Abstract
The high-conductance intracellular calcium (Ca2+) channel RyR2 is essential for the coupling of excitation and contraction in cardiac muscle. Among various modulators, calmodulin (CaM) regulates RyR2 in a Ca2+-dependent manner. Here we reveal the regulatory mechanism by which porcine RyR2 is modulated by human CaM through the structural determination of RyR2 under eight conditions. Apo-CaM and Ca2+-CaM bind to distinct but overlapping sites in an elongated cleft formed by the handle, helical and central domains. The shift in CaM-binding sites on RyR2 is controlled by Ca2+ binding to CaM, rather than to RyR2. Ca2+-CaM induces rotations and intradomain shifts of individual central domains, resulting in pore closure of the PCB95 and Ca2+-activated channel. By contrast, the pore of the ATP, caffeine and Ca2+-activated channel remains open in the presence of Ca2+-CaM, which suggests that Ca2+-CaM is one of the many competing modulators of RyR2 gating.
Similar content being viewed by others
Main
Cardiac muscle contraction is triggered by Ca2+ flux into the cytosol, initially from the extracellular environment, mediated by Cav1.2, and subsequently from the sarcoplasmic reticulum Ca2+ store, mediated by RyR21,2,3. Ryanodine receptors are the largest known ion channels and consist of a homotetramer with a molecular mass of more than 2 megadaltons. More than 80% of the protein folds into a multi-domain cytoplasmic assembly that senses interactions with a variety of modulators, which range from ions to proteins4,5,6. The precise regulation of RyR2 activity is critical for each heartbeat. Aberrant activity of RyR2 is associated with life-threatening cardiac arrhythmias7,8,9,10.
The 17-kDa protein CaM is an essential calcium sensor that has a central role in most calcium signalling events11. CaM consists of roughly symmetrical N- and C-terminal lobes (N- and C-lobes hereafter), joined by a flexible hinge12,13. Each lobe can cooperatively bind to two Ca2+ ions, with a micromolar-range binding affinity, via two EF-hand (helices E and F-hand) motifs. Upon Ca2+ binding, the exposure of several hydrophobic residues in both lobes facilitates CaM binding to the target sequence. CaM interacts directly with ryanodine receptors with a 1:1 stoichiometry of the CaM–RyR protomers14,15 and binding affinity at nanomolar range14.
Regulation of ryanodine receptors by CaM, however, is isoform-specific. CaM shows biphasic regulation of RyR1, acting as a weak activator at nanomolar levels of Ca2+ (apo-CaM) and an inhibitor at micromolar levels of Ca2+ (Ca2+-CaM)14,16. By contrast, apo-CaM has no effect17 or an inhibitory effect on RyR214, whereas Ca2+-CaM inhibits RyR214. CaM has also been shown to facilitate the termination of store-overload-induced Ca2 + release (SOICR)18. Aberrant interactions between CaM and RyR2 are associated with heart failure19,20,21,22, and correction of impaired CaM–RyR2 interactions may serve as a therapy for lethal arrhythmia in pressure-overload-induced heart failure23.
Structural characterization of RyR–CaM complexes has been limited to low-resolution electron microscopy maps that suggest two overlapping, but distinct, binding sites in RyR1 for apo- and Ca2+-CaM24,25,26. A peptide that corresponds to residues 3614–3643 of RyR1 (residues 3581–3612 in the central domain of RyR2) binds to both apo- and Ca2+-CaM15,27. The crystal structure of Ca2+-CaM bound to the peptide revealed hydrophobic anchors in the N and C termini of the peptide that accommodate the C- and N-lobes of Ca2+-CaM, respectively28.
To elucidate the modulation of RyR2 by CaM, we report eight cryo-electron microscopy (cryo-EM) structures of RyR2 that collectively reveal molecular recognition characteristics for different forms of CaM and provide insights into the regulation of RyR2 channel gating by CaM.
Structures of RyR2 under eight conditions
To achieve a better understanding of RyR2 modulation by CaM (Extended Data Fig. 1a, b), we determined the cryo-EM structures of the porcine RyR2 (Extended Data Fig. 1c) under the following eight conditions.
Condition (1) consisted of RyR2 bound to FKBP12.6 and apo-CaM (hereafter FKBP12.6/apo-CaM) and was used to assess the apo-CaM binding site. Condition (2) consisted of RyR2 bound to FKBP12.6 and a Ca2+-binding-deficient CaM mutant that mimics apo-CaM24,29 (CaM-M) in the presence of ATP, caffeine and low [Ca2+] (hereafter FKBP12.6/ATP/caffeine/low-[Ca2+]/CaM-M), this structure was used to investigate the mechanism for the binding-location switch of CaM. Condition (3) consisted of RyR2 bound to FKBP12.6 in the presence of ATP, caffeine and low [Ca2+] (FKBP12.6/ATP/caffeine/low-[Ca2+]), the presence of which maximizes the open state. Condition (4) consisted of RyR2 bound to FKBP12.6 and Ca2+-CaM in the presence of ATP, caffeine and low [Ca2+] (hereafter FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM); this condition was used to examine the effect of Ca2+-CaM on the open RyR2 channel in the presence of FKBP12.6, ATP, caffeine and low [Ca2+]. Conditions (5) and (6) corresponded to conditions (3) and (4), respectively, but were treated with CHAPS and DOPC instead of digitonin. Condition (7) consisted of RyR2 in high [Ca2+] in the presence of FKBP12.6, ATP, caffeine and Ca2+-CaM (hereafter FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM); this environment was used to achieve a better resolution for Ca2+-CaM. Condition (8) consisted of RyR2 bound to Ca2+-CaM in the presence of 2,2′,3,5′,6-pentachlorobiphenyl (PCB95) and low [Ca2+] (hereafter PCB95/low-[Ca2+]/Ca2+-CaM); this condition was used to investigate the effect of Ca2+-CaM on PCB95 and the Ca2+-activated RyR2 channel30. The eight conditions and corresponding structures are summarized in Supplementary Table 1 and Extended Data Table 1, respectively.
All cryo-EM datasets were processed following the same procedure (Extended Data Fig. 2). The FKBP12.6/apo-CaM RyR2 structure was determined at an overall resolution of 3.6 Å, the highest among all of the available RyR2 structures (Fig. 1a and Extended Data Figs. 1i, 3a, 4a–i). The secondary structural elements of apo-CaM were clearly resolved (Fig. 1a and Extended Data Fig. 3b). The FKBP12.6/ATP/caffeine/low-[Ca2+]/CaM-M RyR2 structure was determined at an overall resolution of 4.2 Å, in which the well-resolved CaM-M is positioned similarly to apo-CaM on RyR2 (Fig. 1b and Extended Data Figs. 1i, 3c, d). The densities for both lobes of Ca2+-CaM are visible in the FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM RyR2 structure (3.9 Å resolution) and PCB95/low-[Ca2+]/Ca2+-CaM RyR2 structure (4.4 Å resolution) in which the N-lobe is better-resolved than the C-lobe. By contrast, only one lobe of Ca2+-CaM—the N lobe as judged from the comparison with the structure of FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM at 3.9 Å—is discernible in the FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM structure with a resolution of 4.2 Å (Fig. 1c, d and Extended Data Figs. 1, 3).
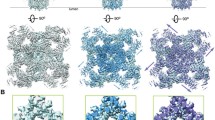
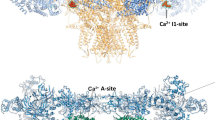
a–d, Overall electron microscopy maps for four indicated complexes. a, RyR2 in the presence of FKBP12.6/apo-CaM (3.6 Å). b, RyR2 in the presence of FKBP12.6/ATP/caffeine/low [Ca2+]/CaM-M (4.2 Å). c, RyR2 in the presence of FKBP12.6/ATP/caffeine/high [Ca2+]/Ca2+-CaM (3.9 Å). d, RyR2 in the presence of PCB95/low-[Ca2+]/Ca2+-CaM (4.4 Å). Insets, cytoplasmic views of the channel gates. The dashed circles indicate the distances between the Cα atoms of the gating residues in the diagonal protomers. See Extended Data Table 1 for details of the structures. SR, sarcoplasmic reticulum. e, The pore of RyR2 remains closed in the FKBP12.6/apo-CaM structure. The ion permeation path, calculated by HOLE41, is illustrated as yellow dots. SF, selectivity filter. f, g, Open pores of RyR2 in FKBP12.6/ATP/caffeine/low-[Ca2+]/CaM-M (f) and FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM (g). h, The corresponding pore radii of RyR2 for the three structures shown in e–g. Electron microscopy maps were generated in Chimera and contoured at levels of 0.027, 0.022, 0.02 and 0.015 for a–d, respectively. All structures were prepared using PyMOL (http://www.pymol.org).
With regard to the gating state of the eight structures reported here, the clearly resolved Ile4868 residues on the S6 helical bundle of FKBP12.6/apo-CaM constitute the constriction site with a radius of approximately 1 Å, which is identical to the previously reported apo-RyR2 in the closed state30 (Fig. 1e, h). The constriction site appears to shift to Gln4864 with an expanded radius of around 3 Å in conditions (2), (3) and (5)–(7) (Fig. 1f–h and Extended Data Fig. 1d, h), similar to that in the open PCB95/low-[Ca2+] structure30. However, the density for the side chain of Gln4864 in FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM is not well-resolved. We, therefore, compared the distances of Cα atoms of the gating residues in the diagonal protomers, which are approximately 16 Å for FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM and 11 Å for FKBP12.6/apo-CaM (Fig. 1a and Extended Data Fig. 1e). The constriction site in PCB95/low-[Ca2+]/Ca2+-CaM is constituted by Ile4868, for which the diagonal distance of the Cα atoms is approximately 11 Å—similar to that in FKBP12.6/apo-CaM (Fig. 1a, d).
As seen in RyR1, the Ca2+-, ATP- and caffeine-binding sites are located at the interfaces between the central and channel domains of RyR231 (Extended Data Fig. 4j–m).
Location of apo-CaM in RyR2
Consistent with the low-resolution structure of RyR1, apo-CaM is located in an elongated cleft formed by the handle, helical and central domains of RyR2 in FKBP12.6/apo-CaM24,25 (Fig. 2a). The N-lobe is stuck in the upper half of the cleft formed by helical domain 1 (HD1), whereas the C-lobe is located at the bottom edge of the cleft surrounded by the handle and central domains of RyR2 (Fig. 2a).
a, Apo-CaM is located in a cleft formed by the handle, helical and central domains of RyR2. One RyR2 protomer is shown in domain-coloured surface view. b, c, Multiple interfaces between RyR2 and apo-CaM. The red, dashed boxes indicate the interfaces. d, A previously unresolved helix α−1 on RyR2 serves as the primary docking site for both lobes of Ca2+-CaM. e, Functional validation of the observed interfaces between mouse RyR2 and human CaM. Open probabilities of single RyR2 channels before (control) and after addition of CaM(WT) (1 μM). Data are mean ± s.e.m. from RyR2(WT) (n = 9), RyR2(Y2156A) (n = 8), RyR2(V3599A) (n = 7), RyR2(W3587A) (n = 8) and RyR2(L3590A) (n = 9 single channels) and analysed by paired, two-sided Student’s t-test (versus its own control) with P values shown in blue and by one-way analysis of variance (ANOVA) with a Dunnett’s post hoc test (versus RyR2(WT) control and RyR2(WT) with CaM(WT), respectively) with adjusted P values shown in black. f, The termination threshold of Ca2+ release in Ryr2WT- and Ryr2-mutant-expressing HEK293 cells transfected with no CaM (control), CaM(WT) or CaM-M. Data are mean ± s.e.m. with the number of independent experiments for each condition shown and analysed by one-way ANOVA with a Dunnett’s post hoc test with adjusted P values shown in blue (versus its own control) and in black (versus RyR2(WT) control).
The FKBP12.6/apo-CaM structure reveals five surface patches on RyR2 that interact with apo-CaM. The N-lobe interacts with RyR2 through three interfaces that are mainly located in HD1 (Fig. 2b and Extended Data Fig. 5a). The most prominent interface is formed between the N terminus of helix 4 (N4) in the N-lobe and the C termini of helices 2b and α1 in HD1, and is mainly mediated by extensive hydrophobic residues. Phe66, Pro67 and Leu70 on N4 probably interact with Tyr2203 in helix 2b and Tyr2157 (human Tyr2156) in helix α1. Ile10 in N1 may also interact with Tyr2157 (Extended Data Fig. 5a, g). The human Y2156C variant is linked to catecholaminergic polymorphic ventricular tachycardia8. The second interface is mediated by charged residues between the N terminus of N1 in the N-lobe and helix α1 in HD1 and the N terminus of helix α0 in the central domain. The third interface is formed between a region rich in acidic residues in N3 of the N-lobe and Lys2558 in helix 8b of HD1 (Fig. 2b and Extended Data Fig. 5a).
The C-lobe interacts with RyR2 through two interfaces (Fig. 2c). One is consistent with previous reports15,27. Residues 3593–3607 in the central domain folds into a newly resolved helix α ‘minus 1’ (helix α−1) that is enclosed by the hydrophobic cavity of the C-lobe, representing the primary interface. Phe3604 serves as a hydrophobic anchor for the hydrophobic cavity of the C-lobe. A minor interface is formed between helix 12 and the C terminus of helix 11 in the handle domain with the C terminus of C1 and the loop between C2 and C3 in the C-lobe, also through hydrophobic interactions (Fig. 2c and Extended Data Fig. 5b, c, h).
Shift of CaM-binding site in RyR2 after Ca2+ loading
CaM markedly slips down along the cleft after Ca2+ loading, making extensive interactions with the central domain (Fig. 2d). The N-lobe is anchored by the central domain and the C-lobe drops beyond the cleft, coordinated only by helix α−1 (Fig. 2d). The limited contact may explain the structural flexibility of the C-lobe.
The binding of Ca2+-CaM with intact RyR2 is similar to that of Ca2+-CaM with the RyR1 peptide28. The N- and C-lobes of CaM interact with the C- and N termini of helix α−1, respectively (Fig. 2d). Phe3604 and Trp3588 anchor the hydrophobic cavities of the N- and C-lobes, respectively (Extended Data Fig. 5d–f, i). An additional interface is formed between the N terminus of N3 and the C terminus of helix α9 in the central domain to further stabilize the binding of N-lobe. Asp51 in N3 probably interacts with Arg2209 in helix 2b of HD1 (Fig. 2d and Extended Data Fig. 5e).
Functional validation of RyR2–CaM interfaces
We assessed the effect of mutations in RyR2 and CaM that are located in the structurally revealed interfaces on CaM regulation of RyR2. Wild-type CaM (CaM(WT)) strongly reduced the open probability of single wild-type mouse RyR2 (RyR2(WT)) channels. The following mutations markedly reduced CaM inhibition of single RyR2 channels: RyR2(Y2156A) (porcine Y2157A) near an interface between the apo-CaM N-lobe and RyR2, RyR2(V3599A) near both the apo-CaM C-lobe–RyR2 and Ca2+-CaM N-lobe–RyR2 interfaces, and RyR2(W3587A) and RyR2(L3590A) near the Ca2+-CaM C-lobe–RyR2 interface (Fig. 2e and Extended Data Fig. 6).
We next examined the effect of these RyR2 mutations on the termination of RyR2-mediated SOICR. As previously shown18, CaM(WT) increased Ca2+ release termination in RyR2(WT)-expressing HEK293 cells, whereas CaM-M reduced Ca2+ release termination (that is, a longer calcium release) (Fig. 2f and Extended Data Fig. 7a–c). Consistent with their effect on single RyR2 channels, all four mutations in RyR2 significantly reduced Ca2+ release termination in HEK293 cells, probably by impairing the effect of endogenous CaM on RyR2 inhibition (Fig. 2f). These RyR2 mutations also reduced or abolished the effect of exogenously expressed CaM(WT) and CaM-M on Ca2+ release termination (Fig. 2f), but had little or no effect on SOICR activation or store capacity (Extended Data Fig. 7d, e).
We also assessed the functional importance of CaM residues near the RyR2–CaM interfaces. Mutations in CaM near the apo-CaM N-lobe–RyR2(K2153/Y2156) interface (CaM(E15A), CaM(F66A) and CaM(L70A)), near the apo-CaM C-lobe–RyR2(V3599) interface (CaM(M110A) and CaM(F142A)), near the Ca2+-CaM N-lobe–RyR2(V3599) interface (CaM(F20A) and CaM(F69A)) and near the Ca2+-CaM C-lobe–RyR2(W3587/L3590) interface (CaM(F93A), CaM(L106A) and CaM(M146A))—as with CaM-M—significantly reduced the effect of CaM on Ca2+ release termination compared to CaM(WT) (Extended Data Fig. 7f). All CaM mutations except for CaM(F20A) and CaM(F142A) had little or no effect on SOICR activation or store capacity (Extended Data Fig. 7g, h). Note that some mutations in CaM may induce conformational changes, thus affecting CaM–RyR2 interactions allosterically. Collectively, these functional studies support the importance of the newly identified RyR2–CaM interfaces in CaM regulation of RyR2.
Ca2+-dependent shift in CaM-binding sites on RyR2
The location and conformation of CaM-M and apo-CaM are identical in the structures (Fig. 3a), which suggests that the positional switch for apo-CaM and Ca2+-CaM results from the distinct conformations of CaM after Ca2+ loading instead of a direct effect of Ca2+ on RyR2. The CaM lobes have previously been reported in three conformations—open, semi-open and closed—when interacting with other proteins32,33,34. Comparative docking analysis of these conformations into the electron microscopy reconstruction for FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM suggests an open conformation for the N-lobe (Extended Data Fig. 8a, b). An important distinction between the open and semi-open or closed C-lobes is the slightly larger helical angle between C1 and C4, both of which were resolved in the map and conformed to the open state (Extended Data Fig. 8c, d). These analyses are consistent with the observations in the crystal structure of RyR1 peptide-bound Ca2+-CaM28. Upon Ca2+ loading, the compact structure of apo-CaM is relaxed to Ca2+-CaM35, which slips down towards the N-terminal part of helix α−1, consistent with previous studies36.
a, Ca2+ binding to RyR2 is not responsible for the positional shift in apo-CaM and Ca2+-CaM. In the presence of 20 μM Ca2+, the binding site for CaM-M remains the same as for apo-CaM. The two structures are superimposed relative to CaM. b, Upon Ca2+ loading, the expansion of CaM structure may lead to steric hindrance between the N-lobe and HD1. The N-lobes are superimposed. Red arrows indicate directions of conformational changes from apo-CaM to Ca2+-CaM. c, The shift in the CaM-binding site is accompanied by marked conformational changes in helices α−1 and 12 in RyR2. Red arrows indicate the conformational changes in RyR2 from FKBP12.6/apo-CaM to FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM. d, In FKBP12.6/apo-CaM, helix α−1 contacts helix α9. e, In FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM, helix α−1 of RyR2 is positioned away from helix α9. Helix N3 in the N-lobe contacts helix α9 instead. f, Helix α−1 of RyR2 serves as an essential anchor for CaM. Red arrow indicates the direction of conformational change of helix α−1 from FKBP12.6/apo-CaM to FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM.
Helix α−1 and helix 12 are two helices that were only resolved in the structures with CaM, probably owing to stabilization of these segments through CaM binding. Pronounced shifts in these two helices are observed between the structures bound to apo-CaM and Ca2+-CaM. In the presence of Ca2+-CaM, helix α−1 swings by around 90° and the N-terminal half of helix 12 also bends by nearly 90° (Fig. 3c). Accordingly, helix α−1, which contacts helix α9 of the central domain in FKBP12.6/apo-CaM, is positioned away from helix α9 in FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM. Now helix α9 is in contact with helix 3 in the N-lobe of Ca2+-CaM (Fig. 3d–f). The CaM-bound RyR2 structures shown here reveal that helix α−1 serves as an essential anchor for CaM (Fig. 3f).
Inhibitory modulation of RyR2 by Ca2+-CaM
Both caffeine and ATP are located at the interfaces between the U-motif and O-ring, locking them into a stable unit that stabilizes the open state (Extended Data Fig. 9a). Caffeine and ATP counteract inhibitory effect of Ca2+-CaM on RyR2, manifested by the lack of intradomain change of individual central domains. The pore remains open (Extended Data Figs. 1e, 9b). Nevertheless, Ca2+-CaM induces an anticlockwise rotation of the central domains in the cytoplasmic view in the same direction as that from the open to the closed state30,37 (Extended Data Fig. 9c and Supplementary Video 1). The central domains undergo similar shifts in the presence of high concentrations of Ca2+ and CaM, but not in the CHAPS plus DOPC condition (Extended Data Fig. 10a, b).
By contrast, RyR2 activated by PCB95 and Ca2+ is closed after addition of Ca2+-CaM (Fig. 1d). Detailed structural examination shows an anticlockwise rotation of the central domains and outward motions of the auxiliary motifs of the individual central domain, including helices α0, α1, α4 and the U-motif, with respect to the centre of the concave surface (Extended Data Figs. 9d, 10c). The motion of the U-motif appears to release the pulling force for the dilation of the S6 helix, resulting in closure of the pore (Extended Data Fig. 9e). Taken together, these results indicate that the inhibitory force of Ca2+-CaM is sufficient to overcome the synergistic activation of RyR2 by PCB95 and Ca2+, but not by the collective effect of ATP, caffeine and Ca2+ (Fig. 4).
The two RyR2 structures on the left were obtained in the presence of CHAPS and DOPC instead of digitonin, which was used for all other structural determinations. Despite a rotation (indicated by red arrows) of the central domains, the pore of ATP, caffeine and Ca2+-activated RyR2 channel remains open in the presence of Ca2+-CaM under these two conditions (left four). By contrast, Ca2+-CaM leads to closure of PCB95 and Ca2+-activated RyR2 channel (right two). Our study demonstrates that the inhibitory force of Ca2+-CaM can overcome the synergistic activation by PCB95 and Ca2+ but not by ATP, caffeine and Ca2+ (bottom right). The top right structure PCB95/low-Ca2+ was obtained from a previous study (RCSB Protein Data Bank (PDB) code 5GOA).
Discussion
Among the reported RyR2 sequences for CaM-binding38, residues 4247–4277 are invisible in our structure. The segment (residues 2023–2039) that contains the protein kinase A phosphorylation site Ser2032 is resolved in our structures, but shows no interaction with CaM even at high concentrations39. The other two sequences (1942–1966 and 3582–3608) bind to CaM in our structures (Extended Data Fig. 10d). Despite the distinct effects of apo-CaM on RyR1 and RyR214, the primary apo-CaM-binding sequences are invariant in these two isoforms (Extended Data Fig. 10e). The molecular determinants for the functional difference are yet to be revealed.
The inhibitory mechanisms by which CaM regulates RyR2 must be investigated on an already opened channel by structural biology. As the presence of micromolar-range Ca2+ is required for opening ryanodine receptors by cryo-EM studies30,31,32,37, it is impractical to obtain an open structure in the absence of micromolar concentrations of Ca2+ (apo-CaM form). Although the location and conformation of CaM-M appear identical to those of apo-CaM, it has previously been reported that modulation of RyR2 by CaM-M is distinct from that by apo-CaM40—although the mechanism needs to be investigated further.
Owing to extensive interactions between the U-motif and O-ring, the two undergo coupled motions during channel gating30,37. The presence of caffeine and ATP locks them into a more rigid structure, probably increasing the energy barrier for inhibiting RyR2 by Ca2+-CaM. By contrast, the PCB95- and Ca2+-activated RyR2 channel can be effectively closed by Ca2+-CaM. Therefore, the gating state of RyR2 is defined by the combined effect of competing stimulatory and inhibitory regulators (Fig. 4). It remains to be investigated whether the conclusions presented here can be recapitulated for other ryanodine receptor isoforms or in lipid bilayers and the relevance to disease-related mutations (Extended Data Fig. 10f, g).
Methods
Expression and purification of GST–FKBP12.6
Because the sequence of porcine FKBP12.6 is not available in the public domain, human FKBP12.6 was applied to pull-down porcine RyR2 (pRyR2)30. The complementary DNA of full-length human FKBP12.6 (also known as FKBP1B) was cloned into the pGEX-4T-2 vector with a C-terminal 6×His tag and an N-terminal glutathione S-transferase (GST) tag. Protein was overexpressed in the Escherichia coli BL21 (DE3) strain at 18 °C for 12–15 h after the addition of 0.2 mM isopropyl-β-d-thiogalactoside (IPTG) to cells with an optical density at 600 nm (OD600) of 1.0. Cells collected by centrifugation were resuspended in lysis buffer (25 mM Tris, pH 8.0, 150 mM NaCl). Cell debris was removed by centrifugation at 22,000g for 1 h, and the supernatant was applied to Ni2+-NTA resin (Qiagen). The resin was washed with both W1 buffer (25 mM Tris, pH 8.0, 500 mM NaCl) and W2 buffer (25 mM Tris, pH 8.0, 20 mM imidazole) and eluted with 25 mM Tris, pH 8.0 and 300 mM imidazole. The elution was further purified by anion-exchange chromatography (SOURCE 15Q, GE Healthcare).
Expression and purification of the wild-type CaM and CaM mutant
In mammals, three independent genes (CALM1–CALM3) with approximately 80% identity42 are transcribed into at least eight mRNAs that encode identical CaM proteins43. It has previously been reported that the first methionine residue of CaM was removed under physiological conditions44. The complementary DNA of human CALM3 without the initial Met was cloned into the pET21 vector with an N-terminal 6×His tag followed by an N-terminal SUMO tag and a stop codon in the C terminus, preventing the translation of a C-terminal 6×His tag in the original pET21 vector (Extended Data Fig. 1a). The expression and purification protocol was similar to that of GST–FKBP12.6 mentioned above. Specifically, the N-terminal 6×His tag and SUMO tag were removed together by the SUMO protease UlP1p45 during purification. The CaM protein was further purified by anion-exchange chromatography (SOURCE 15Q, GE Healthcare) using buffer 1 (25 mM Tris, pH 8.0) and buffer 2 (1 M NaCl, 25 mM Tris, pH 8.0). Finally, the protein was applied to size-exclusion chromatography (SEC; Superdex-200, GE Healthcare) in buffer F (20 mM HEPES, pH 7.4, 200 mM NaCl, 0.1% digitonin, 1.3 μg ml−1 aprotinin, 1 μg ml−1 pepstatin, 5 μg ml−1 leupeptin, 0.2 mM PMSF and 2 mM DTT), which is the same as that used for the last-step purification of RyR2. The N-terminal boundary of wild-type CaM was confirmed by N-terminal sequencing (Extended Data Fig. 1b). The expression and purification of the CaM mutant that is deficient in Ca2+ binding at all four EF-hand Ca2+-binding sites (E32A, E68A, E105A and E141A) (denoted as CaM-M) were the same as for the wild-type CaM.
Preparation of sarcoplasmic reticulum membranes from porcine heart
The procedures for preparing the membranes of the sarcoplasmic reticulum from porcine hearts were similar to previously described procedures30. A single porcine heart was cut into small pieces and then resuspended in five volumes of homogenization buffer A (20 mM HEPES, pH 7.4, 150 mM NaCl, 5 mM EDTA, 1.3 μg ml−1 aprotinin, 1 μg ml−1 pepstatin, 5 μg ml−1 leupeptin and 0.2 mM PMSF). Homogenization was performed in a blender (JYL-C010, Joyoung) for fifteen cycles. The debris was removed by low-speed centrifugation (6,000g) for 10 min. The supernatant was further centrifuged at high speed (20,000g) for 1 h. The pellet was then resuspended in two volumes of homogenization buffer B (20 mM HEPES, pH 7.4, 1 M NaCl, 1.3 μg ml−1 aprotinin, 1 μg ml−1 pepstatin, 5 μg ml−1 leupeptin, 0.2 mM PMSF and 2 mM DTT) and flash-frozen in liquid nitrogen.
Purification of pRyR2 by GST–FKBP12.6
The pRyR2–FKBP12.6 complex was purified based on previously described procedures30 with slight modifications. The membrane of the sarcoplasmic reticulum from a single porcine heart was solubilized at 4 °C for 2 h in homogenization buffer B supplemented with 5% CHAPS and 1.25% soy bean lecithin. After solubilization, the final concentration of NaCl in the system was diluted to 200 mM by homogenization buffer B without NaCl. Approximately 5–6 mg of purified GST–FKBP12.6 was then added to the system and further incubated for 1 h at 4 °C. After ultrahigh-speed centrifugation (200,000g), the supernatant was loaded onto a GS4B column (GE Healthcare). The resin was washed with buffer similar to the homogenization buffer B, except that the NaCl concentration was 200 mM and 0.1% digitonin was added. The complex was eluted by a solution containing 80 mM Tris, pH 8.0, 200 mM NaCl, 10 mM GSH, 0.1% digitonin, 1.3 μg ml−1 aprotinin, 1 μg ml−1 pepstatin, 5 μg ml−1 leupeptin, 0.2 mM PMSF and 2 mM DTT. The eluted protein was further purified through SEC (Superose 6, 10/300 GL, GE Healthcare) in buffer F. The pRyR2–FKBP12.6 complex fractions were concentrated to approximately 0.1 mg ml−1 for electron microscopy sample preparation. Specifically, for the FKBP12.6/apo-CaM sample, 5 mM EDTA, which has no effect on the zinc finger structure of RyR230, was included throughout purification of RyR2. For the CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+] and CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM samples, the proteins were extracted only by CHAPS and washed and eluted by a buffer containing 0.5% CHAPS plus 0.002% DOPC. The eluted proteins were further purified through SEC in buffer F except 0.1% digitonin was replaced by 0.25% CHAPS plus 0.001% DOPC. For the PCB95/low-[Ca2+]/Ca2+-CaM sample, the proteins were purified using GST–FKBP12 as a bait and the RyR2–FKBP12 (containing GST–FKBP12) complex fell apart during SEC purification30.
Cryo-EM sample preparation
The cryo-EM samples of RyR2–CaM complexes were prepared as follows. FKBP12.6/apo-CaM: 5 mM EDTA was added to CaM (in buffer F) before sample preparation, and CaM with a final concentration of 250 μM was added to RyR2 (in buffer F plus 5 mM EDTA). FKBP12.6/ATP/caffeine/low-[Ca2+]/CaM-M: CaM-M (in buffer F) and RyR2 (in buffer F) were separately added with 20 μM Ca2+ (low-Ca2+ concentration), 5 mM ATP and 5 mM caffeine and CaM-M with a final concentration of 250 μM was added to RyR2. FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM: CaM (in buffer F) and RyR2 (in buffer F) were separately added to 20 μM Ca2+, 5 mM ATP and 5 mM caffeine and CaM with a final concentration of 2.5 μM was added to RyR2. The other samples were prepared by the same procedure. Note that 5 mM Ca2+ represents high-Ca2+ concentration. Vitrobot Mark IV (FEI) was used for the preparation of cryo-EM grids. The procedures for preparing the eight samples were the same. Aliquots (3 μl each) of pRyR2 samples were placed on glow-discharged lacey carbon grids (Ted Pella). Grids were blotted for 2 s and flash-frozen in liquid ethane. Owing to the presence of high concentrations of Ca2+ in the filter paper used for blotting, it has previously been reported that the final concentration of free Ca2+ may be much higher than those used during sample preparation46. The low- and high-Ca2+ concentrations presented here only indicate those used during sample preparation and may be lower than the true Ca2+ concentrations.
Cryo-EM image acquisition
With regard to the FKBP12.6/ATP/caffeine/low-[Ca2+], FKBP12.6/ATP/caffeine/low-[Ca2+]/CaM-M, CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+], CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM, FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM and PCB95/low-[Ca2+]/Ca2+-CaM datasets, grids were transferred to a Titan Krios (Thermo Fisher Scientific) electron microscope operating at 300 kV equipped with a Cs-corrector (Thermo Fisher Scientific), Gatan K2 Summit detector and GIF Quantum energy filter. Zero-loss movie stacks were automatically collected using AutoEMationII47,48 with a slit width of 20 eV on the energy filter and a defocus range from −1.3 μm to −1.7 μm in super-resolution mode at a nominal magnification of 105,000×. Each stack was exposed for 5.6 s with an exposure time of 0.175 s per frame, resulting in 32 frames per stack. The total dose was approximately 50 e− Å−2 for each stack. The stacks were motion-corrected with MotionCor249 and binned twofold, resulting in a pixel size of 1.091 Å per pixel. With regard to the FKBP12.6/apo-CaM and FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM datasets, micrographs were collected using a Gatan K2 Summit detector mounted on a Titan Krios electron microscope (FEI Company) operating at 300 kV and equipped with a GIF Quantum energy filter (slit width 20 eV). Micrographs were recorded in the super-resolution mode with a normal magnification of 105,000 ×, resulting in a calibrated pixel size of 0.669 Å. Each stack of 32 frames was exposed for 8 s, with an exposing time of 0.25 s per frame. The total dose rate was about 45.6 e− Å−2 for each stack. All 32 frames in each stack were motion-corrected with MotionCor2 and binned to a pixel size of 1.338 Å. The defocus value of each image was set from −0.8 μm to −1.8 μm. In addition, dose weighting was performed50. The defocus values were estimated with Gctf51.
Image processing
Image-processing procedures were similar to those previously reported30. Diagrams of the procedures used in data processing are presented in Extended Data Fig. 2. For the FKBP12.6/apo-CaM dataset, 1,180,104 particles were picked from 7,800 micrographs by RELION 2.052 using templates low-pass-filtered to 20 Å to limit reference bias. After two rounds of two-dimensional classification, 832,833 particles were selected and subjected to global angular search three-dimensional classification using RELION 2.0 with one class and a step size of 7.5°. The electron microscopy map of the previously published open structure of RyR230, which was low-pass-filtered to 60 Å, was used as the initial model. After global angular search three-dimensional classification, the particles were further subjected to three-dimensional classification with 10 classes and a local angular search step of 3.75°. The local angular search three-dimensional classification was performed several times with the output from different iterations of the global angular search three-dimensional classification as input. After the merging of all good classes and removal of the duplicated particles, the particles were subjected to three-dimensional autorefinement using THUNDER software53. The final particle number for the three-dimensional autorefinement was 208,715, resulting in a 3.6 Å resolution map after post-processing. The same procedures were performed for the other datasets. The resolution was estimated with the gold-standard Fourier shell correlation 0.143 criterion54 with the high-resolution noise-substitution method55.
Model building and structure refinement
The model of the RyR2 open structure (PDB code 5GOA)30 was fitted into the maps of the eight conditions by Chimera56 and manually adjusted in COOT57. FKBP12 from the rabbit RyR1/FKBP12 complex structure (PDB code 3J8H)58 was used for homologous model building of FKBP12.6. The apo-CaM from the crystal structure 3WFN was fitted into the maps obtained in the presence of CaM-M or apo-CaM and manually adjusted in COOT. Similarly, the crystal structure of Ca2+-CaM in complex with the RyR1 peptide (PDB code 2BCX) was fitted into the maps obtained in the presence of Ca2+-CaM and manually adjusted in COOT. Structure refinement was performed using PHENIX59 in real space with restrained secondary structure and geometry. The statistics of the three-dimensional reconstruction and model refinement are summarized in Extended Data Table 1.
Evaluation of the conformations of N- and C-lobes of CaM
Different conformations of N- and C-lobes were docked into the electron microscopy reconstruction for FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM using the ‘Fit in Map’ tool of Chimera, selecting the options that include ‘Real-time correlation’, ‘7-Å resolution of Use map simulated from atoms’, ‘Use only data above contour level from first map’, ‘Optimize correlation’, ‘Correlation calculated about mean data value’, ‘Allow rotation and shift’ and ‘Move whole molecules’.
Site-directed mutagenesis
Point mutations in mouse Ryr2 and in human CALM1 were generated with the overlap extension method using PCR. In brief, a EcoRV/HpaI DNA fragment containing the RyR2(Y2156A) mutation and an AgeI/SalI fragment containing the RyR2(V3599A), RyR2(W3587A) or RyR2(L3590A) mutations were obtained by overlapping PCR and used to replace the corresponding wild-type fragment in the NheI/BsiWI fragment of Ryr2, The mutated NheI/BsiWI fragment was then used to replace the corresponding wild-type fragment in the full-length Ryr2 cDNA in pcDNA5. A HindIII/XhoI full-length CALM1 DNA fragment containing various point mutations was generated by overlapping PCR, which was then subcloned into pcDNA3. All point mutations in Ryr2 and CALM1 were confirmed by DNA sequencing.
Generation of stable, inducible cell lines expressing RyR2(WT) and mutants
Stable, inducible HEK293 cell lines expressing RyR2(WT), RyR2(Y2156A), RyR2(V3599A), RyR2(W3587A) and RyR2(L3590A) were generated using the Flp-In T-REx Core Kit from Invitrogen. These cell lines were not authenticated. These cells tested negative for mycoplasma contamination. In brief, Flp-In T-REx HEK293 cells were co-transfected with the inducible expression vector pcDNA5/FRT/TO containing the Ryr2WT or Ryr2-mutant cDNA and the pOG44 vector encoding the Flp recombinase in 1:5 ratios using the calcium phosphate precipitation method. The transfected cells were washed with phosphate buffered saline (PBS; 137 mM NaCl, 8 mM Na2HPO4, 1.5 mM KH2PO4 and 2.7 mM KCl, pH 7.4) 24 h after transfection followed by a change into fresh medium for 24 h. The cells were then washed again with PBS, collected and plated onto new dishes. After the cells had attached (around 4 h), the growth medium was replaced with a selection medium containing 200 μg ml−1 hygromycin (Invitrogen). The selection medium was changed every 3–4 days until the desired number of cells was grown. The hygromycin-resistant cells were pooled, aliquoted (1 ml) and stored at −80 °C. These positive cells are believed to be isogenic, because the integration of Ryr2 cDNA is mediated by the Flp recombinase at a single FRT site.
Single-cell luminal Ca2+ imaging
Luminal Ca2+ levels in RyR2WT- or Ryr2-mutant-expressing HEK293 cells transfected with or without CaM(WT) or CaM mutants were measured using single-cell Ca2+ imaging and the fluorescence resonance energy transfer (FRET)-based endoplasmic-reticulum luminal Ca2+-sensitive chameleon protein D1ER as previously described60,61. The cells were grown to 95% confluence in a 75-cm2 flask, dissociated with PBS (137 mM NaCl, 8 mM Na2HPO4, 1.5 mM KH2PO4 and 2.7 mM KCl, pH 7.4) and plated on glass coverslips placed on tissue culture dishes at approximately 10% confluence 18–20 h before transfection with cDNA for D1ER and cDNAs for CaM(WT) or CaM mutants using the calcium phosphate precipitation method. After transfection for 24 h, the growth medium was then changed to an induction medium containing 1 μg ml−1 tetracycline. After induction for around 22 h, the coverslip was mounted onto an inverted microscope (Nikon TE2000-S) and the cells on the coverslip were perfused continuously with Krebs–Ringer–HEPES buffer (125 mM NaCl, 5 mM KCl, 1.2 mM KH2PO4, 6 mM glucose, 1.2 mM MgCl2 and 25 mM HEPES, pH 7.4) containing various concentrations of CaCl2 (0, 1 and 2 mM) to induce SOICR, followed by the addition of 1.0 mM tetracaine, which was used to estimate the store capacity, and caffeine (20 mM), which was used to estimate the minimum store level by depleting the endoplasmic-reticulum Ca2+ stores at room temperature (23 °C). Images were captured with Compix Simple PCI 6 software every 2 s using the Nikon TE2000-S inverted microscope equipped with an S-Fluor 20×/0.75 NA objective. The filters used for D1ER imaging were λex = 436 ± 20 nm for CFP and λex = 500 ± 20 nm for YFP, and λem = 465 ± 30 nm for CFP and λem = 535 ± 30 nm for YFP with a dichroic mirror (500 nm). The amount of FRET in individual cells was determined from the ratio of the light emission at 535 and 465 nm. FSOICR is defined as the FRET level at which SOICR occurs, and Ftermi is defined as the FRET level at which SOICR terminates. The maximum FRET signal Fmax is defined as the FRET level after tetracaine treatment. The minimum FRET signal Fmin is defined as the FRET level after caffeine treatment. The termination and activation thresholds of SOICR in individual cells were determined using the equations shown in Extended Data Fig. 7a. The store capacity is calculated by subtracting Fmin from Fmax. Individual data points represent the average measurements of around 10–30 cells from one coverslip in one set of experiment. The number of experiments and coverslips for each condition is used as the sample size for data analyses.
Single-channel recordings in planar lipid bilayers
Recombinant RyR2(WT) and mutant channels were purified from cell lysates prepared from HEK293 cells transfected with the Ryr2WT or Ryr2-mutant (Y2156A, V3599A, W3587A or L3590A) cDNA by sucrose density gradient centrifugation as previously described62. Heart phosphatidylethanolamine (50%) and brain phosphatidylserine (50%) (Avanti Polar Lipids), dissolved in chloroform, were combined and dried under nitrogen gas and resuspended in 30 μl of n-decane at a concentration of 12 mg lipid per ml. Bilayers were formed across a 250-μm hole in a Delrin partition separating two chambers. The trans chamber (800 μl) was connected to the head stage input of an Axopatch 200A amplifier (Axon Instruments). The cis chamber (1.2 ml) was held at virtual ground. A symmetrical solution containing 250 mM KCl and 25 mM HEPES, pH 7.4 was used for all recordings, unless indicated otherwise. A 4-μl aliquot (around 1 μg protein) of the sucrose density gradient-purified recombinant RyR2(WT) or mutant channels was added to the cis chamber. Spontaneous channel activity was always tested for sensitivity to EGTA and Ca2+. The chamber to which the addition of EGTA inhibited the activity of the incorporated channel presumably corresponds to the cytosolic side of the Ca2+ release channel. The direction of single channel currents was always measured from the luminal to the cytosolic side of the channel, unless mentioned otherwise. Recordings were filtered at 2,500 Hz. Data analyses were carried out using the pCLAMP 8.1 software package (Axon Instruments). Free Ca2+ concentrations were calculated using a computer program that has previously been described63.
Statistical analysis
Data are mean ± s.e.m., derived from independent samples or independent experiments. All experiments were performed with at least five biological replicates. The GraphPad Prism 8.1 software was used to test for differences between groups. We used Student’s t-test (paired, two-tailed) or one-way ANOVA with a Dunnett’s post hoc test. P < 0.05 was considered to be statistically significant.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Atomic coordinates and electron microscopy density maps of the following structures have been deposited in the PDB (http://www.rcsb.org) and the Electron Microscopy Data Bank (EMDB https://www.ebi.ac.uk/pdbe/emdb/). FKBP12.6/apo-CaM (PDB, 6JI8; EMDB, EMD-9833), FKBP12.6/ATP/caffeine/low-[Ca2+]/CaM-M (PDB, 6JII; EMDB, EMD-9834), FKBP12.6/ATP/caffeine/low-[Ca2+] (PDB, 6JI0; EMDB, EMD-9831), FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM (PDB, 6JIU; EMDB, EMD-9836), CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+] (PDB, 6JRR; EMDB, EMD-9879), CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM (PDB, 6JRS; EMDB, EMD-9880), FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM (PDB, 6JIY; EMDB, EMD-9837) and PCB95/low-[Ca2+]/Ca2+-CaM (PDB, 6JV2; EMDB: EMD-9889) complexes. Source Data for Fig. 2e, f and Extended Data Figs. 1c, 6f, 7d–h are available in the online version of the paper. All other data are available from the corresponding authors upon reasonable request.
References
Fabiato, A. Calcium-induced release of calcium from the cardiac sarcoplasmic reticulum. Am. J. Physiol. 245, C1–C14 (1983).
Nakai, J. et al. Primary structure and functional expression from cDNA of the cardiac ryanodine receptor/calcium release channel. FEBS Lett. 271, 169–177 (1990).
Otsu, K. et al. Molecular cloning of cDNA encoding the Ca2+ release channel (ryanodine receptor) of rabbit cardiac muscle sarcoplasmic reticulum. J. Biol. Chem. 265, 13472–13483 (1990).
Rodney, G. G., Williams, B. Y., Strasburg, G. M., Beckingham, K. & Hamilton, S. L. Regulation of RYR1 activity by Ca2+ and calmodulin. Biochemistry 39, 7807–7812 (2000).
Timerman, A. P. et al. The ryanodine receptor from canine heart sarcoplasmic reticulum is associated with a novel FK-506 binding protein. Biochem. Biophys. Res. Commun. 198, 701–706 (1994).
Yamaguchi, N., Xu, L., Pasek, D. A., Evans, K. E. & Meissner, G. Molecular basis of calmodulin binding to cardiac muscle Ca2+ release channel (ryanodine receptor). J. Biol. Chem. 278, 23480–23486 (2003).
Laitinen, P. J. et al. Mutations of the cardiac ryanodine receptor (RyR2) gene in familial polymorphic ventricular tachycardia. Circulation 103, 485–490 (2001).
Medeiros-Domingo, A. et al. The RYR2-encoded ryanodine receptor/calcium release channel in patients diagnosed previously with either catecholaminergic polymorphic ventricular tachycardia or genotype negative, exercise-induced long QT syndrome: a comprehensive open reading frame mutational analysis. J. Am. Coll. Cardiol. 54, 2065–2074 (2009).
Priori, S. G. & Chen, S. R. Inherited dysfunction of sarcoplasmic reticulum Ca2+ handling and arrhythmogenesis. Circ. Res. 108, 871–883 (2011).
Priori, S. G. et al. Mutations in the cardiac ryanodine receptor gene (hRyR2) underlie catecholaminergic polymorphic ventricular tachycardia. Circulation 103, 196–200 (2001).
Hoeflich, K. P. & Ikura, M. Calmodulin in action: diversity in target recognition and activation mechanisms. Cell 108, 739–742 (2002).
Babu, Y. S. et al. Three-dimensional structure of calmodulin. Nature 315, 37–40 (1985).
Copley, R. R., Schultz, J., Ponting, C. P. & Bork, P. Protein families in multicellular organisms. Curr. Opin. Struct. Biol. 9, 408–415 (1999).
Balshaw, D. M., Xu, L., Yamaguchi, N., Pasek, D. A. & Meissner, G. Calmodulin binding and inhibition of cardiac muscle calcium release channel (ryanodine receptor). J. Biol. Chem. 276, 20144–20153 (2001).
Moore, C. P. et al. Apocalmodulin and Ca2+ calmodulin bind to the same region on the skeletal muscle Ca2+ release channel. Biochemistry 38, 8532–8537 (1999).
Tripathy, A., Xu, L., Mann, G. & Meissner, G. Calmodulin activation and inhibition of skeletal muscle Ca2+ release channel (ryanodine receptor). Biophys. J. 69, 106–119 (1995).
Fruen, B. R., Bardy, J. M., Byrem, T. M., Strasburg, G. M. & Louis, C. F. Differential Ca2+ sensitivity of skeletal and cardiac muscle ryanodine receptors in the presence of calmodulin. Am. J. Physiol. Cell Physiol. 279, C724–C733 (2000).
Tian, X., Tang, Y., Liu, Y., Wang, R. & Chen, S. R. Calmodulin modulates the termination threshold for cardiac ryanodine receptor-mediated Ca2+ release. Biochem. J. 455, 367–375 (2013).
Hino, A. et al. Enhanced binding of calmodulin to the ryanodine receptor corrects contractile dysfunction in failing hearts. Cardiovasc. Res. 96, 433–443 (2012).
Lavorato, M. et al. Dyad content is reduced in cardiac myocytes of mice with impaired calmodulin regulation of RyR2. J. Muscle Res. Cell Motil. 36, 205–214 (2015).
Yamaguchi, N. et al. Cardiac hypertrophy associated with impaired regulation of cardiac ryanodine receptor by calmodulin and S100A1. Am. J. Physiol. Heart Circ. Physiol. 305, H86–H94 (2013).
Yamaguchi, N., Takahashi, N., Xu, L., Smithies, O. & Meissner, G. Early cardiac hypertrophy in mice with impaired calmodulin regulation of cardiac muscle Ca release channel. J. Clin. Invest. 117, 1344–1353 (2007).
Kato, T. et al. Correction of impaired calmodulin binding to RyR2 as a novel therapy for lethal arrhythmia in the pressure-overloaded heart failure. Heart Rhythm 14, 120–127 (2017).
Huang, X., Fruen, B., Farrington, D. T., Wagenknecht, T. & Liu, Z. Calmodulin-binding locations on the skeletal and cardiac ryanodine receptors. J. Biol. Chem. 287, 30328–30335 (2012).
Samsó, M. & Wagenknecht, T. Apocalmodulin and Ca2+-calmodulin bind to neighboring locations on the ryanodine receptor. J. Biol. Chem. 277, 1349–1353 (2002).
Wagenknecht, T. et al. Locations of calmodulin and FK506-binding protein on the three-dimensional architecture of the skeletal muscle ryanodine receptor. J. Biol. Chem. 272, 32463–32471 (1997).
Yamaguchi, N., Xin, C. & Meissner, G. Identification of apocalmodulin and Ca2+-calmodulin regulatory domain in skeletal muscle Ca2+ release channel, ryanodine receptor. J. Biol. Chem. 276, 22579–22585 (2001).
Maximciuc, A. A., Putkey, J. A., Shamoo, Y. & Mackenzie, K. R. Complex of calmodulin with a ryanodine receptor target reveals a novel, flexible binding mode. Structure 14, 1547–1556 (2006).
Maune, J. F., Klee, C. B. & Beckingham, K. Ca2+ binding and conformational change in two series of point mutations to the individual Ca2+-binding sites of calmodulin. J. Biol. Chem. 267, 5286–5295 (1992).
Peng, W. et al. Structural basis for the gating mechanism of the type 2 ryanodine receptor RyR2. Science 354, aah5324 (2016).
des Georges, A. et al. Structural basis for gating and activation of RyR1. Cell 167, 145–157 (2016).
Wei, R. et al. Structural insights into Ca2+-activated long-range allosteric channel gating of RyR1. Cell Res. 26, 977–994 (2016).
Wang, C. et al. Structural analyses of Ca2+/CaM interaction with NaV channel C-termini reveal mechanisms of calcium-dependent regulation. Nat. Commun. 5, 4896 (2014).
Wang, C., Chung, B. C., Yan, H., Lee, S. Y. & Pitt, G. S. Crystal structure of the ternary complex of a NaV C-terminal domain, a fibroblast growth factor homologous factor, and calmodulin. Structure 20, 1167–1176 (2012).
Jurado, L. A., Chockalingam, P. S. & Jarrett, H. W. Apocalmodulin. Physiol. Rev. 79, 661–682 (1999).
Rodney, G. G. et al. Calcium binding to calmodulin leads to an N-terminal shift in its binding site on the ryanodine receptor. J. Biol. Chem. 276, 2069–2074 (2001).
Bai, X. C., Yan, Z., Wu, J., Li, Z. & Yan, N. The central domain of RyR1 is the transducer for long-range allosteric gating of channel opening. Cell Res. 26, 995–1006 (2016).
Brohus, M., Søndergaard, M. T., Chen, S. R. W., van Petegem, F. & Overgaard, M. T. Ca2+-dependent calmodulin binding to cardiac ryanodine receptor (RyR2) calmodulin-binding domains. Biochem. J. 476, 193–209 (2019).
Xiao, B. et al. Characterization of a novel PKA phosphorylation site, serine-2030, reveals no PKA hyperphosphorylation of the cardiac ryanodine receptor in canine heart failure. Circ. Res. 96, 847–855 (2005).
Fruen, B. R. et al. Regulation of the RYR1 and RYR2 Ca2+ release channel isoforms by Ca2+-insensitive mutants of calmodulin. Biochemistry 42, 2740–2747 (2003).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Fischer, R. et al. Multiple divergent mRNAs code for a single human calmodulin. J. Biol. Chem. 263, 17055–17062 (1988).
Kortvely, E. & Gulya, K. Calmodulin, and various ways to regulate its activity. Life Sci. 74, 1065–1070 (2004).
Sasagawa, T. et al. Complete amino acid sequence of human brain calmodulin. Biochemistry 21, 2565–2569 (1982).
Hirano, H., Kobayashi, J. & Matsuura, Y. Structures of the karyopherins Kap121p and Kap60p bound to the nuclear pore-targeting domain of the SUMO protease Ulp1p. J. Mol. Biol. 429, 249–260 (2017).
Paknejad, N. & Hite, R. K. Structural basis for the regulation of inositol trisphosphate receptors by Ca2+ and IP3. Nat. Struct. Mol. Biol. 25, 660–668 (2018).
Fan, X. et al. Near-atomic resolution structure determination in over-focus with volta phase plate by Cs-corrected cryo-EM. Structure 25, 1623–1630 (2017).
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Hu, M. et al. A particle-filter framework for robust cryo-EM 3D reconstruction. Nat. Methods 15, 1083–1089 (2018).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Yan, Z. et al. Structure of the rabbit ryanodine receptor RyR1 at near-atomic resolution. Nature 517, 50–55 (2015).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Palmer, A. E., Jin, C., Reed, J. C. & Tsien, R. Y. Bcl-2-mediated alterations in endoplasmic reticulum Ca2+ analyzed with an improved genetically encoded fluorescent sensor. Proc. Natl Acad. Sci. USA 101, 17404–17409 (2004).
Jones, P. P. et al. Endoplasmic reticulum Ca2+ measurements reveal that the cardiac ryanodine receptor mutations linked to cardiac arrhythmia and sudden death alter the threshold for store-overload-induced Ca2+ release. Biochem. J. 412, 171–178 (2008).
Jiang, D. et al. Enhanced store overload-induced Ca2+ release and channel sensitivity to luminal Ca2+ activation are common defects of RyR2 mutations linked to ventricular tachycardia and sudden death. Circ. Res. 97, 1173–1181 (2005).
Fabiato, A. & Fabiato, F. Calculator programs for computing the composition of the solutions containing multiple metals and ligands used for experiments in skinned muscle cells. J. Physiol. 75, 463–505 (1979).
Acknowledgements
We thank X. Li for technical support for electron microscopy image acquisition; the Tsinghua University Branch of China National Center for Protein Sciences (Beijing) for providing the cryo-EM facility support; the computational facility support on the cluster of Bio-Computing Platform (Tsinghua University Branch of China National Center for Protein Sciences Beijing) and the ‘Explorer 100’ cluster system of Tsinghua National Laboratory for Information Science and Technology; M. T. Overgaard for providing the CaM protein for single channel studies. This work was funded by the National Key R&D Program (2016YFA0500402) and the National Key Basic Research (973) Program (2015CB910101) from Ministry of Science and Technology of China and the National Natural Science Foundation of China (projects 31621092, 31630017 and 81861138009). N.Y. is supported by the Shirley M. Tilghman endowed professorship from Princeton University. This work was also supported by research grants from the Heart and Stroke Foundation of Canada, the Canadian Institutes of Health Research and the Heart and Stroke Foundation Chair in Cardiovascular Research (S.R.W.C.).
Author information
Authors and Affiliations
Contributions
D.G. and N.Y. conceived the project. D.G., X.C. and G.Z. prepared the electron microscopy samples. D.G., X.C., G.H. and J.L. conducted the cryo-EM analysis. J.W., L.Z. and R.W. performed the functional experiments. All authors contributed to data analysis. D.G., S.R.W.C. and N.Y. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Protein purification and structural determination.
a, Schematic of vector construction for recombinant expression of CaM without the N-terminal Met. b, N-terminal sequencing confirmed removal of the initial Met. c, SEC purification of the affinity-purified complex of pRyR2–FKBP12.6 (containing GST–FKBP12.6). The experiment was repeated five times independently with similar results. Peak fractions were resolved by SDS–PAGE and visualized by Coomassie blue staining (Supplementary Fig. 1). UV, ultraviolet. MWM, molecular weight marker. d, The channel is open in the presence of ATP, caffeine and Ca2+ under both digitonin and CHAPS-and-DOPC (indicated by an asterisk) conditions. The pore of CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM remains open after Ca2+-CaM loading. The ion-conduction path, calculated by HOLE, is illustrated by dots in each structure. A, ATP; C, caffeine; Ca2+-CaM, Ca2+-bound CaM; CaM-M, a Ca2+-binding-deficient CaM mutant that mimics apo-CaM; F, FKBP12.6; L-Ca2+, low Ca2+ concentration. e, Overall electron microscopy map of the FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM complex. Inset, the cytoplasmic view of the channel gate. Because the side chains of the Gln4864-gating residues are not well-resolved, the distance between the Cα atoms of Gln4864 gating in the diagonal protomers is shown in the dashed circle. The density corresponding to CaM was generated from the map that was low-pass-filtered to 5.6 Å with a contour level of 0.015; the other regions were from the 4.2 Å map with a contour level of 0.023. f, Overall electron microscopy map of the CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM complex. The density corresponding to CaM was generated from the map that was low-pass-filtered to 5.6 Å with a contour level of 0.013; the other regions were from the 3.7 Å map with a contour level of 0.021. g, Although the concentrations of Ca2+-CaM are the same in these three conditions, only the N-lobe can be resolved in the FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM and CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM RyR2 structures. The reason may be that the HD2 in these two structures presents a steric hindrance for C-lobe binding. P, PCB95. h, The corresponding pore radii of the three structures are plotted. i, j, Gold-standard Fourier shell correlation curves for electron microscopy maps of the eight datasets. H-Ca2+, high Ca2+ concentration.
Extended Data Fig. 2 Flow chart for cryo-EM data processing.
See Methods for details. a, Data processing of the FKBP12.6/apoCaM, FKBP12.6/ATP/caffeine/low-[Ca2+]/CaM-M, CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+], and PCB95/low-[Ca2+]/Ca2+-CaM datasets. b, Data processing of the FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM, FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM, FKBP12.6/ATP/caffeine/low-[Ca2+], and CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM datasets.
Extended Data Fig. 3 Local resolution maps of the eight reconstructions.
a, c, e, g, i, k, l, n, The local resolution maps estimated with RELION 2.0. All electron microscopy maps were generated in Chimera and contoured at levels of 0.027 (a), 0.022 (c), 0.023 (e), 0.021 (g), 0.02 (i), 0.021 (k), 0.015 (l) and 0.021 (n). b, Electron microscopy map of apo-CaM from the reconstruction shown in a. d, Electron microscopy map of CaM-M. f, h, The electron microscopy densities of Ca2+-CaM were generated from the maps of FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM and CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM that were low-pass-filtered to 5.6 Å with contour levels of 0.015 (f) and 0.013 (h), respectively. j, m, Electron microscopy densities of Ca2+-CaM in FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM (j) and PCB95/low-[Ca2+]/Ca2+-CaM (m). The densities of both lobes were resolved in the map, although the N-lobe was resolved better than the C-lobe.
Extended Data Fig. 4 Representative local electron microscopy maps for FKBP12.6/apo-CaM and densities of the binding sites for Ca2+, ATP and caffeine in FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM.
a–i, The electron microscopy maps for the representative segments of RyR2. All of the maps were contoured at 5.5σ. j, The binding sites for Ca2+, ATP and caffeine in RyR2. The blue, dotted circle indicates the O-ring that is formed by the C-terminal subdomain (CTD), cytoplasmic subdomain in the voltage-sensor-like domain (VSC) and the cytoplasmic portion of S6 (S6C). Ca2+ is located in the cleft that is formed by the central domain and C-terminal subdomain. ATP is located in a pocket formed by the U-motif, C-terminal subdomain and S6C. Caffeine is located at the interface formed by the U-motif, helix α4, C-terminal subdomain and voltage-sensor-like domain. The red letter indicates a disease-causing variant. k–m, The local densities of the Ca2+-, ATP- and caffeine-binding sites. The electron microscopy maps of the FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM (+ATP, caffeine and Ca2+) and FKBP12.6/apo-CaM (−ATP, caffeine and Ca2+) RyR2 structures are shown. All of the electron microscopy maps were contoured at a level of 0.029.
Extended Data Fig. 5 Binding interfaces between CaM and RyR2.
a, Three interfaces are formed between the N-lobe of apo-CaM and RyR2. HD1 serves as the major binding site for the N-lobe. The Cα atom of Lys2558 is shown as a sphere. b, c, Two interfaces are formed between the C-lobe of apo-CaM and RyR2. Helix α−1 is the major binding site of the C-lobe. d, e, The interfaces between the N-lobe of Ca2+-CaM and RyR2. f, The interface between the C-lobe of Ca2+-CaM and RyR2. g, Local densities of the probable interacting residues, Tyr2157, Tyr2203, Arg2206 and Lys2154 in RyR2. The electron microscopy map was contoured at 5.5σ. h, Density of helix α−1 in the FKBP12.6/apo-CaM RyR2 structure. The sequence can be reliably assigned based on the indicated bulky residues. i, Density of helix α−1 in the FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM RyR2 structure. The C-terminal half of helix α−1 is reliably assigned, a few bulky residues facilitate the sequence alignment. As both the N-terminal half of helix α−1 and C-lobe of Ca2+-CaM had a lower resolution, the density shown here may belong to Trp3588. The electron microscopy maps in h and i were contoured at levels of 0.027 and 0.016, respectively.
Extended Data Fig. 6 Effect of RyR2 mutations on CaM regulation of single RyR2 channels.
a–e, Single-channel activities were recorded in a symmetrical recording solution containing 250 mM KCl and 25 mM HEPES, pH 7.4. Representative current traces of single RyR2(WT) (n = 9), RyR2(Y2156A) (n = 8), RyR2(V3599A) (n = 7), RyR2(W3587A) (n = 8) and RyR2(L3590A) (n = 9) channels are shown. The Ca2+ concentration on the cytoplasmic and luminal face of the channel was 440 nM and around 45 nM, respectively. Open probability (Po), mean open time (To) and mean closed time (Tc) of the same channel before and after addition of CaM(WT) (1 μM) are depicted. Baselines are indicated by short bars on the right. f, Percentages of inhibition of channel open probability by CaM. Data are mean ± s.e.m. from single RyR2(WT) (n = 9), RyR2(Y2156A) (n = 8), RyR2(V3599A) (n = 7), RyR2(W3587A) (n = 8) and RyR2(L3590A) (n = 9) channels and analysed by one-way ANOVA with a Dunnett’s post hoc test (versus RyR2(WT)) and adjusted P values are indicated on the graph.
Extended Data Fig. 7 Effect of RyR2 and CaM mutations on the termination of Ca2+ release in HEK293 cells.
HEK293 cell lines that express RyR2(WT) and RyR2 mutants were co-transfected with the FRET-based endoplasmic-reticulum luminal Ca2+-sensing protein D1ER and with no CaM (control), CaM(WT) or the Ca2+-binding-deficient CaM mutant, CaM-M. a–c, Representative single-cell luminal Ca2+ recordings of RyR2(WT) cells transfected with no CaM (a; n = 155 cells), RyR2(WT) cells transfected with CaM(WT) (b; n = 178 cells) and RyR2-WT cells transfected with CaM-M (c; n = 177 cells) are shown. FSOICR indicates the FRET level at which SOICR occurs and Ftermi represents the FRET level at which SOICR terminates. The signal Fmax is defined as the FRET level after tetracaine treatment. The minimum FRET signal maximum FRET Fmin is defined as the FRET level after caffeine treatment. d, The activation threshold was determined as shown in a. e, The store capacity was calculated by subtracting Fmin from Fmax. d, e, Data are mean ± s.e.m. with the number of independent experiments for each condition shown on the graph and analysed by one-way ANOVA with a Dunnett’s post hoc test (versus RyR2(WT) control) and adjusted P values are indicated. f–h, RyR2(WT) cells were co-transfected with the FRET-based endoplasmic-reticulum luminal Ca2+-sensing protein D1ER and CaM(WT) or CaM mutants (CaM-M, CaM(E15A), CaM(F66A), CaM(L70A), CaM(M110A), CaM(F142A), CaM(F20A), CaM(F69A), CaM(F93A), CaM(L106A) and CaM(M146A)). CaM mutations close to a specific CaM–RyR2 interface are grouped and are indicated. The termination threshold (f), activation threshold (g) and store capacity (h) were determined as described in a–e above. f–h, Data are mean ± s.e.m. with the number of independent experiments for each condition shown on the graph and analysed by one-way ANOVA with a Dunnett’s post hoc test (versus CaM(WT)) and adjusted P values are indicated.
Extended Data Fig. 8 Evaluation of the conformations of N- and C-lobes of CaM in the FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM structures.
a, The electron microscopy map (low-pass-filtered to 4.8 Å resolution at a contour level of 0.015) of FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM. Red and blue boxes indicate the N- and C-lobes, respectively. b, Docking of the three reported conformations of the N-lobe of CaM into our electron microscopy map suggests an open conformation of the N-lobe in FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM. CC indicates the cross-correlation coefficient. c, An important distinction between C-lobes in the open and semi-open or closed states is the enlarged angle between helices C1 and C4. d, Docking analysis supports the open conformation of the C-lobe in FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM.
Extended Data Fig. 9 Inhibitory mechanism of RyR2 by Ca2+-CaM.
a, Both caffeine and ATP are located at the interfaces between the U-motif and O-ring, locking these elements into a stable unit. b, There is almost no intradomain rearrangement of the individual central domain between the structures of FKBP12.6/ATP/caffeine/low-[Ca2+] and FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM. c, Ca2+-CaM induces anticlockwise rotation of the overall central domain in FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM when viewed from the cytoplasmic side. The overall tetrameric FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM and FKBP12.6/ATP/caffeine/low-[Ca2+] RyR2 structures are superimposed relative to C-terminal subdomain of the channel domain. Red arrows indicate the conformational changes upon Ca2+-CaM binding. d, Ca2+-CaM induces intradomain shifts of the individual central domain in the PCB95 and Ca2+-activated channel. e, The PCB95 and Ca2+-activated channel closes after Ca2+-CaM loading.
Extended Data Fig. 10 Conformational changes induced by Ca2+-CaM and mapping of previously identified CaM-binding sequences and disease-associated point mutations onto the structures of RyR2–CaM complexes.
a, Compared to FKBP12.6/ATP/caffeine/low-[Ca2+], the four central domains in the FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM RyR2 structure undergo an anticlockwise rotation. The overall tetrameric FKBP12.6/ATP/caffeine/low-[Ca2+] and FKBP12.6/ATP/caffeine/high-[Ca2+]/Ca2+-CaM RyR2 structures are superimposed relative to the C-terminal subdomain of the channel domain. b, Compared to the CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+] RyR2 structure, almost no conformational change was induced by Ca2+-CaM in the CHAPS- and DOPC-treated FKBP12.6/ATP/caffeine/low-[Ca2+]/Ca2+-CaM RyR2 structure. c, Compared to the PCB95/low-[Ca2+] RyR2 structure, the overall central domain in the PCB95/low-[Ca2+]/Ca2+-CaM RyR2 structure undergoes an anticlockwise rotation. The overall tetrameric PCB95/low-[Ca2+] and PCB95/low-[Ca2+]/Ca2+-CaM RyR2 structures are superimposed relative to the C-terminal subdomain of the channel domain. d, Structural mapping of previously reported CaM-binding sequences. Orange, the overlapping binding sequences of apo-CaM and Ca2+-CaM; cyan, the binding sequence of Ca2+-CaM; yellow, the binding sequence of apo-CaM; blue, segments that are not involved in binding in our structures; red, sequences that are invisible in the structures. The residue numbers in brackets that are labelled grey indicate that the sequences are invisible in the structure. e, The primary apo-CaM binding sequences in RyR2 are the same in RyR1. Red residues highlight the key contact residues. f, Mapping of the disease-associated point mutations onto the structure of the RyR2–apo-CaM complex. The mutations in HD1 and apo-CaM are coloured blue and red, respectively. g, Mapping of the CaM disease-associated point mutations onto the structure of the RyR2–Ca2+-CaM complex.
Supplementary information
Supplementary Information
Supplementary Figure 1 and Supplementary Table 1.
41586_2019_1377_MOESM3_ESM.mp4
Video 1 Rotation of the Central domains induced by Ca2+-CaM. Structures of the tetrameric F/A/C/L-Ca2+ and F/A/C/L-Ca2+/Ca2+-CaM are superimposed relative to the CTD of the Channel domain. Shown here are the four Central domains in the cytoplasmic view. The morph was generated in PyMol.
Rights and permissions
About this article
Cite this article
Gong, D., Chi, X., Wei, J. et al. Modulation of cardiac ryanodine receptor 2 by calmodulin. Nature 572, 347–351 (2019). https://doi.org/10.1038/s41586-019-1377-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1377-y
- Springer Nature Limited
This article is cited by
-
Metabolic remodelling in atrial fibrillation: manifestations, mechanisms and clinical implications
Nature Reviews Cardiology (2024)
-
Kinetics and mapping of Ca-driven calmodulin conformations on skeletal and cardiac muscle ryanodine receptors
Nature Communications (2024)
-
Structural basis for ryanodine receptor type 2 leak in heart failure and arrhythmogenic disorders
Nature Communications (2024)
-
Ca2+/calmodulin-mediated desensitization of glutamate receptors shapes plant systemic wound signalling and anti-herbivore defence
Nature Plants (2024)
-
Cysteines 1078 and 2991 cross-linking plays a critical role in redox regulation of cardiac ryanodine receptor (RyR)
Nature Communications (2023)