Abstract
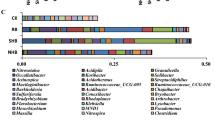
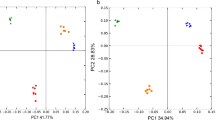
This research work was oriented to outlining the diversity of Gram-negative culturable portion of the bacterial community in three fruit plants rhizosphere. Rhizosphere samples were taken from European chestnut (Castanea sativa Mill), true service tree (Sorbus domestica L.) and cornelian cherry (Cornus mas L.) plants. Experiments were conducted for three years during the vegetation period, and the bacterial community structure was assessed with cultivation-dependent approach. Many Gram-negative isolates (n = 251) from the rhizosphere survived sub culturing and were identified by biochemical tests. A total of 57 species belonging to 29 genera were identified and assigned to four broad taxonomic groups (Bacteroidetes, Alpha-, Beta- and Gamma-proteobacteria). Several specific bacterial cluster communities were identified inside all the three rhizospheres. Most of the species belonged to the genera Moraxella, Pseudomonas, Pantoea, Enterobacter and Acinetobacter. In addition, while, using the plate count analysis, large discrepancies in numbers among physiological groups of bacteria cultured from three rhizosphere samples have not been revealed, more expressive distinctions among bacterial populations were obtained concerning the relative abundance of different genera, different taxonomic groups as well as different diversity indices. Furthermore, the number of cultured bacteria and their taxonomic distribution in the rhizosphere of all three plants changed not only explicitly during vegetation period but continually during the three years of investigation. It seems that rhizosphere bacterial populations of each plant are under the influence of the specific root-released materials.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bowen, G.D. & Rovira, A.D. 1991. The rhizosphere, the hidden half, pp. 641–649. In: Waisel, Y, Eshel, A. & Kafkafi, U. (eds) Plant Roots — The Hidden Half, Marcel Dekker Inc., New York.
Cook, R.J., Thomashow, L.S., Weller, D.M., Fujimoto, D., Mazzola, M., Bangera, G. & Kim, D.S. 1995. Molecular mechanisms of defense by rhizobacteria against root disease. Proc. Natl. Acad. Sci. USA 92: 4197–4201.
Duineveld, B.M., Kowalchuk, G.A., Keijzer, A., van Elsas, J.D. & van Veen, J.A. 2001. Analysis of bacterial communities in the rhizosphere of chrysanthemum via denaturing gradient gel electrophoresis of PCR-amplified 16S rRNA as well as DNA fragments coding for 16S rRNA. Appl. Environ. Microbiol. 67: 172–178.
Ellis, R.J., Morgan, P., Wieghtman, A.J. & Fry, J.C. 2003. Cultivation-dependent and-independent approaches for determining bacterial diversity in heavy-metal-contaminated soil. Appl. Environ. Microbiol. 69: 3223–3230.
Griffiths, B.S., Ritz, K., Ebblewhite, N. & Dobson, G. 1999. Soil microbial community structure: effects of substrate loading rates. Soil Biol. Biochem. 31: 145–153.
Hessenmueller, A. & Zeller, W. 1996. Biological control of soil-borne Phytophthora species on strawberry with bacterial antagonists: 1. Antagonistic effect of colonization of rhizoplane. Z. Pflanzenkrankh. Pflanzenschutz 103: 602–609.
Holmes, B., Pinning, C.A. & Davson, C.A. 1986. A probability matrix for the identification of gramnegative, aerobic, non-fermentative bacteria that grow on nutrient agar. J. Gen. Microbiol. 132: 1827–1842.
Juhásová, G. 1999. Hubové choroby gaštana obyčajného (Gastanea sativa Mill). [Fungal diseases of chestnut (Castanea sativa Mill).] Veda, Bratislava, 189 pp.
Juhásová, G. & Hrubík, P. 1984. Choroby a škodcovia cudzokrajných drevín na Slovensku. [The diseases and evil-doers of foreign woods in Slovakia.] Acta Dendrobiologica, Veda, Bratislava, 164 pp.
Kopřivník, B. & Vychodilová, M. 1975. Kultivace a identifikace mixobakterií. Acta HEM, 4: 17–20. (Cultivation and identification of myxobacteria)
Liljeroth, E., Burgers, S.L.G.E. & van Veen, J.A. 1991. Changes in bacterial populations along roots of wheat seedlings. Biol. Fertil. Soils 10: 276–280.
Landa, B.B., Mavrodi, O.V., Raaijmakers, J.M., McSpadden Gardener, B.B., Thomashow, L.S. & Weller, D.M. 2002. Differential ability of genotypes of 2,4-diacetylphloroglucinol-producing Pseudomonas fluorescens strains to colonize the roots of pea plants. Appl. Environ. Microbiol. 68: 3226–3237.
Lynch, J.M. & Whipps, J.M. 1990. Substrate flow in the rhizosphere. Plant Soil 129: 1–10.
Mitsui, H., Gorlach, K., Lee, H.J., Hattori, R. & Hattori, T. 1997. Incubation time and media requirements of culturable bacteria from different phylogenetic groups. J. Microbiol. Methods 30: 103–110.
Mohan, N. & Mahadevan, A. 2003. Effect of phenol on ultra structure and plasmid DNA of Xanthomonas oryzae pv. oryzae. Indian. J. Exp. Biol. 41: 78–81.
Naseby, D.C. & Lynch, J.M. 1999. Effects of Pseudomonas fluorescens F113 on ecological functions in the pea rhizosphere are dependent on pH. Microb. Ecol. 37: 248–256.
Naseby, D.C. & Lynch, J.M. 2001. Effect of 2,4-diacetylphloroglucinol producing, overproducing, and nonproducing Pseudomonas fluorescens F113 in the rhizosphere of pea. Microb. Ecol. 42: 193–200.
Neal, J.R., Jr., Atkinson, T.G. & Larson, R.I. 1970. Changes in the rhizosphere microflora of spring wheat induced by disomic substitution of a chromosome. Can. J. Microbiol. 16: 153–158.
Neal, J.R., Jr., Larson, R.I. & Atkinson, T.G. 1973. Changes in rhizosphere populations of selected physiological groups of bacteria related to substitution of specific pairs of chromosomes in spring wheat. Plant Soil 39: 209–212.
Ovreås, L. & Torsvik, V. 1998. Microbial diversity and community structure in two different agricultural soil communities. Microb. Ecol. 36: 303–315.
Penaloza-Vazquez, A., Fakhr, M.K., Bailey, A.M. & Bender, C.L. 2004. AlgR functions in algC expression and virulence in Pseudomonas syringae pv. syringae. Microbiology 150: 2727–2737.
Raaijmakers, J.M. & Weller, D.M. 2001. Exploiting genotypic diversity of 2,4-diacetylphloroglucinol-producing Pseudomonas spp.: characterization of superior root-colonizing P. fluorescens strain Q8r1-96. Appl. Environ. Microbiol. 67: 2545–2554.
Sanford, G.B. 1946. Soil-borne diseases in relation to the microflora associated with various crops and soil amendments. Soil Sci. 61: 9–22.
Sliwinski, B.J., Kreuzer, M., Wettstein, H.R. & Machmuller, A. 2002. Rumen fermentation and nitrogen balance of lambs fed diets containing plant extracts rich in tannins and saponins, and associated emissions of nitrogen and methane. Arch. Tierernahr. 56: 379–392.
Sorbo, B.S., Giordano, S., Ricciardi, L., Ferrara, S., Montesano, D., Castaldo Cobianchi, R., Vuotto, M.L. & Ferrara, L. 2000. Antibacterial and allelopathic activity of extract from Castanea sativa leaves. Fitoterapia 71: S110–S116.
Swinnen, J., van Veen, J.A. & Merck, R. 1994. 14-C pulse labeling of field-grown spring wheat: an evaluation of its use in rhizosphere carbon budget estimations. Soil Biol. Biochem. 26: 161–170.
Toyota, K., Yamamoto, K. & Kimura, M. 1994. Isolation and characterization of bacteria responsible for the suppression of Fusarium oxysporum f. sp. raphani on the host rhizoplane. Soil Sci. Plant Nutr. 40: 381–390.
Vidaver, A.K. 2002. Uses of antimicrobials in plant agriculture. Clin. Infect. Dis. 34(Suppl 3): S107–110.
Weindling, R. 1946. Microbial antagonism and disease control. Soil Sci. 61: 23–30.
Yang, C.H., Crowley, D.E., Borneman, J. & Keen, N.T. 2001. Microbial phyllosphere populations are more complex than previously realized. Proc. Natl. Acad. Sci. USA 98: 3889–3894.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Harichová, J., Karelová, E., Chovanová, K. et al. Comparison of culturable Gram-negative bacterial community structures in the rhizosphere of three fruit plants. Biologia 61, 663–670 (2006). https://doi.org/10.2478/s11756-006-0138-1
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.2478/s11756-006-0138-1