Abstract
Aging is a multifactorial process characterized by a gradual loss of physiological functions, which leads to increased vulnerability of the organism to age-related diseases and, finally, to death. Several theories have been proposed to explain the nature of aging, one of which relates aging to the damage to cell structures and DNA caused by free radicals. However, an increasing amount of evidence suggests that the molecular mechanisms of aging are also associated with epigenetic modifications, such as DNA methylation, noncoding RNA and histone changes. In this review, we will analyze the significance of the results of these studies and show how the interrelated effects of oxidative stress and epigenetics can explain the cause of the extinction of physiological functions during aging.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Aging is the process of the gradual extinction of all physiological functions, which leads to an increase in the vulnerability of the body to age-related diseases and, finally, to death [1–3]. This process is still insufficiently studied, although many different theories attempting to explain it have been put forward [1–3]. Unfortunately, none of the theories so far fully explain all aspects of aging, which is a complex biological process. The most famous and best-studied theory is the free radical theory of aging, which was first proposed by D. Harman [39] and again 2 years later by N. Emanuel [10]. This theory associates aging with the accumulation of free radicals produced by the energy metabolism of mitochondria, which leads to the development of cell toxicity [39] and damage to nuclear [15, 16] and mitochondrial DNA (mtDNA) [10, 11], as well as cell membrane structures [83]. According to V.K. Koltover, the author of the concept of “limited reliability of biosystems,” the accumulation of peroxidation products in cells and tissues limits the reliability of biosystems, which explains aging and the prolongation of the life of animals with bioantioxidants [49]. At the same time, there is evidence that aging also depends on the activity of certain genes [6], the molecular mechanisms of regulation of which are associated with epigenetic changes in the cell [52]. We tried to connect these two mechanisms, the accumulation of free radicals and epigenetic modifications, for an exhaustive review of the manner in which a combination of different epigenetic changes and oxidative stress can affect the aging process [75].
EPIGENETIC APPARATUS OF THE CELL
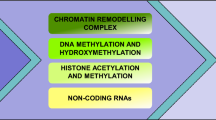
The term “epigenetics” was first used by the English embryologist K. Waddington to describe the variability of the formation of bodily structures during the course of embryogenesis. A key element of Waddington’s epigenetic concept was the idea that the various gene products (proteins, hormones, etc.) are signals that cause genes to change their activity. In accordance with these ideas, Waddington defined epigenetics as “a branch of biology that studies the causal interactions of genes and their products that form the phenotype” [77]. Then, in 1987, R. Holliday revised the term “epigenetics” as “hereditary changes in gene expression that are not associated with changes in the DNA sequence” [44]. Based on these definitions, it can be assumed that the epigenome implies the presence in the cell of certain mechanisms that control gene expression. Indeed, three such mechanisms are known today: DNA methylation, histone modifications, and the effects of noncoding RNAs [29]. Studies have shown that these epigenetic mechanisms together form the “epigenetic landscape” necessary for the regulation of the expression of tissue genes, X-chromatin inactivation, and genomic imprinting [71].
DNA methylation is an essential epigenetic modification that regulates gene expression. When DNA is methylated with DNA methyltransferases, the methyl group is transferred to the carbon atom in the fifth position of cytosine with S-adenosyl-methionine (SAM) as a methyl group donor, resulting in the formation of 5-methylcytosine (5mC); and S-adenosyl-L-homocysteine. In mammalian cells, there are several cytosine methyltransferases: DNMT1, DNMT3a, and DNMT3b. Although DNA methylation is a stable epigenetic sign, it can be removed by passive or active demethylation processes. Passive methylation loss can be achieved via successive cycles of DNA replication in the absence of functional enzymes, such as DNMT1/UHRF1 [80], the inhibition of DNMT enzymes [63], the cytosolic localization of DNMT [48], and the degradation of DNMT recruitment on DNA [16]. At the same time, the active removal of 5mC occurs due to the formation of 5-hydroxymethylcytosine (5hmC) during the oxidation of 5mC under the influence of TET (ten eleven tranclocation) family of enzymes. As for histone modification, it should be noted that eukaryotic DNA is packed in chromatin, which consists of nucleosome units encircled by 147 pairs of DNA nucleotides around the octamer of the four main histones (H2A, H2B, H3, and H4). Between two adjacent nucleosomes, there is a linker histone that binds to the outer side of the nucleosome, fixing a DNA strand on it.
Historically, chromatin was classified as euchromatin or heterochromatin (in accordance with its state of compaction), although there is a whole spectrum of chromatin states, which indicates the high flexibility of its macromolecule. The chromatin structure can be modified with the mechanisms of the recording, reading, and erasing of chromatin, which can reconstruct nucleosomes or modify histones via posttranslational modifications (acetylation of histones, phosphorylation, glycosylation, ubiquitilation, and sumoylation), establishing various states of chromatin transcription [31, 32]. Finally, noncoding RNAs (noncoding RNAs or ncRNAs) can perform their regulatory functions by acting as epigenetic regulators of gene expression and chromatin remodeling. The detailed mechanisms of their functioning have not yet been fully studied, but it is already known that they recruit various histone-modifying enzymes that recognize (read), add (write), delete (erase), and replace chromatin modifications. A good example is the long noncoding RNA (long ncRNA or lncRNA) XIST (X-inactive-specific transcript), which covers one of the X chromosomes, recruiting the repressive complex of the second group polycomb (PRC2 or polycomb repressive complex 2) and causing heterochromatization and transcriptional repression of the entire X chromosome [50, 87]. Therefore, an understanding of the involvement of the dynamics and regulation of various epigenetic modifications in the aging process is of great interest.
OXIDATIVE STRESS
An increase in the formation of free radicals in the body (oxidative stress) and the associated increase in lipid peroxidation processes (lipid peroxidation syndrome) are accompanied by disturbances in the properties of biological membranes and cell functioning [4]. The phrase “oxidative stress” first appeared in the scientific literature in the 1970s and was used by the authors to indicate the impairment caused by the action of a strong oxidizing agent [65]. Subsequently, the term oxidative stress began to be used to refer to the effects of ROS on blood cells. Soon, the concept of oxidative stress as a pathological condition caused by an excess of free radicals and the insufficiency of the antioxidant defense system arose. Thus, oxidative stress is an imbalance between the antioxidant defense of the body and the formation of ROS in favor of the latter [26]. ROS, in turn, are highly reactive molecules that consist of a variety of chemicals, including superoxide anion (\({\text{O}}_{2}^{ - }\)), hydroxyl radical (OH), and hydrogen peroxide (H2O2). ROS are produced as a result of uncontrolled production in interactions with the molecular structures of the cell, such as DNA, proteins, lipids and carbohydrates, which leads to damage to these molecules and a change in the activity of metabolic pathways. These ROS effects ultimately lead to the development of various diseases, such as cancer, neurodegenerative diseases, and diabetes, as well as aging [60].
Mitochondria are the main intracellular source of ROS generation as a result of electron transfer during ATP production [5, 58]. This is due to the fact that electron “leakage” occurs in the respiratory chain, especially from the I and III mitochondrial enzyme complexes [31]. The increase in ROS production can also be caused by exogenous factors, such as radioactivity and ultraviolet radiation. To prevent oxidative stress, the cells are equipped with a network of various antioxidant defense molecules consisting of enzymatic and nonenzymatic mechanisms. These endogenous antioxidant enzymes are glutathione S-transferase P1 (GTSP1), glutathione peroxidase, catalase, SOD, sulfiredoxin, and peroxiredoxin [27]. As for nonenzymatic mechanisms, they consist of a variety of low molecular weight antioxidants, which include glutathione and vitamins C (ascorbic acid), A (retinol), and E (tocopherol) [19]. Both of these systems—enzymatic and nonenzymatic—function in concert, maintaining antioxidant protection at the proper level; however, with excessive ROS production and the absence of an adequate increase in the synthesis of intracellular antioxidant proteins, the normal redox state of the cell changes, causing oxidative stress and damage to cell structures.
RELATIONSHIP BETWEEN EPIGENETICS AND OXIDATIVE STRESS
According to a number of studies [18, 33, 53], oxidative stress increases with age. It arises, as noted above, due to ROS accumulation and is accompanied by a decrease in cell repair mechanisms, which ultimately leads to a wide range of DNA damage, as well as to the impairment of the epigenetic state of the cell. Here are some examples of the close interaction of oxidative stress and the epigenetic landscape. For example, ROS can affect the methyloma (genome methylation profile) via the formation of oxidized DNA lesions due to the hydroxylation of pyrimidines and 5hmC, which can interfere due to structural similarities with epigenetic signals associated with 5hmC [51, 59]. ROS also affect DNA demethylation via the oxidation of DNA and TET-mediated hydroxymethylation [20]. ROS can indirectly modulate the activity of the epigenetic apparatus, since histone-modifying enzymes depend on the intracellular level of major metabolites, such as acetyl coenzyme A, Fe, ketoglutarate, NAD+, and S-adenosylmethionine, which indicates a close relationship between epigenetic changes with the global cellular metabolism and the cell energy level [73]. Therefore, oxidative stress can affect the cell globally, e.g. at several levels—from DNA and histones to histone modifiers, which can cause changes in the epigenetic landscape of the cell [55].
EPIGENETIC MODIFICATIONS AND AGING
Epigenetic changes are a sign of the aging of an organism [52] and, as research results have shown, they are the missing link explaining the difference in the aging structure between two monozygotic twins [34, 73]. The information encoded in the epigenome includes DNA methylation, chromatin remodeling, posttranslational modifications of histone proteins, structural and functional histone variants, and the transcription of noncoding RNA. The combination of all these different types of epigenetic information contains both the function and the fate of all cells and tissues of the body.
DNA METHYLATION AND THE “EPIGENETIC CLOCK” CONCEPT
It was established that aging is accompanied by a global reduction in DNA methylation and the hypermethylation of promoters of certain genes [42, 36]. DNA hypomethylation occurs in transposed DNA repeating elements, including Alu and LINE-1 elements, which leads to an increase in the activity of retrotransposon and genomic instability [79]. Thus, specific CpG islands of regulatory transcription genes [35], apoptosis [61], development, and differentiation [67] demonstrated significant hypomethylation during aging. The variations in the profile of methylation of CpG sites with age were so stable that some researchers used this as a tool to assess the biological age of tissues. For example, S. Bocklandt et al. [13] determined the epigenetic profile of three genes (EDARDD, TOM1L1, and NPTX2) and showed that it linearly changes with age over five decades. An algorithm for calculation of the biological age was developed in another study based on an analysis of more than 8000 tissue and cell samples from people of different ages [45]. This algorithm revealed 353 methylation sites, which, in the end, formed the basis of the concept of “epigenetic clocks.” Analysis of the methylation profile in these areas made it possible to evaluate tissue age with an error of no more than 3 years. Since then, the epigenetic clock has been used to assess the effects of diet, exercise, education, and other lifestyle factors on this molecular measure of aging.
However, it should be noted that DNA methylation is not the only epigenetic biomarker of aging that changes significantly with age. For example, changes in the level of genomic distribution of histone methylation and acetylation profiles of histones with age were reported [12, 64, 70], as well as more significant shifts in the level of histone acetylation in elderly twins as compared with young twins [34]. Consequently, it can be assumed that not only DNA methylation, but also other epigenetic markers, can be associated to a certain extent with the rate of molecular aging, and, taken together, can probably more accurately identify the biological age [1, 3].
HISTONE MODIFICATIONS
Histones undergo a wide range of posttranslational modifications that have a major impact on the global chromatin structure, gene expression, genome stability, and replication. Therefore, histone modifications can apparently affect all biological processes, including aging, which is confirmed by data from studies on the development of an imbalance between activating and repressive histone modifications in the elderly. It was shown that the histone methylations H3K4me3 and H3K27me3, which are epigenetic modifications associated with transcription, are related to the regulation of life span. For example, the inhibition of ASH-2 and SET-2 methyltransferases was associated with a global decrease in H3K4me3 levels, which increased lifespan. Conversely, the inhibition of RBR-2 demethylase reduced life expectancy [38]. The global structure of histone methylation is different for different organisms due to differences in their aging processes; however, there was an increasing trend of the signs of the activation of histone methylation labels affecting chromatin compaction in cells with aging [54]. Other examples are studies of histone deacetylase SIR-2 of C. elegans nematodes, which was associated with an increase in longevity [24, 76]. Conversely, the SIRT-1 homolog decreased with aging (probably due to an increase in ROS level) [46, 62], inhibiting heterochromatin silencing, which could alter gene expression during aging [60]. It is important to note that these data provide convincing evidence of the role epigenetics plays in regulating life expectancy.
NONCODING RNA
Noncoding RNAs, both short (mainly microRNAs, miRNAs) and long (lncRNAs), affect aging by controlling gene expression at the transcriptional and posttranscriptional levels in many ways. The most characteristic example of the role of miRNAs in aging is from a study performed on C. elegans nematodes. A significant component of it is miRNA lin-4 [47], which regulates the aging and expression of the target, the lin-14 transcript according to the feedback rule [14]. Reduced lin-4 activity decreased lifespan and accelerated tissue aging, while the lin-4 overexpression or lin-14 knockdown increased lifespan. Age-related differences in the miRNAs profile were also found in humans [61], and dysregulation in the expression of lncRNAs was found in diseases associated with aging [37]. A good example is lncRNAH19, which controls the imprinting of the conserved cluster of H19 and IGF2 (insulin-like growth factor). The loss of IGF2-H19 imprinting leads to an increase in the expression of IGF2 associated with age-related diseases [21].
OXIDATIVE STRESS AND AGING
Uncontrolled ROS production leads to devastating consequences in the body, and therefore many theories have been proposed to explain the phenomenon of aging from the standpoint of oxidative stress. One of them is the theory of free radicals, which claims that aging is a consequence of the accumulation of harmful effects caused by free radicals [39]. As evidence of this theory, an increase in ROS production by mitochondria is often found in the tissues of old animals [53]. A significant increase in the level of oxidative damage in cells during aging was reported in other studies [18, 33]. The results of these studies were consistent with the statements of free radical theory that oxidative stress leads to the accumulation of damage to lipids, nucleic acids, proteins and carbohydrates, causing cell dysfunction and making the body more vulnerable to harmful external agents. Since mitochondria are the main producer of ROS in mammalian cells, 8-oxo-7,8-dihydro-2'-deoxyguanosine (8-oxodeoxyguanosine), the predominant form of free radical DNA damage, exhibits a higher level in mtDNA than in nuclear DNA, which suggests a higher susceptibility of mtDNA to oxidative damage [8, 9, 56, 66, 68]. Therefore, the mitochondrial theory of aging, which is closely related to the theory of free radicals, postulates that it is oxidative damage that occurs during the oxidative phosphorylation of mitochondrial macro molecules, such as mtDNA, proteins, or lipids, which are responsible for aging [56]. In addition, mitochondria play a decisive role in the regulation of apoptosis; therefore, age-related mitochondrial oxidative stress promotes apoptosis during aging [81]. There is growing experimental data that indicate that antioxidants targeting mitochondria have a beneficial effect during aging [7, 23]. It was shown that they provide protection against oxidative damage in mitochondria to a greater extent than nontargeted cellular antioxidants because of their ability to penetrate the mitochondrial phospholipid bilayer and eliminate ROS at the very beginning of their formation [25, 41] (Fig. 1).
Association of epigenetic mechanisms and oxidative stress with aging. Oxidative stress caused by endogenous or exogenous factors increases the level of ROS, the main producers of which are mitochondria. Uncontrolled ROS production contributes to aging and the development of age-related diseases. In addition, ROS are a factor modulating the activity of the epigenetic apparatus. Epigenetic changes, such as global reduction in DNA methylation or hypomethylation of specific genes, as well as other epigenetic modifications, are associated with aging.
PREMATURE AGING AS A RESULT OF EPIGENETIC EFFECTS AND OXIDATIVE STRESS
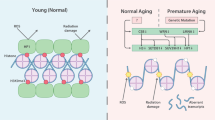
Aging causes deterioration of the epigenetic landscape, as well as an increase in the level of oxidative stress in the cell, which can ultimately contribute to the development of diseases [2]. In nature, there are the so-called progeroid syndromes (from the Greek πϱογέϱος—prematurely aged) syndromes, a group of systemic diseases that are very similar to physiological aging. They serve as a powerful tool for research on the physiological process of aging.
The most famous diseases of premature aging are Hutchinson-Gilford syndrome and Werner syndrome. The first of these syndromes is caused by a mutation in the LMNA gene; it encodes two protein products (lamin A and lamin C), which are the main components of the internal nuclear membrane [30]. This mutation leads to an abnormal version of the lamin A protein called progerin. At the same time, Werner syndrome is caused by mutations in the ATP-dependent helicase WRN gene [85], which is involved in the signaling pathway for DNA recovery. Hutchinson-Gilford syndrome is characterized by a particularly pronounced susceptibility to DNA damage caused by ROS, which indicates a deterioration in the ability to repair DNA damage [17]. Interestingly, it is during these syndromes that epigenetic and chromatin structures are damaged. Thus, profiles of aberrant DNA methylation (increased methylation) are found in hypomethylated regions and, conversely, a loss of methylation in hypermethylated regions [43]. A loss of heterochromatin is also caused by the absence of WRN in Werner syndrome [86] and progerin accumulation in HGPS [72]. The epigenetic tag H3K27me3 is lost on the inactive X chromosome as a result of a decrease in the regulation of the EZH2 methyltransferase enzyme in Hutchinson-Gilford syndrome [82].
Epigenetics and oxidative stress play a significant role in the pathogenesis of other diseases. For example, the organs of the respiratory system are constantly exposed to endogenous (air pollutants, poisonous gases, and cigarette smoke) sources of oxidizing agents. As a result, this large-scale accumulation of ROS directly affects lung cells, reducing their functional activity. This is especially noticeable in smokers and patients with COPD; an increase in their ROS level leads to the deregulated expression of pro-inflammatory genes and a decrease in the enzymatic activity of HDAC2 deacetylase [74, 82]. This histone deacetylase delays cell aging by negatively regulating genes that promote aging, such as p21 and p16 [83]. Therefore, a decrease in histone deacetylase can accelerate cell aging in COPD.
The nervous system is also very sensitive to the effects of oxidative stress. Thus, one of the earliest events in the pathogenesis of Alzheimer’s disease is oxidative damage to DNA [22]; it is caused by the oxidation of guanine to 8-oxodeoxyguanosine, which can alter the binding of transcription factors to DNA and affect epigenetic signals [28]. The evidence for the role of epigenetic modifications and oxidative stress in cardiovascular diseases is increasing. The production of nitric oxide (NO) by NO synthases (NOS) plays an important cardioprotective role by regulating blood pressure and vascular tone, as well as inhibiting platelet aggregation and leukocyte adhesion. However, the interaction of NO with superoxide leads to the formation of peroxynitrite, which reduces its activity and increases oxidative stress. NO inactivation by ROS is also responsible for endothelial dysfunction, which contributes to the development of cardiovascular disease.
A significant role of oxidative stress is also exhibited in cancer. Oncogenic cancer cells have been shown to generate elevated ROS levels as by-products of their metabolic rate to maintain tumorigenicity [78]. It would seem that high ROS levels should cause cell death, but this does not happen. The cancer cells bypass this obstacle by increasing the synthesis of intracellular antioxidant proteins. As a result, a high ROS level and the simultaneous passage of protumorogenic signals that do not lead to cell death are maintained [40, 57, 69, 84]. With nonsmall cell lung cancer, SOD1 is expressed at a relatively high level. The function of the enzyme is to maintain a low level of superoxide in the cytosol in order to protect the cell from oxidative stress and subsequent death. When exposed to this enzyme with a small molecule, ATN-224 (SOD1 inhibitor), tumor growth decreased, which indicates the potential clinical use of this inhibitor for these types of adenocarcinoma [36].
CONCLUSIONS
Thus, the facts above demonstrate how the interrelated effects of oxidative stress and epigenetic changes affect the aging process. In addition, these data expand our understanding of the molecular foundations of aging and open up new paths to influence this process. For example, the creation of mitochondria-targeted antioxidants already makes it possible to protect mitochondria from oxidative damage to a greater extent than nontargeted cellular antioxidants and thereby to slow cell death. Another promising innovation is the invention of epigenetic clocks, which allow the determination of a person’s biological age. For example, if one’s biological age by this “clock” is ahead of one’s chronological age, then this will make it possible to predict life expectancy, the development of diseases associated with aging, and even death with a certain degree of probability. Another question is whether the epigenetic clock is a driver or a simple age marker. In the first case, researchers have exciting prospects for the development of tools to manipulate molecular aging processes.
Change history
29 July 2024
An Erratum to this paper has been published: https://doi.org/10.1134/S2079057024010016
REFERENCES
Anisimov, V.N., Is there an unambigous answer to the question: Whether the aging program exists or not?, Russ. J. Gen. Chem., 2010, vol. 80, no. 7, pp. 1395–1406.
Anisimov, V.N., Molekulyarnye i fiziologicheskie mekhanizmy stareniya (Molecular and Physiological Mechanisms of Senescence), St. Petersburg: Nauka, 2008, in 2 vols.
Anisimov, V.N., Serpov, V.Yu., Finagentov, A.V., and Khavinson, V.K., A new stage of development of gerontology and geriatrics in Russia: problems o creation of a geriatric care system. Part 2. The structure of the system, scientific approach, Adv. Gerontol., 2018, vol. 8, no. 1, pp. 1–11.
Vladimirov, Yu.A., Free radicals in biological systems, Sorosovskii Obraz. Zh., 2000, vol. 6, no. 12, pp. 13–19.
Grivennikova, V.G. and Vinogradov, A.V., Generation of reactive oxygen species by mitochondria, Usp. Biol. Khim., 2013, vol. 53, no. 12, pp. 245–296.
Moskalev, A.A., Starenie i geny (Aging and Genes), St. Petersburg: Nauka, 2008.
Neroev, V.V., Archipova, M.M., Bakeeva, L.E., et al., Mitochondria-targeted plastoquinone derivatives as tools to interrupt execution of the aging program. 4. Age-related eye disease. SkQ1 returns vision to blind animals, Biochemistry (Moscow), 2008, vol. 73, no. 12, pp. 1317–1328.
Skulachev, V.P., Oxygen in a living cell: good and evil?, Sorosovskii Obraz. Zh., 1996, vol. 3, no. 3, pp. 4–10.
Skulachev, V.P., The phenomenon of programmed death. Mitochondria, cells and organs: the role of reactive oxygen species, Sorosovskii Obraz. Zh., 2001, vol. 7, no. 6, pp. 4–10.
Emanuel’, N.M. and Lipchina, L.P., Effect of chain oxidation inhibitors of development of leukemia in mice, Dokl. Akad. Nauk SSSR, 1958, vol. 121, pp. 141–144.
Balaban, R.S., Nemoto, S., and Finkel, T., Mitochondria, oxidants, and aging, Cell, 2005, vol. 120, no. 4, pp. 483–495. https://doi.org/10.1016/j.cell.2005.02.001
Benayoun, B.A., Pollina, E.A., and Brunet, A., Epigenetic regulation of ageing: linking environmental inputs to genomic stability, Nat. Rev. Mol. Cell Biol., 2015, vol. 16, no. 10, pp. 593–610. https://doi.org/10.1038/nrm4048
Bocklandt, S., Lin, W., Sehl, M.E., et al., Epigenetic predictor of age, PLoS One, 2011, vol. 6, art. ID e14821. https://doi.org/10.1371/journal.pone.0014821
Boehm, M. and Slack, F., A developmental timing microRNA and its target regulate life span in C. elegans,Science, 2005, vol. 310, no. 5756, pp. 1954–1957. https://doi.org/10.1126/science.1115596
Bohr, V.A. and Anson, R.M., DNA damage, mutation and fine structure DNA repair in aging, Mutat. Res., 1995, vol. 338, nos. 1–6, pp. 25–34. PMID 7565878
Bostick, M., Kim, J.K., Estève, P.O., et al., UHRF1 plays a role in maintaining DNA methylation in mammalian cells, Science, 2007, vol. 317, no. 5845, pp. 1760–1764. https://doi.org/10.1126/science.1147939
Camozzi, D., Capanni, C., Cenni, V., et al., Diverse lamin-dependent mechanisms interact to control chromatin dynamics: focus on laminopathies, Nucleus, 2014, vol. 5, no. 5, pp. 427–440. https://doi.org/10.4161/nucl.36289
Capel, F., Rimbert, V., Lioger, D., et al., Due to reverse electron transfer, mitochondrial H2O2 release increases with age in human vastuslateralis muscle although oxidative capacity is preserved, Mech. Ageing Dev., 2005, vol. 126, no. 4, pp. 505–511. https://doi.org/10.1016/j.mad.2004.11.001
Cencioni, C., Spallotta, F., Martelli, F., et al., Oxidative stress and epigenetic regulation in aging and age-related diseases, Int. J. Mol. Sci., 2013, vol. 14, no. 9, pp. 17643–17663. https://doi.org/10.3390/ijms140917643
Chia, N., Wang, L., Lu, X., et al., Hypothesis: environmental regulation of 5-hydroxymethylcytosine by oxidative stress, Epigenetics, 2011, vol. 6, no. 7, pp. 853–856. PMID 21617369
Christofor, G., Naik, P., and Hanahan, D., A second signal supplied by insulin-like growth factor II in oncogene-induced tumorigenesis, Nature, 1994, vol. 369, no. 6479, pp. 414–418. https://doi.org/10.1038/369414a0
Coppedè, F. and Migliore, L., Evidence linking genetics, environment, and epigenetics to impaired DNA repair in Alzheimer’s disease, J. Alzheimer’s Dis., 2010, vol. 20, no. 4, pp. 953–966. https://doi.org/10.3233/JAD-2010-1415
Dai, D.-F., Karunadharma, P.P., Chiao, Y.A., et al., Altered proteome turnover and remodeling by short-term caloric restriction or rapamycin rejuvenate the aging heart, Aging Cell, 2014, vol. 13, no. 3, pp. 529–539. https://doi.org/10.1111/acel.12203
Dang, W., Steffen, K.K., Perry, R., et al., Histone H4 lysine 16 acetylation regulates cellular lifespan, Nature, 2009, vol. 459, no. 7248, pp. 802–807. https://doi.org/10.1038/nature08085
Dashdorj, A., Jyothi, K.R., Lim, S., et al., Mitochondria-targeted antioxidant MitoQ ameliorates experimental mouse colitis by suppressing NLRP3 inflammasome-mediated inflammatory cytokines, BMC Med., 2013, vol. 11, no. 178. https://doi.org/10.1186/1741-7015-11-178
Dasuri, K., Zhang, L., and Keller, J.N., Oxidative stress, neurodegeneration, and the balance of protein degradation and protein synthesis, Free Radical Biol. Med., 2013, vol. 62, pp. 170–185. https://doi.org/10.1016/j.freeradbiomed.2012.09.016
Davies, K.J., Oxidative stress, antioxidant defenses, and damage removal, repair, and replacement systems, IUBMB Life, 2000, vol. 50, nos. 4–5, pp. 279–289. https://doi.org/10.1080/713803728
Dizdaroglu, M., Jaruga, P., Birincioglu, M., and Rodriguez, H., Free radical-induced damage to DNA: mechanisms and measurement, Free Radical Biol. Med., 2002, vol. 32, no. 11, pp. 1102–1115.
Egger, G., Liang, G., Aparicio, A., and Jones, P.A., Epigenetics in human disease and prospects for epigenetic therapy, Nature, 2004, vol. 429, no. 6990, pp. 457–463. https://doi.org/10.1038/nature02625
Eriksson, M., Brown, W.T., Gordon, L.B., et al., Recurrent de novo point mutations in lamin A cause Hutchinson-Gilford progeria syndrome, Nature, 2003, vol. 423, no. 6937, pp. 293–298. https://doi.org/10.1038/nature01629
Ernst, J., Kheradpour, P., Mikkelsen, T.S., et al., Mapping and analysis of chromatin state dynamics in nine human cell types, Nature, 2011, vol. 473, no. 7345, pp. 43–49. https://doi.org/10.1038/nature09906
Fisher-Wellman, K.H. and Neufer, P.D., Linking mitochondrial bioenergetics to insulin resistance via redox biology, Trends Endocrinol. Metab., 2012, vol. 23, no. 3, pp. 142–153. https://doi.org/10.1016/j.tem.2011.12.008
Fraga, C.G., Shigenaga, M.K., Park, J.W., et al., Oxidative damage to DNA during aging: 8-hydroxy-2'-deoxyguanosine in rat organ DNA and urine, Proc. Natl. Acad. Sci. U.S.A., 1990, vol. 87, no. 12, pp. 4533–4537. PMID 2352934
Fraga, M.F., Ballestar, E., Paz, M.F., et al., Epigenetic differences arise during the lifetime of monozygotic twins, Proc. Natl. Acad. Sci. U.S.A., 2005, vol. 102, no. 30, pp. 10604–10609. https://doi.org/10.1073/pnas.0500398102
Gentilini, D., Mari, D., Castaldi, D., et al., Role of epigenetics in human aging and longevity: genome-wide DNA methylation profile in centenarians and centenarians’ offspring, Age (Netherlands), 2013, vol. 35, no. 5, pp. 1961–1973. https://doi.org/10.1007/s11357-012-9463-1
Glasauer, A., Sena, L.A., Diebold, L.P., et al., Targeting SOD1 reduces experimental non-small cell lung cancer, J. Clin. Invest., 2014, vol. 124, no. 1, pp. 117–128. https://doi.org/10.1172/JCI71714
Grammatikakis, I., Panda, A.C., Abdelmohsen, K., and Gorospe, M., Long noncoding RNAs (lncRNAs) and the molecular hallmarks of aging, Aging (N.Y.), 2014, vol. 6, no. 12, pp. 992–1009. https://doi.org/10.18632/aging.100710
Greer, E.L., Maures, T.J., Ucar, D., et al., Transgenerational epigenetic inheritance of longevity in Caenorhabditis elegans,Nature, 2011, vol. 479, no. 7373, pp. 365–371. https://doi.org/10.1038/nature10572
Harman, D., Aging: a theory based on free radical and radiation chemistry, J. Gerontol., 1956, vol. 11, pp. 298–300. https://doi.org/10.1093/geronj/11.3.298
Hayes, J.D. and McMahon, M., NRF2 and KEAP1 mutations: permanent activation of an adaptive response in cancer, Trends Biochem. Sci., 2009, vol. 34, no. 4, pp. 176–188. https://doi.org/10.1016/j.tibs.2008.12.008
Hegde, M.L., Mantha, A.K., Hazra, T.K., et al., Oxidative genome damage and its repair: implications in aging and neurodegenerative diseases, Mech. Ageing Dev., 2012, vol. 133, no. 4, pp. 157–168. https://doi.org/10.1016/j.mad.2012.01.005
Heyn, H., Li, N., Ferreira, H.J., et al., Distinct DNA methylomes of newborns and centenarians, Proc. Natl. Acad. Sci. U.S.A., 2012, vol. 109, no. 26, pp. 10522–10527. https://doi.org/10.1073/pnas.1120658109
Heyn, H., Moran, S., and Esteller, M., Aberrant DNA methylation profiles in the premature aging disorders Hutchinson Gilford Progeria and Werner syndrome, Epigenetics, 2013, vol. 8, no. 1, pp. 28–33. https://doi.org/10.4161/epi.23366
Holliday, R., The inheritance of epigenetic defects, Science, 1987, vol. 238, no. 4824, pp. 163–170. PMID 3310230
Horvath, S., DNA methylation age of human tissues and cell types, Genome Biol., 2013. 1456. https://doi.org/10.1186/gb-2013-14-10-r115
Hwang, J., Yao, H., Caito, S., et al., Redox regulation of SIRT1 in inflammation and cellular senescence, Free Radical Biol. Med., 2013, vol. 61, pp. 95–110.https://doi.org/10.1016/j.freeradbiomed.2013.03.015
Ibáñez-Ventoso, C., Ibáñez-Ventoso, C., Yang, M., et al., Modulated microRNA expression during adult lifespan in Caenorhabditis elegans,Aging Cell, 2006, vol. 5, no. 3, pp. 235–246. https://doi.org/10.1111/j.1474-9726.2006.00210.x
Jurkowska, R.Z., Jurkowski, T.P., and Jeltsch, A., Structure and function of mammalian DNA methyltransferases, ChemBioChem, 2011, vol. 12, no. 2, pp. 206–222. https://doi.org/10.1002/cbic.201000195
Koltover, V.K., Theory of reliability in systems biology: aging versus reliability, in Recent Advances in Systems Biology Research, New York: Nova Science, 2014, pp. 109–130.
Lee, J.T., Davidow, L.S., and Warshawsky, D., Tsix, a gene antisense to Xist at the X-inactivation centre, Nat. Genet., 1999, vol. 21, no. 4, pp. 400–404. https://doi.org/10.1038/7734
Lewandowska, J. and Bartoszek, A., DNA methylation in cancer development, diagnosis and therapy-multiple opportunities for genotoxic agents to act as methylome disruptors or remediators, Mutagenesis, 2011, vol. 26, no. 4, pp. 475–487. https://doi.org/10.1093/mutage/ger019
Lόpez-Otίn, C., Blasco, M.A., Partridge, L., et al., The hall marks of aging, Cell, 2013, vol. 153, no. 6, pp. 1194–1217. https://doi.org/10.1016/j.cell.2013.05.039
Maynard, S., Schurman, S.H., Harboe, C., et al., Base excision repair of oxidative DNA damage and association with cancer and aging, Carcinogenesis, 2008, vol. 30, no. 1, pp. 2–10.https://doi.org/10.1093/carcin/bgn250
McCauley, B.S. and Dang, W., Histone methylation and aging: lessons learned from model systems, Biochim. Biophys. Acta, Gene Regul. Mech., 2014, vol. 1839, no. 12, pp. 1454–1462. https://doi.org/10.1016/j.bbagrm.2014.05.008
Mecocci, P., MacGarvey, U., Kaufman, A.E., et al., Oxidative damage to mitochondrial DNA shows marked age-dependent increases in human brain, Ann. Neurol., 1993, vol. 34, no. 4, pp. 609–616. https://doi.org/10.1002/ana.410340416
Miquel, J., Economos, A.C., Fleming, J., and Johnson, J.E., Mitochondrial role in cell aging, Exp. Gerontol., 1980, vol. 15, no. 6, pp. 575–591.
Mitsuishi, Y., Taguchi, K., Kawatani, Y., et al., Nrf2 redirects glucose and glutamine into anabolic pathways in metabolic reprogramming, Cancer Cell, 2012, vol. 22, no. 1, pp. 66–79. https://doi.org/10.1016/j.ccr.2012.05.016
Murphy, M.P., How mitochondria produce reactive oxygen species, Biochem. J., 2009, vol. 417, no. 1, pp. 1–13. https://doi.org/10.1042/BJ20081386
Murphy, T.M., Perry, A.S., and Lawler, M., The emergence of DNA methylation as a key modulator of aberrant cell death in prostate cancer, Endocrinol. Relat. Cancer, 2008, vol. 15, no. 1, pp. 11–25.https://doi.org/10.1677/ERC-07-0208
Newsholme, P., Rebelato, E., Abdulkader, F., et al., Reactive oxygen and nitrogen species generation, antioxidant defenses, and β-cell function: a critical role for amino acids, J. Endocrinol., 2012, vol. 214, no. 1, pp. 11–20. https://doi.org/10.1530/JOE-12-0072
Noren Hooten, N., Abdelmohsen, K., Gorospe, M., et al., MicroRNA expression patterns reveal differential expression of target genes with age, PLoS One, 2010, vol. 5, no. 5, art. ID e10724. https://doi.org/10.1371/journal.pone.0010724
Oberdoerffer, P., Michan, S., McVay, M., et al., SIRT1 redistribution on chromatin promotes genomic stability but alters gene expression during aging, Cell, 2008, vol. 135, no. 5, pp. 907–918. https://doi.org/10.1016/j.cell.2008.10.025
Oda, M., Oxley, D., Dean, W., and Reik, W., Regulation of lineage specific DNA hypomethylation in mouse trophectoderm, PLoS One, 2013, vol. 8, no. 6, art. ID e68846. https://doi.org/10.1371/journal.pone.0068846
Pal, S. and Tyler, J.K., Epigenetics and aging, Sci. Adv., 2016, vol. 2, no. 7. e1600584. https://doi.org/10.1126/sciadv.1600584
Paniker, N.V., Srivastava, S.K., and Beutler, E., Glutathione of the red cells. Effect of glutathione reductase deficiency on the stimulation of hexose monophosphate shunt under oxidative stress, Biochim. Biophys. Acta, Gen. Subj., 1970, vol. 215, no. 3, pp. 456–460.
Richter, C., Park, J.W., and Ames, B.N., Normal oxidative damage to mitochondrial and nuclear DNA is extensive, Proc. Natl. Acad. Sci. U.S.A., 1988, vol. 85, no. 17, pp. 6465–6467. PMID 3413108
Salpea, P., Russanova, V.R., Hirai, T.H., et al., Postnatal development-and age-related changes in DNA-methylation patterns in the human genome, Nucleic Acids Res., 2012, vol. 40, no. 14, pp. 6477–6494. https://doi.org/10.1093/nar/gks312
Santos, J.H., Mandavilli, B.S., and Houten van, B., Mitochondrial DNA, Methods Mol. Biol., 2002, vol. 197, pp. 159–176. https://doi.org/10.1385/1-59259-284-8:159
Schafer, Z.T., Grassian, A.R., Song, L., et al., Antioxidant and oncogene rescue of metabolic defects caused by loss of matrix attachment, Nature, 2009, vol. 461, no. 7260, pp. 109–113. https://doi.org/10.1038/nature08268
Sen, P., Shah, P.P., Nativio, R., and Berger, S.L., Epigenetic mechanisms of longevity and aging, Cell, 2016, vol. 166, no. 4, pp. 822–839. https://doi.org/10.1016/j.cell.2016.07.050
Sharma, S., Kelly, T.K., and Jones, P.A., Epigenetics in cancer, Carcinogenesis, 2010, vol. 31, no. 1, pp. 27–36. https://doi.org/10.1093/carcin/bgp220
Shumaker, D.K., Dechat, T., Kohlmaier, A., et al., Mutant nuclear lamin A leads to progressive alterations of epigenetic control in premature aging, Proc. Natl. Acad. Sci. U.S.A., 2006, vol. 103, no. 23, pp. 8703–8708. https://doi.org/10.1073/pnas.0602569103
Simpson, N.E., Tryndyak, V.P., Pogribna, M., et al., Modifying metabolically sensitive histone marks by inhibiting glutamine metabolism affects gene expression and alters cancer cell phenotype, Epigenetics, 2012, vol. 7, no. 12, pp. 1413–1420. https://doi.org/10.4161/epi.22713
Sundar, I.K., Yao, H., and Rahman, I., Oxidative stress and chromatin remodeling in chronic obstructive pulmonary disease and smoking-related diseases, Antioxid. Redox Signaling, 2013, vol. 18, no. 15, pp. 1956–1971. https://doi.org/10.1089/ars.2012.4863
Skulachev, V.P., SkQ1 treatment and food restriction-two ways to retard an aging program of organisms, Aging (N.Y.), 2011, vol. 3, no. 11, pp. 1045–1050. https://doi.org/10.18632/aging.100410
Viswanathan, W. and Guarente, L., Regulation of Caenorhabditis elegans lifespan by sir-2.1 transgenes, Nature, 2011, vol. 477, no. 7365, pp. E1–E2. https://doi.org/10.1038/nature10440
Waddington, C.H., Preliminary notes on the development of the wings in normal and mutant strains of Drosophila,Proc. Natl. Acad. Sci. U.S.A., 1939, vol. 25, no. 7, pp. 299–307. PMCID PMC1077909
Weinberg, F., Hamanaka, R., Wheaton, W.W., et al., Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity, Proc. Natl. Acad. Sci. U.S.A., 2010, vol. 107, no. 19, pp. 8788–8793. https://doi.org/10.1073/pnas.1003428107
Wilson, A.S., Power, B.E., and Molloy, P.L., DNA hypomethylation and human diseases, Biochim. Biophys. Acta,Rev. Cancer, 2007, vol. 1775, no. 1, pp. 138–162. https://doi.org/10.1016/j.bbcan.2006.08.007
Wu, H. and Zhang, Y., Reversing DNA methylation: mechanisms, genomics, and biological functions, Cell, 2014, vol. 156, nos. 1–2, pp. 45–68. https://doi.org/10.1016/j.cell.2013.12.019
Yamaguchi, R. and Perkins, G., Dynamics of mitochondrial structure during apoptosis and the enigma of Opa1, Biochim. Biophys. Acta,Bioenerg., 2009, vol. 1787, no. 8, pp. 963–972. https://doi.org/10.1016/j.bbabio.2009.02.005
Yao, H. and Rahman, I., Role of histone deacetylase 2 in epigenetics and cellular senescence: implications in lung inflammaging and COPD, Am. J. Physiol. Lung Cell. Mol. Physiol., 2012, vol. 303, no. 7, pp. L557–L566. https://doi.org/10.1152/ajplung.00175.2012
Yeo, E.-J. and Park, S.C., Age-dependent agonist-specific dysregulation of membrane-mediated signal transduction: emergence of the gate theory of aging, Mech. Ageing Dev., 2002, vol. 123, no. 12, pp. 1563–1578.
Young, T.W., Mei, F.C., Yang, G., et al., Activation of antioxidant pathways in ras-mediated oncogenic transformation of human surface ovarian epithelial cells revealed by functional proteomics and mass spectrometry, Cancer Res., 2004, vol. 64, no. 13, pp. 4577–4584. https://doi.org/10.1158/0008-5472.CAN-04-0222
Yu, C.E., Oshima, J., Fu Y.H., et al., Positional cloning of the Werner’s syndrome gene, Science, 1996, vol. 272, no. 5259, pp. 258–262. PMID 8602509
Zhang, W., Li, J., Suzuki, K., et al., Aging stem cells. A Werner syndrome stem cell model unveils heterochromatin alterations as a driver of human aging, Science, 2015, vol. 348, no. 6239, pp. 1160–1163. https://doi.org/10.1126/science.aaa1356
Zhao, J., Sun, B.K., Erwin, J.A., et al., Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome, Science, 2008, vol. 322, no. 5902, pp. 750–756. https://doi.org/10.1126/science.1163045
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by V. Mittova
This article has been retracted. Please see the retraction notice for more detail: https://doi.org/10.1134/S2079057024010016"
About this article
Cite this article
Aitbaev, K.A., Murkamilov, I.T. & Fomin, V.V. RETRACTED ARTICLE: Molecular Mechanisms of Aging: The Role of Oxidative Stress and Epigenetic Modifications. Adv Gerontol 9, 417–425 (2019). https://doi.org/10.1134/S2079057019040027
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S2079057019040027