Abstract
Environmental impacts of xenobiotic compounds released to air, water, and soil have opened a way for bioremediation to emerge as a green technology that can be safely applied to reduce pollutant concentrations to a minimum in a relative short period of time. “Hard to break” molecules such as asphaltenes, celluloses, and dyes are better treated with mycoremediation techniques. Fungi are higher eukaryotic microorganisms that secrete a good quantity of enzymatic complexes to break covalent bonds on these xenobiotics. As mycoremediation analysis grew, a better understanding of fungal metabolism on extreme environmental conditions is needed to deepen bioremediation-related genes and processes. These can be reached by global molecular approaches such as transcriptomic studies. Until now, limited genomic functional annotations and efficient nucleic acid extractions in bioremediation processes are the main delaying issues in the advancing way to the understanding of these interesting fungal metabolic activities. In this section we expose common technical strategies of RNA extraction protocols and comparison of recent transcriptional studies, as a basic introduction to those interested in applying genomic global approaches, in this area in construction of future efficient application of mycoremediation.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
2.1 Introduction: Fungal Bioremediation (Mycoremediation)
Increasing industrial activities around the world is essential to sustain all countries’ economy and provide career options. Nowadays, there is a fourth industrial revolution taking place, and little is known about the details of how this new industrial approach will stablish to manage, contain, and dispose chemical and energy wastes (McMaster 2018). By these days, different kinds of pollutants have forever altered natural environments; it is in these affected areas that bioremediation has proven to be a valuable and adaptable set of techniques that can degrade, reduce, oxidize, or encapsulate hazardous materials such as hydrocarbons, heavy metals, pesticides, and radionuclide elements (Das 2014). Such techniques are focused on the stimulation of either macroorganisms (plants) (Bell et al. 2014) or microorganisms that possess specific metabolic pathways to create enzymes that would interact with the xenobiotic compounds to either cleave C:H, C:N, C:C, and C:S bonds for organic compounds (Ladino-Orjuela et al. 2016) or to reduce, precipitate, accumulate, sorb, or even leach metallic atoms and inorganic molecules by reducing its electrons (Nancharaiah et al. 2016). They do this around their surrounding environments. Microorganisms are vast; we call them to englobe the Protista, Fungi, and Protozoa kingdoms where mostly bacteria and fungi excel at being extensively used because of their good abundancy in soil and water and their efficient remediation rates related to their metabolic plasticity (Whitby 2010). Bioremediation studies had always focused on enhancing these rates by providing specific growth factors applying oxygen, nitrogen, and/or mineral sources and controlling pH, H2O levels, and temperature depending on the pollutant and the details of the environment. Bacteria such as Pseudomonas and Bacillus have proven to be very efficient with degradation of aromatic compounds, BTEX (benzene, toluene, ethylbenzene, and xylene), and short to medium hydrocarbon chains compounds. But they drop degradation rates at pollutants with higher molecular complexity such as crude oil, creosote, and tar. It is in these hard to break pollutants that fungal strains are used to provide better results for their eukaryotic genetics that produce higher enzymatic compounds (20–60 kDa) that can cleave these kinds of molecules (Deshmukh et al. 2016).
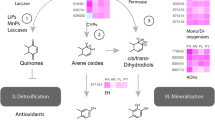
Within the Fungi kingdom, there is an estimation of 1.5 million species (Hawksworth 2001) classified in four or five phyla, being those belonging to the Ascomycota and to the Basidiomycota the most frequent in soil and employed for bioremediation (Tortella et al. 2005; Sankaran et al. 2010). Biological features of these fungi are the secretion of cellulolytic enzymes such as cellobiohydrolases (CBH), evolved beta-galactosidases (EBG), and β-glucosidases. Some of these cellulase enzyme complexes such as the lytic polysaccharide monooxygenases (LMPO) are widely described among the white-, brown-, and soft-rot fungi as the extracellular enzymes that decompose complex nutrients (pollutants) into simpler substances that can be easily assimilated through the cell wall (Obeng et al. 2017). By doing so, the fungus might produce antibiotics or other suppressive metabolites. This is difficult to demonstrate at the scale of individual hyphae, but Burton and Coley-Smith (1993) reported that antibacterial compounds were released by hyphae of Rhizoctonia species that are members of the Basidiomycota known to degrade cellulose. Another special aspect of fungi is that they do not fix nitrogen from the atmosphere; they rather use amino acids or, in extreme polluted soils with ammonia or ammonium (NH4) environments, they use them as a nitrogen source. After uptake, ammonia/ammonium is combined with organic acids, usually to produce either glutamic acid (from α-ketoglutaric acid) or aspartic acid; then the other amino acids needed can be formed by transamination reactions. Heavy metal bioremediation is an extended field as their concentration in mg/kg of many elements like Zn, Ni, Hg, Cr, Pb, Cu, Cd, As, Co, Sn, Au, Pd, Pt, Ag, Ru, Th, U, Am, and Ra could mean a serious environmental pollution, risking ecosystem and human health. Fortunately, organisms such as Saccharomyces cerevisiae and Rhizopus arrhizus have been used as a promising cleaning technology because of its high removal capacity and eco-friendly and cost-effective properties. Such cleaning effect relies on their exopolysaccharide (EPS) production that ends up excluding heavy metals (Saǧ et al. 1995; Ozer and Ozer 2003; Mohite et al. 2017). Other helpful enzymes like laccases, lignin peroxidases (LiP), and manganese peroxidases (MnP) had been reported to interact with other complex aromatic pollutants such as dyes (Vats and Mishra 2018). Nevertheless, all these approaches have not yet explained which metabolic routes are active at every specific moment of the bioremediation process, in order to guide (once well-known) the biodegradation process with elevated efficiency. Before the achievement of this goal, two main objectives must be contemplated: first, an efficient implementation of nucleic acid extraction technique, making it possible for -omics studies to develop this area in the presence of many other recalcitrant substances, and second a precise description of the fungi or fungal communities related with the process (this can be achieved by a metagenomic approach).
Nucleic acid extraction (DNA-RNA) techniques focus on creating a lysis effect on the membranes or cell walls from a pure culture to environmental samples. Fungal cell walls contain chitin, β-(1,3)-β-(1,4)- β-(1,6) glucans, and other proteins like melanin and mannan (Gow et al. 2017); such osmotic pressure effect can be applied on samples to release nucleic acids from such well-protected fungi cells by the use of chemical (SDS or phenol), physical (glass or zirconia beads, microwaves, sonication, or frozen/thaw cycles), and enzymatic methods (lysozyme, proteinase K). They can be performed by preparing all reactives or using commercial kits, like TRI Reagent kit (Sigma) that pre-ensembles this desired solutions in an easy-to-follow procedure (Moré et al. 1994; Boon et al. 2000; Nelson et al. 2007; Jiang et al. 2011). Sampling methodology and sample treatment are essential of a good nucleic acid yielding. Depending on the origin of the sample, we could find many inhibitors such as humic acids, metals, and xenobiotics that interfere with the extraction process (Wintzingerode et al. 1997; He et al. 2009). Sampling procedures for toxic environments may be contemplated (WHO 2004) to then proceed with the nucleic acid extraction protocols mentioned before. The best quality (less impurities) and quantity (μg of nucleic acid yield) of DNA and RNA come from commercial kits; nevertheless fungal nucleic acid may be better obtained from enzymatic/phenol-chloroform extraction methods as it is shown in the study by Guillén-Navarro et al. (2015) as they compare different extraction procedures. Preferred RNA extraction protocol could be decided by the sample circumstance, ribonucleic acid from a fresh mycelium on an enriched medium may be well obtained from a classic liquid nitrogen-buffer-phenol-chloroform extraction, but the ribonucleic acid from living fungi in a polluted environmental sample may get a better yield by any commercial RNA extraction kits.
2.2 Molecular Approaches in Fungal Bioremediation
It is clear that microbial metabolism can achieve specific pollutant bioremediation, thus enhancing polluted site diagnoses is vital to stablish the most adequate treatment. For accidental spillage of pollutants or catastrophic incidents that end up with impacted waters, atmosphere, and soils of natural or urban environments, most countries have specific contingency and normativity to be followed (National Research Council 2014). The use of microorganisms that can degrade and restore wellness is allowed following all protocols that clearly stipulate the use of native microbial communities or, if a biological product may be applied, is necessary to demonstrate the avoidance of GMOs; even these have yield excellent results in lab conditions (Ruta et al. 2017; Zhang et al. 2015). Recently mentioned genomic DNA extraction protocols can be applied at recent or longtime polluted sites to get a lecture of gene targets most commonly used for sequence-based fungal identification; these are the internal transcribed spacer regions ITS-1 and ITS-2, which are variable regions (the ITS region) and can be located between conserved genes encoding the 18S, 5.8S, and 28S ribosomal subunits. Another variable region is called the D1/D2 region, located toward the 5′ end of the large nuclear ribosomal subunit (28S rDNA). These regions can be amplified using ITS-1, ITS-4, NL-1, and NL-4 primers (Romanelli et al. 2014). Total microbial communities are seriously deplenished when toxic pollutants get at pristine water and soils (Rodrigues et al. 2015; Morais et al. 2016). If there is no good mixture of helpful microorganisms, then biological products may be of use; there are plenty of fungal products in the market (http://www.epa.gov/emergencies/content/ncp/index.htm). To mention some examples, there are the US patent numbers, 6,143,549; 6,383,800; 6,387,689; 6,387,691; 6,495,134; 6,664,102; 6,727,087; and 6,972,169, that belong to fungal product inventions to target pollutants like chromated copper arsenate (CCA), pentachlorophenol, ammoniacal copper quat (ACQ), and creosote. The modus operandi is that the fungal inoculum is applied to the polluted site and maintained in an aerated and hydrated environment having temperature conditions sufficient to allow the inoculum to grow and metabolize the xenobiotic compounds (Lamar et al. 2000; Illman et al. 2002a, b, c, d, 2003, 2004, 2005). Also, it is worthy to mention that fungal community, if it is the case, can change among the bioremediation process, so it is a good idea to follow up these changes associated with the expression of genes related to biodegradation process. If a single fungal strain is used, this description can be avoided.
For general bioremediation process then, having efficient molecular approaches for both cases, time and cost, can give us a deep understanding of how microbial communities shift depending on the molecules that are on treatment. So we could be in the correct status to start an expression analysis. Expanding the understanding of how we can fix biological treatments to our favor, several molecular tools can be used with a total nucleic acid from a polluted site, like using specific microorganism probes, and if there is an RNA extraction instead, we can scale it up to a complete new scale of genomic possibilities, like transcriptomic analysis (Fig. 2.1).
RNA extraction protocols mentioned before set an advent of next-generation sequencing methods. This has made possible to sequence whole microbial genomes of polluted sites at a much lower cost providing opportunities to examine gene expression patterns under adverse environmental conditions. Genomic and transcriptomic analyses have recently been used to characterize bacteria and fungi that have the ability to degrade multiple xenobiotic compounds (Maghsoudi et al. 2016). Strategies in which this is applied can be described in two fundamentally different ways: (a) sequence-based (involving RNA extraction to then synthesize sequenciable cDNA) and (b) function-based. The basis for both approaches has been the construction of metagenomic clone libraries (Warnecke and Hess 2009). Several new technologies in sequencing equipment had to progress and lower its cost so scientist could have access to equipment like Roche’s 454, Illumina’s Solexa, and ABI’s SOLiD to mention some examples and begin to build genomic libraries that we could relay to compare and identify metabolic profiles of natural environments. Some of these investigations are addressed in Table 2.1.
The most well-known yeast Saccharomyces cerevisiae has already been proven to have positive cleaning effects in water and soil treatments against heavy metals (Soares and Soares 2012). Transcriptome studies on these universal biological models have revealed that its production of sulfur compounds comes from the expression of glutathione (GSH) pathways under As (III) exposure. This was proven by RNA isolation, cDNA synthesis, microarray hybridization, and analysis of the transcriptional response to arsenite (Thorsen et al. 2007). Saccharomyces genome expression still needs to be proven when confronting other recalcitrant molecules. In particular, one example of the use of brewing yeast strains to remediate heavy metal pollution can be taken into account; due to the autoaggregation properties that this fungus possesses, they can be quickly and easily separated from the treated effluent. This intrinsic property avoids the use of cell immobilization techniques or solid-liquid separation processes. After treating the effluent, the convenient management of the contaminated biomass and the selective recovery of metals to ensure the minimization of waste production and low operating costs become important (Soares and Soares 2012). Cellulose and hemicellulose compounds are very persistent in natural habitats, and human’s excessive use of recipient products had resulted in much garbage generated and disposed at landfills (Mathews et al. 2015). Cyathus bulleri, Postia placenta, and Phanerochaete chrysosporium are white rot fungi that show promising results in providing diverse glucanases, glycosyl hydrolases, carbohydrate-active enzyme (CAZymes), and laccase isoforms that work in depolymerization (Rineau et al. 2012). These complex molecules make significant reduction in the number of side chains of the pollutant molecule. This was observed during transcriptome analysis of these substrates (like wood) submerged into liquid medium composed of other trace elements (Kameshwar and Qin 2017). Other hazardous pollutants that have been addressed are petrochemical products such as anthracene, anthrone, petroleum asphaltenes, and even crude oil; for these substances fungal species like Neosartorya fischeri, Phanerochaete chrysosporium, Aspergillus niger, Trichoderma harzianum, Talaromyces purpurogenum, and Aspergillus flavus have good bioremediation activity. RNA extraction procedures at different times of treatments with these fungal species which were carried out by using cetyltrimethylammonium bromide (CTAB) or TRI Reagent kit (Sigma) have led to the possibility to evaluate transcriptomic responses of catalase, peroxidase, laccase, and cytochrome P450 monooxygenases encoding genes that could present beneficial activities in accidental oil spillage by stimulating the inhabiting rhizospheric fungal strains (Hernández-López et al. 2015; Asemoloye et al. 2018). A major xenobiotic compound like dichlorvos (2,2-dichlorovinyl dimethyl phosphate) is an organophosphate widely used as an insecticide, to control and protect houses and stored product from arthropods. It has been commercially available since 1961, and its use has become controversial because of its prevalence in urban waterways and the fact that its toxicity extends well beyond insects (Gao et al. 2009; Kazemi et al. 2012). Exploration of possible resolutions to this impact relies on the use of Trichoderma atroviride; this filamentous cosmopolitan fungus commonly found in soil has shown the expression of 5382 differential genes in response to exposure of dichlorvos stress for 2, 6, and 24 h in lab conditions. At these times, RNA extraction with cold 5% trichloroacetic acid was performed to show a transcriptional profile in which metabolic pathways were found to be regulated by the super families hex1 encoding cytochrome P450, glutathione-S-transferase, flavoprotein, Hsp70, and Hsp90. After reads mapping and gene clustering analysis, it was found that ABC transporters were affected by the disruption of hex1 gene. This deletion was made in order to prove that ABC transporter genes might play a vital role in the tolerance process. Expression patterns of seven selected ABC transporter genes were confirmed by qRT-PCR of Trichoderma atroviride and a T. atroviride hex1-deleted mutant (Zhang et al. 2015).
2.3 Fungal Transcriptomic Perspectives
Further research in other environmental liability sites will be a powerful resolutive tool that could position fungal bioremediation treatments as most effective and safe suggested options of restoration procedures as the fourth industrial age settles. Recent and longtime polluted environments can be first evaluated by a DNA and RNA extraction procedure to screen native microbial communities (DNA sequencing) and their metabolic activity (transcriptomic). In this way there could be evidence that a polluted site already possesses genomic potential for self-cleaning with an appropriate bioremediation technique. If a microbial community does not outright this, there can always be other potential biological options like many fungal species that can be inoculated. Their use can be forever preferred around commercial bioremediation products mentioned before. Transcriptomics and metatranscriptomics are influential techniques that let us know what are the active genes essential for the continuance of microbial populations in adverse conditions, highlighting the genomic potential of a particular organism or a microbial community using this discipline of applied genomics (Warnecke et al. 2007). Other various expression profile studies in other species have been performed over the last decades to focus on central interrogates in fields like medicine and microbiology, providing valuable insight into the understanding of how certain phenotypes such as radiation resistance , pathogenesis, or heat-shock resistance are correlated with gene expression (Qiu et al. 2008; Liu et al. 2003; Audia et al. 2008). These transcriptomic studies were usually performed using DNA microarrays. Initially these experiments were restricted to a single organism grown in pure culture but were soon expanded to target several organisms at once (You et al. 2008; Parro et al. 2007; Bulow et al. 2008). A disadvantage of the DNA microarray-based expression profiling is that gene or even complete genome sequences and the corresponding annotations are required before a functional microarray chip can be manufactured (Darby and Hall 2008). Hence, DNA microarrays are rather ineffective for the discovery of novel biocatalysts from the environment. Other challenges in using DNA-based microarrays include their limited detection sensitivity and quantification reliability (Darby and Hall 2008). Bioremediation techniques stimulate microorganisms with genes that are transcribed under specific environmental conditions. For this, direct extraction techniques of bacterial, archaeal, and eukaryotic mRNA from environmental samples were reported recently (Bailly et al. 2007; Poretsky et al. 2005). In both studies, cDNA libraries from environmental RNA were constructed and 400 and 119 clones sequenced, respectively. Most of the obtained sequences had no significant hit to any protein sequence that was previously deposited in public databases. Thus, these studies demonstrate the potential of discovering novel proteins. Furthermore, less sequencing capacity is required for transcripts as compared to the analysis of genomic or metagenomic DNA. This is especially true for the low coding density of eukaryotic genomes . Substantial progress for the efficient analysis of more complex expression profiles has become available with the development of next-generation sequencing technologies mentioned before. These new technologies allow not only the direct sequencing of DNA or likewise cDNA without any cloning step (Medini et al. 2008) but also increased the throughput in terms of numbers of base pairs sequenced per run and decreased cost per sequenced base. Although all three of these new technologies produced extremely short reads when they initially entered the market, the performance of the third generation of the 454 pyrosequencer increased substantially. An average read length of ∼400 bp has been reported for the recently released 454FLX Titanium platform (Hugenholtz and Tyson 2008), and it is not unlikely that future releases will produce reads comparable to those produced with traditional dye terminator sequencing technology (i.e., also referred to as Sanger methodology). The first report of using 454 pyrosequencing for studying the metatranscriptome of a complex microbial community was published in 2006. Using this highly parallelized sequencing technology, Leininger et al. (2006) could show that archaeal transcripts of the key enzyme (amoA) for ammonia oxidation were several magnitudes more abundant in soils than the bacterial version of it, suggesting archaea as the numerical dominant ammonia oxidizers in soil (Leininger et al. 2006). The obtained 25 Mbp of sequence data of this transcriptome study were further analyzed by Urich et al. (2008). They reported that 8% of the initial >250 k reads could be identified as mRNA tags. In 2008, Frias-Lopez et al. produced >50 Mbp by 454 pyrosequencing, still using the first generation of this technology (Frias-Lopez et al. 2008). Gilbert et al. in 2008 followed shortly afterward with >300 Mbp of sequence data now using the second generation called GS-FLX (Gilbert et al. 2008). All three studies demonstrated how high-throughput sequencing technologies can be applied with ease to access information stored in known and unknown transcripts that have been isolated directly from complex environments such as marine and soil microbial communities.
2.4 Concluding Remarks
A continual progress of the genomic fields ensuring metabolic profiles of native or inducted microbial communities during bioremediation process is essential for coming pollution adversities and definition of environmental authorities’ procedures. One of the main problems to overcome in transcriptomic studies is the correct ribonucleic acid extraction protocol, as interference molecules like humic acids, metals, and xenobiotic compounds may compromise the quality and quantity obtained; thus a different extraction protocol may be chosen depending on the integrity of the fungal bioremediation technique. RNA from fungi exposed to pollutant-enriched media in lab conditions may go well by a traditional liquid nitrogen-buffer-phenol-chloroform extraction, but in situ bioremediation treatments of soil and water may contemplate the use of a commercial kit as these products had proven to be efficient against interference molecules. Once there is a good RNA recovery from fungal biological treatments, transcriptomic studies will help to continue the expansion of database transcripts. This is an urgent matter as fungal genomes are not quite well annotated, and there is a lack of understanding on how fungal genes become functional even though there are predicted gene function suggestions for Saccharomyces and Aspergillus. Thus, transcriptomics applied in mycoremediation treatments succeed when known genes demonstrate new functions against xenobiotic compounds and also when structural or functional unknown genes are discovered. In this case, transcriptomic studies may greatly benefit science by describing new metabolic pathways to later build punctual bioremediation strategies. Deep understanding of fungal genomic structure and expressed metabolism pathways is a key concept to be approached in time. Scientific community had made significant efforts to have bigger and well-annotated databases as there are ongoing projects like 1000 fungal genomes (http://1000.fungalgenomes.org/home). This initiative promotes and facilitates user community participation to gather contributions from taxonomists, mycologists, and those focusing on fungal genomics contributing to a better understanding on how fungal genomes behave under recalcitrant chemical exposure like heavy metals, hydrocarbons, and pesticides. It is motivating that all around the globe, many research groups accept the challenge of immersing themselves in the analysis of transcriptomic metadata in order to advance in the description of degradation processes of polluting compounds, which brings us closer to the installation of industrial processes of generalized bioremediation.
References
Asemoloye MD, Ahmad R, Jonathan SG (2018) Transcriptomic responses of catalase, peroxidase and laccase encoding genes and enzymatic activities of oil spill inhabiting rhizospheric fungal strains. Environ Pollut 235:55–64
Audia JP, Patton MC, Winkler HH (2008) DNA microarray analysis of the heat shock transcriptome of the obligate intracytoplasmic pathogen Rickettsia prowazekii. Appl Environ Microbiol 74:7809–7812
Bailly J, Fraissinet-Tachet L, Verner MC, Debaud JC, Lemaire M, Wesolowski-Louvel M, Marmeisse R (2007) Soil eukaryotic functional diversity, a metatranscriptomic approach. ISME J 1:632
Bell TH, Joly S, Pitre FE, Yergeau E (2014) Increasing phytoremediation efficiency and reliability using novel omics approaches. Trends Biotechnol 32(5):271–280
Boon N, Marlé C, Top EM, Verstraete W (2000) Comparison of the spatial homogeneity of physico-chemical parameters and bacterial 16S rRNA genes in sediment samples from a dumping site for dredging sludge. Appl Microbiol Biotechnol 53:742–747
Bulow SE, Francis CA, Jackson GA, Ward BB (2008) Sediment denitrifier community composition and nirS gene expression investigated with functional gene microarrays. Environ Microbiol 10:3057–3069
Burton RJ, Coley-Smith JR (1993) Production and leakage of antibiotics by Rhizoctonia cerealis, R. oryzae-sativae and R. tuliparum. Mycol Res 97:86–90
Chigu NL, Hirosue S, Nakamura C, Teramoto H, Ichinose H, Wariishi H (2010) Cytochrome P450 monooxygenases involved in anthracene metabolism by the white-rot basidiomycete Phanerochaete chrysosporium. Appl Microbiol Biotechnol 87(5):1907–1916
Darby AC, Hall N (2008) Fast forward genetics. Nat Biotechnol 26:1248–1249
Das S (ed) (2014) Microbial biodegradation and bioremediation. Elsevier, Rourkela Odisha, pp 167–201
Deshmukh R, Khardenavis AA, Purohit HJ (2016) Diverse metabolic capacities of fungi for bioremediation. Indian J Microbiol 56(3):247–264
Frias-Lopez J, Shi Y, Tyson GW, Coleman ML, Schuster SC, Chisholm SW, Delong EF (2008) From the cover: microbial community gene expression in ocean surface waters. Proc Natl Acad Sci U S A 105:3805–3810
Gao J, Liu L, Liu X, Zhou H, Lu J, Huang S, Wang Z (2009) The occurrence and spatial distribution of organo phosphorous pesticides in Chinese surface water. Bull Environ Contam Toxicol 82:223–229
Gilbert JA, Field D, Huang Y, Edwards R, Li W, Gilna P, Joint FI (2008) Detection of large numbers of novel sequences in the metatranscriptomes of complex marine microbial communities. PLoS One 3:3042
Gow NA, Latge JP, Munro CA (2017) The fungal cell wall: structure, biosynthesis, and function. Microbiol Spectr 5:1–25
Guillén-Navarro K, Herrera-López D, López-Chávez MY, Cancino-Gómez M, Reyes-Reyes AL (2015) Assessment of methods to recover DNA from bacteria, fungi and archaea in complex environmental samples. Folia Microbiol 60(6):551–558
Hawksworth DL (2001) The magnitude of fungal diversity: the 1.5 million species estimate revisited. Mycol Res 105(12):1422–1432
He Y, Zhao Y, Zhou G, Huang M (2009) Evaluation of extraction and purification methods for obtaining PCR-amplifiable DNA from aged refuse for microbial community analysis. Word J Microbiol Biotechnol 25(11):2043–2051
Hernández-López EL, Ramírez-Puebla ST, Vazquez-Duhalt R (2015) Microarray analysis of Neosartorya fischeri using different carbon sources, petroleum asphaltenes and glucose-peptone. Genom Data 5:235–237
Hugenholtz P, Tyson GW (2008) Microbiology: metagenomics. Nature 455:481
Illman BL, Yang VW, Ferge LA (2002a) US Patent No. 6,383,800. US Patent and Trademark Office, Washington, DC
Illman BL, Yang VW, Ferge LA (2002b) US Patent No. 6,387,689. US Patent and Trademark Office, Washington, DC
Illman BL, Yang VW, Ferge LA (2002c) US Patent No. 6,387,691. US Patent and Trademark Office, Washington, DC
Illman BL, Yang VW, Ferge LA (2002d) US Patent No. 6,495,134. US Patent and Trademark Office, Washington, DC
Illman BL, Yang VW, Ferge LA (2003) US Patent No. 6,664,102. US Patent and Trademark Office, Washington, DC
Illman BL, Yang VW, Ferge LA (2004) US Patent No. 6,727,087. US Patent and Trademark Office, Washington, DC
Illman BL, Yang VW, Ferge LA (2005) US Patent No. 6,972,169. US Patent and Trademark Office, Washington, DC
Jiang YX, Wu JG, Yu KQ, Ai CX, Zou F, Zhou HW (2011) Integrated lysis procedures reduces extraction biases of microbial DNA from mangrove sediments. J Biosci Bioeng 111(2):153–157
Kameshwar AKS, Qin W (2017) Metadata analysis of Phanerochaete chrysosporium gene expression data identified common CAZymes encoding gene expression profiles involved in cellulose and hemicellulose degradation. Int J Biol Sci 13(1):85
Kazemi M, Tahmasbi A, Valizadeh R, Naserian A, Soni A (2012) Organophosphate pesticides: a general review. Agric Sci Res J 2:512–522
Ladino-Orjuela G, Gomes E, da Silva R, Salt C, Parsons JR (2016) Metabolic pathways for degradation of aromatic hydrocarbons by bacteria. Rev Environ Contam Toxicol 237:105–121
Lamar RT, Lestan D, Smith CE, Dietrich DM (2000) US Patent No. 6,143,549. US Patent and Trademark Office, Washington, DC
Leininger S, Urich T, Schloter M, Schwark L, Qi J, Nicol GW, Prosser JI, Schuster SC, Schleper C (2006) Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nature 442:806
Liu Y, Zhou J, Omelchenko MV, Beliaev AS, Venkateswaran A, Stair J, Wu L, Thompson DK, Xu D, Rogozin IB, Gaidamakova EK, Zhai M, Makarova KS, Koonin EV, Daly MJ (2003) Transcriptome dynamics of Deinococcus radiodurans recovering from ionizing radiation. Proc Natl Acad Sci U S A 100:4191–4196
Maghsoudi E, Fortin N, Greer C, Maynard C, Pagé A, Duy SV, Dorner S (2016) Cyanotoxin degradation activity and mlr gene expression profiles of a Sphingopyxis sp. isolated from Lake Champlain, Canada. Environ Sci Process Impact 18(11):1417–1426
Mathews SL, Pawlak J, Grunden AM (2015) Bacterial biodegradation and bioconversion of industrial lignocellulosic streams. Appl Microbiol Biotechnol 99(7):2939–2954
McMaster R (2018) Is the fourth industrial revolution relevant to you? Nurs Health Sci 20(2):139–141
Medini D, Serruto D, Parkhill J, Relman DA, Donati C, Moxon R, Falkow S, Rappuoli R (2008) Microbiology in the post-genomic era. Nat Rev Microbiol 6:419
Mohite BV, Koli SH, Narkhede CP, Patil SN, Patil SV (2017) Prospective of microbial exopolysaccharide for heavy metal exclusion. Appl Biochem Biotechnol 183(2):582–600
Morais D, Pylro V, Clark IM, Hirsch PR, Tótola MR (2016) Responses of microbial community from tropical pristine coastal soil to crude oil contamination. Peer J 4:1733
Moré MI, Herrick JB, Silva MC, Ghiorse WC, Madsen EL (1994) Quantitative cell lysis of indigenous microorganisms and rapid extraction of microbial DNA from sediment. Appl Environ Microbiol 60(5):1572–1580
Nancharaiah YV, Mohan SV, Lens PNL (2016) Biological and bioelectrochemical recovery of critical and scarce metals. Trends Biotechnol 34(2):137–155
National Research Council (2014) Review of EPA’s integrated risk information system (IRIS) process. National Academies Press, Washington, DC
Nelson DM, Ohene-Adjei S, Hu FS, Cann IKO, Mackie RI (2007) Bacterial diversity and distribution in the holocene sediments of a northern temperate lake. Microbial Ecol 54(2):252–263
Obeng EM, Adam SNN, Budiman C, Ongkudon CM, Maas R, Jose J (2017) Lignocellulases: a review of emerging and developing enzymes, systems, and practices. Bioresour Bioprocess 4(1):16
Ozer A, Ozer D (2003) Comparative study of the biosorption of Pb(II), Ni(II) and Cr(VI) ions onto S. cerevisiae: determination of biosorption heats. J Hazard Mater 100:219–229
Parro V, Moreno-Paz M, Gonzalez-Toril E (2007) Analysis of environmental transcriptomes by DNA microarrays. Environ Microbiol 9:453–464
Poretsky RS, Bano N, Buchan A, Lecleir G, Kleikemper J, Pickering M, Pate WM, Moran MA, Hollibaugh JT (2005) Analysis of microbial gene transcripts in environmental samples. Appl Environ Microbiol 71:4121–4126
Qiu J, Guo Z, Liu H, Zhou D, Han Y, Yang R (2008) DNA microarray-based global transcriptional profiling of Yersinia pestis in multicellularity. J Microbiol 46:557–563
Rineau F, Roth D, Shah F, Smits M, Johansson T, Canbäck B, Grigoriev IV (2012) The ectomycorrhizal fungus Paxillus involutus converts organic matter in plant litter using a trimmed brown-rot mechanism involving Fenton chemistry. Environ Microbiol 14(6):1477–1487
Rodrigues EM, Kalks KH, Fernandes PL, Tótola MR (2015) Bioremediation strategies of hydrocarbons and microbial diversity in the Trindade Island shoreline. Mar Pollut Bull 101(2):517–525
Romanelli AM, Fu J, Herrera ML, Wickes BL (2014) A universal DNA extraction and PCR amplification method for fungal rDNA sequence-based identification. Mycoses 57(10):612–622
Ruta LL, Kissen R, Nicolau I, Neagoe AD, Petrescu AJ, Bones AM, Farcasanu IC (2017) Heavy metal accumulation by Saccharomyces cerevisiae cells armed with metal binding hexapeptides targeted to the inner face of the plasma membrane. Appl Microbiol Biotechnol 101(14):5749–5763
Saǧ Y, Özer D, Kutsal T (1995) A comparative study of the biosorption of lead(II) ions to Z. ramigera and R. arrhizus. Process Biochem 30:169–174
Sankaran S, Khanal SK, Jasti N, Jin B, Pometto AL, Van Leeuwen JH (2010) Use of filamentous fungi for wastewater treatment and production of high value fungal byproducts: a review. Crit Rev Environ Sci Technol 40(5):400–449
Sato S, Feltus FA, Iyer P, Tien M (2009) The first genome-level transcriptome of the wood-degrading fungus Phanerochaete chrysosporium grown on red oak. Curr Genet 55(3):273–286
Soares EV, Soares HM (2012) Bioremediation of industrial effluents containing heavy metals using brewing cells of Saccharomyces cerevisiae as a green technology: a review. Environ Sci Pollut R 19(4):1066–1083
Thorsen M, Lagniel G, Kristiansson E, Junot C, Nerman O, Labarre J, Tamás MJ (2007) Quantitative transcriptome, proteome, and sulfur metabolite profiling of the Saccharomyces cerevisiae response to arsenite. Physiol Genomics 30(1):35–43
Tortella GR, Diez MC, Durán N (2005) Fungal diversity and use in decomposition of environmental pollutants. Crit Rev Microbiol 31(4):197–212
Urich T, Lanzen A, Qi J, Huson DH, Schleper C, Schuster FSC (2008) Simultaneous assessment of soil microbial community structure and function through analysis of the meta-transcriptome. PLoS One 3:2527
Vats A, Mishra S (2018) Identification and evaluation of bioremediation potential of laccase isoforms produced by Cyathus bulleri on wheat bran. J Hazard Mater 344:466–479
Verma S, Verma PK, Meher AK, Dwivedi S, Bansiwal AK, Pande V, Chakrabarty D (2016) A novel arsenic methyltransferase gene of Westerdykella aurantiaca isolated from arsenic contaminated soil: phylogenetic, physiological, and biochemical studies and its role in arsenic bioremediation. Metallomics 8(3):344–353
Warnecke F, Hess M (2009) A perspective: metatranscriptomics as a tool for the discovery of novel biocatalysts. J Biotechnol 142(1):91–95
Warnecke F, Luginbühl P, Ivanova N, Ghassemian M, Richardson TH, Stege JT, Cayouette M, Mchardy AC, Djordjevic G, Aboushadi N, Sorek R, Tringe SG, Podar M, Garcia-Martin H, Kunin V, Dalevi D, Madejska J, Kirton E, Platt D, Szeto E, Salamov A, Barry K, Mikhailova N, Kyrpides NC, Matson EG, Ottesen EA, Zhang X, Hernández M, Murillo C, Acosta LG, Rigoutsos I, Tamayo G, Green BD, Chang C, Rubin EM, Mathur EJ, Robertson DE, Hugenholtz P, Leadbetter JR (2007) Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 450:560–565
Whitby C (2010) Microbial naphthenic acid degradation. Adv Appl Microbiol 70:93–125
Wintzingerode FV, Göbel UB, Stackebrandt E (1997) Determination of microbial diversity in environmental samples: pitfalls of PCR-based rRNA analysis. FEMS Microbiol Rev 21(3):213–229
World Health Organization (2004) Guidelines for drinking-water quality: recommendations, vol 1. World Health Organization, Geneva
Wymelenberg AV, Gaskell J, Mozuch M, Sabat G, Ralph J, Skyba O, Kersten PJ (2010) Comparative transcriptome and secretome analysis of wood decay fungi Postia placenta and Phanerochaete chrysosporium. Appl Environ Microbiol 76(11):3599–3610
You Y, Fu C, Zeng X, Fang D, Yan X, Sun B, Xiao D, Zhang J (2008) A novel DNA microarray for rapid diagnosis of enteropathogenic bacteria in stool specimens of patients with diarrhea. J Microbiol Methods 75:566–571
Zhang T, Tang J, Sun J, Yu C, Liu Z, Chen J (2015) Hex1-related transcriptome of Trichoderma atroviride reveals expression patterns of ABC transporters associated with tolerance to dichlorvos. Biotechnol Lett 37(7):1421–1429
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Ledezma-Villanueva, A., Adame-Rodríguez, J.M., Aréchiga-Carvajal, E.T. (2018). Transcriptomics as a First Choice Gate for Fungal Biodegradation Processes Description. In: Prasad, R., Aranda, E. (eds) Approaches in Bioremediation. Nanotechnology in the Life Sciences. Springer, Cham. https://doi.org/10.1007/978-3-030-02369-0_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-02369-0_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-02368-3
Online ISBN: 978-3-030-02369-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)