Abstract
O-GlcNAc is the attachment of β-N-acetylglucosamine to the hydroxyl group of serine and threonine in nuclear and cytoplasmic proteins. It is generally not further elongated but exists as a monosaccharide that can be rapidly added or removed. Thousands of proteins involved in gene transcription, protein translation, and degradation as well as the regulation of signal transduction contain O-GlcNAc. Brain is one of the tissues where O-GlcNAc is most highly expressed and deletion of neuronal O-GlcNAc leads to death early in development. O-GlcNAc is also important for normal adult brain function, where dynamic processes like learning and memory at least in part depend on the modification of specific proteins by O-GlcNAc. Conversely, too much or too little O-GlcNAc on other proteins participates in neurodegenerative processes underlying diseases such as Alzheimer’s and Parkinson’s. In this chapter, we describe the expression and regulation of O-GlcNAc in the nervous system.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- O-linked N-acetylglucosamine
- O-GlcNAc
- Learning and memory
- Neurodegeneration
- Alzheimer’s disease
- Signaling
- Nutrient sensing
- Post-translational modifications
16.1 Introduction
O-GlcNAc is the attachment of β-N-acetylglucosamine to the hydroxyl group of serine and threonine in nuclear and cytoplasmic proteins (Torres and Hart 1984). It is generally not further elongated but exists as a monosaccharide that can be rapidly added or removed (Hart et al. 2011). Thousands of proteins involved in gene transcription, protein translation, and degradation as well as the regulation of signal transduction contain O-GlcNAc (Trinidad et al. 2012; Alfaro et al. 2012). Brain is one of the tissues where O-GlcNAc is most highly expressed and deletion of neuronal O-GlcNAc leads to death early in development (Kreppel et al. 1997; O'Donnell et al. 2004). O-GlcNAc is also important for normal adult brain function, where dynamic processes like learning and memory at least in part depend on the modification of specific proteins by O-GlcNAc (Tallent et al. 2009; Rexach et al. 2012). Conversely, too much or too little O-GlcNAc on other proteins participates in neurodegenerative processes underlying diseases such as Alzheimer's and Parkinson's (Arnold et al. 1996; Liu et al. 2004; Yuzwa et al. 2012; Wang et al. 2010a, b; Marotta et al. 2012). In this chapter, we describe the expression and regulation of O-GlcNAc in the nervous system.
16.2 O-GlcNAc Is a Ubiquitous Monosaccharide That Cycles onto and off Serine and Threonine
16.2.1 O-GlcNAc Is Not Elongated to Yield Complex Oligosaccharides
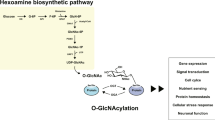
O-GlcNAc is the covalent modification of nuclear and cytoplasmic proteins by β-N-acetylglucosamine (Torres and Hart 1984). O-GlcNAc is formed as a derivative of glucose through the hexosamine biosynthesis pathway (HBP). In the HBP, the oxygen on the second carbon of fructose-6-phosphate is exchanged for nitrogen forming GlcN-6-P, prior to acetylation of the nitrogen to yield GlcNAc-6-P. This is then coupled to the high-energy molecule uridine diphosphate (UDP), UDP-GlcNAc (Fig. 16.1). Upon modification of proteins by O-GlcNAc, the GlcNAc is cleaved from the UDP and attached in β-position to the hydroxyl group of serine or threonine (O-β-GlcNAc, O-GlcNAc). The reaction is catalyzed by the O-GlcNAc transferase, OGT. The removal of GlcNAc is catalyzed by the O-GlcNAc hydrolase, O-GlcNAcase (OGA).
O-GlcNAc cycling is regulated by two enzymes, OGT and OGA, and total O-GlcNAc levels depend on several metabolic pathways. The donor substrate for O-GlcNAc, UDP-GlcNAc, is produced by the hexosamine biosynthesis pathway (HBP). Many metabolites feed into the HBP and thereby modulate UDP-GlcNAc production. In the nucleus and cytosol of cells, the O-GlcNAc transferase (OGT) cleaves the UDP-GlcNAc and adds the GlcNAc to serine or threonine on proteins. The GlcNAc can then be removed from the protein by another nucleocytoplasmic protein, the O-GlcNAc hydrolase, OGA
Unlike “classical” O- and N-linked protein glycosylation the GlcNAc is generally not elongated but exists as a monosaccharide. In fact, when O-GlcNAc is artificially capped by galactose, its biological function is lost (Fang and Miller 2001). Although, O-GlcNAc is smaller than complex oligosaccharides, it is still much larger than many other protein modifications, such as protein methylation or protein phosphorylation (Hart et al. 2011).
16.2.2 O-GlcNAc Is Mostly Expressed on the Inside of Cells in Multicellular Organisms
O-GlcNAc is a highly conserved posttranslational modification. It has been found in evolutionary distinct clades like plantae, fungi, and animalia (Kreppel et al. 1997; Webster et al. 2009). In multicellular organisms, all types of cells investigated so far contain O-GlcNAc (Hart et al. 2011). O-GlcNAc has also been identified in some unicellular organisms, e.g. giardia—the oldest eukaryote, and inside several types of virus (Banerjee et al. 2009; Benko et al. 1988; Caillet-Boudin et al. 1989). Nonetheless, most studies on unicellular organisms fail to report the presence of β-O-GlcNAc. Protozoans modify proteins by O-linked GlcNAc but primarily on extracellular proteins and in α-linkage rather than in β-linkage. Yeasts appear to lack O-GlcNAc entirely. Bacteria are also largely devoid of cytoplasmic O-GlcNAc even though some bacterial proteins have been shown to carry O-linked GlcNAc (Schirm et al. 2004; Fredriksen et al. 2012). Interestingly, the bacterium Clostridium novyi exploits O-GlcNAc by encoding an O-GlcNAc transferase that modifies small G-proteins in the infected cell (Selzer et al. 1996; Hart et al. 2007, 2011).
O-GlcNAc is expressed almost exclusively on the inside of cells (Torres and Hart 1984). Until the discovery of O-GlcNAc protein glycosylation was known to occur only on proteins exposed to the extracellular matrix, or in cellular organelles topographically similar to the outside of the cell such as the endoplasmatic reticulum (ER) and the Golgi apparatus. In contrast, nearly all proteins that contain O-GlcNAc are expressed in the cytosolic or nuclear fraction of the cell. Proteins anchored to the cell membrane are modified with O-GlcNAc but usually only on parts stretching into the cytosol. This comes as no surprise as the O-GlcNAc transferase, OGT, is mainly nucleocytoplasmic rather than present in the Golgi or ER as other glycosyltransferases (there are at least two O-GlcNAc transferases with their active sites in the lumen of the ER, called, eOGTs, but these enzymes are distinct from the enzyme regulating nucleocytoplasmic O-GlcNAc. O-GlcNAc has been detected on extracellular domains of a handfull of proteins, e.g. Notch) (see Sect. 16.2.2; Alfaro et al. 2012). Also the hexosaminidase removing O-GlcNAc, OGA, is cytosolic and active at neutral pH (see Sect. 16.2.3). By comparison, cellular glycosidases breaking down glycoconjugates retrieved from the cell surface are primarily found in the lysosome and prefer an acidic milieu.
Importantly, the concentration of O-GlcNAc is not uniform across the cell. Some parts, like the nuclear membrane, are heavily modified whereas other parts, like the mitochondria, contain O-GlcNAc but to a lesser degree. All major organelles and other cytosolic substructures, e.g. the proteasome and ribosome, express O-GlcNAc (Holt and Hart 1986; Zhang et al. 2003; Zeidan et al. 2010). The precise level varies over time and is finely tuned to meet the conditions of the cell (see Sects. 16.2.3, 16.3 and 16.4).
16.2.3 O-GlcNAc Can Be Dynamically Attached and Removed
Whether or not a protein is modified by O-GlcNAc varies substantially over time. On many proteins, including the heat-shock protein αB-crystallin, the O-GlcNAc half-life is much shorter than the half-life of the peptide backbone (Roquemore et al. 1996). In fact, studies using selective inhibitors of the enzyme that removes O-GlcNAc from proteins, OGA, show that cycling rates are often on the order of minutes, making O-GlcNAc more akin to protein phosphorylation than “classical” protein glycosylation. “Classical” N- and O-linked glycosylation of proteins, glycosylation of proteins exposed to the extracellular matrix or within the secretory pathway, is, largely, stable once the mature glycan has been attached. There are examples of proteins, e.g. the nucleoporins that form pores through the nuclear membrane, where the O-GlcNAc does not appear to turnover faster than the protein itself (Holt et al. 1987; Miller et al. 1999).
It has been proposed that O-GlcNAc cycling works like a light switch with only two modes of operation—either “on” or “off.” One argument in favor of this idea is the fact that there are only two enzymes that add and remove O-GlcNAc from proteins, OGT and OGA, respectively (see Sect. 16.2). Indeed, in some situations cells react by either elevating or suppressing global O-GlcNAc levels. For example, abundant nutrient supply leads to a general increase in O-GlcNAc and scant supply to a general decrease (see Sects. 16.2.2 and 16.4.3). Likewise, cellular stress is associated with raised O-GlcNAc throughout the cell (Zachara et al. 2004). All the same, early studies showed that activation of lymphocytes causes O-GlcNAc levels in the cytosol to go up while they go down in the nucleus (Kearse and Hart 1991). A recently developed FRET (fluorescence resonance energy transfer) reporter that measures OGT activity in real-time demonstrated further that during serum stimulation of transformed cell lines, OGT was activated manifold in some parts of the cytosol whereas in nearby areas OGT activity remained at baseline (Carillo et al. 2011). Work on the regulation of signal transduction by O-GlcNAc describes the same picture; upon stimulation, in a single pathway there can be proteins that become better substrates for OGT but also proteins that become worse substrates for OGT (Whelan et al. 2010). Thus, despite there being only two enzymes that add and remove O-GlcNAc, changes in O-GlcNAc can occur “locally” within the cell. For example, by forming dynamic multipartner complexes OGT and OGA can be directed towards select targets among a broader range of available substrates (see Sects 16.2.2–16.3; Whelan et al. 2008; Cheung and Hart 2008; Housley et al. 2009).
The spatiotemporal regulation of O-GlcNAc cycling is complex and occurs on many levels. While nutrients and stress can cause global changes in O-GlcNAc, binding partners to OGT and OGA tune O-GlcNAc occupancy locally. Below we will discuss in detail how O-GlcNAc cycling is controlled in the nervous system and how the dynamic modification of proteins by O-GlcNAc helps the brain to develop and respond to challenges in the environment.
16.3 O-GlcNAc Is Added to Proteins by OGT and Removed by OGA
16.3.1 Only Two Enzymes Regulate the Cycling of O-GlcNAc
O-GlcNAc exists as a monosaccharide on nuclear and cytoplasmic proteins and can cycle rapidly and repeatedly over the lifetime of the polypeptide chain (see Sect. 16.1). In a single cell, including neurons, thousands of proteins carry O-GlcNAc (Trinidad et al. 2012; see Sect. 16.3.2). Change in O-GlcNAc can happen globally throughout the cell but also locally on individual proteins or sites within a protein (see Sect. 16.2.3). In mammals, there are only two enzymes that add and remove O-GlcNAc. The O-GlcNAc transferase (OGT) adds O-GlcNAc to proteins (Haltiwanger et al. 1990; Kreppel et al. 1997). The O-GlcNAc glycosidase, O-GlcNAcase (OGA), removes O-GlcNAc from proteins (Dong and Hart 1994; Gao et al. 2001). As we will see in Sects. 16.4 and 16.5 loss or deregulation of O-GlcNAc cycling leads to severe developmental brain defects, impaired brain function in the adult and risk for many neurodegenerative disorders, e.g. Alzheimer's disease. In this section we will discuss how OGT and OGA are both promiscuous in order to accept a broad range of targets while at the same time specific to ensure that O-GlcNAc cycles on the correct site at the correct time and place.
16.3.2 O-GlcNAc Transferase; A Highly Conserved Glycosyltransferase Present in the Nucleus and Cytosol
In mammals, O-GlcNAc transferase (OGT) is encoded by a single gene. The gene is highly conserved and lies close to the centromeric region of the X chromosome (Xq13) (Shafi et al. 2000; Nolte and Muller 2002; Kreppel et al. 1997). It spans about 45 kilobase pairs (kb) and its locus is linked to Parkinsonian dystonia, a neurodegenerative movement disorder (Nolte and Muller 2002; Muller et al. 1998). In most organs there are five major OGT transcripts ranging from 4.2 kb to 9.5 kb. The transcripts undergo alternative splicing and two 4 kb transcripts may arise from an internal promoter. In brain the larger 9.5 and 6.4 kb transcripts, which include exons located 5′ of the internal promoter, dominate (Hanover et al. 2003; Nolte and Muller 2002). Most studies so far argue that the total expression of OGT is stable during most conditions. However, little is known about the regulation of the OGT gene and at least in neuroblastoma cells OGT mRNA increases after depriving the cells of glucose (Cheung and Hart 2008).
The protein encoded by OGT contains two major domains. The N-terminal half is comprised of several tetratricopeptide (TPR) repeats and the C-terminal half binds UDP-GlcNAc and harbors the glycosyltransferase activity. The TPR repeats form a flexible superhelix that can accommodate many protein–protein interactions (Lyer et al. 2003; Kreppel et al. 1997; Kreppel and Hart 1999; Jinek et al. 2004). The C-terminus exhibits a more compact structure and resembles members of the GT-B superfamily of glycosyltransferases (Gtfs) but adopts some unique folds as well (Lazarus et al. 2011). While most OGT-interacting proteins are believed to bind the TPR repeats the C-terminus includes regions that are necessary and sufficient for some interactions, e.g. to the mitogen-activated kinase (MAPK) p38 in neurons (Cheung and Hart 2008). The C-terminus also mediates translocation to the cell membrane upon insulin stimulation, probably via a cluster of lysines that pairs electrostatically with negatively charged phosphatylinositol (3,4,5)-triphosphate (PIP3) (Yang et al. 2008).
Catalysis occurs by an ordered sequential mechanism where OGT binds first UDP-GlcNAc and then the substrate. A hydroxyl group on the incoming substrate performs a direct nucleophilic attack on the anomeric carbon on GlcNAc, thereby inverting the glycosidic bond from UDP-α-GlcNAc to (serine/threonine-) O-β-GlcNAc. In the catalytic groove, there is a general base (histidine 498) and a general acid (probably lysine 842) that catalyze the nucleophilic attack by activating the hydroxyl group (Lazarus et al. 2011 Banerjee et al. 2013). In vitro experiments have shown that if the TPR domain is removed from OGT, the C-terminus alone can modify peptides with O-GlcNAc. In contrast, for protein substrates to be modified, the TPR domain is required (Kreppel and Hart 1999; Lyer and Hart 2003). It has been hypothesized based on computer simulations that the TPRs induce a conformational change in the substrate that enables the O-GlcNAc site to dock at the catalytic groove. Most likely, OGT goes through a conformational change as well; the TPR helix pivots around the intervening region between the TPRs and the C-terminus exposing the entrance to the groove (Trinidad et al. 2012; Lazarus et al. 2011.
Three major isoforms of OGT have been described (Hanover et al. 2003). All share the same catalytic domain but differ in their N-terminus. Nucleocytoplasmic OGT (ncOGT) is the full-length variant and includes 11–12 TPRs, depending on the species. Mitochondrial OGT (mOGT) starts with a mitochondrial targeting sequence, followed by a membrane-spanning region and continues with the last nine TPRs found in ncOGT. The third, and shortest, isoform is soluble OGT (sOGT). It contains only three TPRs. According to most studies, only ncOGT is present in total brain lysate (Kreppel et al. 1997; Marz et al. 2006). However, sOGT may become upregulated in older animals and little is known about whether mOGT or sOGT is present in specific regions of the brain or in specific subcellular compartments (Liu et al. 2012). Apart from mOGT, which is anchored to the inner membrane of the mitochondria, almost all OGT activity is found in the nucleocytoplasm or as a nonintegral membrane protein associated with the cytosolic face of cell membranes (Hanover et al. 2003; Haltiwanger et al. 1990)
In vivo, OGT oligomerizes into a dimer or trimer (Kreppel et al. 1997; Marz et al. 2006). OGT was first purified from rat liver and described as a heterotrimer consisting of two ncOGT and one 78 kDa unit. The 78 kDa unit is enriched in certain tissues and may correspond to sOGT, but may also be a proteolytic fragment of ncOGT (Haltiwanger et al. 1992; Kreppel et al. 1997). The major form of OGT expressed in brain is more likely a homodimer of ncOGT (Marz et al. 2006). Dimerization occurs over an evolutionary conserved hydrophobic region in the TPR domain (TPR 6). Dimerization is stable even in very high salt concentrations and probably not subject to posttranslational regulation (Kreppel and Hart 1999; Jinek et al. 2004).
OGT is exquisitely regulated adding O-GlcNAc only to particular sites at any given time and place. In vitro, OGT exhibits sequence specificity. If purified OGT is mixed with UDP-GlcNAc and a peptide that contains several possible O-GlcNAc sites, often only one or a few sites become significantly modified. Likewise, substrate peptides derived from different proteins are often modified with different efficiency (Kreppel and Hart 1999). Also UDP-GlcNAc levels influence peptide substrate specificity (Kreppel and Hart 1999; Shen et al. 2012). Nevertheless, there is no absolute substrate consensus sequence for OGT. Indeed, the catalytic site of OGT interacts primarily with the peptide backbone of the substrate and not particular side chains (Lazarus et al. 2011). In vivo, O-GlcNAc sites concentrate on disordered regions of proteins and close to proline and valine, the so called PVS motifs (Alfaro et al. 2012; Trinidad et al. 2012; Hart et al. 2011). Presumably, the reason is that such regions can be made to fit OGT's catalytic groove more easily.
Primary sequence plays a role in determining the major O-GlcNAc sites on a given protein. However, in cells, whether a particular protein will be modified depends also on OGT-binding proteins. OGT operates as a holoenzyme, where its interacting proteins direct OGT to its substrates (Whelan et al. 2008; Cheung and Hart 2008; Cheung et al. 2008; Housley et al. 2009; Marz et al. 2006; Yang et al. 2002). For example, in neurons, neurofilaments become O-GlcNAcylated only after activated p38 binds OGT (Cheung et al. 2008). OGT-interacting proteins can enhance OGT activity towards peptide substrates as well (Marz et al. 2006). Moreover, OGT is multiple phosphorylated and it modifies itself with O-GlcNAc (Kreppel et al. 1997; Song et al. 2008; Whelan et al. 2008). Phosphorylation of OGT can both activate OGT, e.g. CaMKIV during neuronal depolarization, and alter OGT's substrate specificity (Whelan et al. 2008; Song et al. 2008; Bullen and Hart Forthcoming paper). OGT is strongly inhibited by free UDP but, unlike most other glycosyltransferases that utilize a UDP-bound sugar as donor, OGT does not require divalent cations (Haltiwanger et al. 1990). For a summary of the regulation of OGT, see Fig. 16.2.
The regulation of the O-GlcNAc transferase, OGT, is complex. In cells, OGT forms an oligomer that interacts with many other proteins. The composition of the oligomer depends on what OGT isoforms are transcribed and probably also on proteolytic processing. The interacting proteins direct OGT to its substrates at specific points in time and space. OGT activity can also be regulated by posttranslational modifications such as O-GlcNAc and O-phosphate and UDP-GlcNAc abundance
OGT has emerged as a key cellular nutrient sensor. The donor for O-GlcNAc, UDP-GlcNAc, is produced by the hexosamine biosynthesis pathway, the HBP (see Sect. 16.2.1). UDP-GlcNAc is an abundant high-energy small molecule and ranges in concentrations within cells from 0.1 to 1 mM. Almost all metabolic pathways feed into the HBP and contribute to the production of UDP-GlcNAc, including fatty acids, nitrogen and 2–5 % of all cellular glucose. A rich energy supply elevates UDP-GlcNAc, and, subsequently, protein O-GlcNAcylation whereas a scarce supply is associated with reduced levels of both UDP-GlcNAc and O-GlcNAc (Hart et al. 2011; Hart et al. 2007). Also in the brain fasting has been shown to decrease O-GlcNAc generally, a change reversed upon re-feeding (Liu et al. 2004; Li et al. 2006). Interestingly, depending on the substrate, the Km for UDP-GlcNAc can vary more than 20-fold but most become better substrates in the presence of higher concentrations of UDP-GlcNAc (Shen et al. 2012; Kreppel and Hart 1999). Therefore, although metabolic flux is associated with global changes in O-GlcNAc, some proteins are more sensitive to nutrient supply than others. In Sects. 16.4 and 16.5 we will discuss how OGT as a nutrient sensor is involved in the pathology behind neurodegenerative disease.
16.3.3 O-GlcNAcase; A Cytosolic O-β-GlcNAc Hydrolase with Neutral pH Optima
In accord with the O-GlcNAc transferase, the enzyme that removes O-GlcNAc from proteins, the O-GlcNAcase (OGA), is expressed from a single gene. The gene was first cloned as an antigen associated with meningioma and it is localized to the long arm of chromosome 10 (10q24), a locus tightly linked to late-onset Alzheimer's disease (Heckel et al. 1998; Bertram et al. 2000). In addition, alternatively spliced OGA transcripts have been associated with sporadic cases of Alzheimer’s disease (Twine et al. 2011).
OGA hydrolyses GlcNAc from proteins using substrate-assisted catalysis. Two aspartates activate the acetamido group on GlcNAc's second carbon to perform a nucleophilic attach on its anomeric carbon. This results in the displacement of GlcNAc from the protein (Macauley et al. 2005; Dennis et al. 2006). Contrary to many other hexosaminidases, OGA has neutral pH optima and is not inhibited by GalNAc. Nor does it remove GalNAc from proteins but is specific for β-linked O-GlcNAc (Dong and Hart 1994). OGA is also genetically and immunogenically distinct from other glycosidases (Gao et al. 2001).
OGA has been identified in all tissues so far investigated. It is expressed in the nucleus and cytoplasm of cells and the highest expression is found in brain (Gao et al. 2001). OGA is highly conserved but contains a middle, intrinsically unfolded region that exhibits more variability (Gao et al. 2001; Heckel et al. 1998; Butkinaree et al. 2008). Like OGT, OGA consists of two major domains. The glycosidase domain is C-terminal. The N-terminus contains a histone acetyl transferase (HAT) domain. While the HAT domain was reported to be functionally active (Toleman et al. 2004; Toleman et al. 2006), other groups have not been able to repeat this finding (Butkinaree et al. 2008). Caspase 3 cleaves OGA between the glycosidase domain and the HAT domain during apoptosis without any loss of O-GlcNAcase activity (Butkinaree et al. 2008).
Very little is known about the regulation of OGA. It is believed that OGA forms multipartner complexes that direct OGA to its substrates, much like how OGT is regulated (Hart et al. 2011; see above). Recently, highly specific pharmacological inhibitors of OGA have been developed (Yuzwa et al. 2008; Yuzwa et al. 2012). In Sects. 16.4 and 16.5 we will learn more about the way in which OGA is important for the brain's ability to learn and form memories and serves as a drug target for neurodegenerative disorders.
16.4 O-GlcNAc Is Highly Expressed in the Nervous System
16.4.1 O-GlcNAc is Found Throughout the Brain
Brain is one of the organs where O-GlcNAc is the most abundant. OGT and OGA are expressed in comparatively high levels in the brain (Kreppel et al. 1997; Lubas et al. 1997; Gao et al. 2001). Both enzymes are present across brain regions, including cortex, the cerebellum, and subcortical nuclei such as the amygdala (Liu et al. 2012; Rexach et al. 2012). Within neurons, O-GlcNAc has been identified in all subcellular structures investigated, albeit to differing degrees. Biochemical fractionation was used to show that O-GlcNAc transferase activity was present in synapses (Cole and Hart 2001). Immunoelectron microscopy indicated that OGT and O-GlcNAc were present in both the post- and presynapse. In the presynapse, OGT is concentrated on synaptic vesicles storing neurotransmitter (Akimoto et al. 2003). Most research on O-GlcNAc in the brain has focused on O-GlcNAc's role in neurons of the central nervous system. Less is known about its expression in neurons of the peripheral nervous system or in glia.
16.4.2 Thousands of Neuronal Proteins Are Modified by O-GlcNAc
One difficulty in studying the role of O-GlcNAc for brain function is its vast abundance. Technical breakthroughs have allowed identification of O-GlcNAc sites en masse by coupling selective enrichment of O-GlcNAc-modified proteins with electron capture dissociation (ECD) or electron transfer dissociation (ETD) mass spectrometry (types of mass spectrometry that can fragment peptides without losing the GlcNAc) (Wang et al. 2010a, b; Trinidad et al. 2012). According to some estimates, about 40 % of all neuronal proteins are modified by O-GlcNAc (Trinidad et al. 2012). Some proteins, like bassoon, which is important for neurotransmitter release, have more than a dozen O-GlcNAc sites. On other proteins, e.g. CaMKII, only a single site has been detected (Trinidad et al. 2012). As discussed in Sect. 16.2.2 O-GlcNAc sites concentrate on structurally disordered regions of proteins and there is some preference for a sequence context rich in proline and valine. For any particular site, O-GlcNAc is usually present in substochiometric levels (Hart et al. 2007). The absolute occupancy is dynamic and depends on the activity of the neuron, possibly due to CaMK-dependent stimulation of OGT (Khidekel et al. 2007; Rexach et al. 2010; Song et al. 2008).
Proteins modified by O-GlcNAc belong to all functional classes (Fig. 16.3). Many O-GlcNAc proteins are shared across cell types and underlie constitutive cellular functions such as transcription, translation, and protein degradation. Nevertheless, the role played by O-GlcNAc on these factors is often neuron-specific. For example, in neurons, the common transcription factor cyclic AMP-response element binding protein (CREB) is modified by O-GlcNAc on serine 40 upon cell depolarization. Once modified, CREB is prevented from binding to its coactivator CRTC (CREB-regulated transcription coactivator). This inhibits CREB and leads to an inactivation of several transcription pathways involved in synaptic plasticity (see Sect. 16.4.2; Rexach et al. 2012). Other O-GlcNAc proteins are particular to neurons, including certain scaffolding proteins, signaling proteins, and cytoskeletal proteins. Many of these are proteins found in the synapse. One interesting case is CaMKII-α. CaMKII-α is modified by O-GlcNAc in a small region known to contain one phosphorylation site that can activate the enzyme and another phosphorylation site that deactivates it (Trinidad et al. 2012). The regulation of CaMKII-α activity is crucial for synaptic events that underlie learning and memory (Lisman et al. 2002). Therefore, elucidation of how O-GlcNAc may fine tune CaMKII-α activity is important not only to our understanding of synapse biology but also higher-order brain function. Moreover, in the synapse, there is a significant overrepresentation of kinases, the enzymes that add O-phosphate to proteins, among the proteins that are modified by O-GlcNAc (Trinidad et al. 2012).
O-GlcNAc modified proteins in neurons belong to all functional classes. In neurons, more than a 1,000 proteins carry O-GlcNAc. The O-GlcNAc-modified proteins include chromatin, transcription factors, cytoskeletal proteins, and enzymes like kinases. Many of these are present in the synapse. The figure shows the distribution of O-GlcNAc in brain across cell compartments, modified from Alfaro et al. Proc Natl Acad Sci USA. (2012) 109(19):7280–5
Several proteins that contribute to the development of neurocognitive disease carry O-GlcNAc. The transcription factor methyl CpG binding protein 2 (MeCP2) coordinates activity-dependent gene transcription and is modified by O-GlcNAc. Loss of MeCP2 causes Rett syndrome, a developmental disorder classically associated with repetitive and stereotypical hand movements and mental retardation. It has been shown that only the MeCP2 molecules that do not carry O-GlcNAc were activated in response to neuronal depolarization (Wang et al. 2010a, b; Rexach et al. 2012). Seemingly on MeCP2, O-GlcNAc serves as a checkpoint for turning on or off activity-dependent gene transcription. Other O-GlcNAc proteins that are intimately involved in neurocognitive disease include tau (Alzheimer’s disease) and α-synuclein (Parkinson’s disease) (Arnold et al. 1996; Wang et al. 2010a, b).
16.4.3 O-GlcNAc Regulates Diverse Cellular Processes
Proteins modified by O-GlcNAc regulate diverse cellular processes such as gene transcription, protein translation and degradation and signal transduction. The function of the O-GlcNAc modification depends on the specific protein that is modified as well as the site on that protein that is modified (Hart et al. 2007; Hart et al. 2011). Here we will briefly discuss different mechanisms by which O-GlcNAc underlies normal cell function.
16.4.3.1 Gene Transcription
RNA polymerase II. O-GlcNAc may directly affect basal transcription by inactivating RNA polymerase II. RNA polymerase II in the so-called pre-initiation complex is heavily modified by O-GlcNAc in its carboxyl terminal domain (CTD). However, the RNA polymerase II in the so-called elongation complex is devoid of O-GlcNAc. Instead, upon transcription initiation the CTD of RNA polymerase II becomes phosphorylated. It has been suggested that the loss of O-GlcNAc is required before it can be phosphorylated and thereby activated (Kelly et al. 1993; Comer and Hart 1999; Ranuncolo et al. 2012).
Histones. Histones are modified by multiple posttranslational modifications, including O-GlcNAc (Sakabe et al. 2010). The combination of these modifications is hypothesized to form a “code” that either facilitates or prevents the access of transcription factors to DNA. The O-GlcNAc on serine 112 enhances the (mono-) ubiquitination of histone 2B thereby activating gene transcription (Fujiki et al. 2011).
Transcription factors. O-GlcNAc takes advantage of several different mechanisms to regulate the activity of transcription factors toward gene transcription (Ozcan et al. 2010). O-GlcNAc activates NeuroD by promoting its shuttling from the cytosol to the nucleus and protects p53 by saving it from ubiquitin-mediated degradation (Andrali et al. 2007). O-GlcNAc on serine 228 on Oct4 does not affect its general activity but influences its promoter specificity (Jang et al. 2012).
16.4.3.2 Protein Translation
O-GlcNAc plays several distinct roles during the translation of messenger RNA to polypeptide chains. OGT and OGA bind very tightly to the ribosome and overexpression of OGT was shown to facilitate ribosome assembly. Several ribosomal proteins are modified by O-GlcNAc, including the mammalian target of rapamycin (mTOR) pathway protein S6 (Zeidan et al. 2010). Also associated translational factors are modified by O-GlcNAc. The eukaryotic initiation factor 2 (eIF2) initiates translation by forming a complex with p67. It has been suggested that the O-GlcNAc on p67 is required for p67’s ability to protect eIF2 from phosphorylation and thereby inactivation (Datta et al. 1989. Ray et al. 1992).
16.4.3.3 Protein and Vesicle Trafficking
The subcellular distribution of multiple proteins has been shown to depend on O-GlcNAc (Sayat et al. 2008; Geng et al. 2012). For instance, E-cadherin is a cell junction protein that, among other things, is important for inhibitory synapse formation (Fiederling et al. 2011). If the O-GlcNAc on E-cadherin's cytosolic tail is not removed, E-cadherin becomes trapped in the endoplasmatic reticulum (ER) and cannot be transported further to the Golgi apparatus (Geng et al. 2012). O-GlcNAc may also play a role in the transport of microvesicles within cells, e.g. vesicles that mediate the release of neurotransmitter. There is no direct evidence showing that O-GlcNAc regulates microvesicle trafficking per se. However, proteins such as adaptor protein complex 2 (AP-2) that mediates clathrin-dependent endocytosis and N-ethylmaleimide-sensitive fusion protein (NSF), which is an ATPase involved in vesicle fusion, are modified by O-GlcNAc (Clark et al. 2008).
16.4.3.4 Protein Degradation
O-GlcNAc is intimately involved in the control of protein degradation. O-GlcNAc sites often fall in regions with high PEST-scores (the Pro-Glu-Ser-Thr sequence is associated with rapid degradation of proteins) and an increase in global O-GlcNAc leads to an increased ubiquitination of proteins whereas a decrease in global O-GlcNAc decreases protein ubiquitination (Hart et al. 2007; Guinez et al. 2008). In addition, O-GlcNAc can directly inhibit the proteasome by modifying the 26S and 19S proteasomes (Zang et al. 2003).
16.4.3.5 Signaling and the Crosstalk Between O-GlcNAc and O-Phosphate
An emerging theme in the regulation of signal transduction pathways is the crosstalk between O-GlcNAc and O-phosphate. Many O-phosphorylated proteins are also O-GlcNAc proteins (Trinidad et al. 2012). O-GlcNAc and O-phosphate can directly and reciprocally inhibit each other by sharing the same site, as in the case of threonine 58 in the trans-activation domain of the transcription factor c-myc (Chou et al. 1995a; Chou et al. 1995b). On other proteins the crosstalk is indirect and can be both negative and positive. Removal of O-GlcNAc from threonine 57/serine 58 on CaMKIV, for example, prevents the phosphorylation of threonine 200, the main activation site on CaMKIV. At the same time, loss of O-GlcNAc on serine 189 facilitates threonine 200 phosphorylation (Dias et al. 2009). In fact, on a global level, the occupancy of most dynamic phosphorylation sites is affected by acute inhibition of O-GlcNAc cycling (Wang et al. 2008). As a group, kinases, the enzymes that add O-phosphate to proteins, are more often modified by O-GlcNAc than other kinds of proteins (Dias et al. 2012; Trinidad et al. 2012). Furthermore, as we discussed in Sect. 16.2.2, phosphorylation of OGT can activate OGT and influence its substrate specificity.
16.5 O-GlcNAc Is Essential for Brain Function
In Sects. 16.1–16.4 we have learned that O-GlcNAc dynamically modifies a vast array of neuronal proteins involved in many cellular processes, such as gene transcription and signal transduction. In the following section we will discuss how the regulation of neuronal function by O-GlcNAc underlies normal brain function and contributes to neurocognitive disease (summarized in Fig. 16.4).
O-GlcNAc underlies normal brain development and function and contributes to the development of neurocognitive disease by regulating diverse cellular processes. (A) O-GlcNAc is expressed in the brain from early development until late in life. O-GlcNAc is essential for brain development and modulates many normal functions in the adult, e.g. learning and memory. Disturbed O-GlcNAc cycling may contribute to the pathology behind many neurodegenerative diseases. How O-GlcNAc impacts neuron function is exemplified in (B). (a) When the transcription factor CREB is modified by O-GlcNAc, it cannot bind the coactivator CRTC. This leads to the repression of many genes that are involved in synaptic plasticity. (b) Tau is normally modified by O-GlcNAc. In Alzheimer’s disease, tau loses its O-GlcNAc and precipitates into cytotoxic aggregates. (c) It is believed that dynamic changes in synaptic function underlie learning and memory. At presynaptic terminals, O-GlcNAc may regulate synaptic plasticity by affecting the recycling of neurotransmitter-containing vesicles, possibly through affecting the function of the vesicle-binding protein synapsin I. In the postsynapse, the O-GlcNAc effect on learning and memory may be related to modulation of neurotransmitter receptor trafficking. (d) One mechanism by which O-GlcNAc impacts normal development is its regulation of axonal growth and branching
16.5.1 Early and Late Brain Development Depends Upon O-GlcNAc Cycling
In vertebrates, O-GlcNAc is essential for normal development. In many different cell types, it has been shown that perturbations of O-GlcNAc cycling interrupt the cell cycle by preventing cytokinesis (Slawson et al. 2005; Slawson et al. 2008; Wang et al. 2010a, b). In the embryo, this leads to failure of stem cell proliferation and subsequent death at the single cell stage (Shafi et al. 2000). Studies taking advantage of partial depression of OGT expression, where cell proliferation is preserved, indicated that renewal of stem cell pluripotency and stem cell differentiation depends upon O-GlcNAc. In fact, two members of the core pluripotency network in embryonic stem cells, Oct4 and Sox2, are modified by O-GlcNAc. Loss of O-GlcNAc on threonine 228 of Oct4 alters its promoter specificity disrupting transcription of dozens of genes, including some that are involved in neuronal differentiation (e.g. Nanog) (Jang et al. 2012; Wang et al. 2012).
Later in development, O-GlcNAc plays a role in brain morphogenesis and patterning. Loss of OGT decreases brain size, while overexpression of OGT blurs the organization and distinction of hind-, mid-, and forebrain regions in zebra fish (Webster et al. 2009). Likewise, when OGT was deleted from neurons specifically by crossing floxed OGT mice with mice expressing Cre recombinase under the synapsin I promoter, viability until term was reduced. The pups that survived failed to develop any locomotor activity and died within 10 days (O'Donnell et al. 2004). In postmitotic neurons, the O-GlcNAc glycosylation of CREB underlies both axonal and dendritic growth and may, at least partly, explain the impaired development of brain function (Rexach et al. 2012). O-GlcNAc is also known to affect axonal branching (Francisco et al. 2009).
16.5.2 O-GlcNAc Underlies Learning and Memory
Learning and memory are fundamental properties of the brain. Over the past decades it has been shown that the brain is a highly malleable organ that dynamically responds to challenges in the environment. It is believed that memories are encoded by neurons through changes in their synaptic communications with other neurons, so-called synaptic plasticity. A prevailing model for how such changes occur is long-term potentiation (LTP) and depression (LTD), occurring mainly at the postsynapse (see, e.g., Hanley 2008; Lynch 2004). Multiple disorders involving mental retardation such as Fragile X and Rett syndrome have been linked to proteins underlying synaptic plasticity (Verpelli and Sala 2012; Grant 2012). OGT is present in the synapse and many postsynaptic proteins involved in LTP and LTD carry O-GlcNAc (see Sects. 16.3.1–2 and 16.4.2–3). Using electrophysiology, it was further established that O-GlcNAc cycling is necessary for normal expression of LTP; pharmacological inhibition of OGT diminished LTP whereas OGA inhibition elevated LTP (Tallent et al. 2009). The effect may relate to the synaptic relocation of the α-amino-3-hydroxy-5-methyl-4-isoxazoleproprionic (AMPA) receptor, the glutamate neurotransmitter receptor responsible for most fast synaptic transmission in the central nervous system (CNS). O-GlcNAc modifies CaMKII in close proximity to a phosphorylation site that regulates its activity (see Sect. 16.3.2). CaMKII, in turn, is one of the major pathways that can induce the synaptic insertion of the AMPA receptor (Tallent et al. 2009; Shepherd and Huganir 2007). Interestingly, an OGT-dependent increase in AMPA receptor conductivity was absent in AMPA receptors where the main CaMKII phosphorylation site was mutated to alanine (Kanno et al. 2010).
O-GlcNAc cycling is important for dynamic synaptic function at the presynapse. Acute OGA inhibition in vivo decreases paired-pulse facilitation (PPF). PPF is a measurement of vesicle release of neurotransmitter. O-GlcNAc may regulate PPF via synapsin I. The presynaptic protein synapsin I is extensively modified by O-GlcNAc and inhibition of OGA increases the phosphorylation of synapsin I (Tallent et al. 2009).
The studies investigating the effect of O-GlcNAc on learning and memory have so far relied heavily upon pharmacological manipulation of OGT and OGA. Unfortunately, some of the drugs used are known to cause off-target effects in cells. This may explain some apparent contradictions in published results (e.g. Tallent et al. 2009 argues that O-GlcNAc does not affect basal synaptic transmission whereas Kanno et al. 2010 does see a large effect on basal AMPA receptor function). For future research, it will be important to develop mouse models of O-GlcNAc function and focus on mechanisms for the way in which O-GlcNAc may underlie learning and memory. One good example is a recent study that compared wild-type CREB with an O-GlcNAc loss-of-function mutant of CREB. This study overexpressed these proteins in vivo and showed that the O-GlcNAc mutant enhanced CREB-dependent transcription of several synaptic plasticity associated genes as well as short-term memory in mice (Rexach et al. 2012).
16.5.3 Impaired O-GlcNAc Cycling Contributes to Neurodegenerative Disease
Neurodegenerative disease is a collection of disorders that are characterized by general cognitive decline and loss of neurons and neuronal synapses. They become more common with age but early-onset variants exist (Koffie et al. 2011; Schulz-Schaeffer 2010). Impaired O-GlcNAc cycling is implicated in the development of several types of neurodegenerative disease, including Alzheimer's, Parkinsonism, and Huntington's (Arnold et al. 1996; Dias and Hart 2007; Wang et al. 2012; Hart et al. 2011). The loci for the genes encoding OGT and OGA are linked to Parkinsonian dystonia and Alzheimer's disease, respectively. In addition, OGA transcripts are alternatively spliced in the brain in patients with sporadic Alzheimer’s (see Sects. 16.2.2–16.3).
Two pathological hallmarks of Alzheimer’s disease are neurofibrillary tangles and amyloid plaques (Koffie et al. 2011; Dias and Hart 2007). The neurofibrillary tangles consist of paired helical filaments of tau, a protein that is important for microtubuli stability. Tau is extensively modified by O-GlcNAc, including in the region that binds to microtubuli (Arnold et al. 1996; Wang et al. 2010a, b). Results from both in vitro and in vivo experiments indicated that increasing O-GlcNAc on tau protected it from filament precipitation (Yuzwa et al. 2012). O-GlcNAc may save tau from precipitating either directly or indirectly by protecting it from hyperphosphorylation (Yuzwa et al. 2012; Yuzwa et al. 2008). Alzheimer’s patients present with hyperphosphorylated and hypo-O-GlcNAc glycosylated tau. No O-GlcNAc can be found on the tau tangles isolated from these patients (Liu et al. 2004). Importantly, in mouse models of neurodegenerative disease resulting from expressing mutant forms of tau prone to precipitate, elevating global O-GlcNAc, including O-GlcNAc on tau, by selectively inhibiting OGA pharmacologically, slowed symptom progression (Yuzwa et al. 2012). Interestingly, the main contributor amyloid plaques, amyloid precursor protein (APP), is also modified by O-GlcNAc (Griffith et al. 1995).
Alzheimer's disease is intimately associated with dysregulated glucose metabolism and insulin resistance (Biessels et al. 2006; de la Monte and Wands 2008). Due to the regulation of O-GlcNAc by nutrient supply and metabolic hormones, OGT has emerged as a key nutrient sensor in cells (see Sect. 16.2.2). Much of the toxicity associated with excessive intake of glucose is mediated by the hexosamine biosynthesis pathway, the same pathway that produces the O-GlcNAc donor substrate UDP-GlcNAc. Many forms of insulin resistance are also related to OGT (Hart et al. 2007). Overexpression of OGT can independently lead to insulin resistance (Yang et al. 2008). From this perspective, impaired O-GlcNAc signaling represents a model, and pharmacological target, for how metabolic dysfunction may result in neurodegenerative disease.
O-GlcNAc may also be involved in neurodegenerative diseases other than Alzheimer’s. For example, in transgenic models where a polyglutamine expansion of huntingtin (the protein that causes Huntington's disease) was overexpressed, deletion of OGT attenuated neuron loss while deletion of OGA exacerbated neuron loss (Wang et al. 2012). In addition, α-synuclein, the protein found in lesions overrepresented in Parkinson's disease, carries O-GlcNAc in the region known to induce self-aggregation (Wang et al. 2010a, b). It should also be noted that O-GlcNAc in other cell types becomes elevated from many different types of stress and that deletion of OGT makes cells more vulnerable to stress (Zachara et al. 2004; Kazemi et al. 2010).
16.6 Summary and Outlook
Every time we think, act, and feel information is passed between neurons in the brain. Neurons are extraordinary cells with complex morphology and a lifespan that matches the lifespan of the organism. The posttranslational modification O-GlcNAc, the attachment of β-N-acetylglucosamine to serine and threonine, is expressed in all compartments of the neuron. By the help of only two enzymes, OGT and OGA, O-GlcNAc regulates thousands of proteins underlying diverse cellular processes such as gene transcription and signal transduction. The big challenge for the future will be to derive a mechanistic picture for how specific O-GlcNAc sites affect protein function and thereby synaptic transmission. Recently, several new techniques have been developed that improve and simplify the detection of O-GlcNAc. Nevertheless, more tools such as site-specific antibodies for O-GlcNAc are needed to make the O-GlcNAc field more accessible to the broader neuroscience research community. Understanding the way O-GlcNAc works in the nervous system will not only afford fundamental principles of brain function but also provide novel targets for the treatment of neurocognitive disease.
References
Akimoto Y, Comer FI, Cole RN, Kwakami AKH, Hirano H, Hart GW. Localization of the O-GlcNAc transferase and O-GlcNAc-modified proteins in rat cerebellar cortex. Brain Res. 2003;966(2):194–205.
Alfaro JF, Gong CX, Monroe ME, Aldrich JT, Clauss TR, Purvine SO, et al. Tandem mass spectrometry identifies many mouse brain O-GlcNAcylated proteins including EGF domain-specific O-GlcNAc transferase targets. Proc Natl Acad Sci U S A. 2012;109(19):7280–5.
Andrali SS, Qian Q, Ozcan S. Glucose mediates the translocation of NeuroD1 by O-linked glycosylation. J Biol Chem. 2007;282(21):15589–96.
Arnold SC, Johnson GVW, Cole RN, Dong DLY, Lee M, Hart GW. The microtubule-associated protein tau is extensively modified with O-linked N-acetylglucosamine. J Biol Chem. 1996;271(46):28741–4.
Banerjee S, Robbins PW, Samuelson J. Molecular characterization of nucleocytosolic O-GlcNAc transferases of Giardia lamblia and Cryptosporidium parvum. Glycobiology. 2009;19(4):331–6.
Banerjee PS, Hart GW, Cho JW. Chemical approaches to study O-GlcNAcylation. Chem Soc Rev. 2013;42(10):4345–57.
Benko DM, Haltiwanger RS, Hart GW, Gibson W. Virion basic phosphoprotein from human cytomegalovirus contains O-linked N-acetylglucosamine. Proc Natl Acad Sci U S A. 1988;85(8):2573–7.
Bertram L, Blacker D, Mullin K, Keeney D, Jones J, Basu S, et al. Evidence for genetic linkage of Alzheimer’s disease to chromosome 10q. Science. 2000;290(5500):2302–3.
Biessels GJ, Staekenborg S, Brunner E, Brayne C, Scheltens P. Risk of dementia in diabetes mellitus: a systematic review. Lancet Neurol. 2006;5(1):64–74.
Bullen, Hart. AMPK regulates OGT substrate specificity. Forthcoming paper.
Butkinaree C, Cheung WD, Park S, Park K, Barbr M, Hart GW. Characterization of beta-N-acetylglucosaminidase cleavage by caspase-3 during apoptosis. J Biol Chem. 2008;283(35):23557–66.
Caillet-Boudin ML, Strecker G, Michalski JC. O-linked GlcNAc in serotype-2 adenovirus fibre. Eur J Biochem. 1989;184(1):205–11.
Carillo LD, Froemming JA, Mahal LK. Targeted in vivo O-GlcNAc sensors reveal discrete compartment-specific dynamics during signal transduction. J Biol Chem. 2011;286(8):6650–8.
Cheung WD, Hart GW. AMP-activated protein kinase and p38 MAPK activate O-GlcNAcylation of neuronal proteins during glucose deprivation. J Biol Chem. 2008;283(19):13009–20.
Cheung WD, Sakabe K, Housley MP, Dias WB, Hart GW. O-linked beta-N-acetylglucosaminyltransferase substrate specificity is regulated by myosin phosphatase targeting and other interacting proteins. J Biol Chem. 2008;283(49):33935–41.
Chou TY, Dang CV, Hart GW. Glycosylation of the c-Myc transactivation domain. Proc Natl Acad Sci U S A. 1995a;92(10):4417–21.
Chou TY, Hart GW, Dang CV. c-Myc is glycosylated at threonine 58, a known phosphorylation site and a mutational hot spot in lymphomas. J Biol Chem. 1995b;270(32):18961–5.
Clark PM, Dweck JF, Mason DE, Hart CR, Buck SB, Peters EC, et al. Direct in-gel fluorescence detection and cellular imaging of O-GlcNAc-modified proteins. J Am Chem Soc. 2008;130(35):11576–7.
Cole RN, Hart GW. Cytosolic O-glycosylation is abundant in nerve terminals. J Neurochem. 2001;79(5):1080–9.
Comer FI, Hart GW. O-GlcNAc and the control of gene expression. Biochim Biophys Acta. 1999;1473(1):161–71.
Datta B, Ray MK, Chakrabarti D, Wylie DE, Gupta NK. Glycosylation of eukaryotic peptide chain initiation factor 2 (eIF-2)-associated 67-kDa polypeptide (p67) and its possible role in the inhibition of eIF-2 kinase-catalyzed phosphorylation of the eIF-2 alpha-subunit. J Biol Chem. 1989;264(34):20620–4.
de la Monte SM, Wands JR. Alzheimer’s disease is type 3 diabetes-evidence reviewed. J Diabetes Sci Technol. 2008;2(6):1101–13.
Dennis RJ, Taylor EJ, Macauley MS, Stubbs KA, Turkenburg JP, Hart SJ, et al. Structure and mechanism of a bacterial beta-glucosaminidase having O-GlcNAcase activity. Nat Struct Mol Biol. 2006;13(4):365–71.
Dias WB, Hart GW. O-GlcNAc modification in diabetes and Alzheimer’s disease. Mol Biosyst. 2007;3(11):766–72.
Dias WB, Cheung WD, Wang Z, Hart GW. Regulation of calcium/calmodulin-dependent kinase IV by O-GlcNAc modification. J Biol Chem. 2009;284(32):21327–37.
Dias WB, Cheung WD, Hart GW. O-GlcNAcylation of kinases. Biochem Biophys Res Commun. 2012;422(2):224–8.
Dong DL, Hart GW. Purification and characterization of an O-GlcNAc selective N-acetyl-beta-D-glucosaminidase from rat spleen cytosol. J Biol Chem. 1994;269(30):19321–30.
Fang B, Miller MW. Use of galactosyltransferase to assess the biological function of O-linked N-acetyl-d-glucosamine: a potential role for O-GlcNAc during cell division. Exp Cell Res. 2001;263(2):243–53.
Fiederling A, Ewert R, Andreyeva A, Jungling K, Gottmann K. E-cadherin is required at GABAergic synapses in cultured cortical neurons. Neurosci Lett. 2011;501(3):167–72.
Francisco H, Kollins K, Varghis N, Vocadlo D, Vosseller K, Gallo G. O-GLcNAc post-translational modifications regulate the entry of neurons into an axon branching program. Dev Neurobiol. 2009;69(2–3):162–73.
Fredriksen L, Mathiesen G, Moen A, Bron PA, Kleerebezem M, Eijsink VG, et al. The major autolysin Acm2 from Lactobacillus plantarum undergoes cytoplasmic O-glycosylation. J Bacteriol. 2012;194(2):325–33.
Fujiki R, Hashiba W, Sekine H, Yokoyama A, Chikanishi T, Ito S, et al. GlcNAcylation of histone H2B facilitates its monoubiquitination. Nature. 2011;480(7378):557–60.
Gao Y, Wells L, Comer FI, Parker GJ, Hart GW. Dynamic O-glycosylation of nuclear and cytosolic proteins: cloning and characterization of a neutral, cytosolic beta-N-acetylglucosaminidase from human brain. J Biol Chem. 2001;276(13):9838–45.
Geng F, Zhi W, Anderson RA, Leber B, Andrews DW. Multiple post-translational modifications regulate E-cadherin transport during apoptosis. J Cell Sci. 2012;125(Pt 11):2615–25.
Grant SG. Synaptopathies: diseases of the synaptome. Curr Opin Neurobiol. 2012;22(3):522–9.
Griffith LS, Mathes M, Schmitz B. Beta-amyloid precursor protein is modified with O-linked N-acetylglucosamine. J Neurosci Res. 1995;41(2):270–8.
Guinez C, Mir AM, Dehennaut V, Cacan R, Harduin-Lepers A, Michalski JC, et al. Protein ubiquitination is modulated by O-GlcNAc glycosylation. FASEB J. 2008;22(8):2901–11.
Haltiwanger RS, Holt GD, Hart GW. Enzymatic addition of O-GlcNAc to nuclear and cytoplasmic proteins. Identification of a uridine diphospho-N-acetylglucosamine:peptide beta-N-acetylglucosaminyltransferase. J Biol Chem. 1990;265(5):2563–8.
Haltiwanger RS, Blomberg MA, Hart GW. Glycosylation of nuclear and cytoplasmic proteins. Purification and characterization of a uridine diphospho-N-acetylglucosamine:polypeptide beta-N-acetylglucosaminyltransferase. J Biol Chem. 1992;267(13):9005–13.
Hanley JG. AMPA receptor trafficking pathways and links to dendritic spine morphogenesis. Cell Adh Migr. 2008;2(4):276–82.
Hanover JA, Song Y, Lubas WB, Shin SH, Ragano-Caracciola M, Kochran J, et al. Mitochondrial and nucleocytoplasmic isoforms of O-linked GlcNAc transferase encoded by a single mammalian gene. Arch Biochem Biophys. 2003;409(2):287–97.
Hart GW, Housley MP, Slawson C. Cycling of O-linked beta-N-acetylglucosamine on nucleocytoplasmic proteins. Nature. 2007;26(7139):1017–22.
Hart GW, Slawson C, Ramirez-Correa G, Lagerlof O. Cross talk between O-GlcNAcylation and phosphorylation: roles in signaling, transcription, and chronic disease. Annu Rev Biochem. 2011;80:825–58.
Heckel D, Comtesse N, Brass N, Blin N, Zang KD, Meese E. Novel immunogenic antigen homologous to hyaluronidase in meningioma. Hum Mol Genet. 1998;7(12):1859–72.
Holt GD, Hart GW. The subcellular distribution of terminal N-acetylglucosamine moieties. Localization of a novel protein-saccharide linkage, O-linked GlcNAc. J Biol Chem. 1986;26(17):8049–57.
Holt GD, Snow CM, Senior A, Haltiwanger RS, Gerace L, Hart GW. Nuclear pore complex glycoproteins contain cytoplasmically disposed O-linked N-acetylglucosamine. J Cell Biol. 1987;104(5):1157–64.
Housley MP, Udeshi ND, Rodgers JT, Shabanowitz J, Puigserver P, Hunt DF, et al. A PGC-1alpha-O-GlcNAc transferase complex regulates FoxO transcription factor activity in response to glucose. J Biol Chem. 2009;284(8):5148–57.
Jang H, Kim TW, Yoon S, Choi SY, Kang TW, Kim SY, et al. O-GlcNAc regulates pluripotency and reprogramming by directly acting on core components of the pluripotency network. Cell Stem Cell. 2012;11(1):62–74.
Jinek M, Rehwinkel J, Lazarus BD, Izaurralde E, Hanover JA, Conti E. The superhelical TPR-repeat domain of O-linked GlcNAc transferase exhibits structural similarities to importin alpha. Nat Struct Mol Biol. 2004;11(10):1001–7.
Kanno T, Yaguchi T, Nagata T, Mukasa T, Nishizaki T. Regulation of AMPA receptor trafficking by O-glycosylation. Neurochem Res. 2010;35(5):782–8.
Kazemi Z, Chang H, Haserodt S, McKen C, Zachara NE. O-linked beta-N-acetylglucosamine (O-GlcNAc) regulates stress-induced heat shock protein expression in a GSK-3beta-dependent manner. J Biol Chem. 2010;285(50):39096–107.
Kearse KP, Hart GW. Lymphocyte activation induces rapid changes in nuclear and cytoplasmic glycoproteins. Proc Natl Acad Sci U S A. 1991;88(5):1701–5.
Kelly WG, Dahmus ME, Hart GW. RNA polymerase II is a glycoprotein. Modification of the COOH-terminal domain by O-GlcNAc. J Biol Chem. 1993;268(14):10416–24.
Khidekel N, Ficarro SB, Clark PM, Bryan MC, Swaney LD, Rexach JE, et al. Probing the dynamics of O-GlcNAc glycosylation in the brain using quantitative proteomics. Nat Chem Biol. 2007;3(6):339–48.
Koffie RM, Hyman BT, Spires-Jones TL. Alzheimer’s disease: synapses gone cold. Mol Neurodegener. 2011;6(1):63.
Kreppel LK, Hart GW. Regulation of a cytosolic and nuclear O-GlcNAc transferase. Role of the tetratricopeptide repeats. J Biol Chem. 1999;274(45):32015–22.
Kreppel LK, Blomberg MA, Hart GW. Dynamic glycosylation of nuclear and cytosolic proteins. Cloning and characterization of a unique O-GlcNAc transferase with multiple tetratricopeptide repeats. J Biol Chem. 1997;272(14):9308–15.
Lazarus MB, Nam Y, Jian J, Sliz P, Walker S. Structure of human O-GlcNAc transferase and its complex with a peptide substrate. Nature. 2011;469(7331):564–7.
Li X, Lu F, Wang JZ, Gong CX. Concurrent alterations of O-GlcNAcylation and phosphorylation of tau in mouse brains during fasting. Eur J Neurosci. 2006;23(8):2078–86.
Lisman J, Schulman H, Cline H. The molecular basis of CaMKII function in synaptic and behavioural memory. Nat Rev Neurosci. 2002;3(3):175–90.
Liu F, Iqbal K, Grundke-Iqbal I, Hart GW, Gong CX. O-GlcNAcylation regulates phosphorylation of tau: a mechanism involved in Alzheimer’s disease. Proc Natl Acad Sci U S A. 2004;101(29):10804–9.
Liu Y, Li X, Yu Y, Shi J, Liang Z, Run X, et al. Developmental regulation of protein O-GlcNAcylation, O-GlcNAc transferase and O-GlcNAcase in mammalin brain. PLoS One. 2012;7(8):e43724.
Lubas WA, Frank DW, Krause M, Hanover JA. O-Linked GlcNAc transferase is a conserved nucleocytoplasmic protein containing tetratricopeptide repeats. J Biol Chem. 1997;272(14):9316–24.
Lyer SPN, Hart GW. Roles of the tetratricopeptide repeat domain in O-GlcNAc transferase targeting and protein substrate specificity. J Biol Chem. 2003;278(27):24608–16.
Lyer SPN, Akimoto Y, Hart GW. Identification and cloning of a novel family of coiled-coil domain proteins that interact with O-GlcNAc transferase. J Biol Chem. 2003;278(7):5399–409.
Lynch MA. Long-term potentiation and memory. Physiol Rev. 2004;84(1):87–136.
Macauley MS, Whitworth GE, Debowski AW, Chin D, Vocadlo DJ. O-GlcNAcase uses substrate-assisted catalysis: kinetic analysis and development of highly selective mechanism-inspired inhibitors. J Biol Chem. 2005;280(27):25313–22.
Marotta NP, Cherwien CA, Abeywardana T, Pratt MR. O-GlcNAc modification prevents peptide-dependent acceleration of α-synuclein aggregation. Chembiochem. 2012;13(18):2665–70.
Marz P, Stetefeld J, Bendfeldt K, Nitsch C. Reinstein, Shoeman RL et al. Ataxin-10 interacts with O-linked beta-N-acetylglucosamine transferase in the brain. J Biol Chem. 2006;281(29):20263–70.
Miller MW, Caracciolo MR, Berlin WK, Hanover JA. Phosphorylation and glycosylation of nucleoporins. Arch Biochem Biophys. 1999;367(1):51–60.
Muller U, Steinberger D, Nemeth AH. Clinical and molecular genetics of primary dystonias. Neurogenetics. 1998;1(3):165–77.
Nolte D, Muller U. Human O-GlcNAc transferase (OGT): genomic structure, analysis of splice variants, fine mapping in Xq13.1. Mamm Genome. 2002;13(1):62–4.
O'Donnell N, Zachara NE, Hart GW, Marth JD. Ogt-dependent X-chromosome-linked protein glycosylation is a requisite modification in somatic cell function and embryo viability. Mol Cell Biol. 2004;24(2):1680–90.
Ozcan S, Andrali SS, Cantrell JE. Modulation of transcription factor function by O-GlcNAc modification. Biochim Biophys Acta. 2010;1799(5–6):353–64.
Ranuncolo SM, Ghosh S, Hanover JA, Hart GW, Lewis BA. Evidence of the involvement of O-GlcNAc-modified human RNA polymerase II CTD in transcription in vitro and in vivo. J Biol Chem. 2012;287(28):23549–61.
Ray MK, Datta B, Chakraborty A, Chattopadhyay A, Meza-Keuthen S, Gupta NK. The eukaryotic initiation factor 2-associated 67-kDa polypeptide (p67) plays a critical role in regulation of protein synthesis initiation in animal cells. Proc Natl Acad Sci U S A. 1992;89(2):539–43.
Rexach JE, Rogers CJ, Yu SH, Tao J, Sun YE, Hsieh-Wilson LC. Quantification of O-glycosylation stoichiometry and dynamics using resolvable mass tags. Nat Chem Biol. 2010;6(9):645–51.
Rexach JE, Clark PM, Mason DE, Neve RL, Peters EC, Hsieh-Wilson LC. Dynamic O-GlcNAc modification regulates CREB-mediated gene expression and memory formation. Nat Chem Biol. 2012;89(3):253–61.
Roquemore EP, Chevrier MR, Cotter RJ, Hart GW. Dynamic O-GlcNAcylation of the small heat shock protein alpha B-crystallin. Biochemistry. 1996;35(11):3578–86.
Sakabe K, Wang Z, Hart GW. Beta-N-acetylglucosamine (O-GlcNAc) is part of the histone code. Proc Natl Acad Sci U S A. 2010;107(46):19915–20.
Sayat R, Leber B, Grubac V, Wiltshire L, Persad S. O-GlcNAc-glycosylation of beta-catenin regulates its nuclear localization and transcriptional activity. Exp Cell Res. 2008;314(15):2774–87.
Schirm M, Kalmokoff M, Aubry AP, Thibault P, Sandoz M, Logan SM. Flagellin from Listeria monocytogenes is glycosylated with beta-O-linked N-acetylglucosamine. J Bacteriol. 2004;186(20):6721–7.
Schulz-Schaeffer WJ. The synaptic pathology of alpha-synuclein aggregation in dementia with Lewy bodies, Parkinson’s disease and Parkinson’s disease dementia. Acta Neuropathol. 2010;120(2):131–43.
Selzer J, Hofmann F, Rex G, Wilm M, Mann M, Just I, Aktories K. Clostridium novyi alpha-toxin-catalyzed incorporation of GlcNAc into Rho subfamily proteins. J Biol Chem. 1996;271(41):25173–7.
Shafi R, Lyer SPN, Ellies LG, O’Donnell N, Marek KW, Chui D, et al. The O-GlcNAc transferase gene resides on the X chromosome and is essential for embryonic stem cell viability and mouse ontogeny. Proc Natl Acad Sci U S A. 2000;97(11):5735–9.
Shen DL, Gloster TM, Yuzwa SA, Vocadlo DJ. Insights into O-linked N-acetylglucosamine ([0-9]O-GlcNAc) processing and dynamics through kinetic analysis of O-GlcNAc transferase and O-GlcNAcase activity on protein substrates. J Biol Chem. 2012;287(19):15395–408.
Shepherd JD, Huganir RL. The cell biology of synaptic plasticity: AMPA receptor trafficking. Annu Rev Cell Dev Biol. 2007;23:613–43.
Slawson C, Zachara NE, Vosseller K, Cheung WD, Lane MD, Hart GW. Perturbations in O-linked beta-N-acetylglucosamine protein modification cause severe defects in mitotic progression and cytokinesis. J Biol Chem. 2005;280(38):32944–56.
Slawson C, Lakshmanan T, Knapp S, Hart GW. A mitotic GlcNAcylation/phosphorylation signaling complex alters the posttranslational state of the cytoskeletal protein vimentin. Mol Biol Cell. 2008;19(10):4130–40.
Song M, Kim HS, Park JM, Kim SH, Kim IH, Ryu SH, et al. o-GlcNAc transferase is activated by CaMKIV-dependent phosphorylation under potassium chloride-induced depolarization in NG-108-15 cells. Cell Signal. 2008;20(1):94–104.
Tallent MK, Varghis N, Skorobogatko Y, Hernandez-Cuebas L, Whelan K, Vocadlo DJ, et al. In vivo modulation of O-GlcNAc levels regulates hippocampal synaptic plasticity through interplay with phosphorylation. J Biol Chem. 2009;284(1):174–81.
Toleman C, Paterson AJ, Whisenhunt TR, Kudlow JE. Characterization of the histone acetyltransferase (HAT) domain of a bifunctional protein with activable O-GlcNAcase and HAT activities. J Biol Chem. 2004;279(51):53665–73.
Toleman CA, Paterson AJ, Kudlow JE. The histone acetyltransferase NCOAT contains a zinc finger-like motif involved in substrate recognition. J Biol Chem. 2006;281(7):3918–25.
Torres CR, Hart GW. Topography and polypeptide distribution of terminal N-acetylglucosamine residues on the surfaces of intact lymphocytes. Evidence for O-linked GlcNAc. J Biol Chem. 1984;259(5):3308–17.
Trinidad JC, Barkan DT, Gulledge BF, Thalhammer A, Sali A, Shoepfer R, et al. Global identification and characterization of both O-GlcNAcylation and phosphorylation at the murine synapse. Mol Cell Proteomics. 2012;11(8):215–29.
Twine NA, Janitz K, Wilkins MR, Janitz M. Whole transcriptome sequencing reveals gene expression and splicing differences in brain regions affected by Alzheimer’s disease. PLoS One. 2011;6(1):e16266.
Verpelli C, Sala C. Molecular and synaptic defects in intellectual disability syndromes. Curr Opin Neurobiol. 2012;22(3):530–6.
Wang Z, Gucek M, Hart GW. Cross-talk between GlcNAcylation and phosphorylation: site-specific phosphorylation dynamics in response to globally elevated O-GlcNAc. Proc Natl Acad Sci U S A. 2008;105(37):13793–8.
Wang Z, Udeshi ND, O’Malley M, Shabanowitz J, Hunt DF, Hart GW. Enrichment and site mapping of O-linked N-acetylglucosamine by a combination of chemical/enzymatic tagging, photochemical cleavage, and electron transfer dissociation mass spectrometry. Mol Cell Proteomics. 2010a;9(1):153–60.
Wang Z, Namrata UD, Slawson C, Compton PD, Sakabe K, Cheung WD, et al. Extensive crosstalk between O-GlcNAcylation and phosphorylation regulates cytokinesis. Sci Signal. 2010b;3(104):ra2.
Wang P, Lazarus BD, Forsythe ME, Love DC, Krause MW, Hanover JA. O-GlcNAc cycling mutants modulate proteotoxicity in Caenorhabditis elegans models of human neurodegenerative diseases. Proc Natl Acad Sci U S A. 2012;109(43):17669–74.
Webster DM, Teo CF, Sun Y, Wloga D, Gay S, Klonowski KD, et al. O-GlcNAc modifications regulate cell survival and epiboly during zebrafish development. BMC Dev Biol. 2009;9:28.
Whelan SA, Lane MD, Hart GW. Regulation of the O-linked beta-N-acetylglucosamine transferase by insulin signaling. J Biol Chem. 2008;283(31):21411–7.
Whelan SA, Dias WB, Thiruneelakantapillai L, Lane MD, Hart GW. Regulation of insulin receptor substrate 1 (IRS-1)/AKT kinase-mediated insulin signaling by O-Linked beta-N-acetylglucosamine in 3T3-L1 adipocytes. J Biol Chem. 2010;285(8):5204–11.
Yang X, Zhang F, Kudlow JE. Recruitment of O-GlcNAc transferase to promoters by corepressor mSin3A: coupling protein O-GlcNAcylation to transcriptional repression. Cell. 2002;110(1):69–80.
Yang X, Ongusaha PP, Miles PD, Havstad JC, Zhang F, So WV, et al. Phosphoinositide signalling links O-GlcNAc transferase to insulin resistance. Nature. 2008;451(7181):964–9.
Yuzwa SA, Macauley MS, Heinonen JE, Shan X, Dennis RJ, He Y, et al. A potent mechanism-inspired O-GlcNAcase inhibitor that blocks phosphorylation of tau in vivo. Nat Chem Biol. 2008;4(8):483–90.
Yuzwa SA, Shan X, Macauley MS, Clark T, Skorobogatko Y, Vosseller K, et al. Increasing O-GlcNAc slows neurodegeneration and stabilizes tau against aggregation. Nat Chem Biol. 2012;8(4):393–9.
Zachara NE, O’Donnell N, Cheung WD, Mercer JJ, Marth JD, Hart GW. Dynamic O-GlcNAc modification of nucleocytoplasmic proteins in response to stress. A survival response of mammalian cells. J Biol Chem. 2004;279(29):30133–42.
Zeidan Q, Wang Z, De Maio A, Hart GW. O-GlcNAc cycling enzymes associate with the translational machinery and modify core ribosomal proteins. Mol Biol Cell. 2010;21(12):1922–36.
Zhang F, Su K, Yang X, Bowe DB, Paterson AJ, Kudlow JE. O-GlcNAc modification is an endogenous inhibitor of the proteasome. Cell. 2003;115(6):715–25.
Conflicts of Interest
The authors declare no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this chapter
Cite this chapter
Lagerlöf, O., Hart, G.W. (2014). O-GlcNAcylation of Neuronal Proteins: Roles in Neuronal Functions and in Neurodegeneration. In: Yu, R., Schengrund, CL. (eds) Glycobiology of the Nervous System. Advances in Neurobiology, vol 9. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-1154-7_16
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1154-7_16
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-1153-0
Online ISBN: 978-1-4939-1154-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)