The development of SSRs designed for rose has accelerated in the last several years and is now a focal point of the map development especially as we move towards developing a consensus map to combine the mapping data from the various maps that have been developed. In the past 5 years, there have been reports from Europe (Esselink et al., 2003; Debener, personal communication; Yan et al., 2005a; Hibrand-Saint Oyant et al., 2007), the USA (Zhang et al., 2006), and Japan (Kimura et al., 2006) describing the development of both genomic (256 primer pairs) and EST (44 primer pairs) based microsatellites.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 DNA Markers Available and Future Developments
The development of SSRs designed for rose has accelerated in the last several years and is now a focal point of the map development especially as we move towards developing a consensus map to combine the mapping data from the various maps that have been developed. In the past 5 years, there have been reports from Europe (Esselink et al., 2003; Debener, personal communication; Yan et al., 2005a; Hibrand-Saint Oyant et al., 2007), the USA (Zhang et al., 2006), and Japan (Kimura et al., 2006) describing the development of both genomic (256 primer pairs) and EST (44 primer pairs) based microsatellites. Of these, 168 of the genomic primer pair sequences are proprietary and are available via a MTA (material transfer agreement) with restrictions on their commercial use in breeding or in variety protection (Esselink et al., 2003; Yan et al., 2005a; Debener, personal communication, ConCipio) (Table 1).
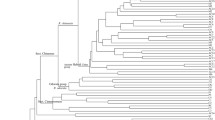
Mapping work has shown that each of these major groups of microsatellites (Rh, Rw, and EST-SSRs) map to all 7 of the linkage groups of rose (Yan et al., 2005a; Zhang et al., 2006; Hibrand-Saint Oyant et al., 2007). The only set of SSRs mapped to several maps are those developed in the USA (Zhang et al., 2006). These 30 SSRs have been mapped to the 90–69 map (Zhang et al., 2006), the 97/7 map (Zhang, 2003), and the Hw map (Hibrand-Saint Oyant et al., 2007). Using this data, a preliminary partial consensus map was developed with 5 of the 7 linkage groups (LG) by Sriyani Rajapakse (Byrne et al., PAG talk, Jan 2007) (Fig. 1). Current work in the USA, Europe and Israel is working towards rose maps with sufficient overlapping microsatellites to enable the development of a consensus map for the rose genome.
With a primary interest in the use of molecular markers to aid in the development of highly disease resistant plant varieties (Byrne, 2007) several research groups have developed, characterized, and mapped RGAs (resistance gene analogues) and PKs (protein kinase) as an approach to find markers for the loci controlling resistance to powdery mildew and black spot (Xu et al., 2005; Yan et al., 2005a; Linde et al., 2006; Hattendorf and Debener, 2007b). Thus far, RGAs have been mapped onto 3 mapping populations. This includes a few selected RGA derived RFLPs near the CRPM1 locus on the Rosa roxburghii map (Xu et al., 2005) and about 50 RGA loci mapped on both the 94/1 and 97/7 populations (Yan et al., 2005a; Linde et al., 2006). In both of the latter populations the RGAs clustered heavily on LGs 1 and 7 with few or none on the LGs 3 and 6. Interestingly there were no RGAs located close to the major powdery mildew QTL clusters or the Rpp1 loci on LGs 3 and 4 although several co-localized with some minor QTLs on other LGs (Linde et al., 2006; Hattendorf and Debener, 2007b). The closest RGA locus to the Rdr1 locus was about 20 cM away (Hattendorf and Debener, 2007a). The 24 PK loci mapped on the 94/1 population were not associated with disease resistance either (Yan et al., 2005a).
SNPs or single nucleotide polymorphisms represent the third generation of molecular markers. SNPs are the most abundant co-dominant markers class. In comparison to SSRs, due to their abundance, they can easily be detected by direct sequencing of genomic regions or EST. However as they are mainly bi-allelic, they are less polymorphic than SSRs. In plants, they can be used for diverse applications such as marker assisted selection, germplasm fingerprinting, gene mapping, population structure analysis (as LD analysis), and genome wide analysis. Furthermore different techniques have been developed in human to genotype with high throughput SNPs protocol (SNPlex, Illumina®, chip based technologies, Amplifluor®, TaqMan®). As rose is a highly heterozygous plant, SNPs can be easily detected, and represent a large new sources of markers. In rose, SNPs have been reported to exist every 64–68 bp in the PR2 and PR5 genes in Rosa roxburghii which is comparable to the frequency seen in rice (170 bp), maize (48 bp) (Xu et al., 2007) and Prunus mume (55 bp) (Fang et al., 2005). In a study to discover new markers for resistance to powdery mildew, SNPs were searched for within the PR2 (pathogenesis-related gene family class 2: B-1,3-glucanase) and PR5 (pathogenesis-related gene family class 5: osmotin) genes as created in rose via degenerate primers. The 23 SNPs discovered were converted to SNAPs (single nucleotide amplified polymorphism) to create a simple efficient marker that is scored as absent or present. The SNAP primer pairs were tested and 19 gave good products and 17 were polymorphic and mapped in the R. roxburghii mapping population. Sixteen mapped to the same linkage group and one (Glu7) was linked to a minor QTL for powdery mildew (Xu et al., 2007). This approach has promise as an additional procedure for finding markers for targeted regions of the genome.
2 Use of Markers for Identification and Studies of Rose Population Structure, Taxonomy, Domestication, and Diversity Studies
Since the early 1990s, the papers published using various molecular markers in the study of rose varieties, germplasm, and species has increased dramatically from 0 to 1 per year to 5–6 per year. This increase corresponded to the development of the PCR based markers which were less expensive per unit of information than either isozymes or genomic probe technology such as RFLPs, mini satellites or microsatellites.
Varietal identification has been an important field of inquiry especially as it pertains to patent rights of roses and other plants. UPOV (International Union for the Protection of New Varieties) introduced the concept of essentially derived varieties (EDV) in 1991 to protect the rights of breeders (UPOV, 1991). In the case of roses, this would be the right of the original breeder to any commercially useful sports (somatic mutations) of the varieties that they develop. It has been shown repeatedly that either RAPDs (Torres et al., 1993; Gallego and Martinez, 1996; Matsumoto and Fukui, 1996; Debener et al., 1997, 2000; Benedetti et al., 2001; Mohapatra and Rout, 2005) , AFLPs (Debener et al., 2000; Zhang et al., 2001; Vosman et al., 2004; Wen and Deng, 2005) or SSRs (Esselink et al., 2004; Rusanov et al., 2005; Scariot et al., 2006; Babaei et al., 2007) are very useful to distinguish among varieties and their seedlings but not necessarily between their sports . Consequently it is a straightforward process to determine whether a plant is a sport of a variety. Of these three markers, RAPDS is the least consistent among different laboratories (Jones et al., 1997) and consequently is the least desirable choice. The most consistent among laboratories and easiest to genetically interpret are SSRs and ISSRs which were recently improved (Laurent Crespel, personal communication). Nevertheless, AFLPs have been shown to be sufficiently consistent among laboratories to easily distinguish among sports and seedlings of rose varieties (Vosman et al., 2004) and thus useful for EDV evaluations.
These three markers, beyond being used for varietal identification are versatile and have been used effectively in assessing the segregation patterns in dog roses (Werlemark et al., 1999; Nybom et al., 2004, 2006), quantifying the heterozygosity of roses (Crespel et al., 2001), in bulked segregant analysis in search of a marker for the Rdr1 gene (Malek et al., 2000), the identification of outcrosses (Debener et al., 2003), and in a range of diversity and phenetic studies. The early studies used RAPDs and mostly examined a few varieties and/or species (Debener et al., 1996; Millan et al., 1996; Walker and Werner, 1997; Olsson et al., 2000). A few topics such as the relationships among rose species (Matsumoto et al., 1998; Jan et al., 1999; Wissemann and Ritz, 2005; Koopman et al., manuscript submitted) and the diversity of Rosa damascena (Iwata et al., 2000; Baydar et al., 2004; Rusanov et al., 2005; Tabaei-Aghdaei et al., 2006; Babaei et al., 2007; Kiani et al., 2007) have been studied with a variety of molecular markers all of which support the same conclusions confirming the usefulness of molecular markers.
3 Current Situation for Linkage Maps
Genetic mapping for rose has been done or is being developed in both diploid and tetraploid rose populations (Tables 2 and 3). The diploids consist of three groups of crosses. The German populations (94/1 and 97/7) that are based on an introgression population combining traits of R. multiflora with those of Garden roses (Debener and Mattiesch, 1999; Linde and Debener, 2003; Yan et al., 2005b; Linde et al., 2006). The second group involves various crosses with R. wichurana. This includes the population developed in France (Hw) (Crespel et al., 2002), in Spain (Dugo et al., 2005) and in the USA (WOB) (Shupert et al., 2007). It is also interesting to note that the R. wichurana genotype used in both the Spain and the USA rose mapping populations is the same genotype, ‘Basye’s Thornless’. The third mapping population is a population of Rosa roxburghii, developed to study the inheritance of powdery mildew resistance (Xu et al., 2005).
On the tetraploid level, a map has been developed with the 90-69 F2 population which is a complex hybrid that involves parentage of cultivated roses and both tetraploid (R. carolina and R. virginiana) and diploid (R. rugosa rubra and R. wichurana Basye’s Thornless’) species in its recent breeding history (Rajapakse et al., 2001; Zhang et al., 2006). In the mapping efforts for the black spot resistance gene, Rdr1, three tetraploid populations, all crosses with the rose 91/100–5 which is heterozygous for the Rdr1 gene, were used in a bulked segregant analysis study to locate markers that could be mapped onto the diploid map. The third tetraploid population, a cross between the cut flower varieties ‘Golden Gate’ and ‘Fragrant Cloud’ was made to study the complex traits of fragrance and post harvest quality (Zamir, personal communication).
Given the highly heterozygous nature of the parents in most of these crosses, the general approach to their analysis has been using the pseudo test cross approach first described by Grattapaglia and Sederoff (1994) for use in forest trees. To further complicate mapping, from 10 to 39% of the markers have exhibited distorted segregation (Table 2). This segregation distortion could be caused by the interspecific nature of the crosses used, self incompatibility segregating in some of the populations, gametophytic selection by sub-lethal genes conferring low viability on the zygote, embryo, or seedling level or maybe competitive differences in the performance of genetically variable pollen. Whatever the reason, high levels of segregation distortion limit the quality of the map by limiting the number of markers that are used in making the initial frame work map as the markers with distorted segregation ratios are only added in the last mapping step.
The tetraploid mapping requires larger populations as compared to the diploids for the same quality map and are more difficult to map because the mapping programs are designed for the diploid situation. Thus the mapping is generally restricted to loci that are behaving as diploids.
The lengths of the diploid maps reported vary from 238 to 545 cM on 6 to 8 linkage groups and the tetraploid maps vary from 674 to 920 cM on 14 to 23 linkage groups (Table 2). Thus the total length of the diploid is about 500 cM with an average chromosome length between 70 and 80 M (Yan et al., 2005a). The density of the maps ranges from low of 5.83 cM/marker (Dugo et al., 2005) to a high of 1.9 cM/marker (Yan et al., 2005a) with the majority being at medium density of 2.5–4.0 cM/marker (Table 2).
The earliest maps used RAPDs and AFLPs (Debener and Mattiesch, 1999; Rajapakse et al., 2001) and consisted of male and female parent maps as these markers do not lend themselves for map integration. Later as other markers such as RFLPs, SSRs, RGAs, PKs, SCARs, CAPs (Table 2) and most recently SNP derived SNAPs (Xu et al., 2007) were developed they have been and are being incorporated into the maps. At the present time there are several hundred microsatellites available for rose (Esselink et al., 2003; Yan et al., 2005a; Kimura et al., 2006; Zhang et al., 2006) and mapping of these have allowed the integration of the male and females maps (Yan et al., 2005a; Linde et al., 2006; Zhang et al., 2006; Hibrand-Saint Oyant et al., 2007). With microsatellite mapping data a preliminary consensus map for 5 of the 7 chromosomes has been constructed for the maps of 97/7, 90-69 F2, and the Hw (Rajapaske, personal communication; Byrne et al., 2007 – PAG meeting abstract) (Fig. 1). As the various research groups place other SSRs and codominant markers on the maps, a rose consensus map will be developed.
The objective of mapping is to better understand the organization of the genome in question and to find markers for specific traits that might facilitate the breeding process. Consequently, a range of qualitative and quantitative trait loci have been placed on the maps. These include qualitative traits such as loci that control pink/white flower color, single/double flower corolla, white petal stripes, black spot resistance, powdery mildew resistance, recurrent flowering, and stem and petiole prickles. Among the QTLs mapped include those controlling powdery mildew resistance, plant vigor, stem prickle density and size, leaf size, flower size, days to flowering or to 50% bloom, and petal number (Table 4). Unfortunately, although these QTL studies are interesting, most used a small number of progeny which limits the accuracy of the mapping and thus the usefulness of the markers identified for the specific traits. In these mapping studies, a number of close linkages between a trait and a marker such as a RAPD or AFLP have been reported as well as a SCAR (SCM10) and two CAP (Rd1, RdY) markers for Rdr1 (Malek et al., 2000; Kaufmann et al., 2003; Zhang, 2003), two SCAR markers for Rpp1 (Linde et al., 2004) and two RGAs (RGA22C, Glu 7 RGA) for a major and minor QTL for powdery mildew resistance (Xu et al., 2005, 2007).
4 Inheritance of Known Characters
Roses have been cultivated for several thousand years but only since the late part of the 19th century have breeders been doing controlled crosses in the development of new varieties as opposed to planting out open pollinated seed and selecting among the seedlings (Krüssmann, 1981). At present, there are a few public breeding efforts, hundreds of small private breeders and scores of large breeding operations that name and release several hundred new varieties of roses every year throughout the world. In spite of all this breeding effort, the inheritance of the major traits is poorly understood due to the lack of public research into the breeding and genetics of the species. Nevertheless, in recent years in conjunction with the development of linkage maps of the rose this knowledge has increased several fold (Table 4).
The earliest reports on the inheritance of traits in the USA focussed on ornamental characteristics and were not necessarily well documented but revealed the breeder’s conclusions from a few documented cases and his breeding experience (Hurst, 1941; Lammerts, 1945; Morey, 1954). This was followed with additional work in the1960s on the inheritance of red color (Lammerts, 1960, 1964), 1970s on the inheritance of the recurrent bloom trait (Semeniuck, 1971a, b), in the 1980s in Holland, Canada and India, and finally in the last 20 years a flurry of activity connected with the development of genetic linkage maps for a variety of rose mapping populations in the USA, Asia (China), and Europe (Spain, Germany, Holland, and France).
5 Flower Traits
The most important trait in roses is the recurrent blooming trait, a probable vernalization mutation which reduces the juvenility period and allows the plant to continuously bloom without any need for a vernalization period. This trait was introduced from China during the period of exploration and was initially integrated into European roses by random crosses from gardens where both types grew. This transformed the rose into the most popular ornamental plant in the world. As early as 1941, Hurst indicated that the recurrent trait was recessive to the seasonal bloom but it was not established until 1971 by Semeniuck (1971a, b) and then further confirmed by several others (Vries and Dubois, 1984; Debener, 1999; Crespel et al., 2002; Dugo et al., 2005). Recently it has been mapped on the Hw map (Crespel et al., 2002) (Tables 2 and 3).
The inheritance of the flower form (single vs. double flowers) was first reported in 1999 (Debener, 1999; Debener and Mattiesch, 1999) as a dominant allele in the locus conferring the double corolla/flower state and has since been confirmed in other rose progenies (Crespel et al., 2002; Yan et al., 2005b; Linde et al., 2006; Shupert et al., 2007. This locus has been mapped on several rose populations with different names: Blfo, d6, Dc, NP (Debener and Mattiesch, 1999; Crespel et al., 2002; Zhang, 2003; Linde et al., 2006; Hibrand-Saint Oyant et al., 2007). Preliminary data using SSR makers showed that Blfo and NP/d6 are on the same homologous linkage group (Hibrand-Saint Oyant, personal communication). Within the double corolla/flower category there is a wide range of petals from 5 to several hundred. Genetic analysis in several populations indicates that the number of petals within the double flower is additive and controlled by several QTLs (Debener et al., 2001; Zhang, 2003; Hibrand-Saint Oyant et al., 2007).
Flower size (flower diameter, petal length) appears to be additively inherited with a high (h 2 > 0.70) heritability (Lal et al., 1982; Dugo et al., 2005; Shupert et al., 2007). Mapping of this trait indicated that there were 4 major QTL loci, 2 each on the first two linkage groups of each of the parents (Dugo et al., 2005).
Flower production has not been well studied. A small study in India indicated that floriferousness (number of flowers a plant produced) was highly heritable with a broad sense heritability of 0.96. More recent work focussed on flower production during the warm summer months and mapped a single gene (Rb = repeat bloom) on the D2 linkage group of the 97/7 progeny (Zhang, 2003).
Flower color in rose is extremely important and has been studied repeatedly beginning with early studies which indicated which colors were dominant over the others and suggestions on the breeding course to obtain specific colors (Lammerts, 1945; Morey, 1954). Subsequent studies by Lammerts (1960; 1964) with tetraploid roses concluded that there was a gene A/a which controlled the presence of cyanidin (red) and a gene M/m which controlled the magenta color. With the magenta color gene, the color was dependent on the gene dose with dark purple being homozygous dominant (MMMM), a solerino purple corresponding to a duplex condition (MMmm) and a homozygous recessive state (mmmm) giving a non fading currant red (assuming a red background with dominant A allele present). Since the pink to red color is controlled by a range of anthocyanins, the inheritance of specific anthocyanins (quercetin, kaempferol, cyanidin, peonin and pelargonidin) was examined in tetraploid roses. Both these studies came to the conclusion that these pigments were quantitatively inherited (Vries et al., 1980; Marshall et al., 1983). Recent work with diploid populations has identified a locus (Bfla) on the A2/B2 linkage group of the 94/1 map which controls pink and white color. In this case, pink is dominant over white although in one population (WOB), the heterozygous plants were a lighter pink as compared to the homozygous pink plants (Debener, 1999; Debener and Mattiesch, 1999; Yan et al., 2005b; Shupert et al., 2007). Finally in the 97/7 population, when the flower color was rated into 4 categories (Crimson, dark pink, pink, and pale) several QTLs were located on the 4th linkage group of this map (Zhang, 2003).
Beyond anthocyanins, the color in roses is determined by a range in carotenoids which give colors in the yellow/orange range. Lammerts (1945; 1960) hypothesized from his breeding experience that deep yellow was recessive to light yellow. The only other study describes a general breeding procedure to develop yellow everblooming roses utilizing R. foetida as the source of yellow flower colour. Although the authors never hypothesize its mode of inheritance, the breeding procedure indicates that yellow is dominant as it is expressed in the F1 and segregates in the back cross progenies (Vries and Dubois, 1978).
The presence or absence of white stripes on the petals, although it did not segregate as expected for a single dominant trait, was nevertheless mapped onto the 97/7 rose map on the 3rd linkage group along with the loci for double flowers and prickles (Linde et al., 2006).
QTLs for flowering time (50% bloom) were located on two chromosomes by Dugo et al. (2005) and another QTL (first bloom) was located on another chromosome by Hibrand-Saint Oyant et al., 2007). It has been suggested that the location of the QTLs were different because the flowering times were measured differently, and thus the two groups measured different aspects of this complex trait (Hibrand-Saint Oyant et al., 2007).
Flower fragrance is a very complex trait which is controlled by a multitude of chemical pathways as well as environmentally sensitive (Verhoeven et al., 2003) but has been bred for successfully over the years as evidenced by the presence of wonderfully fragrant varieties. The inheritance appears to be quantitative (Lammerts, 1945) but recent work looking at the monoterpene alcohol and aldehyde pathway that is responsible for the typical European rose fragrance indicates that there is also a major gene is responsible for turning the pathway on and off (Cherri-Martin et al., 2007) in addition to an array of genes that modify the quantity of the various scent components. Current work in the functional genomics of fragrance in rose is beginning to identify the key enzymes that create and regulate the various fragrance components and promises to lead to the development of better tools and understanding of this essential trait (Vainstein et al., 2003)
6 Plant Traits
In studies of growth type, the climbing growth types was found to be dominant over non-climbing growth types (Lammerts, 1945; Morey, 1954), and the dwarf type was found to be dominant to non dwarf types (Dubois and Vries, 1987). It is not known if these two traits are controlled at the same locus. In a recent mapping study, when a spreading bush growth type was crossed with a sprawling ground cover growth type the resultant F1 plant was of an intermediate growth type. Upon selfing the F1 plant, the F2 segregated for all three types in a ratio to suggest that these forms were inherited co dominantly (Rajapakse et al., 2001).
Other growth type related traits such as the length of the flowering stem (Lammerts, 1945), the branch number, plant height and plant spread (Lal et al., 1982) all were inherited as polygenic characters with the last three with a high broad sense heritability (h 2 = 0.88–0.95) (Lal et al., 1982).
Plant vigor, an important trait in cut rose breeding, was measured using 10 parameters (number of nodes, stem thickness, shoot length, chlorophyll content, leaf area, specific leaf weight, leaf dry weight, stem dry weight, total dry weight and growth rate) in a replicated progeny at two sites. The additive h 2 for each individual trait ranged from moderate to moderately high (0.48–0.71). QTL analysis identified 42 QTLs which clustered mainly on three chromosomes (I-2, I-6, I-7 of 94/1 population). Very importantly it was shown that although a GxE interaction exists it was much smaller than the main genetic effect of the trait (Yan et al., 2007).
Leaf size whether measured as leaf width, leaf area, or leaflet number was found to be quantitative in nature (Lammerts, 1945; Zhang, 2003; Shupert et al., 2007)
There is a major locus controlling the presence or absence of stem prickles with the presence being the dominant allele (Debener, 1999; Linde et al., 2006; Shupert et al., 2007). Another independent locus controls the presence or absence of petiole prickles (Rajapakse et al., 2001). Among germplasm with the dominant gene for stem prickles, the density and size of the prickles is quantitatively inherited (Lammerts, 1945; Lal et al., 1982; Crespel et al., 2002; Zhang, 2003).
Additional traits that have been analyzed include the male sterility, dwarfness, the moss trait (dominant with modifiers), MDH-2 (co-dominant), foliage glossiness (glossy dominant over dull), flower neck strength (quantitative), and the ability of rose callus to form somatic embryos (additive) (Lammerts, 1945; Lewis and Basye, 1961; Vries and Dubois, 1984; Rajapakse et al., 2001; Burrell et al., 2006).
7 Adaptation Traits
There have been numerous reports on the varietal differences in resistance to black spot (Wenefrida and Spencer, 1993; Carlson-Nilsson, 2000), powdery mildew (Mence and Hildebrandt, 1966; Atkiss, 1978; Ferrero et al., 2001), nematodes (Wang et al., 2004a), and cold hardiness (Lehmushovi, 1987; Karam and Sullivan, 1991). As might have been expected for black spot and powdery mildew both major gene and quantitative inheritance has been reported.
Two major genes (Rdr1, Rbs) have been mapped for black spot resistance within 10 cM on the same linkage group (Malek and Debener, 1998; Debener et al., 2003; Zhang, 2003; Yan et al., 2005b). The Rdr1 locus was found tightly linked to an AFLP marker (M10) which was converted into a SCAR (SCM10), several RFLPs (BMA1-4) (Malek et al., 2000) and several BAC end markers (Kaufmann et al., 2003). Subsequently one of the RFLP fragments (BMA3) was converted into CAP marker (RdrY) (Zhang, 2003). These markers may be useful in MAS (marker assisted selection).
Powdery mildew resistance was first reported as a dominant gene in 1945 by Lammerts and more recently the dominant resistance genes Rpp1 (Linde and Debener, 2003; Linde et al., 2004) and Rpm (Zhang, 2003) as well as a major resistance QTL, CRPM1 (Xu et al., 2005) have been mapped. A SCAR marker has been developed for the Rpp1 loci and a RGA (Resistance Gene Analogue) derived RFLP marker (RGA22C) has been located close to the CRPM1 locus (Xu et al., 2005) which should facilitate MAS (Linde et al., 2004). Other QTLs for powdery mildew have also been located on maps of 3 rose mapping populations (Zhang, 2003; Dugo et al., 2005; Xu et al., 2005, 2007; Linde et al., 2006). Unfortunately, at this point it is not known how these powdery mildew resistance major genes or QTLs are related.
Resistance to both nematodes (Meloidogyne hapla) and winter cold have been reported to be polygenic (Svejda, 1977b, a, 1979; Wang et al., 2004b).
8 Physical Maps, BAC Libraries and Current Situation for Positional Cloning
The construction of BAC libraries is the first step towards the construction of a physical map and the positional cloning of genes of interest in rose. Thus far two BAC libraries have been constructed and one is under construction (Kaufmann et al., 2003; Hess et al., 2007). Thus far there is a BAC library constructed from a colchicine-induced tetraploid of Rosa rugosa var alba cv C5-23 which has been used for chromosome walking and contig assembly for the Rdr1 black spot resistance gene (Kaufmann et al., 2003). The other BAC library was constructed from the Rosa chinensis cv ‘Old Blush’ with the objective of using it for positional cloning of the recurrent blooming gene (Hess et al., 2007). The two libraries are complementary as they are constructed from genetically diverse genotypes and used different restriction enzymes (HindII versus BamHI) for the partial digestion step. A third library from the multiflora derived black spot resistant 88/124–46 rose comprises 60,000 clones with an average insert size of 46 kb. This is being done with pCLC04541 cosmid that allows the transmission of small inserts via Agrobacterium. The objective of this library is to achieve complementation of the black spot resistance gene Rdr1(Kaufmann et al., 2003; Debener, personal communication).
In the construction of both BAC libraries, the basic procedure describes by Zhang (2003) was modified to minimize the interference of the high levels of polysaccharides and phenols in rose tissue which tend to lower the resultant DNA quality. Although both the German (Kaufmann et al., 2003) and the American (Hess et al., 2007) laboratories grew the plants in the greenhouse and put the leaves through a dark treatment for 3–8 days before extraction, they differed in the subsequent steps. With the R. rugosa library, high levels of PVP 40 (4%), 0.13% DIECA, 0.1% ascorbic acid, and 0.5% 2-mercaptoethanol was used in the extraction buffer to counteract the high phenolic concentration of the initial extraction (Kaufmann et al., 2003). The approach with the ‘Old Blush’ library was to heavily fertilize and prune the rose bushes to induce rapid growth in the dark followed by an increased number of wash/filter steps (from 1–2 to 6) of the nuclei before they were embedded into the LMP agarose plugs (Hess et al., 2007).
Although a contig of the Rdr1 locus was constructed and the several markers (AFLP, CAPs and BAC end markers) were mapped within a cM of the Rdr1 locus, the high heterozygosity of the rose genome created the situation of having allelic forms of the same locus. As compared to creating a BAC library of the homozygous plant, this situation in a heterozygous plant essentially doubles the work involved in the development of a contig and positional cloning of a gene (Kaufmann et al., 2003).
9 Synteny to Other Rosaceous Crops
The comparative genome studies will allow the transfer of genetic and genomic information between species (e.g. orthologous gene controlling disease resistance or flowering) and may help to elucidate the genome relationships and evolution within the Rosoideae tribe (Rosa, Fragaria and Rubus, mainly) and more distant genus within the Rosaceae family (such Malus, Pyrus or Prunus).
The alignment of the genetic maps reveals a high synteny between species of a same genus, for instance, it was proposed that Prunus genome can be treated as a single genetic entity (Arus et al., 2006), with a single exception of a reciprocal translocation. In rose, the first step was to develop consensus markers with other Rosaceae species. The first attempt was done using heterologous microsatellite primer pairs from Malus, Prunus, and Fragaria Of the 31 apple, 22 peach and 8 sour cherry SSRs screened on rose DNA only a two apple, 6 peach, and none of the sour cherry SSRs were mapped to a rose map. (Rajapakse et al., 2001; Dugo et al., 2005; Zhang et al., 2006; Hibrand-Saint Oyant et al., 2007). Recently, the screening of strawberry primers on rose DNA revealed a much higher rate of good amplification and polymorphism, seven new consensus polymorphic SSR markers were developed (Hibrand-Saint Oyant et al., 2007). Anonymous SSRs are not convenient markers for genome comparison in Rosaceae. Their transferability is too low (problem of amplification and present of the SSR) between the tribes. However, within the tribe, in the case of SSR in coding sequence (as EST-SSR) the transferability is higher, and those markers can be used in synteny studies (Rousseau et al., 2006). SSRs from Fragaria were transferred to Rosa (and vice versa) not by looking for SSR polymorphism in the homologue sequences but by detecting SNP using the SSCP technique. By such a method the transferability was increased to 10%. Furthermore, strawberry homologues of candidate genes for floral initiation in rose were isolated in strawberry (Rousseau et al., 2006). Twenty four candidate genes were tested and 25% can be amplified and were polymorphic in strawberry.
From these new consensus markers, Rousseau et al. (2006) started a macrosynteny study between Rosa and Fragaria. The 24 markers provided anchor points to link the 7 homologous linkage groups. Preliminary results indicate numerous rearrangements between Rosa and Fragaria (Hibrand Saint Oyant L. and Denoyes, unpublished). New consensus markers need to be developed, and those markers have also to be tested in other species from the Rosoideae tribe (Rubus) and Rosaceae family. Possible good consensus markers are marker based on coding sequences that are single or low copy. This is the case for COS (Conserved Orthologue Set) previously developed in tomato (Fulton et al., 2002) or the Gene Pair Haplotype developed in strawberry (Davis et al., 2007).
Interestingly, macrosynteny between the strawberry and rose diploids reveal that the locus controlling recurrent blooming and perpetual flowering in rose and strawberry respectively are homologous (Hibrand Saint Oyant L. and Denoyes B., unpublished). This raised the question whether the RB locus has the same function as genes for seasonal/recurrent flowering in strawberry (Albani et al., 2004) and primocane bearing in blackberry and raspberry (Lopez-Medina and Moore, 1999; Lim and Knight, 2000).
To date, QTL mapping has been the method of choice for localizing the genetic basis of ornamental traits in rose. However QTL mapping presents limitations as the need for large progenies (especially in perennial species as rose), the time and space necessary for their phenotyping, and the fact that traits depend on the environment. Now genetics are moving towards LD (Linkage Disequilibrium) mapping to identify genomic regions contributing to specific phenotypes. LD mapping identifies correlations between a trait and a marker (reviewed by Mackay and Powell, 2007). According to LD decay, LD mapping may help to validate candidate genes or detect new QTLs in genome wide associations. The major advantage of LD mapping is that you do not need crosses as it relies on population samples. Development of LD mapping will need an international effort for the development of adequate populations (determination of the number of individual, population structure) and high density marker map to identify genome wide associations. The development of high density, based on SSR or SNP map will be feasible with the complete genome sequencing of rose.
References
Albani, M.C., Battey, N.H., and Wilkinson, M.J. (2004). The development of ISSR-derived SCAR markers around the SEASONAL FLOWERING LOCUS (SFL) in Fragaria vesca. Theoretical and Applied Genetics 109, 571–579.
Arus, P., Yamamoto, T., Dirlewanger, E., and Abbott, A.G. (2006). Synteny in the Rosaceae. Plant Breeding Reviews 27, 175–211.
Atkiss, L. (1978). Disease resistant rose varieties. American Rose Annual 63, 99–104.
Babaei, A., Tabaei-Aghdaei, S.R., Khosh-Khui, M., Omidbaigi, R., Naghavi, M.R., Esselink, G.D., and Smulders, M.J.M. (2007). Microsatellite analysis of Damask rose (Rosa damascena Mill.) accessions from various regions in Iran reveals multiple genotypes. BMC Plant Biology 7, (8 March 2007).
Baydar, N.G., Baydar, H., and Debener, T. (2004). Analysis of genetic relationships among Rosa damascena plants grown in Turkey by using AFLP and microsatellite markers. Journal of Biotechnology 111, 263–267.
Benedetti, L.d., Mercuri, A., Bruna, S., Burchi, G., and Schiva, T. (2001). Genotype identification of ornamental species by RAPD analysis. Acta Horticulturae 546, 391–394.
Burrell, A.M., Lineberger, R.D., Rathore, K.S., and Byrne, D.H. (2006). Genetic variation in somatic embryogenesis of rose. HortScience 41, 1165–1168.
Byrne, D.H. (2007). Molecular marker use in perennial plant breeding in fruit and ornamental crops. Acta Horticulturae, in press.
Carlson-Nilsson, B.U. (2000). Resistance to Marssonina rosae in Rosa L. seedlings obtained from controlled crosses including germplasm L83. Acta Agriculturae Scandinavica. Section B, Soil and Plant Science 50, 176–182.
Cherri-Martin, M., Jullien, F., Heizmann, P., and Baudino, S. (2007). Fragrance heritability in Hybrid Tea roses. Scientia Horticulturae 113, 177–181.
Crespel, L., Zhang, D., Meynet, J., and Gudin, S. (2001). Estimation of heterozygosity in two botanic rose species using AFLP markers. Acta Horticulturae 546, 187–191.
Crespel, L., Chirollet, M., Durel, C.E., Zhang, D., Meynet, J., and Gudin, S. (2002). Mapping of qualitative and quantitative phenotypic traits in Rosa using AFLP markers. Theoretical and Applied Genetics 105, 1207–1214.
Davis, T.M., Folta, K.M., Shields, Z., and Zhang, Q. (2007). Gene Pair Markers: An Innovative Tool for Comparative Linkage Mapping. In Proceedings 6th North American Strawberry Symposium, pp. in press.
Debener, T. (1999). Genetic analysis of horticulturally important morphological and physiological characters in diploid roses. Gartenbauwissenschaft 64, 14–20.
Debener, T., and Mattiesch, L. (1999). Construction of a genetic linkage map for roses using RAPD and AFLP markers. Theoretical and Applied Genetics 99, 891–899.
Debener, T., Bartels, C., and Mattiesch, L. (1996). RAPD analysis of genetic variation between a group of rose cultivars and selected wild rose species. Molecular Breeding 2, 321–327.
Debener, T., Bartels, C., and Spethmann, W. (1997). Parentage analysis in interspecific crosses between rose species with RAPD markers. Gartenbauwissenschaft 62, 180–184.
Debener, T., Janakiram, T., and Mattiesch, L. (2000). Sports and seedlings of rose varieties analysed with molecular markers. Plant Breeding 119, 71–74.
Debener, T., Mattiesch, L., and Vosman, B. (2001). A molecular marker map for roses. Acta Horticulturae 547, 283–287.
Debener, T., Dohm, A., and Mattiesch, L. (2003). Use of diploid self incompatible rose genotypes as a tool for gene flow analyses in roses. Plant Breeding 122, 285–287.
Dubois, L.A.M., and Vries, D.P.d. (1987). On the inheritance of the dwarf character in polyantha x Rosa chinensis minima (Sims) Voss F1-populations. Euphytica 36, 535–539.
Dugo, M.L., Satovic, Z., Millan, T., Cubero, J.I., Rubiales, D., Cabrera, A., and Torres, A.M. (2005). Genetic mapping of QTLs controlling horticultural traits in diploid roses. TAG Theoretical and Applied Genetics 111, 511–520.
Esselink, G.D., Smulders, M.J.M., and Vosman, B. (2003). Identification of cut rose (Rosa hybrida) and rootstock varieties using robust sequence tagged microsatellite site markers. Theoretical and Applied Genetics 106, 277–286.
Esselink, G.D., Nybom, H., and Vosman, B. (2004). Assignment of allelic configuration in polyploids using the MAC-PR (microsatellite DNA allele counting-peak ratios) method. Theoretical and Applied Genetics 109, 402–408.
Fang, J., Devanand, P.S., and Chao, C.T. (2005). Practical strategy for identification of single nucleotde polymorphisms in fruiting mei (Prunus mume Sied. et Zucc.) from amplified fragment lenght polymorphism fragments. Plant Molecular Biology 23, 227–239.
Ferrero, F., Cadour-Marvaldi, P., Guilotea, E., Jacob, Y., Coudret, A., and Sallanon, H. (2001). Evaluation of the resistance to powdery mildew, Sphaerotheca pannosa var. rosae, of rose-tree species and hybrids. B. First exploration of the resistance biodiversity in a crossing program resistance. Acta Horticulturae 547, 379–381.
Fulton, T.M., Hoeven, R.v.d., Eannetta, N.T., and Tanksley, S.D. (2002). Identification, analysis, and utilization of conserved ortholog set markers for comparative genomics in higher plants. Plant Cell 14, 1457–1467.
Gallego, F.J., and Martinez, I. (1996). Molecular typing of rose cultivars using RAPDs. Journal of Horticultural Science 71, 901–908.
Grattapaglia, D., and Sederoff, R. (1994). Genetic linkage maps of Eucalyptus grandis and Eucalyptus urophylla using a pseudo-testcross: mapping strategy and RAPD markers. Genetics 137, 1121–1137.
Hattendorf, A., and Debener, T. (2007a). NBS-LRR-RGAs in roses: Diversity, genomic organization, expression and chromosomal location. Acta Horticulturae, in press.
Hattendorf, A., and Debener, T. (2007b). Molecular characterization of NBS-LRR-RGAs in the rose genome. Physiologia Plantarum 129, 775–786.
Hess, G., Scheuring, D., Byrne, D.H., and Zhang, D. (2007). Towards positional cloning of the everblooming gene in plants: A BAC library of Rosa chinensis cv. Old Blush. Acta Horticulturae, in press.
Hibrand-Saint Oyant, L., Crespel, L., Zhang, L., Rajapakse, S., and Foucher, F. (2007). Genetic linkage map of Rose with new microsatellite markers to identify QTL controlling flowering traits. Tree Genetics and Genomes, accepted.
Hurst, C.C. (1941). Notes on the origin and evolution of our garden roses. Journal of Royal Horticultural Society 66, 73–82.
Iwata, H., Kato, T., and Ohno, S. (2000). Triparental origin of Damask roses. Gene 259, 53–59.
Jones, C.J., Edwards, K.J., Castaglione, S., Winfield, M.O., Sala, F., Wiel, C.v.d., Bredemeijer, G., Vosman, B., Matthes, M., Daly, A., Brettschneider, R., Bettini, P., Buiatti, M., Maestri, E., Malcevschi, A., Marmiroli, N., Aert, R., Volckaert, G., Rueda, J., Linacero, R., Vazquez, A., and Karp, A. (1997). Reproducibility testing of RAPD, AFLP and SSR markers in plants by a network of European laboratories. Molecular Breeding 3, 381–390.
Karam, F.H., and Sullivan, J.A. (1991). A rapid method for detection of cold hardiness in roses. HortScience 26, 59–60.
Kaufmann, H., Mattiesch, L., Lorz, H., and Debener, T. (2003). Construction of a BAC library of Rosa rugosa Thunb. and assembly of a contig spanning Rdr1, a gene that confers resistance to blackspot. Molecular Genetics and Genomics 268, 666–674.
Kiani, M., Zamani, Z., Khalighi, A., Fatahi, R., and Byrne, D.H. (2007). Wild genetic diversity of Rosa damascena Mill. germplasm in Iran revealed by RAPD analysis. Scientia Horticulturae, in press.
Kimura, T., Nishitani, C., Iketani, H., Ban, Y., and Yamamoto, T. (2006). Development of microsatellite markers in rose. Molecular Ecology Notes 6, 810–812.
Koopman, W.J.M., Vosman, B., Sabatino, G.J.H., Visser, D., Van Huylenbroeck, J., De Riek, J., De Cock, K., Wisseman, V., Ritz, C.M., Maes, B., Werlemark, G., Nybom, H., Debener, T., Linde, M., and Smulders, M.J.M. (manuscript submitted). AFLP markers as a tool to reconstruct complex relationships in the genus Rosa (Rosaceae).
Krüssmann, G. (1981). The complete book of roses. (Portland, Ore.: Timber Press).
Lal, S.D., Seth, J.N., Yadav, J.P., and Danu, N.S. (1982). Genetic variability and correlation studies in rose. I. Phenotypic variability, heritability and genetic advance. Progressive Horticulture 14, 234–236.
Lammerts, W.E. (1945). The scientific basis of rose breeding. American Rose Annual 30, 70–79.
Lammerts, W.E. (1960). Inheritence of magenta color in roses. American Rose Annual 45, 119–125.
Lammerts, W.E. (1964). Inheritance of the scarlet-vermillon signal red colors. American Rose Annual 49, 167–172.
Lehmushovi, A. (1987). Rose varieties in the experimental filed 1981–1984. Annales Agriculturae Fenniae 26, 4.
Lewis, W.H., and Basye, R.E. (1961). Analysis of nine crosses between diploid Rosa species. Proceedings of the American Society for Horticultural Science 78, 572–579.
Lim, Y.K., and Knight, V.H. (2000). The successful transfer of primocane fruiting expression from raspberry to Rubus hybrid berry. Euphytica 116, 257–263.
Linde, M., and Debener, T. (2003). Isolation and identification of eight races of powdery mildew of roses (Podosphaera pannosa) (Wallr.: Fr.) de Bary and the genetic analysis of the resistance gene Rpp1. Theoretical and Applied Genetics 107, 256–262.
Linde, M., Mattiesch, L., and Debener, T. (2004). Rpp1, a dominant gene providing race-specific resistance to rose powdery mildew (Podosphaera pannosa): molecular mapping, SCAR development and confirmation of disease resistance data. Theoretical and Applied Genetics 109, 1261–1266.
Linde, M., Hattendorf, A., Kaufmann, H., and Debener, T. (2006). Powdery mildew resistance in roses: QTL mapping in different environments using selective genotyping. TAG Theoretical and Applied Genetics 113, 1081–1092.
Lopez-Medina, J., and Moore, J.N. (1999). Chilling enhances cane elongation and flowering in primocane-fruiting blackberries. HortScience 34, 638–640.
Mackay, I., and Powell, W. (2007). Methods for linkage disequilibrium mapping in crops. Trends in Plant Science 12, 57–63.
Malek, B.v., and Debener, T. (1998). Genetic analysis of resistance to blackspot (Diplocarpon rosae) in tetraploid roses. Theoretical and Applied Genetics 96, 228–231.
Malek, B.v., Weber, W.E., and Debener, T. (2000). Identification of molecular markers linked to Rdr1, a gene conferring resistance to blackspot in roses. Theoretical and Applied Genetics 101, 977–983.
Marshall, H.H., Campbell, C.G., and Collicutt, L.M. (1983). Breeding for anthocyanin colors in Rosa. Euphytica 32, 205–216.
Matsumoto, S., and Fukui, H. (1996). Identification of rose cultivars and clonal plants by random amplified polymorphic DNA. Scientia Horticulturae 67, 49–54.
Mence, M.J., and Hildebrandt, A.C. (1966). Resistance to powdery mildew in rose. Annals of Applied Biology 58, 309–320.
Millan, T., Osuna, F., Cobos, S., Torres, A.M., and Cubero, J.I. (1996). Using RAPDs to study phylogenetic relationships in Rosa. Theoretical and Applied Genetics 92, 273–277.
Mohapatra, A., and Rout, G.R. (2005). Identification and analysis of genetic variation among rose cultivars using random amplified polymorphic DNA. Zeitschrift fur Naturforschung 60, 611–617.
Morey, D. (1954). Observation on the genetics of the mutant climbing factor in hybrid tea roses. American Rose Annual 39, 89–97.
Nybom, H., Esselink, G.D., Werlemark, G., and Vosman, B. (2004). Microsatellite DNA marker inheritance indicates preferential pairing between two highly homologous genomes in polyploid and hemisexual dog-roses, Rosa L. sect. Caninae DC. Heredity 92, 139–150.
Nybom, H., Esselink, G.D., Werlemark, G., Leus, L., and Vosman, B. (2006). Unique genomic configuration revealed by microsatellite DNA in polyploid dogroses, Rosa sect. Caninae. Journal of Evolutionary Biology 19, 635–648.
Olsson, A., Nybom, H., and Prentice, H.C. (2000). Relationships between Nordic dogroses (Rosa L. sect. Caninae, Rosaceae) assessed by RAPDs and elliptic Fourier analysis of leaflet shape. Systematic Botany 25, 511–521.
Rajapakse, S., Byrne, D.H., Zhang, L., and erson, N., Arumuganathan, K., and Ballard, R.E. (2001). Two genetic linkage maps of tetraploid roses. Theoretical and Applied Genetics 103, 575–583.
Rousseau, M., Hibrand-Saint Oyant, L., Foucher, F., Barrot, L., Lalanne, L., Sargent, D., Simpson, D., Laigret, F., and Desnoyes-Rothan, B. (2006). Comparative mapping in the Rosoideae tribe: Rosa and Fragaria. Napier, NZ.
Rusanov, K., Kovacheva, N., Vosman, B., Zhang, L., Rajapakse, S., Atanassov, A., and Atanassov, I. (2005). Microsatellite analysis of Rosa damascena Mill. accessions reveals genetic similarity between genotypes used for rose oil production and old Damask rose varieties. TAG Theoretical and Applied Genetics 111, 804–809.
Scariot, V., Akkak, A., and Botta, R. (2006). Characterization and genetic relationships of wild species and old garden roses based on microsatellite analysis. Journal of the American Society for Horticultural Science 131, 66–73.
Semeniuck, P. (1971a). Inheritance of recurrent blooming in Rosa wichuraiana. The Journal of Heridity 62.
Semeniuck, P. (1971b). Inheritance of recurrent and non recurrent in ‘Goldilocks’ x Rosa wichuraiana progeny. The Journal of Heridity 62.
Shupert, D.A., Byrne, D.H., and Pemberton, H.B. (2007). The inheritance of flower traits, leaflet number and prickles in rose. Acta Horticulturae, in press.
Svejda, F. (1977a). Breeding for improvement of flowering attributes of winterhardy Rosa rugosa hybrids. Euphytica 26, 697–701.
Svejda, F. (1977b). Breeding for improvement of flowering attributes of winterhardy Rosa kordesii Wulff hybrids. Euphytica 26, 703–708.
Svejda, F. (1979). Inheritance of winterhardiness in roses. Euphytica 28, 309–314.
Tabaei-Aghdaei, S.R., Monfared, H.H., Fahimi, H., Ebrahimzade, H., Jebelly, M., Naghavi, M.R., and Babaei, A. (2006). Genetic variation analysis of different populations of Rosa damascena in NW. Iran using RAPD markers. Iranian Journal of Botany 12, 121–127.
Torres, A.M., Millan, T., and Cubero, J.I. (1993). Identifying rose cultivars using random amplified polymorphic DNA markers. HortScience 28, 333–334.
Vainstein, A., Zamir, D., and Weiss, D. (2003). Fragrance and pigments/Functional genomics. In Encyclopedia of rose science, A.V. Roberts, T. Debener, and S. Gudin, eds (Oxford: Elsevier), pp. 263–265.
Verhoeven, H.A., Blass, J., and Brandenbourg, W.A. (2003). Fragrance profiles of wild and cultivated roses. In Encyclopedia of rose science, A.V. Roberts, T. Debener, and S. Gudin, eds (Oxford: Elsevier), pp. 240–248.
Vosman, B., Visser, D., Voort, J.R.v.d., Smulders, M.J.M., and Eeuwijk, F.v. (2004). The establishment of ‘essential derivation’ among rose varieties, using AFLP. Theoretical and Applied Genetics 109, 1718–1725.
Vries, D.P.d., and Dubois, L.A.M. (1978). On the transmission of the yellow flower colour from Rosa foetida to recurrent flowering hybrid tea-roses. Euphytica 27, 205–210.
Vries, D.P.d., and Dubois, L.A.M. (1984). Inheritance of the recurrent flowering and moss characters in F1 and F2 hybrid tea x R. centifolia muscosa (Aiton) Seringe populations. Gartenbauwissenschaft 49, 97–100.
Vries, D.P.d., Garretsen, F., Dubois, L.A.M., and Keulen, H.A.v. (1980). Breeding research on rose pigments. II. Combining ability analyses of variance of four flavonoids in F1 populations. Euphytica 29, 115–120.
Walker, C.A., Jr., and Werner, D.J. (1997). Isozyme and randomly amplified polymorphic DNA (RAPD) analyses of Cherokee rose and its putative hybrids ‘Silver Moon’ and ‘Anemone’. Journal of the American Society for Horticultural Science 122, 659–664.
Wang, D., Fan, J., and Ranu, R.S. (2004a). Cloning and expression of 1-aminocyclopropane-1-carboxylate synthase cDNA from rosa (Rosa x hybrida). Plant Cell Reports 22, 422–429.
Wang, X., Jacob, Y., Mastrantuono, S., Bazzano, J., Voisin, R., and Esmenjaud, D. (2004b). Spectrum and inheritance of resistance to the root-knot nematode Meloidogyne hapla in Rosa multiflora and R. indica. Plant Breeding 123, 79–83.
Wen, X.P., and Deng, X.X. (2005). Micropropagation of chestnut rose (Rosa roxburghii Tratt) and assessment of genetic stability in in vitro plants using RAPD and AFLP markers. Journal of Horticultural Science and Biotechnology 80, 54–60.
Wenefrida, I., and Spencer, J.A. (1993). Marssonia rosae variants in Mississippi and their virulence on selected rose cultivars. Plant Disease 77, 246–248.
Werlemark, G., Uggla, M., and Nybom, H. (1999). Morphological and RAPD markers show a highly skewed distribution in a pair of reciprocal crosses between hemisexual dog rose species, Rosa sect. Caninae. Theoretical and Applied Genetics 98, 557–563.
Xu, Q., Wen, X., and Deng, X. (2005). Isolation of TIR and nonTIR NBS-LRR resistance gene analogues and identification of molecular markers linked to a powdery mildew resistance locus in chestnut rose (Rosa roxburghii Tratt). TAG Theoretical and Applied Genetics 111, 819–830.
Xu, Q., Wen, X., and Deng, X. (2007). Cloning of two classes of PR genes and the development of SNAP markers for powdery mildew resistance loci in chestnut rose (Rosa roxburghii Tratt). Molecular Breeding 19, 179–191.
Yan, Z., Bai, Y., and Silva, J.A.T.d. (2006). Molecular markers and their use in genetic studies in rose. In Floriculture, ornamental and plant biotechnology, pp. 498–503.
Yan, Z., Visser, P.B., Hendriks, T., Prins, T.W., Stam, P., and Dolstra, O. (2007). QTL analysis of variation for vigour in rose. Euphytica 154, 53–62.
Yan, Z., Denneboom, C., Hattendorf, A., Dolstra, O., Debener, T., Stam, P., and Visser, P.B. (2005a). Construction of an integrated map of rose with AFLP, SSR, PK, RGA, RFLP, SCAR and morphological markers. TAG Theoretical and Applied Genetics 110, 766–777.
Yan, Z.F., Dolstra, O., Hendriks, T., Prins, T.W., Stam, P., and Visser, P.B. (2005b). Vigour evaluation for genetics and breeding in rose. Euphytica 145, 339–347.
Zhang, D., Besse, C., Cao, M.Q., and Gandelin, M.H. (2001). Evaluation of AFLPS for variety identification in modern rose (Rosa hybrida L.). Acta Horticulturae 546, 351–357.
Zhang, L.H. (2003). Genetic linkage map in tetraploid and diploid rose (Clemson, SC: Clemson University).
Zhang, L.H., Byrne, D.H., Ballard, R.E., and Rajapakse, S. (2006). Microsatellite marker development in rose and its application in tetraploid mapping. Journal of the American Society for Horticultural Science 131, 380–387.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Springer Science+Business Media, LLC
About this chapter
Cite this chapter
Byrne, D.H. (2009). Rose Structural Genomics. In: Folta, K.M., Gardiner, S.E. (eds) Genetics and Genomics of Rosaceae. Plant Genetics and Genomics: Crops and Models, vol 6. Springer, New York, NY. https://doi.org/10.1007/978-0-387-77491-6_17
Download citation
DOI: https://doi.org/10.1007/978-0-387-77491-6_17
Publisher Name: Springer, New York, NY
Print ISBN: 978-0-387-77490-9
Online ISBN: 978-0-387-77491-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)