Abstract
The most well-known bioremediation technology is the decomposition and purification of recalcitrant petroleum-based hydrocarbon contaminants by specific microorganisms. Here, first of all, we will learn how they have evolved ingenious enzyme systems. In order to overcome the rate-limiting step of the initial decomposition reaction, the oxygen attracted to the iron-coordinated active center attacks the stable carbon–carbon covalent bond and hydroxylates it brilliantly. Following this step a series of energy consuming reaction continues in the first half. However, since product compounds can be finally used as respiratory substrates, cells can acquire energy in the end. Secondly, it will be demonstrated that formation of a microbial biofilm is a potential bioaugmentation technology that is advantageous in survival competition with robust indigenous microorganisms at a contaminated site. Finally, we will see how microorganisms have elegantly developed projectiles for effective dispersion and emulsification of water-immiscible hydrocarbon compounds.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Microbial Degradation of Stable Hydrocarbon Pollutants
The most significant aspect in microbial metabolisms is their marvelously wide acceptability of substrate electron donors and acceptors in order to obtain chemical energy from the environments. This feature makes microorganisms nature based attractive players for environmental remediation technology in terms of degradation of harmful recalcitrant compounds, including stable hydrocarbons. Microbial degradation of hydrocarbons has been reported in the temperature range from −5 to 70°C [1,2,3]. Transformation of highly reduced and inert hydrocarbon compounds is with no doubt a challenging and difficult biochemical reaction for a single enzyme. However, several multi-component enzyme systems enable microorganisms to utilize hydrocarbons as carbon and energy (electron) sources. Initial biological attack to hydrocarbons is generally the oxygenation/hydroxylation that requires molecular dioxygen as a co-substrate. Dioxygen, an excellent terminal electron acceptor, also contributes to the ring cleavage reaction of homo- and hetero-cyclic aromatic hydrocarbons. Although the dioxygen molecule is omnipresent and highly soluble in water, activation and splitting this triplet ground-state molecule to wed with inert hydrocarbons need special devices. In the world of enzymes, biocatalysts, non-heme iron, heme iron, or flavin nucleotide are designated as a fantastic hidden dagger for this purpose [4]. Donation of electron to these catalytic centers is supported by a series of partner proteins in the multi-component enzyme system, including NADH (Nicotinamide Adenine Dinucleotide) dehydrogenase to yield two electrons and an electron carrier protein ferredoxin or rubredoxin connecting to dioxygenase or monooxygenase components (Figs. 1 and 2).
Alkane monooxygenase complex is composed of three proteins, NADH/Rubredoxin reductase (AlkT), Rubredoxin (AlkG), and Alkane monooxygenase (AlkB). AlkB is embedded into cell membrane. Electron from NADH is transmitted to dinuclear Fe in AlkB through 2Fe-2S cluster in AlkT and AlkG. Dioxygen molecule binds to 2Fe in AlkB followed by single oxygen molecule attacking the terminal carbon of alkane
Reaction mechanisms of three-component naphthalene dioxygenase (NahAa/Ab/AcAc) FAD(H): Flavin adenine dinucleotide https://doi.org/10.1038/nchembio.71
A group of metalloenzyme families, carrying an iron coordinating active site whichever heme or non-heme type, play crucial roles for the binding and activation of dioxygen or monooxygen in various oxidative transformation of environmental pollutants. These include hydroxylation of hydrocarbons and also cleavage of aromatic ring structure. Dioxygen or monooxygen molecule binds to the iron active sites of monooxygenase or dioxygenase component, AlkB or NahAcAd and generates competent iron-peroxo and iron-oxo intermediates to generate a substrate-based radical. These reactive intermediate species allow the enzyme to insert oxygen molecule to the substrate compounds. The product alcohol compounds are further oxidized, for example, to aldehydes and fatty acids by dehydrogenase enzymes and finally degraded completely to CO2 and H2O through metabolic pathways. The catechol (diol) ring structure is split by catechol dioxygenase before transformation to aldehydes or fatty acids.
Environmental pollution by halogenated hydrocarbons among polychlorinated aromatic compounds such as polychlorinated biphenyls, PCB, and dibenzodioxins is of highly concerned. These exert dermal toxicity, immunotoxicity, reproductive deficits, teratogenicity, endocrine toxicity, and carcinogenicity/tumor promotion [5]. However, a wide range of bacteria such as Arthrobacter, Bacillus, Corynebacterium, Luteibacter, Microbacterium, Pseudomonas, Rhodococcus, and Williamsia sp. have been reported thankfully to be capable of utilizing toxic PCBs as carbon and electron sources [6]. Initial oxidation by these bacteria is generally the reaction by biphenyl dioxygenase which functions as a three-component enzyme system similar to naphthalene dioxygenase enzyme as described below (Fig. 2).
On the other hand, another group of halogenated hydrocarbons are degraded by not oxidation but reduction reaction, the so-called halorespiration which is one of the unique anaerobic respirations of bacteria. This means that the target compounds are not used as electron donors for microorganisms but as electron acceptors. These reactions proceed under anoxic conditions in the presence of organic electron donor compounds including methanol. Dehalococcoides ethenogens is the most popular dehalogenating bacteria that consistently dominates at the contaminated sites by tetrachloroethylene, PCE. D. ethenogens converts PCE to trichloroethylene, dichloroethylene, and finally to ethene (ethylene) that is further degraded to CO2 and H2O by aerobic respiration by other symbiotic microorganisms. This reductive dehalogenase also adopts [Fe-S] cluster, [4Fe-4S] at the catalytic site for transferring electron. It has been also indicated that a cobalamin (B12) molecule exists at proximate position of [4Fe-4S] cluster and the coordinated Co atom of the cobalamin directly binds to halogen atom suggesting its contribution to dehalogenation reaction [7]. Here, we are surprised again wonderful tactic of enzyme evolution to complete its difficult mission to detoxify non-natural and recalcitrant pollutants. It may also worth to note that bacteria, prokaryotic microorganisms, never transform pollutants kindly for saving biosphere environments including human health but they need to do so for their living.
1.1 Alkane Monooxygenase/Hydroxylase
Alkane monooxygenase/hydroxylase is a three-component system comprising a soluble mononuclear iron and FAD containing NADH/rubredoxin reductase (AlkT), a [2Fe-2S] soluble rubredoxin (AlkG), and an integral-membrane diiron oxygenase (AlkB) (Fig. 1). This type of multi-component enzyme system is widely distributed in bacteria [8]. AlkB adopts the oxygen rebound mechanism in order to hydroxylate alkanes but preferably from C5 to C16 alkanes. This mechanism involves homolytic cleavage of the C–H bond by an electrophilic oxo-iron intermediate to generate a substrate-based radical. Diiron ligand site of AlkB is composed of eight histidine motifs. These histidine residues are also potential ligands for the diiron atoms contained within alkane monooxygenase. This protein family has hydrophobic six alpha helices that would be capable of spanning the membrane bilayer. Unfortunately, active site structure of AlkB has not been solved yet, however, spectroscopic and genetic evidence suggests a nitrogen-rich coordination environment located near the inner surface of the cytoplasmic membrane with as many as eight histidines coordinating two irons and a carboxylate residue bridging these two metals. A particular amino acid residue located in the middle of transmembrane helix-2 of AlkB has been shown to be important to determine the alkyl length of the substrate. When this amino acid has a bulky side chain like tryptophan, the long-chain alkyl groups (C13<) are not acceptable in the substrate binding cleft.
1.2 Rieske Dioxygenase
Rieske dioxygenase catalyzes the primary cis-dihydroxylation of arene (monocyclic and polycyclic aromatic hydrocarbon) substrates, which is the initial step of many bacterial degradation pathways. Rieske protein was first reported by John Rieske et al. [9], which has a characteristic [2Fe-2S] cluster with 2-His-2-Cys bidentate ligand. Besides a mononuclear iron active site, Rieske dioxygenases carry a dinuclear [2Fe-2S] cluster in which one iron (Fe1) is coordinated by two histidines while the other iron (Fe2) is coordinated by two cysteines. Fe2 remains in a ferric state regardless of the reduction state of the cluster, while Fe1 is converted from a ferric state to a ferrous state when reduced during the reaction. The reaction of the three-component type Rieske dioxygenase requires two electrons from an NAD(P)H and consecutively transferred to the terminal dioxygenase component through a ferredoxin (monomer) and a reductase (monomer). In a single large subunit, the Rieske-type cluster and the mononuclear iron center are too far apart to allow for electron transfer at a distance ~43 Å. However, the quaternary structure (trimer of hetero-dimer or homo-trimer type with three-fold symmetry) allows for electron transfer from a Rieske-type cluster to a mononuclear iron center from a neighboring subunit, which is only 5 Å apart (Fig. 2, [10]). A key role in this electron transfer has been ascribed to an absolutely conserved aspartic acid residue Asp205 in NahAc that bridges between the two metal sites. Structural studies also implicate a side-on binding of a dioxygen to form catalytically active iron(III)-peroxide intermediate which is subsequently converted to a high-valent iron(IV)-oxo or iron(V)-oxo-hydroxo intermediate. After cis-dihydroxylation of the substrate, catalytic mononuclear iron will return to iron(II) resting-state configuration [11, 12].
2 Efficacy of Biofilm Formation by Naphthalene Degrading Bacteria for Bioremediation
Among the strategies to clean up pollutants it is widely recognized that biological treatments, the so-called bioremediation technologies, have the advantage over chemical and physical treatments in terms of their compatibility with natural system thus minimizing environmental impacts [13]. Active on-site bioremediation includes technologies that activate indigenous microbial populations, biostimulation, or introduce specific competent foreign bacteria to the contaminated site, bioaugmentation. Although bioaugmentation is expected to be a quicker and more effective technology than biostimulation, low fitness and poor colonization of the introduced bacteria to the contaminated sites often make efficacy of bioaugmentation poor [14, 15]. The activity and viability of the introduced bacteria often decreases at the contaminated sites when compared with those observed under laboratory conditions, probably due to diverse environmental stresses [16, 17]. These include predation by protozoa, competition with other bacteria, unfavorable pH and temperature conditions, and poor availability of nutrients and oxygen. Catabolite repression by other organic compounds also decreases the expression level of the degradation genes. However, bioaugmentation can become more effective if both microbial ecology and population sizes are taken into account [15, 18, 19]. One of the options for effective clean-up methods of contaminated sites is the use of carrier materials so as to maintain sufficient activity of inoculants during prolonged periods. There are several reports demonstrating that immobilized or encapsulated cells effectively degrade pollutants at the contaminated sites [20, 21]. However, the cellular and molecular mechanisms underlying these technologies are largely unknown.
Biofilms are a multicellular structure of microorganisms formed on surfaces that exhibit sociality under control of quorum sensing signaling molecules [22]. Biofilms are often encased in sticky extracellular polymeric substances such as exopolysaccharides. Forming biofilms is considered a natural strategy of microorganisms to construct and maintain a favorable niche in stressful environments [17, 23]. We propose that application of biofilms to bioproduction and bioaugmentation processes is a natural and rational choice [24]. Indeed, there are several reports that demonstrate the efficacy of biofilm formation by useful bacteria and their use in bioremediation and biotransformation technologies [25,26,27,28]. We have demonstrated benefits of biofilm formation by hydrocarbon-degrading bacterium for stable bioremediation. Shimada et al. compared degradation activities and its persistence in contaminated soils inoculated with biofilms or planktonic cells of naphthalene degrading bacterium Pseudomonas stutzeri T102 [29]. When compared with artificially immobilized and encapsulated cells, the advantage of biofilms is that the high density and tolerance of the constituent cell is naturally achieved. Moreover, biofilms can be introduced to contaminated sites free of supports that might cause additional environmental pollution. The secondary purpose of the study was to examine the potential of model biofilms toward developing bioaugmentation technology.
Pseudomonas stutzeri are ubiquitous and useful environmental bacteria with their high degradation abilities for harmful chemical compounds including biocides, halogenated alcohols, and hydrocarbons [30]. Among the tested laboratory strains, P. stutzeri T102, which was isolated from sludge of an oil reservoir tank in Okinawa [31], formed the biofilms on the surface of various materials including polypropyrene, polystyrene, polyethylene terephthalate, acrylic, and glass bottles. P. stutzeri T102 is capable of degrading PAHs, such as naphthalene and dibenzothiophene. When using a batch culture system for biofilm formation, the amount of T102 biofilms continued increasing even after 120 h (Fig. 3a). Thus, we chose T102 as a model bacterial strain for further experiments. Gross et al. (2007) tested 69 bacterial strains for biofilm forming capacity and showed that many of the best biofilm formers belonged to Pseudomonas [32]. Our experimental result is consistent with their result. Since T102 is a facultative aerobic strain, it formed ring-shape biofilms near the air–liquid interface on the inner surface of glass bottles (Fig. 3b). However, they formed rather uniform biofilms in small plastic tubes, such as 1.5 mL polypropyrene tubes and 96 well polystyrene plates probably due to the small depth of the liquid phase, better aeration condition, reduced hydrophobicity and zeta potential of the tube materials (Fig. 3c). It may be worse to note that a major outer membrane protein, OmpA exhibited opposite effects on the biofilm formation in glass bottles and plastic tubes where surface hydrophobicity is different [33]. This suggests that each environmental bacterium has different surface condition for its optimal biofilm formation. Scanning electron microscopy demonstrated that the macroscopically uniform biofilms of T102 are also not flat but highly structured (Fig. 3d, e).
(a) Biofilm formation by hydrocarbon degrading bacteria. Biofilms were formed at the indicated temperatures in 1.5 mL polypropyrene tubes containing 0.5 mL Y medium. Data points are the averages of triplicate assays and error bars represent standard deviations. Symbols are as follows: closed diamond, P. stutzeri T102 (30˚C); closed square, Gordonia sp. C3 (30˚C); closed circle, Oleomonas sagaranensis HD1 (30˚C); open triangle, Rhodococcus sp. TMP2 (20˚C); open diamond, Rhodococcus sp. T12 (20˚C); cross, Shewanella sp. SIB1 (20˚C); open circle, Arthrobacter sp. CAB1 (30˚C). The latter four strains did not produce biofilms in the culture systems. (b) Crystal violet staining of P. stutzeri T102 biofilms in a 20 mL glass bottle; (c) in an 1.5 mL polypropyrene test tube. (d, e) Scanning electron micrographs. All biofilms were formed at 30˚C for 48 h. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article)
2.1 Naphthalene Degradation by T102 Biofilms and Planktonic Cells
There are reports that biofilm-associated cells are more dormant and inactive than planktonic cells [34,35,36]. This feature of biofilms partly explains their high resistance to environmental stresses [37]. It was thus a concern that T102 biofilms might exhibit less naphthalene degradation activity than planktonic cells. In order to examine this possibility, the activities of T102 biofilm and planktonic cells were compared in a pure liquid-culture system (Fig. 4). Biofilms of T102 were formed in glass bottles. After cultivation at 30°C for 24 h without shaking, free planktonic cells were carefully removed from the bottles and the biofilm formed inside walls (ca 2 × 107 CFUs) was rinsed once with sterile water. The biofilms were soaked in a mineral medium containing 20 ppm naphthalene and the bottles were tightly sealed with butyl rubber stoppers and incubated at 30°C without shaking. Planktonic cells were separately grown in a shaking flask. Mid-exponential phase cells were harvested by centrifugation and washed and suspended with a small amount of medium. BM-medium containing the planktonic cells at the same CFU value (ca 2 × 107 CFUs) as the biofilm samples was prepared in glass bottles with 20 ppm naphthalene added. The bottles were sealed and kept at 30°C as described previously. All the samples were prepared in triplicate for each sampling time and a negative control included that contained no cells. Sample bottles were taken every 4 h until 16 h and then every 8 h until 48 h. Remaining amounts of naphthalene in the samples were analyzed by gas chromatography.
Comparison of naphthalene degradation activity of T102 biofilm and planktonic cell cultures. Data points are the average of triplicate assays; error bars represent standard deviations. Initial CFUs for biofilm and planktonic cell cultures were 21,470,000 and 23,360,000, respectively. Symbols are as follows: no cells, triangle (filled triangle); T102 biofilm cell culture, square (filled square); T102 planktonic cell culture, circle (filled circle)
Experimental results revealed that T102 biofilms did not degrade naphthalene during the first 4 h but after that time they degraded naphthalene faster than planktonic cells (Fig. 4). This interesting observation was reproducible in several independent experiments. T102 biofilms degraded 14 ppm or 70% of initial naphthalene (20 ppm) in 12 h and exhibited a maximum degradation rate of 1.7 ppm h−1 between 4 and 12 h. On the other hand, T102 planktonic cells started to degrade naphthalene immediately after incubation. The degradation rate was almost constant, about 0.5 ppm h−1 for 16 h, and it took 28 h for the degradation of 14 ppm of naphthalene. About 1.6 ppm (8%) of the initial naphthalene was absorbed to the butyl rubber septum of the bottle cap and remained un-degradable. This means that naphthalene was almost completely eliminated from the culture of T102 biofilms after 32 h. We hypothesized that initial delay for degradation by biofilms may be the time that it takes the naphthalene to penetrate through the extracellular matrix of the biofilm so that the genes responsible for degradation of naphthalene might be induced.
2.2 Comparison of Expression Levels of nahA in T102 Biofilms and Planktonic Cells
It is known that a set of genes nahAa, nahAb, nahAc, and nahAd form an operon and encode a ferredoxin, ferredoxin oxidoreductase, and naphthalene dioxygenase large and small subunits, respectively (Fig. 2; [4, 38]). All these genes are essential for the aerobic degradation of naphthalene and related compounds. Thus, we analyzed the expression level of nahAc, which encodes the large subunit of naphthalene dioxygenase, in T102 biofilms and planktonic cells (Fig. 5). Total RNA was extracted from T102 cells and cDNA was synthesized from DNase-treated RNA as previously described [39]. RT-PCR amplification was performed using a primer set targeting nahAc in T102. The PCR products were separated on 1.5% agarose gels and stained by ethidium bromide. Contrary to our hypothesis, nahAc in T102 biofilms expressed nahAc at constant levels from the start of incubation through to 40 h (Fig. 5b). Constant gene expression was also confirmed for nahAa, nahAb, and nahAd (results not shown). These results suggest that the initial lag and subsequent high naphthalene degradation periods exhibited by T102 biofilms are not due to changing gene expression levels of nahAabcd. The nahAabcd operon in Pseudomonas putida has been reported to be inducible under regulation of nahR [40]. In this experiment expression of nahA genes was observed at 0 h, before naphthalene was added to the medium. This suggests that nahR does not exist or does not function to regulate naphthalene degradation in P. stutzeri T102, which has been isolated from the bottom sludge of a petroleum reservoir tank where naphthalene is always abundant.
(a) Naphthalene degradation activity of T102 biofilm (left) and planktonic (right) cell cultures, respectively. Initial CFUs were 18,850,000 (biofilm culture) and 22,120,000 (planktonic cell culture). Data points are the average of triplicate assays; error bars represent standard deviations. (b) RT-PCR analyses of nahAc and 16S rRNA gene expression in the T102 biofilms (left) and planktonic cells (right), respectively
When we carefully observed the culture, we noticed that a part of T102 detached, dispersed and grew planktonically in the culture bottle containing T102 biofilms (Fig. 6a). Thus, we carefully separated these detached cells from biofilms, and examined the expression level of nahAc. It was found that the expression level of nahAc in the detached cells was clearly and significantly higher than that in biofilms from 12 to 20 h (Fig. 6b). This result allowed us to conclude that degradation of naphthalene was largely due to the activity of detached cells rather than biofilms. The naphthalene degradation activities of T102 biofilms, detached, and planktonic cells, were determined as 0.02, 1.1, and 0.3 pg CFUs−1 h−1, respectively. The degradation activity of the biofilms was much lower than planktonic cells probably due to depletion of oxygen in biofilms where cells were densely packed and due to poor naphthalene penetration through the matrix that encases biofilm-associated cells.
(a) Naphthalene degradation activity (open square) and the amount of detached cells (closed diamonds) in a T102 biofilm flask. Data points are the average of triplicate assays and error bars represent standard deviations. (b) RT-PCR analyses of nahAc and 16S rRNA gene expression in the detached cells. Only the cells sample at 0 h was taken from biofilms
2.3 Fitness of T102 Biofilms and Planktonic Cells in Petroleum Contaminated Soils
The observation that T102 biofilms were capable of producing highly active detached cells prompted us to test their performance in soils contaminated with natural petroleum. Petroleum contaminated soils were taken from Ishikari petroleum field (Hokkaido, Japan) and used for following experiments without sterilization. For biofilm samples, T102 biofilms were formed in advance at 30°C for 24 h in a screw cap polypropyrene tubes containing 0.4 mL medium. The liquid culture containing planktonic cells was removed from the tubes leaving biofilms inside wall. Contaminated soils, 0.5 g, and the same amount of filtered cell free spent medium from planktonic cell cultures were added to each tube. Then, two different conditions were set for biofilm samples, one was the “intact biofilm samples” with no treatment, the other was “dispersed biofilm samples” which were vortexed for 2 s to detach and disperse the biofilm pieces into the soils. For planktonic cell samples, 0.5 g of the soils in 2 mL tubes were supplemented with mid-log phase planktonic cell cultures containing similar colony forming units, CFUs, to above biofilm samples. A soil sample with no inoculation of T102 cells was also prepared as a negative control. The soil samples inoculated with “intact biofilms,” “dispersed biofilms,” “planktonic cells,” or no inoculation were incubated at 30°C using the naphthalene vapor exposure method [41]. Naphthalene vapor was continuously supplied by placing a small quantity of naphthalene crystals in a sample tube that was loosely capped and placed on wet paper to avoid desiccation. The Petri dishes containing sample tubes and naphthalene crystals were sealed with parafilm, which is permeable to oxygen.
To estimate the fitness of P. stutzeri T102 in the soils, we used the community DNA fingerprinting method denaturant gradient gel electrophoresis (DGGE) to assess variation in bacterial community composition among samples (Fig. 7). Although this method is semi-quantitative due to different efficiency of DNA extraction and PCR depending on the bacterial strains, it is useful to analyze population changes of single strains over the time. Moreover, distribution of the DNA bands in DGGE gels provides a rough overview of the bacterial community dynamics upon introduction of T102 biofilm and planktonic cells to the soils. Total DNA was prepared from the culture and PCR amplified 16S rRNA gene fragments were separated on an 8% polyacrylamide gel with a denaturing gradient of urea and formamide ranging from 20 to 50%. The density of DNA bands corresponding to P. stutzeri T102 16S rRNA genes was estimated using imaging software (Fig. 7a).
(a) Comparison of the fitness of T102 cells inoculated differently in contaminated soils. Relative density of 16S rRNA-PCR products band of T102 in each DGGE gel is plotted in the graph. Symbols are as follows: T102 dispersed biofilms (triangle, filled triangle); T102 intact biofilms (square, filled square); T102 planktonic cells (circle, filled circle) DGGE gel data for the bacterial community analyses from 0 to 10 weeks. (b) native community without T102 inoculation; (c) Inoculation of T102 planktonic cells; (d) Inoculation of T102 intact biofilm; (e) Inoculation of T102 dispersed biofilm. The arrows indicate position of the 16S rRNA-PCR band of T102. Asterisks * indicate the position of indigenous naphthalene degrader
We observed that the “dispersed biofilm” sample maintained a rather dense T102 DNA bands for 10 weeks and kept over 40% of its original density level for 8 weeks (Fig. 7e). The density of T102 DNA bands of “planktonic cells” and “intact biofilms” samples decreased more rapidly than that of “dispersed biofilms,” and they almost disappeared (ca 2% remained) after 10 weeks (Fig. 7c, d). The decrease in the amount of T102 DNA bands was more significant within the first 3 weeks than later for all experimental conditions. The amount of T102 DNA in the “intact biofilms” sample continuously decreased over the period. This may be because the nutrients and oxygen were more rapidly depleted around the sessile “intact biofilms” than “planktonic cells” and “dispersed biofilms.” The duration for which each sample kept over 20% of the initial density of T102 DNA was 10, 5, and 2 weeks for “dispersed biofilms,” “intact biofilms,” and “planktonic cells” inoculates, respectively (Fig. 7a). These results indicate that “dispersed biofilms” are more tolerant and stable than “intact biofilms” and “planktonic cells” in the petroleum contaminated soils. A band of increasing intensity, indicated by an asterisk, in all samples may be an indigenous naphthalene degrader since it also appeared in the sample without T102 inoculation (Fig. 7a).
The above experimental results are consistent with previous knowledge that biofilm-associated cells are generally more tolerant to environmental stresses than their planktonic counterparts. The environmental robustness of biofilm-associated cells could be attributed to both specific gene expression and physicochemical toughness provided to the densely packed cells by extracellular matrices [42,43,44]. Next, we examined the naphthalene degradation activities of contaminated soils containing either T102 biofilm or planktonic cells.
2.4 Naphthalene Degradation Activity of Soils Containing T102 Biofilms and Planktonic Cells
As shown above, the liquid-culture system inoculated with T102 biofilms exhibited naphthalene degradation activity comparable to or even higher than with T102 planktonic cells (Fig. 4). PCR-DGGE analysis suggested that populations of T102 remained higher in soils inoculated with “dispersed biofilms” rather than the “planktonic cells” during incubation for 10 weeks in contaminated soils. Thus, we expected that the soil sample containing “dispersed biofilms” should exhibit higher degradation activities than “planktonic cells” over the period. But it was difficult to measure naphthalene degradation directly in the contaminated soil samples because the soils contained large amounts of various hydrocarbons and the signal/noise ratio in GC analysis was quite low. We decided to measure naphthalene degradation activity of each soil sample in BM-medium containing an additional 100 ppm naphthalene (Fig. 8). The entire soil sample, 0.5 g, in the 1.5 mL polypropyrene tube was transferred into a 50 mL screw capped glass bottle containing 20 mL of BM-medium and 100 ppm naphthalene in methanol solution. Bottles were tightly closed and incubated at 30°C for 36 h. Extraction of remaining naphthalene and GC/FID analysis were performed as previously described. It was demonstrated that “dispersed biofilms” and “planktonic cells” initially degraded 48 and 52 ppm naphthalene in 36 h, respectively. Their degradation activities gradually decreased as the incubation time increased. When their activities were compared after 9 weeks of incubation, “dispersed biofilms” still degraded 19 ppm naphthalene while “planktonic cells” and no inoculation samples degraded only <0.1 ppm. Petroleum contaminated soils that were taken from Ishikari oil field contained various hydrocarbon compounds such as straight-, branched- and cyclo-alkanes, and PAHs including naphthalene. Thus, it was not surprising that the soils contained significant amounts of naphthalene degrading bacteria. The degradation of naphthalene with no inoculation could be attributed to these indigenous bacteria. Densitometry of the T102 bands of “dispersed biofilms” and “planktonic cells” in DGGE gel revealed that 35 and 9% of the initial band intensities remained after 9 weeks, respectively. These results suggest that the specific naphthalene degradation activity of “dispersed biofilms” including detached cells was much higher than “planktonic cells” in the petroleum contaminated soils after 9 weeks.
2.5 Summary
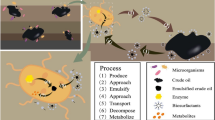
Biofilms confer microbial cells with high resistance to environmental stresses and facilitate their survival in complex microenvironments that help generate diverse cellular heterogeneity and activities. Naphthalene degradation rate of P. stutzeri T102 biofilms was initially low but later became higher than that of T102 planktonic cells. The rapid degradation activity of biofilm cultures could be attributed to producing detached cells. It was shown that “biofilms act as aircraft carriers that launch super-active fighter cells” which may contribute to the continued degradation of harmful environmental contaminants. T102 cells were also shown to be more durable and active in the petroleum contaminated soils when they are introduced as “dispersed biofilms” rather than as “planktonic cell” suspensions. These experimental results suggest that inoculation of contaminated sites with pollutant degrading biofilms may enhance bioaugmentation as a durable and effective bioremediation technology.
3 Biosurfactants
Biosurfactants (biological surfactants), namely BS, are a group of surfactants produced by living organisms. Surfactants are generally considered as chemical products such as detergents and emulsifiers that are abundantly used in various industries. Surfactants are also useful compounds for environmental remediation by accelerating microbial degradation. People may think of them as harmful substances that are far from living organisms, but this is not the case. For example, phospholipids, a major component of cell membranes in all living organisms, are biomolecules with amphiphilicity can also function as surfactants. Furthermore, glycolipids such as cerebroside and ganglioside, which are abundant in the brain, are not exception. In other words, biological complex lipids mostly can function as BS in the broad sense. Thus, surfactants are molecular species that are closely related to living organisms.
A group of microorganisms are known to produce and secrete complex lipids with strong surfactant activity outside the cells, which are generally called biosurfactants, BS [45]. The physiological significance of BS to the producing bacteria is still unclear, but it is natural that for microorganisms growing in hydrophobic environments such as oil fields, the ability to emulsify hydrocarbon compounds for efficient uptake as carbon and energy sources is advantageous for survival. In fact, microorganisms with BS production activity are widely distributed in oil fields, including Bacillus, Pseudomonas, and Rhodococcus [46, 47]. It is a rational idea to activate the domestic or introduce foreign BS-producing bacteria to the site of oil contaminated soils and oceans [48,49,50,51,52,53]. In the case of Serratia, a plant pathogen, BS are secreted to improve the affinity with the wax-covered leaf surface useful for invasion [54]. On the other hand, some BS have antimicrobial activity, which may also be of great benefit not only to the producing bacteria but also to the host plants [55]. In addition, BS have been shown to be important in the formation of three-dimensional structures of microorganisms (so-called biofilms) formed by microorganisms attached to solid surfaces and interfaces, and it can be said that BS are one of the tools that microorganisms have devised to secure their ecological niche while resisting various environmental stresses [56, 57]. The development and effective use of such special natural products is expected to contribute to the realization of a sustainable society with low environmental impact [58].
3.1 Isolation of BS-Producing Bacteria
Isolation of BS-producing bacteria can be performed by using the accumulation of an emulsified layer or the decrease in surface tension of the culture solution as an index. Emulsified layer is observed when culturing bacteria with water-insoluble hydrophobic substance such as a hydrocarbon compound as a carbon source. There is also a simpler method which is called as a plate test. It is convenient to use a blood agar medium on the market, and hemolytic spots due to BS activity are formed around the colonies of the producing bacteria [59]. The authors accidentally found that an agar plate medium prepared by spreading a small amount of crude oil to form thin membrane on the top, prepared for the purpose of isolating hydrocarbon-degrading bacteria, was very useful for selecting biosurfactant-producing bacteria. That is, first, a sample containing microorganisms is spread on this medium to form colonies, and second the surface of the plate medium is observed under the reflected light of a fluorescent lamp and a circular zone in which the oil film is excluded can be observed around the colonies of the biosurfactant-producing bacteria (Fig. 9, [60]). At first, we thought that I had discovered a novel crude oil-degrading bacterium which is capable of evaporating liquid hydrocarbons by directly cutting C–C bond to yield hydrocarbon gasses such as methane or ethane, but later it was revealed that this oil exclusion circle was due to BS activity. Since the size of this oil film exclusion zone is directly proportional to the BS molecular activity and the amount of BS production. Finally, excellent BS-producing bacteria can be obtained by picking up colonies that form a large zone.
Easy BS halo assay for detecting BS production. There are three colonies of BS producing bacteria surrounded by oil displacement zone, BS halo. BS halo can be visualized by different reflection of the light. The size of BS halo directly correlates with the amount and activity of BS [59]
3.2 Production and Purification of BS
When the BS-producing bacteria are grown in liquid-culture using a hydrophobic substrate such as a hydrocarbon compound or vegetable oil, significant amount of emulsified layer is formed by the function of produced BS that cells and hydrocarbon compounds cannot be easily removed by centrifugation. However, in the case of sufficiently highly BS-producing yeasts, whose cell size is much bigger than bacteria, the hydrophobic substrate is almost completely consumed. A bottom white cell layer and biosurfactant containing brown layer are formed under the aqueous culture solution after simply left stand for 1 day. On the other hand, Bacillus and some Pseudomonas bacteria show relatively good biosurfactant productivity even without hydrophobic substrates. That is, the BS can be recovered by acid precipitation or calcium precipitation by adding hydrochloric acid or calcium chloride to the supernatant obtained by precipitating and removing the bacterial cells from culture solution by centrifugation. In addition, since BS form giant micelles in an aqueous solution, they can be concentrated using an ultrafiltration membrane having an excluded molecular weight of about 10 kDa, which is used for protein concentration [59]. In addition, since BS are involved in biofilm formation by microorganisms, good productivity is often seen in solid cultures using soybean meal or okara, and in this case, hexane or ethyl acetate can directly solubilize and extract BS instead of from an aqueous solution [61]. This method enables us to concentrate BS compound easily by evaporation of the organic solvent.
3.3 Types and Structures of BS
Biosurfactant molecules are also composed of hydrophobic and hydrophilic part similarly to chemical surfactants. Since the hydrophobic part of BS is commonly fatty acid esters, their classification can be mainly divided into (1) glycolipid type, (2) lipopeptide type, (3) fatty acid type, and (4) polymeric type based on the structure of the hydrophilic part, but the first two types are most used in industry.
3.3.1 Glycolipid-Type BS
Although there are various types of glycolipid BS depending on the sugar, only the structures of popular rhamnolipids and sophorolipids are shown here.
3.3.1.1 Rhamnolipid
Rhamnolipid is a glycolipid-type biosurfactant produced mainly by bacteria of the genus Pseudomonas, and was first reported as an antibacterial substance against tuberculosis-causing Mycobacterium tuberculosis [62]. Since then, six types of homologous compounds with different monosaccharides, disaccharides, or fatty acid chain lengths (RL1–RL6) have been identified. Figure 10 shows the structures of typical RL2 and RL6. Hisatsuka, K. et al. reported in 1971, when “petroleum (utilizing) fermentation” was in its heyday, that hydrocarbon-degrading Pseudomonas aeruginosa produced rhamnolipids, which was involved in its sound growth [63].
3.3.1.2 Sophorolipid
In 1961, Gorin, P.A. et al. found sophorolipids in the fermented products of the yeast Torulopsis magnoliae and Torulopsis bombicola (later Candida bombicola, now Starmerella bombicola) [64]. As mentioned earlier, it has been found that the productivity of the biosurfactant sometimes is greatly increased by feeding vegetable oil as a carbon source along with glucose. This is probably due to the bacterium that produces lipase, and the fatty acids, which are degradation products of vegetable oil, are rapidly utilized for BS synthesis. This compound is characterized by the ether linkage of hydroxy fatty acids to the sugar (sophorose) backbone and the intramolecular condensation of the carboxyl terminus of the fatty acid with the hydroxyl group of the sugar to form a lactone (Fig. 10). The acid and lactone forms are produced in mixture in the culture medium, and the lactone can be converted to the more water-soluble acid form by ring-opening under alkaline conditions if necessary.
3.3.2 Lipopeptide-Type Biosurfactant
The best known lipopeptide-type biosurfactant is surfactin, which was reported as a thrombolytic agent produced by Bacillus subtilis [65]. Its structure is also unique: a peptide, consisting of seven amino acid residues including two D-form amino acid residues, forms a lactone via amide and ester bonds with fatty acids (Fig. 11). Since then, a series of cyclic lipopeptides with different structures have been reported from bacteria of the genus Bacillus, including lichenysin, fengycin, plipastatin, iturin, and bacillomycins [66]. Serratia spp. and Pseudomonas spp. have also been reported to produce cyclic lipopeptides serawettin, arthrofactin, and syringomycin. Arthrofactin is a cyclic lipopeptide consisting of 11 amino acids, including five D-form amino acid residues, which was discovered by the authors in 1993 from a soil bacterium in Shizuoka Prefecture [59]. Arthrofactin is one of the most effective BS with CMC (critical micelle concentration) value ~0.01 mM and minimum surface tension reduced to 24 mN/m.
These lipopeptides are synthesized without ribosomes by a huge non-ribosomal peptide synthetase (NRPS), in which substrate amino acids are activated and linked one by one. NRPS has a modular structure with an activation domain and a condensation domain before and after the domain that binds the substrate amino acid, thiolation domain. Interestingly, the gene structure of NRPS has a co-linearity rule with that of the product peptide. Moreover, a novel cyclic lipopeptide can be synthesized by replacing the gene module.
3.4 Structure-Activity Relationships of BS
BS, especially lipopeptide-type, have very diverse and complex structures. There is yet a limited knowledge available to understand rationality of its structure. In order to investigate the structural inevitability of arthrofactin and surfactin, we have attempted several structural modifications [67]. The first question was both have two acidic amino acids in common. The experimental results showed that either amidation or methylation of the Asp and Glu residues which eliminates negative charge of the molecule resulted in an increase in surfactant activity but also lost water solubility (Fig. 12). On the other hand, sulfonation to make the molecule more strongly acidic maintained the water solubility but drastically reduced the surface activity. Furthermore, when the lactone was saponified and cleaved lactone ring to form a linear structure, the activity was reduced to one-third in both cases (Fig. 13). These results allow us to conclude that lactone formation can increase the biosurfactant activity by three folds. When the activity was carefully measured by HPLC fractionating the arthrofactin according to their fatty acid chain length, relative biosurfactant activity (/A210) of the most major product was found to be the highest (Fig. 14). Furthermore, the three-dimensional structure of arthrofactin in DMSO solution was investigated using high-performance NMR. It was found that arthrofactin formed a unique helmet-like structure, with hydrophobic amino acids, Ile and Leu, oriented on the upper outer side and hydrophilic amino acids, Asp, Ser, allo-Thr, on the lower inner side (Fig. 15). Higher surface activity of BS than chemical surfactants can be probably attributed to this steric distribution of hydrophilic and hydrophobic part in a biosurfactant molecule which enables to occupy larger interfacial area. Our experimental results demonstrated that the complex BS structure successfully harmonized high surfactant activity and water solubility in a perfect manner. Here, we could see a wonderful aspect of rational natural selection over million years.
Chemical modification of surfactin. Methyl form was obtained by keeping surfactin overnight in anhydrous methanol with conc. HCl. Amide form was obtained from surfactin by keeping two hours at 22° C in methanol containing 5.5 M NH4Cl (pH 5.0) with 0.1 M 1-ethyl-3-(3-dimethylaminopropyl)-carbodiimide (EDC). Sulfonic acid form was obtained by keeping in aminomethane sulfonic acid (pH 8.0) and solubilized by the addition of appropriate amount of acetonitrile and EDC up to 0.1 M
Separation and relative activity of arthrofactin (AF) structural family. (a) Elution pattern of AF family on reverse-phase HPLC with MS detector. (b) Chemical structure and relative oil displacement activity of AF family. Relative activity was determined as follows: the oil displacement circle area was divided by A210 units and each value was compared with the major AF (C10)
3.5 Synthetic Mechanisms of Arthrofactin and Encoding Gene Cluster
Gram-positive Bacillus and Gram-negative Pseudomonas strains produce a variety of lipopeptides with remarkable surface and biological activities. In contrast to the structural diversity of these lipopeptides, their biosynthetic mechanism is basically conserved. They are synthesized nonribosomally by a mega-peptide synthetase unit, non-ribosomal peptide synthetase, NRPS which is composed of several cooperating multifunctional modules, each capable of performing one cycle of peptide elongation. To become an active form, they are post-translationally modified by a phosphopantetheinyl transferase. However, recent analysis of the lipopeptide synthetases suggests that there are several variants of NRPS architecture.
Biosynthesis of arthrofactin is catalyzed by the arthrofactin synthetase (Arf), which consists of three non-ribosomal peptide synthetase, NRPS protein subunits, ArfA (234 kDa), ArfB (474 kDa), and ArfC (648 kDa), which contain two, four, and five functional modules, respectively (Fig. 16, [57]). An additional C-domain was identified in the first module of ArfA, suggesting that the first amino acid could be initially acylated with a fatty acid. Site-directed mutagenesis changing the histidine residue of conserved core motif (HHXXXDG) to alanine impairs arthrofactin production (N. Roongsawang and M. Morikawa, unpublished data). This result suggested that the first C-domain is essential for biosynthesis of lipopeptide. Indeed, the β-hydroxydecanoyl thioester may be coupled to the activated leucine by the action of this C-domain to yield β-hydroxydecanoyl-L-Leu as the initial intermediate. A phylogenetic tree showed that the first C-domain of Arf belongs to N-acyl groups that use fatty acyl-CoA as their starter unit [68]. Although seven out of the 11 amino acid residues in arthrofactin are in the D-form, Arf contains no E-domains, as found in syringomycin and syringopeptin. The A-domain of D-Leu1 specifically recognizes only L-Leu in vitro. Based on these observations, we initially hypothesized that an external racemase may be responsible for incorporation of the D-amino acids in arthrofactin. Different amino acid sequences downstream of a conserved core motif [FFELGGHSLLA(V/M)] in the T-domains were expected to reflect the recognition by external racemase. However, Balibar and coworkers demonstrated in 2005 that Arf contains unique dual C/E domains, which contribute to the conversion of L-amino acids to the D-form [69]. This novel C/E domain is cryptically embedded with the C-domain located downstream of the D-amino acid–incorporating modules. Dual C/E domains can be recognized by an elongated His motif (HHI/LXXXXGD). This feature was also identified in the Syr and Syp synthetases. Another unique characteristic of Arf is the presence of C-terminal tandem Te-domains like syringopeptin. By site-directed mutagenesis, the first Te-domain (ArfC-Te1) was shown to be essential for the completion of macrocyclization and the release of the final product. The second Te-domain (ArfC-Te2) was suggested to be involved in the evolution of Arf to improve the macrocyclization efficiency. Moreover, we found that the gene encoding putative ArfA/B/C exists in the genome sequence of Pseudomonas fluorescens Pf0-1 (YP_347943/YP_347944/YP_347945). Arf represents a novel NRPS architecture that features tandem Te-domains and dual C/E domains. Interestingly, another type of NRPS involved in biosynthesis of a siderophore, pyoverdine, was also identified in arthrofactin-producing Pseudomonas sp. MIS38. A gene encoding NRPS for the chromophore part of pyoverdine contains a conventional E-domain [70]. This observation shows that different NRPS systems with dual C/E domains and a conventional E-domain are both functional in Pseudomonas spp.
The arthrofactin biosynthesis assembly line. Arthrofactin synthetase is composed of ArfA, ArfB, and ArfC enzyme complex. Each enzyme contains several sets of domains responsible for adding aminoacylated substrate amino acid residues. The last dual thioesterase Te1/Te2 releases and cyclizes arthrofactin
Our current knowledge is still not enough to understand the evolutionary history of biosurfactants. On the other hand, modification of NRPS by genetic engineering of the encoding genes is a potential method to produce useful variants. Accumulation of genetic information for lipopeptide synthetases should contribute to design biosurfactants with higher surface activity and/or novel features [71]. Moreover, understanding their biosynthetic pathways and genetic regulation mechanisms will facilitate not only uncovering the evolution of NRPS mechanisms, but also the development of cost-effective methods for large-scale production of useful lipopeptides.
4 Conclusion
Bioremediation is one of the excellent functions that have been naturally acquired in the evolution of microorganisms, whose history is more than one billion years. Therefore, there is no doubt that it is an environmentally friendly technology. However, it has a disadvantage of requiring time much longer than physicochemical methods. When considering the return on investment on the scale of human time, it has been not yet practically used in many occasions. In the future, in order to realize a sustainable biosphere without being bound by human self-convenience in a short span of view, research and development to further improve bioremediation while understanding its characteristics is desired.
References
Kato T, Haruki M, Imanaka T, Morikawa M, Kanaya S (2001a) Isolation and characterization of psychrotrophic bacteria from oil reservoir water and oil sands. Appl Microbiol Biotechnol 55:794–800
Kato T, Haruki M, Imanaka T, Morikawa M, Kanaya S (2001b) Isolation and characterization of long-chain-alkane degrading Bacillus thermoleovorans from deep subterranean petroleum reservoirs. J Biosci Bioeng 91(1):64–70
Kunihiro N, Haruki M, Takano K, Morikawa M, Kanaya S (2005) Isolation and characterization of Rhodococcus sp. strains TMP2 and T12 that degrade 2,6,10,14-tetramethylpentadecane (pristane) at moderately low temperatures. J Biotechnol 115:129–136
Morikawa M (2010) Dioxygen activation responsible for oxidation of aliphatic and aromatic hydrocarbon compounds: current state and variants. Appl Microbiol Biotechnol 87:1596–1603
Ahlborg UG, Becking GC, Birnbaum LS, Brouwer A, Derks HJGM, Feeley M, Color G, Hanberg A, Larsen JC, Liem AKD, Safe SH, Schlatter C, Wvern F, Younes M, Yrjainheikki E (1994) Toxic equivalency factors for dioxin-like PCBs. Chemosphere 28(6):1049–1067
Leigh MB, Prouzová P, Macková M, Macek T, Nagle DP, Fletcher JS (2006) Polychlorinated biphenyl (PCB)-degrading bacteria associated with trees in a PCB-contaminated site. Appl Environ Microbiol 72(4):2331–2342
Payne KAP, Quezada CP, Fisher K, Dunstan MS, Collins FA, Sjuts H, Levy C, Hay S, Rigby SEJ, Leys D (2015) Reductive dehalogenase structure suggests a mechanism for B12-dependent dehalogenation. Nature 517:513–516
van Beilen JB, Funhoff EG (2007) Alkane hydroxylases involved in microbial alkane degradation. Appl Microbiol Biotechnol 74:13–21
Rieske JS, MacLennan DH, Coleman R (1964) Isolation and properties of an iron-protein from the (reduced coenzyme Q)-cytochrome C reductase complex of the respiratory chain. Biochem Biophys Res Commun 15(4):338–344
Parales RE (2003) The role of active-site residues in naphthalene dioxygenase. J Ind Microbiol Biotechnol 30(5):271–278
Karlsson A, Parales JV, Parales RE, Gibson DT, Eklund H, Ramaswamy S (2003) Crystal Structure of naphthalene dioxygenase: side-on binding of dioxygen to iron. Science 299(5609):1039–1042
Bugg TDH, Ramaswamy S (2008) Non-heme iron-dependent dioxygenases: unravelling catalytic mechanisms for complex enzymatic oxidations. Curr Opin Chem Biol 12(2):134–140
Alexander M (1999) Biodegradation and bioremediation.2nd edn. Academic Press
Bouchez T, Patureau D, Dabert P, Juretschko S, Doré J, Delgenès P, Moletta R, Wagner M (2000) Ecological study of a bioaugmentation failure. Environ Microbiol 2:179–190
El-Fantroussi S, Agathos SN (2005) Is bioaugmentation a feasible strategy for pollutant removal and site remediation? Curr Opin Microbiol 8:268–275
van Veen JA, van Overbeek LS, van Elsas JD (1997) Fate and activity of microorganisms introduced into soil. Microbiol Mol Biol Rev 61:121–135
Thompson IP, van-der-Gast CJ, Ciric L, Singer AC (2005) Bioaugmentation for bioremediation: the challenge of strain selection. Environ Microbiol 7:909–915
Dejonghe W, Boon N, Seghers D, Top EM, Verstraete W (2001) Bioaugmentation of soils by increasing microbial richness: missing links. Environ Microbiol 3:649–657
Silva E, Fialho AM, Sá-Correia I, Burns RG, Shaw LJ (2004) Combined bioaugmentation and biostimulation to cleanup soil contaminated with high concentrations of atrazine. Environ Sci Technol 38:632–637
Moslemy P, Neufeld RJ, Giot SR (2002) Biodegradation of gasoline by gellan gum-encapsulated bacterial cells. Biosci Bioeng 80:175–184
Mrozik A, Piotrowska-Seget Z (2010) Bioaugmentation as a stratergy for cleaning up soils contaminated with aromatic compounds. Microbiol Res 182:2675–2679
Watnick P, Kolter R (2000) Biofilms, city of microbes. J Bacteriol 13:20–26
Shemesh M, Kolter R, Losick R (2010) The biocide chlorine dioxide stimulates biofilm formation in Bacillus subtilis by activation of the histidine kinase KinC. J Bacteriol 192:6352–6356
Morikawa M (2006) Beneficial biofilm formation by industrial bacteria Bacillus subtilis and related species. J Biosci Bioeng 101:1–8
Iijima S, Washio K, Okahara R, Morikawa M (2009) Biofilm formation and proteolytic activities of Pseudoalteromonas bacteria that were isolated from fish farm sediments. J Microbial Biotechnol 2:361–369
Qureshi N, Annous BA, Ezeji TC, Karcher P, Maddox IS (2005) Biofilm reactors for industrial bioconversion processes: employing potential of enhanced reaction rates. Microb Cell Fact 4:24
Singh R, Paul D, Jain RK (2006) Biofilms: implications in bioremediation. Trends Microbiol 14:389–397
Yamaga F, Washio K, Morikawa M (2010) Sustainable biodegradation of phenol by Acinetobacter calcoaceticus P23 isolated from the rhizosphere of duckweed Lemna aoukikusa. Environ Sci Technol 44:6470–6474
Shimada K, Itoh Y, Washio K, Morikawa M (2012) Efficacy of forming biofilms by naphthalene degrading Pseudomonas stutzeri T102 toward bioremediation technology and its molecular mechanisms. Chemosphere 87(3):226–233
Lalucat J, Bennasar A, Bosch R, García-Valdés E, Palleroni NJ (2006) Biology of Pseudomonas stutzeri. Microbiol Mol Biol Rev 70:510–547
Hirano S, Kitauchi F, Haruki M, Imanaka T, Morikawa M, Kanaya S (2004) Isolation and characterization of Xanthobacter polyaromaticivorans sp. nov. 127W that degrades polycyclic and heterocyclic aromatic compounds under extremely low oxygen conditions. Biosci Biotechnol Biochem 68:557–564
Gross R, Hauer B, Otto K, Schmid A (2007) Microbial biofilms: new catalysts for maximizing productivity of long-term biotransformations. Biotechnol Bioeng 98:1123–1134
Ma Q, Wood TK (2009) OmpA influences Escherichia coli biofilm formation by repressing cellulose production through the CpxRA two-component system. Environ Microbiol 11:2735–2746
Werner E, Roe F, Bugnicourt A, Franklin MJ, Heydorn A, Molin S, Pitts B, Stewart PS (2004) Stratified growth in Pseudomonas aeruginosa biofilms. Appl Environ Microbiol 70:6188–6196
Lewis K (2007) Persister cells, dormancy and infectious disease. Nat Rev Microbiol 5:48–56
Rani SA, Pitts B, Beyenal H, Veluchamy RA, Lewandowski Z, Davison WM, Buckingham-Meyer K, Stewart PS (2007) Spatial patterns of DNA replication, protein synthesis, and oxygen concentration within bacterial biofilms reveal diverse physiological states. J Bacteriol 189:4223–4233
Gilbert P, Collier PJ, Brown MR (1990) Influence of growth rate on susceptibility to antimicrobial agents: biofilms, cell cycle, dormancy, and stringent response. Antimicrob Agents Chemother 34:1865–1868
Ensley BD, Gibson DT, Laborde AL (1982) Oxidation of naphthalene by a multicomponent enzyme system from Pseudomonas sp. strain NCIB 9816. J Bacteriol 149:948–954
Takei D, Washio K, Morikawa M (2008) Identification of alkane hydroxylase genes in Rhodococcus sp. strain TMP2 that degrades a branched alkane. Biotechnol Lett 30:1447–1452
Park W, Jeon CO, Madsen EL (2002) Interaction of NahR, a LysR-type transcriptional regulator, with the alpha subunit of RNA polymerase in the naphthalene degrading bacterium, Pseudomonas putida NCIB 9816-4. FEMS Microbiol Lett 213:159–165
Park JW, Crowley DE (2006) Dynamic changes in nahAc gene copy numbers during degradation of naphthalene in PAH-contaminated soils. Appl Microbiol Biotechnol 72:1322–1329
Chang WS, van-de-Mortel M, Nielsen L, Nino-de-Guzman G, Li X, Halverson LJ (2007) Alginate production by Pseudomonas putida creates a hydrated microenvironment and contributes to biofilm architecture and stress tolerance under water-limiting conditions. J Bacteriol 189:8290–8299
Bossé JT, Sinha S, Li MS, O’Dwyer CA, Nash JH, Rycroft AN, Kroll JS, Langford PR (2010) Regulation of pga operon expression and biofilm formation in Actinobacillus pleuropneumoniae by sigma E and H-NS. J Bacteriol 192:2414–2423
Halan B, Schmid A, Buehler K (2011) Real-time solvent tolerance analysis of Pseudomonas sp. strain VLB120δC catalytic biofilms. Appl Environ Microbiol 77:1563–1571
Lang S (2002) Biological amphiphiles (microbial biosurfactants). Curr Opin Colloid Interface Sci 7(1–2):12–20
Ron EZ, Rosenberg E (2001) Natural roles of biosurfactants. Environ Microbiol 3(4):229–236
Jahan R, Bodratti AM, Tsianou M, Alexandridis P (2020) Biosurfactants, natural alternatives to synthetic surfactants: physicochemical properties and applications. Adv Colloid Interface Sci 275:102061
Zhang Y, Maier WJ, Miller RM (1997) Effect of rhamnolipids on the dissolution, bioavailability, and biodegradation of phenanthrene. Environ Sci Technol 31:2211–2217
Calvo C, Manzanera M, Silva-Castro GA, Uad I, González-Lópe J (2009) Application of bioemulsifiers in soil oil bioremediation processes. Future prospects. Sci Total Environ 407(12):3634–3640
Sriram MI, Gayathiri S, Gnanaselvi U, Jenifer PS, Raj SM, Gurunathan S (2011) Novel lipopeptide biosurfactant produced by hydrocarbon degrading and heavy metal tolerant bacterium Escherichia fergusonii KLU01 as a potential tool for bioremediation. Bioresour Technol 102(19):9291–9295
Cameotra SS, Bollag J-M (2003) Biosurfactant-enhanced bioremediation of polycyclic aromatic hydrocarbons. Crit Rev Environ Sci Technol 33(2):111–126
Bezza FA, Chirwa EMN (2017) The role of lipopeptide biosurfactant on microbial remediation of aged polycyclic aromatic hydrocarbons (PAHs)-contaminated soil. Chem Eng J 309:563–576
Lamichhane S, Krishna KCB, Sarukkalige R (2017) Surfactant-enhanced remediation of polycyclic aromatic hydrocarbons. J Environ Manage 199:46–61
Matsuyama T, Kaneda K, Nakagawa Y, Isa K, Hara-Hotta H, Yano I (1992) A novel extracellular cyclic lipopeptide which promotes flagellum-dependent and -independent spreading growth of Serratia marcescens. J Bacteriol 174(6):1769–1776
Nielsen TH, Sørensen D, Tobiasen C, Andersen JB, Christophersen C, Givskov M, Sørensen J (2002) Antibiotic and biosurfactant properties of cyclic lipopeptides produced by fluorescent Pseudomonas spp. from the sugar beet rhizosphere. Appl Environ Microbiol 68(7):3416–3423
Pamp SJ, Tolker-Nielsen T (2007) Multiple roles of biosurfactants in structural biofilm development by Pseudomonas aeruginosa. J Bacteriol 189(6):2531–2539
Roongsawang N, Hase K, Haruki M, Imanaka T, Morikawa M, Kanaya S (2003) Cloning and characterization of the gene cluster encoding arthrofactin synthetase from Pseudomonas sp. MIS38. Chem Biol 10(9):869–880
Marchant R, Banat IM (2012) Biosurfactants: a sustainable replacement for chemical surfactants? Biotechnol Lett 34(9):1597–1605
Morikawa M, Daido H, Takao T, Murata S, Shimonishi Y, Imanaka T (1993) A new lipopeptide biosurfactant produced by Arthrobacter sp. strain MIS38. J Bacteriol 175(20):6459–6466
Youssef NH, Duncan KE, Nagle DP, Savage KN, Knapp RM, McInerney MJ (2004) Comparison of methods to detect biosurfactant production by diverse microorganisms. J Microbiol Methods 56(3):339–347
Ohno A, Ano T, Shoda M (1995) Production of a lipopeptide antibiotic, surfactin, by recombinant Bacillus subtilis in solid state fermentation. Biotechnol Bioeng 47(2):209–214
Jarvis FG, Johnson MJ (1949) A glyco-lipid produced by Pseudomonas aeruginosa. J Am Chem Soc 71(12):4124–4126
Hisatsuka K, Nakahara T, Sano N, Yamada K (1971) Formation of rhamnolipid by Pseudomonas aeruginosa and its function in hydrocarbon fermentation. Agric Biol Chem 35(5):686–692
Gorin PAJ, Spencer JFT, Tullock AP (1961) Hydroxy fatty acid glycosides of sophorose from Torulopsis magnoliae. Can J Chem 39:846–895
Arima K, Kakinuma A, Tamura G (1968) Surfactin, a crystalline peptidelipid surfactant produced by Bacillussubtilis: Isolation, characterization and its inhibition of fibrin clot formation. Biochem Biophys Res Commun 31(3):488–494
Raaijmakers JM, Bruijn ID, Nybroe O, Ongena M (2010) Natural functions of lipopeptides from Bacillus and Pseudomonas: more than surfactants and antibiotics. FEMS Microbiol Rev 34(6):1037–1062
Morikawa M, Hirata Y, Imanaka T (2000) A study on the structure–function relationship of lipopeptide biosurfactants. Biochim Biophys Acta 1488(3):211–218
Roongsawang N, Lim SP, Washio K, Takano K, Kanaya S, Morikawa M (2005) Phylogenetic analysis of condensation domains in the nonribosomal peptide synthetases. FEMS Microbiol Lett 252(1):143–151
Balibar CJ, Vaillancourt FH, Walsh CT (2005) Generation of D amino acid residues in assembly of arthrofactin by dual condensation/epimerization domains. Chem Biol 12(11):1189–1200
Lim SP, Roongsawang N, Washio K, Morikawa M (2007) Functional analysis of a pyoverdine synthetase from Pseudomonas sp. MIS38. Biosci Biotechnol Biochem 71(8):2002–2009
Roongsawang N, Washio K, Morikawa M (2011) Diversity of nonribosomal peptide synthetases involved in the biosynthesis of lipopeptide biosurfactants. Int J Mol Sci 12(1):141–172
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd
About this chapter
Cite this chapter
Morikawa, M. (2022). Bioremediation: From Key Enzymes to Practical Technologies. In: Tanaka, S., Kurasaki, M., Morikawa, M., Kamiya, Y. (eds) Design of Materials and Technologies for Environmental Remediation. The Handbook of Environmental Chemistry, vol 115. Springer, Singapore. https://doi.org/10.1007/698_2021_828
Download citation
DOI: https://doi.org/10.1007/698_2021_828
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-5235-7
Online ISBN: 978-981-19-5236-4
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)