+ flux measurements using Ca2+ indicator dyes or radioactive 45Ca2+, single channel recordings, and specific ligand ([3H]ryanodine) binding assays [37, 40, 41]. All three mammalian RyR isoforms are activated by ∼0.5–5 μM Ca2+ depending on assay conditions. Several potential Ca2+ binding sites were initially identified using truncated RyR1 proteins and 45Ca2+ overlays [42,43,44]. Subsequently, site-directed mutagenesis showed that E3987 in RyR2 (E4032 in RyR1) was critical for Ca2+-dependent activation of RyRs [45, 46]. The mutant RyRs showed impaired Ca2+ dependent activation in single channel recordings and [3H]ryanodine binding assay. E4032A-RyR1 expressing myotubes were impaired in caffeine-induced Ca2+ release, but the aberrant function was restored in the presence of ryanodine [47]. Recently, near-atomic level cryo-electron microscopy analysis of RyR1 (∼4 Å resolution) determined the open and closed state conformations of RyR1 [30]. The structure of RyR1 with 30 μM Ca2+, which is optimal for RyR activation, identified a new Ca2+ binding site in RyR1. The Ca2+ binding site is formed by 3 essential amino acids, E3893, E3967, and T5001, together with two auxiliary amino acids, H3895 and Q3970, for secondary coordination of the Ca2+ sphere [30]. In this structural model, E4032 is distal from the bound calcium ion, but forms an interface with carboxyl terminal tail where T5001 locates. This suggests that E4032 contribute to stabilize the conformation of Ca2+ bound RyR1 [30]. Murayama and colleagues introduced point mutations on RyR2 amino acids corresponding to the RyR1 E3893, E3967, and Q3970, and found that the mutations altered Ca2+-dependent activation of RyR2 [48]. These functional results support the idea that the identified Ca2+ binding site serves as a functional Ca2+ regulatory site. Further detail analysis including assessments of other amino acids combined in the presence of other channel agonists and antagonists will further advance structure and function relationship of Ca2+-dependent activation of RyRs. Another ∼6 Å resolution cryo-electron micrograph of RyR1 suggested that 10 mM Ca2+ changed the conformation of the EF hand-type Ca2+ binding domain of RyR1; therefore, it was proposed as a Ca2+ activation site [24]. However, studies with recombinant proteins including the EF hand domain showed Ca2+ affinity was >60 μM [49, 50], which is much higher than the RyR-activating Ca2+ concentration. Also, functional study scrambling of the EF hand sequence in RyR1 and deletion of the entire EF hand in RyR2 did not affect the Ca2+ activation of RyRs [51, 52]. Considering that the structural analysis was determined with 10 mM Ca2+, the EF hand site is likely to be a Ca2+ inactivation site [53]. We also found that the EF hand domain contributes to the isoform-specific regulation of RyRs by calmodulin (see below) [54].
Ca2+-dependent activation of RyR1 and RyR2 are similar in single channel recordings and flux measurements in the SR vesicles; however, Ogawa and colleagues pointed out that RyR2 in rat ventricular SR or as a recombinant form exhibited a suppressed activity at 10–100 μM Ca2+ using [3H]ryanodine equilibrium binding assay [55]. Similar suppressed RyR2 activities were observed in our own study with rabbit recombinant RyR2 using the same technique [56]. Surprisingly, this suppressed activity was restored by ∼1 mM Mg2+ [55], which is usually considered to be an inhibitor of RyR channel activity by competing off Ca2+ at the Ca2+ activation site or binding to the Ca2+ inhibitory site [40, 57]. RyR2 in the rabbit ventricular SR showed this suppression only when AMP or caffeine, RyR activators, were added, suggesting that the suppressed effects depend on the type of RyR2 sample. One possibility for this mechanism is therefore that regulatory factors were removed during the sample preparations. Another possible explanation is that the RyR2 conformation is not very stable under long time (>8 h) equilibrium conditions in the [3H]ryanodine binding assays. We found that replacement of the RyR1-EF hand domain with corresponding RyR2 sequence or the introduction of point mutations in the cytoplasmic loop between the second and the third transmembrane segments (S2-S3 loop) of RyR1 resulted in suppressed activity at 10–100 μM Ca2+ [53, 58]. The results suggested that the EF hand and S2-S3 cytoplasmic loop of RyRs are involved in the conformational stability and Ca2+-dependent regulation (activation/inhibition) of RyR channels.
13.3 Inhibition by Cytoplasmic Ca2+
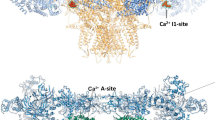
While RyRs are activated by micromolar cytosolic Ca2+, higher concentrations of Ca2+ (>1 mM) inhibit RyR channel activities. Thus, RyRs have a high affinity Ca2+ activation site and a low affinity Ca2+ inactivation site (A and I sites, respectively in Fig. 13.1). These sites are also implicated in Mg2+ inhibition, namely submillimolar Mg2+ competes with activating Ca2+ at A site and millimolar Mg2+ binds to the I site for inhibitory effect [57, 59]. Although the physiological significance of RyR inactivation by millimolar levels of Ca2+ has been questioned, local rise of cytosolic Ca2+ around the RyRs may be sufficient to inhibit RyR channel activity. Single channel recording showed that Ca2+ flux from the lumen to the cytosolic side resulted in a decrease of open probability of both the RyR1 and RyR2 channel, supporting Ca2+-dependent inactivation of RyRs by the released Ca2+ in intact tissues [60, 61]. All three mammalian isoforms of RyR are inhibited by high concentrations of Ca2+; however, affinity for inhibitory Ca2+ in RyR1 is 5–10 times higher than those in RyR2 and RyR3 [6, 53, 62]. Deletion of 52 amino acids including a large cluster (42 amino acids) of negatively charged amino acids in RyR1 resulted in a threefold decrease in Ca2+ inactivation affinity [63]; yet, this change in the local electrostatics property may have caused a large conformational change. Construction and characterization of RyR1/RyR2 chimeras highlighted differences of Ca2+-inactivation affinity between the two RyR isoforms. Chimeric RyRs showed that RyR isoform specific Ca2+ inactivation depends on the sequence of the carboxyl-terminal quarter (∼1000 amino acids) [62, 64, 65]. Further characterization suggested that the second transmembrane segment (S2) and EF hand type Ca2+ binding motifs are involved in the isoform-specific Ca2+-dependent inactivation of RyRs [53]. In agreement with these observations, scrambling one EF hand sequence (EF1) in RyR1 resulted in a twofold reduction in the affinity of Ca2+-dependent inhibition [51]. In near-atomic level cryo-electron microscopy, the EF hand domain and S2-S3 cytoplasmic loop are in close proximity [25]. In another structural model, 10 mM Ca2+ changed the conformation of the EF hand domain [24]. Site-directed mutagenesis of the S2-S3 loop of RyR1 impaired the affinity for Ca2+-dependent inactivation, and resulted in RyR2-type Ca2+-dependent activity profiles [58]. Considering the Ca2+ affinity of the recombinant EF hand domain (60 μM-4 mM) [49, 50], the Ca2+ inactivation site of the RyR is the EF hand motif. One possible mechanism is that the S2-S3 loop transduces the signal of Ca2+ binding to the EF hand domain to the channel pore region including S2 [58]. It should be noted that a point mutation in G4733 of RyR1, which is in close proximity to the EF hand domain (Fig. 13.2), significantly suppressed Ca2+-dependent inactivation [58].
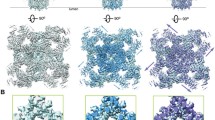
High resolution cryo-electron microscopy structure of RyR1. The closed state of RyR1 (Protein Data Bank Accession 5TB0 [30]) is presented by UCSF Chimera program (https://www.cgl.ucsf.edu/chimera/) [121]. (Left panel) Structure of tetrameric RyR1. TM denotes transmembrane region. (Right panel) Enlargement of region marked with red circle in left panel. The EF hand domain (red) is shown to be adjacent to the S2-S3 loop (blue) in the neighboring subunit [25]. In this structure, Gly4733 and neighboring amino acids are located in close proximity to the EF hand domain, and point mutations on these amino acid residues altered Ca2+-dependent inactivation of RyR1 [58]. Thus, the S2-S3 loop of RyR may transduce its Ca2+-dependent inhibitory signal through the EF hand domain. Note that the S2 transmembrane (green) has also been shown to be involved in Ca2+-dependent inactivation of RyRs [53]
13.4 Regulation by Luminal Ca2+
RyRs could also be regulated by SR luminal Ca2+, as during Ca2+ release the junctional SR Ca2+ concentration drastically drops. This suggests that RyR channel gating can be regulated directly by luminal Ca2+; e.g. SR Ca2+ filling status regulates RyR channel opening and closing. It is known that the SR Ca2+ store with a certain level of Ca2+ exhibits spontaneous Ca2+ release in mammalian cardiac muscle cells [66, 67]. Chen and colleagues found that the store overload-induced Ca2+ release (SOICR) was observed in heterologous cells expressing recombinant RyR channels; therefore, it is likely an intrinsic property of RyRs. [68]. SOICR mechanisms were implicated in the aberrant Ca2+ signaling found in RyR mutation-related skeletal and cardiac muscle diseases [68,69,70]. The muscular disease-associated RyR mutations reduce the threshold for SOICR; therefore, spontaneous Ca2+ release (Ca2+ spills) occurs when the SR Ca2+ store loading is increased by the triggers of pathologies such as catecholamine release. The luminal Ca2+-sensing gate of RyRs was investigated by site-directed mutagenesis, revealing that E4872 on the inner pore helix (S6 transmembrane segment) of RyR2 is essential for luminal Ca2+ activation of RyR2 and SOICR [71]. Knock-in mice harboring the E4872Q-RyR2 mutation were resistant to Ca2+-dependent ventricular tachycardia, suggesting that SOICR is a critical mechanism for arrhythmogenesis [71].
It also should be noted that luminal Ca2+ can also access cytosolic Ca2+ activation and inactivation sites [60, 61] (Fig. 13.1). In single channel measurements of RyR1 and RyR2, luminal Ca2+ passed through RyRs to the cytosolic side in conjunction with potassium ions under a voltage gradient, and activated and inhibited the same RyR channels depending on luminal Ca2+ concentration [60, 61], which suggests that during excitation-contraction coupling, local cytoplasmic Ca2+ concentrations can reach millimolar levels and are sufficient for Ca2+-dependent inactivation of RyRs.
13.5 Regulation by Calmodulin and S100A1
Calmodulin (CaM) is a 16.7-kDa protein that possesses 2 EF hand-type Ca2+ binding sites on both the amino and carboxyl-terminal. Thus, CaM works as a Ca2+ sensing subunit of multiple ion channels [72]. CaM modifies RyR channel function independently from regulation by Ca2+; therefore, RyRs have “dual” cytosolic Ca2+ dependent regulatory mechanisms (direct and indirect). RyRs are regulated by not only the Ca2+ bound form of CaM, but also by CaM at cellular resting Ca2+ concentrations (∼0.1 μM). Ca2+ bound CaM inhibits all three mammalian isoforms of RyR, while CaM activates RyR1 and RyR3 and inhibits RyR2 at submicromolar Ca2+ concentrations [73,74,75,76,77], suggesting that CaM constitutively binds to RyRs to regulate their channel activities by sensing cytoplasmic Ca2+ concentrations. In vitro experiments also showed that CaM regulation of the RyR depends on redox state. Affinities for CaM regulation of RyR channel activity at the oxidized condition are 2–20 fold lower than at the reduced condition [77, 78]. The results are consistent with observations that CaM is dissociated from RyR2, resulting in a Ca2+ leak from SR in failing hearts [79], in which the redox balance possibly shifts to the more oxidized condition [80, 81].
Purified RyR1 and RyR2 as well as the recombinant RyR3 bind 4 CaM per tetrameric RyR, i.e. one RyR subunit binds one CaM [56, 77, 78, 82]. The CaM binding and regulatory domain was identified by trypsin digestion, binding of synthetic RyR1 peptides, and site directed mutagenesis of RyR1 amino acids 3614–3643 [82,83,84]. This domain was confirmed to be conserved in RyR2 and RyR3 by site-directed mutagenesis [56, 78]. Crystal structure analysis of a synthetic RyR1 peptide (amino acids 3614–3643) and CaM complex revealed that the carboxyl-terminal lobe of CaM binds to the peptide, while the amino-terminal lobe binds with low affinity or is free [85], which may explain that multiple RyR domain peptides or fusion proteins can bind to CaM [44, 86,87,88]. Point mutations in RyR1 3614–3643 or the corresponding RyR2 and RyR3 domains eliminated CaM binding and regulation of channel activities [56, 78, 82, 89]; thus, this conserved domain likely plays a primary role for CaM-dependent regulation. Although the primary CaM regulatory domain is well conserved, RyR isoform-specific CaM regulation at submicromolar (cellular resting level) Ca2+ concentrations, namely activation of RyR1 and RyR3 versus inhibition of RyR2, was investigated using RyR1/RyR2 chimera channels. Replacing the flanking regions of the RyR2 CaM binding domain with the RyR1 sequence abolished CaM regulation of RyR2 at submicromolar Ca2+ concentrations [90]. More recently, the EF hand domain and large N-terminal region were shown to be important for isoform-specific CaM regulation of RyRs [54]. These domains possibly mediate long-range interaction between the CaM binding domain and the functional effects on the channel, as the CaM binding domain is ∼10 nm apart from the RyR channel pore region in cryo-electron micrographs [91].
In vivo significance of CaM regulation of RyR1 and RyR2 was studied with genetically modified mice. Knock-in mice carrying point mutations in the RyR2 CaM regulatory domain (W3587A/L3591D/F3603A: ADA mutations) were impaired in CaM binding and regulation of cardiac RyR2 [89]. The mice showed rapidly developing cardiac hypertrophy and died 2–3 weeks after birth. Cardiomyocytes isolated from the mutant mouse hearts exhibited long durations of the spontaneous Ca2+ transients or Ca2+ sparks, indicating that CaM inhibition of RyR2 contributes to the termination of SR Ca2+ release, which is important for heart physiology and growth [89, 92]. The knock-in ADA mice were impaired in CaM regulation of RyR2 at both diastolic (submicromolar) and systolic (micromolar) Ca2+ concentrations, while knock-in mice with a single mutation (L3591D), were only impaired in CaM regulation of RyR2 during diastole and showed more modest levels of cardiac hypertrophy, suggesting that CaM regulation of RyR2 at systolic Ca2+ levels plays a major role in vivo [93]. The corresponding RyR1 mutation (RyR1-L3624D) attenuated both CaM activation and inhibition at submicromolar and micromolar Ca2+ concentrations [82]. However, knock-in mice carrying RyR1-L3624D showed only modest effects on skeletal muscle excitation-contraction coupling without lethality, suggesting that CaM regulation of RyR1 plays a minor role in skeletal physiology [94]. More recently, missense mutations in calmodulin genes were identified in patients with catecholaminergic polymorphic ventricular tachycardia (CPVT) [95], in which CaM mutations likely alter RyR2 regulation [96,97,98]. Thus, the CaM-RyR2 interaction can be a good therapeutic target for the cardiac pathologies [99].
S100A1, another EF hand type Ca2+ binding protein, is also expressed in skeletal and cardiac muscle and regulates intracellular Ca2+ signaling by interacting with multiple Ca2+ handling proteins including RyRs [100,101,102,103]. Competitive binding experiments showed that S100A1 shares a common binding site on RyR1 with CaM [104, 105]. Consistently, the L3624D-RyR1 mutation impaired both CaM and S100A1 activation of RyR1 at submicromolar Ca2+ concentrations in single channel recordings [94]. On the other hand, the corresponding L3591D-RyR2 mutation abolished CaM regulation only at submicromolar Ca2+ level, while S100A1 regulation of the mutant RyR2 was impaired at both submicromolar and micromollar Ca2+ concentrations [93]. The mutations and functional experiments suggested that S100A1 and CaM do not share exactly the same binding site in RyR2. Recent FRET experiments also showed that S100A1 interacts allosterically with the CaM binding site in RyR1 and RyR2 rather than through direct binding [106].
13.6 Regulation by Calsequestrin
On the SR luminal side, the low affinity but high capacity Ca2+ binding protein, calsequestrin (CSQ), localizes to the junctional SR [107, 108]. RyR-associated proteins, triadin and junctin appear to anchor CSQ to the junctional SR through charge interactions [109,110,111] (Fig. 13.1). In addition, it was recently shown that cardiac CSQ could also directly bind to the luminal side of RyR2 [112]. Two isoforms, CSQ1 and CSQ2, are dominantly expressed in skeletal and cardiac muscle, respectively. Direct regulation of RyR channel activities by CSQ have been investigated by planar bilayer single channel recordings, where luminal conditions can be controlled. CSQ regulates the RyR channel in a luminal Ca2+ concentration dependent manner. With high luminal Ca2+ concentrations, CSQ was dissociated from the RyR accessory proteins, triadin and junctin, while CSQ inhibited the RyR channel through the accessory proteins at the intermediate Ca2+ concentration [113,114,115].
Gene knockout of the CSQ in mice demonstrated both its physiological and pathological significance. CSQ1 (Casq1) knockout mice were viable and fertile; however, modest structural and functional changes were observed in the fast twitch skeletal muscle. Ablation of CSQ1 resulted in slightly slower force development and relaxation of the fast twitch muscle. Structural analysis showed that CSQ1 knockout muscle exhibited low SR volume and high mitochondria density, suggesting that CSQ1 is important for muscle development [116]. CSQ2 has been implicated in cardiac pathology. Missense mutations in human CASQ2 gene, resulting in gene knockout or single amino acid substitutions, were found in patients afflicted with catecholaminergic polymorphic ventricular tachycardia [117, 118]. Both mouse models exhibited arrhythmogenesis under the stress conditions of exercise or catecholamine infusion [119, 120]. Consistently, intracellular Ca2+ handling was altered by catecholamine in mutant cardiomyocytes isolated from mouse hearts. These results indicate that CSQ2 regulation of SR Ca2+ and RyR2 channels is pathologically important.
13.7 Closing Remarks
Almost 50 years have passed since Ca2+-induced Ca2+ release, that we now know is associated with Ca2+-dependent activation of RyR, was first reported [33, 34]. In the last 20 years molecular biology and genetic techniques greatly advanced our understanding of the structure/function relationship of RyR channel regulation by small molecules and proteins. More recently, high resolution three dimensional structural analyses have revealed the detailed protein conformations of the RyR channel complexes under different conditions corresponding to the open/closed channels [24,25,26,27, 30]. Combining these approaches and using computational modeling will provide more detailed molecular insights into RyR regulation by Ca2+ and Ca2+ binding proteins at near atomic levels.
References
Clapham DE (2007) Calcium signaling. Cell 131(6):1047–1058
Bootman MD (2012) Calcium signaling. Cold Spring Harb Perspect Biol 4(7):a011171
Berridge MJ (2016) The inositol trisphosphate/calcium signaling pathway in health and disease. Physiol Rev 96(4):1261–1296
Franzini-Armstrong C, Protasi F (1997) Ryanodine receptors of striated muscles: a complex channel capable of multiple interactions. Physiol Rev 77(3):699–729
Lanner JT, Georgiou DK, Joshi AD, Hamilton SL (2010) Ryanodine receptors: structure, expression, molecular details, and function in calcium release. Cold Spring Harb Perspect Biol 2(11):a003996
Meissner G (2017) The structural basis of ryanodine receptor ion channel function. J Gen Physiol 149(12):1065–1089
Inui M, Saito A, Fleischer S (1987) Purification of the ryanodine receptor and identity with feet structures of junctional terminal cisternae of sarcoplasmic reticulum from fast skeletal muscle. J Biol Chem 262(4):1740–1747
Imagawa T, Smith JS, Coronado R, Campbell KP (1987) Purified ryanodine receptor from skeletal muscle sarcoplasmic reticulum is the Ca2+−permeable pore of the calcium release channel. J Biol Chem 262(34):16636–16643
Lai FA, Erickson HP, Rousseau E, Liu QY, Meissner G (1988) Purification and reconstitution of the calcium release channel from skeletal muscle. Nature 331(6154):315–319
Anderson K, Lai FA, Liu QY, Rousseau E, Erickson HP, Meissner G (1989) Structural and functional characterization of the purified cardiac ryanodine receptor-Ca2+ release channel complex. J Biol Chem 264(2):1329–1335
Takeshima H, Nishimura S, Matsumoto T, Ishida H, Kangawa K, Minamino N et al (1989) Primary structure and expression from complementary DNA of skeletal muscle ryanodine receptor. Nature 339(6224):439–445
Zorzato F, Fujii J, Otsu K, Phillips M, Green NM, Lai FA et al (1990) Molecular cloning of cDNA encoding human and rabbit forms of the Ca2+ release channel (ryanodine receptor) of skeletal muscle sarcoplasmic reticulum. J Biol Chem 265(4):2244–2256
Otsu K, Willard HF, Khanna VK, Zorzato F, Green NM, MacLennan DH (1990) Molecular cloning of cDNA encoding the Ca2+ release channel (ryanodine receptor) of rabbit cardiac muscle sarcoplasmic reticulum. J Biol Chem 265(23):13472–13483
Nakai J, Imagawa T, Hakamata Y, Shigekawa M, Takeshima H, Numa S (1990) Primary structure and functional expression from cDNA of the cardiac ryanodine receptor/calcium release channel. FEBS Lett 271(1–2):169–177
Hakamata Y, Nakai J, Takeshima H, Imoto K (1992) Primary structure and distribution of a novel ryanodine receptor/calcium release channel from rabbit brain. FEBS Lett 312(2–3):229–235
Takeshima H (1993) Primary structure and expression from cDNAs of the ryanodine receptor. Ann N Y Acad Sci 707:165–177
Furuichi T, Furutama D, Hakamata Y, Nakai J, Takeshima H, Mikoshiba K (1994) Multiple types of ryanodine receptor/Ca2+ release channels are differentially expressed in rabbit brain. J Neurosci 14(8):4794–4805
Conti A, Gorza L, Sorrentino V (1996) Differential distribution of ryanodine receptor type 3 (RyR3) gene product in mammalian skeletal muscles. Biochem J 316(Pt 1):19–23
Murayama T, Ogawa Y (1997) Characterization of type 3 ryanodine receptor (RyR3) of sarcoplasmic reticulum from rabbit skeletal muscles. J Biol Chem 272(38):24030–24037
Oyamada H, Murayama T, Takagi T, Iino M, Iwabe N, Miyata T et al (1994) Primary structure and distribution of ryanodine-binding protein isoforms of the bullfrog skeletal muscle. J Biol Chem 269(25):17206–17214
Percival AL, Williams AJ, Kenyon JL, Grinsell MM, Airey JA, Sutko JL (1994) Chicken skeletal muscle ryanodine receptor isoforms: ion channel properties. Biophys J 67(5):1834–1850
Ottini L, Marziali G, Conti A, Charlesworth A, Sorrentino V (1996) Alpha and beta isoforms of ryanodine receptor from chicken skeletal muscle are the homologues of mammalian RyR1 and RyR3. Biochem J 315(Pt 1):207–216
Du GG, Sandhu B, Khanna VK, Guo XH, MacLennan DH (2002) Topology of the Ca2+ release channel of skeletal muscle sarcoplasmic reticulum (RyR1). Proc Natl Acad Sci U S A 99(26):16725–16730
Efremov RG, Leitner A, Aebersold R, Raunser S (2015) Architecture and conformational switch mechanism of the ryanodine receptor. Nature 517(7532):39–43
Zalk R, Clarke OB, des Georges A, Grassucci RA, Reiken S, Mancia F et al (2015) Structure of a mammalian ryanodine receptor. Nature 517(7532):44–49
Yan Z, Bai X, Yan C, Wu J, Li Z, Xie T et al (2015) Structure of the rabbit ryanodine receptor RyR1 at near-atomic resolution. Nature 517(7532):50–55
Peng W, Shen H, Wu J, Guo W, Pan X, Wang R et al (2016) Structural basis for the gating mechanism of the type 2 ryanodine receptor RyR2. Science 354(6310):aah5324
Zhong X, Liu Y, Zhu L, Meng X, Wang R, Van Petegem F et al (2013) Conformational dynamics inside amino-terminal disease hotspot of ryanodine receptor. Structure 21(11):2051–2060
Yuchi Z, Yuen SM, Lau K, Underhill AQ, Cornea RL, Fessenden JD et al (2015) Crystal structures of ryanodine receptor SPRY1 and tandem-repeat domains reveal a critical FKBP12 binding determinant. Nat Commun 6:7947
des Georges A, Clarke OB, Zalk R, Yuan Q, Condon KJ, Grassucci RA et al (2016) Structural basis for gating and activation of RyR1. Cell 167(1):145–157. e17
Schneider MF, Chandler WK (1973) Voltage dependent charge movement of skeletal muscle: a possible step in excitation-contraction coupling. Nature 242(5395):244–246
Nabauer M, Callewaert G, Cleemann L, Morad M (1989) Regulation of calcium release is gated by calcium current, not gating charge, in cardiac myocytes. Science 244(4906):800–803
Ford LE, Podolsky RJ (1970) Regenerative calcium release within muscle cells. Science 167(3914):58–59
Endo M, Tanaka M, Ogawa Y (1970) Calcium induced release of calcium from the sarcoplasmic reticulum of skinned skeletal muscle fibres. Nature 228(5266):34–36
Brum G, Rios E, Stefani E (1988) Effects of extracellular calcium on calcium movements of excitation-contraction coupling in frog skeletal muscle fibres. J Physiol 398:441–473
Nakai J, Dirksen RT, Nguyen HT, Pessah IN, Beam KG, Allen PD (1996) Enhanced dihydropyridine receptor channel activity in the presence of ryanodine receptor. Nature 380(6569):72–75
Endo M (2009) Calcium-induced calcium release in skeletal muscle. Physiol Rev 89(4):1153–1176
Rios E (2018) Calcium-induced release of calcium in muscle: 50 years of work and the emerging consensus. J Gen Physiol 150(4):521–537
Murayama T, Kurebayashi N, Ogawa H, Yamazawa T, Oyamada H, Suzuki J et al (2016) Genotype-phenotype correlations of malignant hyperthermia and central Core disease mutations in the central region of the RYR1 channel. Hum Mutat 37(11):1231–1241
Meissner G (1994) Ryanodine receptor/Ca2+ release channels and their regulation by endogenous effectors. Annu Rev Physiol 56:485–508
Coronado R, Morrissette J, Sukhareva M, Vaughan DM (1994) Structure and function of ryanodine receptors. Am J Physiol 266(6 Pt 1):C1485–C1504
Chen SR, Zhang L, MacLennan DH (1992) Characterization of a Ca2+ binding and regulatory site in the Ca2+ release channel (ryanodine receptor) of rabbit skeletal muscle sarcoplasmic reticulum. J Biol Chem 267(32):23318–23326
Chen SR, Zhang L, MacLennan DH (1993) Antibodies as probes for Ca2+ activation sites in the Ca2+ release channel (ryanodine receptor) of rabbit skeletal muscle sarcoplasmic reticulum. J Biol Chem 268(18):13414–13421
Chen SR, MacLennan DH (1994) Identification of calmodulin-, Ca2+-, and ruthenium red-binding domains in the Ca2+ release channel (ryanodine receptor) of rabbit skeletal muscle sarcoplasmic reticulum. J Biol Chem 269(36):22698–22704
Chen SR, Ebisawa K, Li X, Zhang L (1998) Molecular identification of the ryanodine receptor Ca2+ sensor. J Biol Chem 273(24):14675–14678
Li P, Chen SR (2001) Molecular basis of Ca2+ activation of the mouse cardiac Ca2+ release channel (ryanodine receptor). J Gen Physiol 118(1):33–44
Fessenden JD, Chen L, Wang Y, Paolini C, Franzini-Armstrong C, Allen PD et al (2001) Ryanodine receptor point mutant E4032A reveals an allosteric interaction with ryanodine. Proc Natl Acad Sci U S A 98(5):2865–2870
Murayama T, Ogawa H, Kurebayashi N, Ohno S, Horie M, Sakurai T (2018) A tryptophan residue in the caffeine-binding site of the ryanodine receptor regulates Ca2+ sensitivity. Commun Biol 1:98
Xiong H, Feng X, Gao L, Xu L, Pasek DA, Seok JH et al (1998) Identification of a two EF-hand Ca2+ binding domain in lobster skeletal muscle ryanodine receptor/Ca2+ release channel. Biochemistry 37(14):4804–4814
Xiong L, Zhang JZ, He R, Hamilton SL (2006) A Ca2+-binding domain in RyR1 that interacts with the calmodulin binding site and modulates channel activity. Biophys J 90(1):173–182
Fessenden JD, Feng W, Pessah IN, Allen PD (2004) Mutational analysis of putative calcium binding motifs within the skeletal ryanodine receptor isoform, RyR1. J Biol Chem 279(51):53028–53035
Guo W, Sun B, Xiao Z, Liu Y, Wang Y, Zhang L et al (2016) The EF-hand Ca2+ binding domain is not required for cytosolic Ca2+ activation of the cardiac ryanodine receptor. J Biol Chem 291(5):2150–2160
Gomez AC, Yamaguchi N (2014) Two regions of the ryanodine receptor calcium channel are involved in Ca2+-dependent inactivation. Biochemistry 53(8):1373–1379
Xu L, Gomez AC, Pasek DA, Meissner G, Yamaguchi N (2017) Two EF-hand motifs in ryanodine receptor calcium release channels contribute to isoform-specific regulation by calmodulin. Cell Calcium 66:62–70
Chugun A, Sato O, Takeshima H, Ogawa Y (2007) Mg2+ activates the ryanodine receptor type 2 (RyR2) at intermediate Ca2+ concentrations. Am J Physiol Cell Physiol 292(1):C535–C544
Yamaguchi N, Xu L, Pasek DA, Evans KE, Meissner G (2003) Molecular basis of calmodulin binding to cardiac muscle Ca2+ release channel (ryanodine receptor). J Biol Chem 278(26):23480–23486
Laver DR, Baynes TM, Dulhunty AF (1997) Magnesium inhibition of ryanodine-receptor calcium channels: evidence for two independent mechanisms. J Membr Biol 156(3):213–229
Gomez AC, Holford TW, Yamaguchi N (2016) Malignant hyperthermia-associated mutations in the S2-S3 cytoplasmic loop of type 1 ryanodine receptor calcium channel impair calcium-dependent inactivation. Am J Physiol Cell Physiol 311(5):C749–C757
Murayama T, Kurebayashi N, Ogawa Y (2000) Role of Mg2+ in Ca2+-induced Ca2+ release through ryanodine receptors of frog skeletal muscle: modulations by adenine nucleotides and caffeine. Biophys J 78(4):1810–1824
Tripathy A, Meissner G (1996) Sarcoplasmic reticulum lumenal Ca2+ has access to cytosolic activation and inactivation sites of skeletal muscle Ca2+ release channel. Biophys J 70(6):2600–2615
Xu L, Meissner G (1998) Regulation of cardiac muscle Ca2+ release channel by sarcoplasmic reticulum lumenal Ca2+. Biophys J 75(5):2302–2312
Du GG, MacLennan DH (1999) Ca2+ inactivation sites are located in the COOH-terminal quarter of recombinant rabbit skeletal muscle Ca2+ release channels (ryanodine receptors). J Biol Chem 274(37):26120–26126
Hayek SM, Zhu X, Bhat MB, Zhao J, Takeshima H, Valdivia HH et al (2000) Characterization of a calcium-regulation domain of the skeletal-muscle ryanodine receptor. Biochem J 351(Pt 1):57–65
Nakai J, Gao L, Xu L, Xin C, Pasek DA, Meissner G (1999) Evidence for a role of C-terminus in Ca2+ inactivation of skeletal muscle Ca2+ release channel (ryanodine receptor). FEBS Lett 459(2):154–158
Du GG, Khanna VK, MacLennan DH (2000) Mutation of divergent region 1 alters caffeine and Ca2+ sensitivity of the skeletal muscle Ca2+ release channel (ryanodine receptor). J Biol Chem 275(16):11778–11783
Fabiato A, Fabiato F (1979) Calcium and cardiac excitation-contraction coupling. Annu Rev Physiol 41:473–484
Orchard CH, Eisner DA, Allen DG (1983) Oscillations of intracellular Ca2+ in mammalian cardiac muscle. Nature 304(5928):735–738
Jiang D, Xiao B, Yang D, Wang R, Choi P, Zhang L et al (2004) RyR2 mutations linked to ventricular tachycardia and sudden death reduce the threshold for store-overload-induced Ca2+ release (SOICR). Proc Natl Acad Sci U S A 101(35):13062–13067
Jiang D, Wang R, Xiao B, Kong H, Hunt DJ, Choi P et al (2005) Enhanced store overload-induced Ca2+ release and channel sensitivity to luminal Ca2+ activation are common defects of RyR2 mutations linked to ventricular tachycardia and sudden death. Circ Res 97(11):1173–1181
Jiang D, Chen W, Xiao J, Wang R, Kong H, Jones PP et al (2008) Reduced threshold for luminal Ca2+ activation of RyR1 underlies a causal mechanism of porcine malignant hyperthermia. J Biol Chem 283(30):20813–20820
Chen W, Wang R, Chen B, Zhong X, Kong H, Bai Y et al (2014) The ryanodine receptor store-sensing gate controls Ca2+ waves and Ca2+-triggered arrhythmias. Nat Med 20(2):184–192
Saimi Y, Kung C (2002) Calmodulin as an ion channel subunit. Annu Rev Physiol 64:289–311
Tripathy A, Xu L, Mann G, Meissner G (1995) Calmodulin activation and inhibition of skeletal muscle Ca2+ release channel (ryanodine receptor). Biophys J 69(1):106–119
Buratti R, Prestipino G, Menegazzi P, Treves S, Zorzato F (1995) Calcium dependent activation of skeletal muscle Ca2+ release channel (ryanodine receptor) by calmodulin. Biochem Biophys Res Commun 213(3):1082–1090
Chen SR, Li X, Ebisawa K, Zhang L (1997) Functional characterization of the recombinant type 3 Ca2+ release channel (ryanodine receptor) expressed in HEK293 cells. J Biol Chem 272(39):24234–24246
Fruen BR, Bardy JM, Byrem TM, Strasburg GM, Louis CF (2000) Differential Ca2+ sensitivity of skeletal and cardiac muscle ryanodine receptors in the presence of calmodulin. Am J Physiol Cell Physiol 279(3):C724–C733
Balshaw DM, Xu L, Yamaguchi N, Pasek DA, Meissner G (2001) Calmodulin binding and inhibition of cardiac muscle calcium release channel (ryanodine receptor). J Biol Chem 276(23):20144–20153
Yamaguchi N, Xu L, Pasek DA, Evans KE, Chen SR, Meissner G (2005) Calmodulin regulation and identification of calmodulin binding region of type-3 ryanodine receptor calcium release channel. Biochemistry 44(45):15074–15081
Ono M, Yano M, Hino A, Suetomi T, Xu X, Susa T et al (2010) Dissociation of calmodulin from cardiac ryanodine receptor causes aberrant Ca2+ release in heart failure. Cardiovasc Res 87(4):609–617
Terentyev D, Gyorke I, Belevych AE, Terentyeva R, Sridhar A, Nishijima Y et al (2008) Redox modification of ryanodine receptors contributes to sarcoplasmic reticulum Ca2+ leak in chronic heart failure. Circ Res 103(12):1466–1472
Oda T, Yang Y, Uchinoumi H, Thomas DD, Chen-Izu Y, Kato T et al (2015) Oxidation of ryanodine receptor (RyR) and calmodulin enhance Ca release and pathologically alter, RyR structure and calmodulin affinity. J Mol Cell Cardiol 85:240–248
Yamaguchi N, Xin C, Meissner G (2001) Identification of apocalmodulin and Ca2+-calmodulin regulatory domain in skeletal muscle Ca2+ release channel, ryanodine receptor. J Biol Chem 276(25):22579–22585
Moore CP, Rodney G, Zhang JZ, Santacruz-Toloza L, Strasburg G, Hamilton SL (1999) Apocalmodulin and Ca2+ calmodulin bind to the same region on the skeletal muscle Ca2+ release channel. Biochemistry 38(26):8532–8537
Rodney GG, Moore CP, Williams BY, Zhang JZ, Krol J, Pedersen SE et al (2001) Calcium binding to calmodulin leads to an N-terminal shift in its binding site on the ryanodine receptor. J Biol Chem 276(3):2069–2074
Maximciuc AA, Putkey JA, Shamoo Y, Mackenzie KR (2006) Complex of calmodulin with a ryanodine receptor target reveals a novel, flexible binding mode. Structure 14(10):1547–1556
Menegazzi P, Larini F, Treves S, Guerrini R, Quadroni M, Zorzato F (1994) Identification and characterization of three calmodulin binding sites of the skeletal muscle ryanodine receptor. Biochemistry 33(31):9078–9084
Guerrini R, Menegazzi P, Anacardio R, Marastoni M, Tomatis R, Zorzato F et al (1995) Calmodulin binding sites of the skeletal, cardiac, and brain ryanodine receptor Ca2+ channels: modulation by the catalytic subunit of cAMP-dependent protein kinase? Biochemistry 34(15):5120–5129
Lau K, Chan MM, Van Petegem F (2014) Lobe-specific calmodulin binding to different ryanodine receptor isoforms. Biochemistry 53(5):932–946
Yamaguchi N, Takahashi N, Xu L, Smithies O, Meissner G (2007) Early cardiac hypertrophy in mice with impaired calmodulin regulation of cardiac muscle Ca2+ release channel. J Clin Invest 117(5):1344–1353
Yamaguchi N, Xu L, Evans KE, Pasek DA, Meissner G (2004) Different regions in skeletal and cardiac muscle ryanodine receptors are involved in transducing the functional effects of calmodulin. J Biol Chem 279(35):36433–36439
Samso M, Wagenknecht T (2002) Apocalmodulin and Ca2+-calmodulin bind to neighboring locations on the ryanodine receptor. J Biol Chem 277(2):1349–1353
Arnaiz-Cot JJ, Damon BJ, Zhang XH, Cleemann L, Yamaguchi N, Meissner G et al (2013) Cardiac calcium signalling pathologies associated with defective calmodulin regulation of type 2 ryanodine receptor. J Physiol 591(17):4287–4299
Yamaguchi N, Chakraborty A, Huang TQ, Xu L, Gomez AC, Pasek DA et al (2013) Cardiac hypertrophy associated with impaired regulation of cardiac ryanodine receptor by calmodulin and S100A1. Am J Physiol Heart Circ Physiol 305(1):H86–H94
Yamaguchi N, Prosser BL, Ghassemi F, Xu L, Pasek DA, Eu JP et al (2011) Modulation of sarcoplasmic reticulum Ca2+ release in skeletal muscle expressing ryanodine receptor impaired in regulation by calmodulin and S100A1. Am J Physiol Cell Physiol 300(5):C998–C1012
Nyegaard M, Overgaard MT, Sondergaard MT, Vranas M, Behr ER, Hildebrandt LL et al (2012) Mutations in calmodulin cause ventricular tachycardia and sudden cardiac death. Am J Hum Genet 91(4):703–712
Hwang HS, Nitu FR, Yang Y, Walweel K, Pereira L, Johnson CN et al (2014) Divergent regulation of ryanodine receptor 2 calcium release channels by arrhythmogenic human calmodulin missense mutants. Circ Res 114(7):1114–1124
Sondergaard MT, Tian X, Liu Y, Wang R, Chazin WJ, Chen SR et al (2015) Arrhythmogenic calmodulin mutations affect the activation and termination of cardiac ryanodine receptor-mediated Ca2+ release. J Biol Chem 290(43):26151–26162
Vassilakopoulou V, Calver BL, Thanassoulas A, Beck K, Hu H, Buntwal L et al (2015) Distinctive malfunctions of calmodulin mutations associated with heart RyR2-mediated arrhythmic disease. Biochim Biophys Acta 1850(11):2168–2176
Liu B, Walton SD, Ho HT, Belevych AE, Tikunova SB, Bonilla I et al (2018) Gene transfer of engineered calmodulin alleviates ventricular arrhythmias in a calsequestrin-associated mouse model of catecholaminergic polymorphic ventricular tachycardia. J Am Heart Assoc 7(10):e008155
Treves S, Scutari E, Robert M, Groh S, Ottolia M, Prestipino G et al (1997) Interaction of S100A1 with the Ca2+ release channel (ryanodine receptor) of skeletal muscle. Biochemistry 36(38):11496–11503
Most P, Remppis A, Pleger ST, Loffler E, Ehlermann P, Bernotat J et al (2003) Transgenic overexpression of the Ca2+-binding protein S100A1 in the heart leads to increased in vivo myocardial contractile performance. J Biol Chem 278(36):33809–33817
Volkers M, Rohde D, Goodman C, Most P (2010) S100A1: a regulator of striated muscle sarcoplasmic reticulum Ca2+ handling, sarcomeric, and mitochondrial function. J Biomed Biotechnol 2010:178614
Prosser BL, Hernandez-Ochoa EO, Schneider MF (2011) S100A1 and calmodulin regulation of ryanodine receptor in striated muscle. Cell Calcium 50(4):323–331
Prosser BL, Wright NT, Hernandez-Ochoa EO, Varney KM, Liu Y, Olojo RO et al (2008) S100A1 binds to the calmodulin-binding site of ryanodine receptor and modulates skeletal muscle excitation-contraction coupling. J Biol Chem 283(8):5046–5057
Wright NT, Prosser BL, Varney KM, Zimmer DB, Schneider MF, Weber DJ (2008) S100A1 and calmodulin compete for the same binding site on ryanodine receptor. J Biol Chem 283(39):26676–26683
Rebbeck RT, Nitu FR, Rohde D, Most P, Bers DM, Thomas DD et al (2016) S100A1 protein does not compete with calmodulin for ryanodine receptor binding but structurally alters the ryanodine receptor.Calmodulin complex. J Biol Chem 291(30):15896–15907
Saito A, Seiler S, Chu A, Fleischer S (1984) Preparation and morphology of sarcoplasmic reticulum terminal cisternae from rabbit skeletal muscle. J Cell Biol 99(3):875–885
Franzini-Armstrong C, Kenney LJ, Varriano-Marston E (1987) The structure of calsequestrin in triads of vertebrate skeletal muscle: a deep-etch study. J Cell Biol 105(1):49–56
Guo W, Campbell KP (1995) Association of triadin with the ryanodine receptor and calsequestrin in the lumen of the sarcoplasmic reticulum. J Biol Chem 270(16):9027–9030
Zhang L, Kelley J, Schmeisser G, Kobayashi YM, Jones LR (1997) Complex formation between junctin, triadin, calsequestrin, and the ryanodine receptor. Proteins of the cardiac junctional sarcoplasmic reticulum membrane. J Biol Chem 272(37):23389–23397
Kobayashi YM, Alseikhan BA, Jones LR (2000) Localization and characterization of the calsequestrin-binding domain of triadin 1. Evidence for a charged beta-strand in mediating the protein-protein interaction. J Biol Chem 275(23):17639–17646
Handhle A, Ormonde CE, Thomas NL, Bralesford C, Williams AJ, Lai FA et al (2016) Calsequestrin interacts directly with the cardiac ryanodine receptor luminal domain. J Cell Sci 129(21):3983–3988
Beard NA, Sakowska MM, Dulhunty AF, Laver DR (2002) Calsequestrin is an inhibitor of skeletal muscle ryanodine receptor calcium release channels. Biophys J 82(1 Pt 1):310–320
Gyorke I, Hester N, Jones LR, Gyorke S (2004) The role of calsequestrin, triadin, and junctin in conferring cardiac ryanodine receptor responsiveness to luminal calcium. Biophys J 86(4):2121–2128
Beard NA, Casarotto MG, Wei L, Varsanyi M, Laver DR, Dulhunty AF (2005) Regulation of ryanodine receptors by calsequestrin: effect of high luminal Ca2+ and phosphorylation. Biophys J 88(5):3444–3454
Paolini C, Quarta M, Nori A, Boncompagni S, Canato M, Volpe P et al (2007) Reorganized stores and impaired calcium handling in skeletal muscle of mice lacking calsequestrin-1. J Physiol 583(Pt 2):767–784
Lahat H, Pras E, Olender T, Avidan N, Ben-Asher E, Man O et al (2001) A missense mutation in a highly conserved region of CASQ2 is associated with autosomal recessive catecholamine-induced polymorphic ventricular tachycardia in bedouin families from Israel. Am J Hum Genet 69(6):1378–1384
Postma AV, Denjoy I, Hoorntje TM, Lupoglazoff JM, Da Costa A, Sebillon P et al (2002) Absence of calsequestrin 2 causes severe forms of catecholaminergic polymorphic ventricular tachycardia. Circ Res 91(8):e21–e26
Knollmann BC, Chopra N, Hlaing T, Akin B, Yang T, Ettensohn K et al (2006) Casq2 deletion causes sarcoplasmic reticulum volume increase, premature Ca2+ release, and catecholaminergic polymorphic ventricular tachycardia. J Clin Invest 116(9):2510–2520
Song L, Alcalai R, Arad M, Wolf CM, Toka O, Conner DA et al (2007) Calsequestrin 2 (CASQ2) mutations increase expression of calreticulin and ryanodine receptors, causing catecholaminergic polymorphic ventricular tachycardia. J Clin Invest 117(7):1814–1823
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC et al (2004) UCSF chimera–a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612
Acknowledgements
I would like to appreciate Dr. Gerhard Meissner for his mentorship on RyR structure/function analysis when I was working in his laboratory and for his continuous advices and encouragements on my current studies. I am very thankful to Dr. Martin Morad, the director of Cardiac Signaling Center, for providing me with wonderful environment to study Ca2+ signaling, and valuable suggestions and discussions on my studies. I am also grateful to Angela C. Gomez and Jordan S. Carter for their contributions to the RyR mutagenesis studies in my laboratory and for the comments on this manuscript. This study was supported by National Institutes of Health Grants R03AR061030, P20GM103499, and UL1TR001450.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Yamaguchi, N. (2020). Molecular Insights into Calcium Dependent Regulation of Ryanodine Receptor Calcium Release Channels. In: Islam, M. (eds) Calcium Signaling. Advances in Experimental Medicine and Biology, vol 1131. Springer, Cham. https://doi.org/10.1007/978-3-030-12457-1_13
Download citation
DOI: https://doi.org/10.1007/978-3-030-12457-1_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-12456-4
Online ISBN: 978-3-030-12457-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)