Abstract—
Microbiomes of strongly skeletal residual calcareous pelozems (Skeletal Leptosols (Loamic)), carbopetrozems (Calcaric Leptosols (Protic)), petrozems (Skeletal Leptosols (Protic)), and cryozems (Oxyaquic Cryosols (Loamic)) in the north of Novaya Zemlya were studied by the methods of molecular biology. The number of copies of 16S rRNA genes was small and ranged from 2.30 × 107 to 1.63 × 109 gene copies/g soil for archaea and from 3.47 × 108 to 2.26 × 1011 gene copies/g soil for bacteria; the number of copies of ribosomal genes ITS rRNA of fungi varied from 8.87 × 106 to 7.56 × 109 gene copies/g soil. The content of copies of ribosomal genes of all groups of microorganisms sharply decreased down the soil profiles. Bacteria predominated among prokaryotes (up to 90%). The greatest abundance (20%) was manifested by the phyla Proteobacteria, Actinobacteria, and Acidobacteria; the abundances of Bacteroidetes, Firmicutes, Verrucomicrobia, Gemmatimonadetes, and Chloroflexi were about 1–10%. The Archaea domain represented mainly by the Ferroplasma genus (phylum Euryarchaeota), accounted for ≤4% of prokaryotes. The taxonomic diversity of prokaryotes increased down the soil profiles and reached maximum values in the suprapermafrost horizons, where the number of candidate phyla typical of marine ecosystems—Latescibacteria, Tectomicrobia, Parcubacteria, Saccaribacteria, Hydrogenedentes, Peregrinibacteria, Ignavibacteria, and Gracilibacteria—was high.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
INTRODUCTION
Studies of polar regions are becoming increasingly relevant because of the extremely rapid warming in Arctic and Antarctic ecosystems compared to other territories of the Earth. From the end of the 20th century, air temperature beyond the Arctic Circle has increased by an average of 2°C, and for the rest of the Earth, by 0.8°C [32]. In this regard, the phenomenon of “Arctic greening” is increasingly being noted, when lifeless landscapes of barrens are rapidly covered with tundra vegetation [41]. Apparently, this phenomenon leads to an increase in the biological activity of soils, greenhouse gas emissions into the atmosphere [16], and microbial diversity [31]. Soil microorganisms play the key role in the biogeochemical cycles nutrient provision to the ecosystem as a whole [25]. The study of soil microbiome in the polar regions makes it possible to assess the degree of their tolerance toward climate change and ecosystem productivity [20, 36]. Information about the structure of the microbiome allows us to draw conclusions about the specificity of soil formation and the zonality of the soil cover as a whole [9, 42]. The composition and activity of soil microbiome is a highly sensitive indicator of the state of ecosystems in the conditions of the climate warming [16].
Novaya Zemlya (NZ) is the largest, but also the least studied archipelago of the European Arctic due to the nuclear weapons tests on its territory in the middle of the 20th century [10]. Novaya Zemlya is one of the white spots for the biogeography of microorganisms and soil biota in general [23]. Scientific research of this archipelago was greatly facilitated by the Arctic Floating University project, which provided opportunity to collect soil samples considered in this paper. Only two articles on the taxonomic diversity of microorganisms on the Severnyi (Northern) Island of NZ have been published so far [10, 15]. One of them is devoted to soil microbiome [10]. The information presented in it was obtained using the inoculation method, which could only detect about 1–2% of all microorganisms [38]. For the most complete quantitative and taxonomic characterization of the soil microbiome, these data need to be supplemented with the results of molecular biological testing [12].
The aim of this paper is to present quantitative and qualitative taxonomic characterization of soil microbiomes in the northern part of NZ using molecular biological methods.
OBJECTS AND METHODS
The research was carried out in the northern part of the Severnyi (Northern) Island of NZ in the areas of the Russkaya Gavan’ (harbor), Ledyanaya Gavan’, and Blagopoluhiya bays and Cape Zhelaniya. The conditions of collection and storage of samples, as well as a detailed description of vegetation and soil pits and their photos were presented earlier [9]. The coordinates of the key points, the classification position of the soils, and data on some physicochemical properties of the studied samples are presented in Table S1.
Quantitative assessment of the content of ribosomal genes of microorganisms was carried out by the quantitative real-time polymerase chain reaction (PCR). Primers for the 16S rRNA gene were used to account for archaea and bacteria, and primers for the ITS region were used to account for fungi. The reaction was carried out in a Real-Time CFX96 Touch (Bio-Rad) amplifier. The reaction mixture was prepared from the preparation SuperMix Eva Green (Bio-Rad). Solutions of cloned fragments of ribosomal operon of Escherichia coli strain K12 were used as quantitative standards for the concentration of 16S rRNA genes for bacteria, and of strain FG-07 Halobacterium salinarum for archaea; for fungi, yeast strain Saccharomyces cerevisiae Meyen 1B-D1606 was applied. For each sample, the reaction was carried out in three replicates. The concentration of genes was calculated using CFX Manager software, recalculating the number of genes in DNA preparations per 1 g of dry soil, taking into account dilutions and weight of the samples.
The taxonomic structure of the prokaryotic community was determined by the Next Generation Sequencing (NGS) using the Illumina MiSeq platform by pair–terminal reading (2 × 300 base pairs) via generating no less than 10 000 paired reads per sample from the sequences of the V3–V4 16S rRNA hypervariable region gene. The samples were prepared by two–stage PCR: V3–V4 amplification of 16S rRNA, and then PCR amplification of the product for the purpose of barcoding the library. The amplicons obtained after purification on magnetic particles and concentration measurement by the fluorimetric method were ready-made DNA libraries.
The processing of sequencing data was carried out using the automated QIIME algorithm [17], which included forward and reverse reads, removal of technical sequences, filtering of sequences with low reliability of individual nucleotides (quality <Q20), and filtering of chimeric sequences. To subdivide sequences into operational taxonomic units (OTUs), an algorithm with an open reference classification threshold of 97% was used. The alignment of readings on the 16S rRNA sequence and the distribution of sequences by taxonomic units was carried out using the Silva database, version 132 [33].
The diversity and similarity of bacterial communities of the studied substrates were evaluated using α‑diversity indices calculated when combining sequences in OTUs with the 97% similarity of the nucleotide composition of sequences. The following indices were used: the Shannon diversity index (H = Σpi log2pi, where pi is the share of the ith species in the community) and the Pielou’s evenness index, which is the normalization of the Shannon index between 0 and 1.
RESULTS
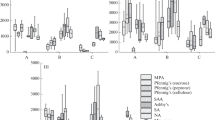
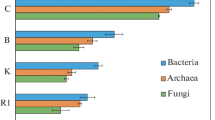
Quantitative assessment of ribosomal gene content by real-time PCR. The number of copies of ribosomal genes of 16S rRNA of archaea is small and varies from 2.30 × 107 to 1.63 × 109 gene copies/g soil. The minimum of archaea was detected in the Rca mineral horizon of the residual-calcareous strongly skeletal petrozem on Cape Zhelaniya (profile CJ-24-25), and the maximum was found in the surface OT-W horizon of the mucky-humus residual calcareous strongly skeletal pelozem studied in the Ledyanaya Gavan’ (profile LG-15-17) (Fig. 1). For most of the studied profiles, the abundance of archaea sharply decreased from the surface to the lower horizons in the profile. However, in the raw-humus residual calcareous cryozem of the Blagopoluchiya Bay area (profile BB-9-10), the cryoturbated residual calcareous strongly skeletal protohumus pelozem of the Ledyanaya Gavan’ area (profile LG-12-14), and the gleyic skeletal cryozem of the Russkaya Gavan’ area (profile RG-1-3), the proportion of archaea was maximal in the subsurface (4–11 cm) CR horizon. Among all the analyzed NZ locations, the smallest number of copies of archaea genes was specific for Cape Zhelaniya, and the largest number was for the Ledyanaya Gavan’ area.
The number of copies of the 16S rRNA genes of bacteria varied from 3.47 × 109 to 2.26 × 1011 genes copies/g soil. The minimum values were found in the Rca mineral horizon of the residual-calcareous strongly skeletal petrozem at Cape Zhelaniya (profile CJ-24-25), and the maximum values were found in the upper Wca horizon of the residual calcareous strongly skeletal pelozem (profile LG-12-14) of the Ledyanaya Gavan’ area. For most of the studied profiles, the amount of bacterial genetic material decreased from the surface horizons to the suprapermafrost horizon. However, in the raw-humus residual calcareous cryozem of the Blagopoluchiya Bay (profile BB-9-10), the abundance of bacteria was maximal in the lower CR mineral horizon (10–19 cm). Among all the analyzed locations, the smallest number of copies of bacterial genes was characteristic of Cape Zhelaniya and the largest number was in the soils of the Ledyanaya Gavan’ area.
The number of ribosomal genes ITS rRNA of fungi varied from 8.87 × 106 to 7.56 × 109 gene copies/g soil (Fig. 2). The lowest values were found in the Rca mineral horizon of the residual-calcareous strongly skeletal petrozem at Cape Zhelaniya (profile CJ-24-25), and the maximum values were found in the upper moss layer of the gleyic skeletal cryozem in the Russkaya Gavan’ area (profile RG-1-3). For most of the analyzed soils, the number of ribosomal genes ITS rRNA of fungi was 108–109 gene copies/g soil. Ribosomal genes of fungi were not detected in the petrozem of the Ledyanaya Gavan’ area (profile RG-4), the surface OT–W horizon of the residual calcareous pelozem of the Ledyanaya Gavan’ area (profile LG-15-17), and the surface protohumus WC horizon of the residual calcareous strongly skeletal cryozem at Cape Zhelaniya (profile CJ-24-25). For almost all of the profiles, the amount of genetic material of fungi decreased down the profile. However, in the raw-humus residual calcareous cryozem of the Blagopoluchiya Bay area (profile BB-9-10) and the cryoturbated residual calcareous strongly skeletal pelozem of the Ledyanaya Gavan’ area (profile LG-12-14), the proportion of mycobiota was maximal in the subsurface horizon. Among the analyzed locations, the smallest number of copies of archaea genes was found in the Ledyanaya Gavan’ area, and the maximum was found in the Russkaya Gavan’ area.
As well as for archaea, bacteria, and fungi, the minimum number of copies of ribosomal genes was noted in the Rca mineral horizon of the residual-calcareous strongly skeletal petrozem at Cape Zhelaniya (profile CJ-24-25). In the raw-humus residual calcareous cryozem of the Blagopoluchiya Bay (profile BB-9-10), all the groups of microorganisms have maximum abundance in the subsurface horizon.
The taxonomic structure of the prokaryotic community was determined by high-throughput sequencing (barcoding of the 16SrRNA gene) for two profiles from the Ledyanaya Gavan’ (LG-12-14 and LG-15-16). The number of detected phyla of prokaryotes increased from the surface organic to deep mineral horizons. The phyla of Proteobacteria, Actinobacteria, and Acidobacteria dominated (Fig. 3, Table 1). The contents of phyla of Bacteroidetes, Firmicutes, Verrucomicrobia, Gemmatimonadetes and Chloroflexi was lower. Minor components of the prokaryotic community in all studied samples (<1% abundance) were represented by Planctomycetes, Chlorobia, Nitrospirae, Cyanobacteria, and Gemmatimonadetes. Sample LG-13 with a domination of Nitrospirae (95%), low content of Firmicutes (4%), and the numbers of Actinobacteria, Acidobacteria, and Bacteroidetes less than 1% was an exception.
Taxonomic structure of the bacterial community (% of the total number of sequences): (1) Actinobacteria, (2) Proteobacteria, (3) Firmicutes, (4) Verrucomicrobia, (5) Chloroflexi, (6) Acidobacteria, (7) Chlorobia, (8) Bacteroidetes, (9) Nitrospirae, (10) Gemmatimonadetes, (11) Cyanobacteria, and (12) other.
Alphaproteobacteria predominated among the phylum of Proteobacteria in most of the samples. There were fewer representatives of the Betaproteobacteria class, which was present mainly in mineral horizons. Representatives of the Deltaproteobacteria class were only found in organic horizons.
Cyanobacteria, whose content in the studied samples did not exceed 0.1–0.2%, were mainly represented by the Melainabacteria group.
The maximum diversity (up to 14 phyla) of bacterial candidate phyla (Latescibacteria, Tectomicrobia, Parcubacteria, Saccaribacteria, Hydrogenedentes, Peregrinibacteria, Ignavibacteria, Gracilibacteria, TM6, BRC1, GAL15, WWE3, WS2, SR1) was found in samples LG-14 and LG-16 of the mucky-humus residual calcareous pelozem.
The content of archaea in the studied soils was insignificant, varying from 0.1 to 3.8% of the number of all prokaryotes, except for the LG-13 sample, where their content reached 36% (Fig. 4). Three phyla were detected among archaea: Euryarchaeota, Thaumarchaeota, and Woesearchaeota. Moreover, representatives of the Euryarchaeota phylum were most often found, the content of which ranged from 60 to 100% (Fig. 5). The Euryarchaeota phylum was mainly represented by the genus Ferroplasma.
DISCUSSION
Real-time PCR Quantitative assessment of ribosomal gene content. The number of copies of ribosomal genes 16S rRNA of bacteria in the studied soils is in agreement with available data on peat soils of the Bolshezemel’skaya tundra [42] and tundra soils of Alaska [27, 39]. This confirms numerous evidences of the increased tolerance of bacteria to extremely low temperatures and oligotrophic conditions of the polar regions [4, 14, 29]. At the same time, the number of copies of ribosomal genes 16S rRNA of archaea in the analyzed samples is two orders of magnitude lower in comparison with that in other Arctic territories [11, 39, 42]. Representatives of this domain are usually characterized by adaptation only to specific environmental conditions and are extremely sensitive to their changes [21], which, apparently, puts archaea in the minor positions of the soil microbiome of the northern Novaya Zemlya.
A relatively low (by an order of magnitude lower) content of copies of the ribosomal genes ITS rDNA of fungi seems strange compared to the results obtained on more northern territories of the Franz Josef Land archipelago [11]. Probably, this can be explained by the locally high content of organic matter in some soils of Franz Josef Land [7, 11] as compared with NZ soils.
The maximum abundance of copies of ribosomal genes of all groups of microorganisms seems to be natural in the surface organic horizons, since they contain the maximum amount of organic carbon and total nitrogen. The exceptions are the profiles with the maximum of copies of ribosomal genes of microorganisms in subsurface and suprapermafrost horizons. This phenomenon can be explained by the avoidance of negative abiotic factors (extreme temperatures, increased levels of ultraviolet radiation, strong winds, etc.) by the biota, as well as by a relatively low content of organic matter even in the surface horizon. The high Arctic and Antarctic regions are characterized by a similar effect of accumulation of microorganisms in the subsurface layers of ahumic soils, desert pavements, and regoliths [4–6].
The number of copies of the 16S rRNA genes of bacteria is two orders of magnitude higher than that of archaea and fungi. This ratio is typical for soil microorganisms of various natural zones [3, 8]. The low number of archaea may be associated with the increasing competition of representatives of this domain with bacteria in conditions of limited food and energy resources [12] characteristic of polar ecosystems. A small number of fungi in the analyzed soils is apparently due to the predominance of dormant forms (spores, conidia, etc.), the content of genetic material in which is less than in mycelium cells [22].
The results obtained in this study on the number of copies of ribosomal bacterial genes correlate with the previously identified indicators of the number of prokaryotic cells in the studied soil profiles of the north of NZ [9]. This fact confirms the hypothesis about the comparability of the results for the characteristics of the soil microbiome obtained using quantitative real-time PCR and luminescent microscopy [1, 2].
Taxonomic structure of the prokaryotic community. In the analyzed samples, Proteobacteria and Actinobacteria are most abundant, their content reaches 30–40%. Acidobacteria, Verrucomicrobia, Bacteroidetes, Chloroflexi, Gemmatimonadetes, Firmicutes, and Nitrospirae are present in lower amounts (no more than 10–14%). Planctomycetes, Fibrobacters, Fusobacteres, Armatimonadetes, and Cyanobacteria are present in minimal amounts (<1%). According to molecular genetic studies, soil bacterial community is mainly composed of nine phyla: Proteobacteria, Acidobacteria, Actinobacteria, Verrucomicrobia, Bacteroidetes, Chloroflexi, Planctomycetes, Gemmatimonadetes, and Firmicutes [24, 26]; the content of representatives of other phyla, as a rule, does not exceed tenths of a percent [30].
Attention is drawn to the taxonomic composition of the prokaryotic community of sample LG13 (the Cca horizon of the residual calcareous pelozem), where the phylum Nitrospirae predominates among bacteria, and Euryarchaeota (represented by the only one Ferroplasma genus) predominate among archaea. It can be assumed that iron transformation processes are active in the studied locus, as indicated by the high content of the genus Ferroplasma [18]. Also, we can suppose the activity of nitrification processes being carried out by bacteria of the phylum Nitrospirae [28].
The archaea of the Euryarchaeota and Taumarchaeota phyla are most often found in the residual calcareous pelozem, and the uncultivated Woesarchaeota phylum has been found in samples LG-15 and LG-16. Prokaryotes are present in the horizons, where the processes of iron transformation (ocherous and gleyed mottles) are visually detected: representatives of the Archaea domain, the Euryarchaeota phylum (genus Ferroplasma). It is known that representatives of the genus Ferroplasma of the phylum Euryarchaeota are chemolithoautotrophs that oxidize Fe2+ to Fe3+ ions to obtain energy [18].
The detection of the phylum of bacteria Nitrospirae and the phylum of archaea Taumarchaeota in the studied samples of pelozems suggests the possibility of participation of these organisms in the transformation of nitrogen compounds. Nitrospira are active participants in the nitrification of nitrogen [28], and thaumarchaeotes (anammox bacteria) participate in anaerobic oxidation of ammonium [37].
The maximum diversity of prokaryotic phyla was revealed in the suprapermafrost mineral horizons, where, in addition to other phyla specific for the soil, a significant number (from 9 to 14) of candidate phyla were found: Latescibacteria, Tectomicrobia, Parcubacteria, Saccaribacteria, Hydrogenedentes, Peregrinibacteria, Ignavibacteria, and Gracilibacteria. Interestingly, many of them were previously found in marine ecosystems [19, 34, 35]. Apparently, the presence of these phyla is associated with the proximity of the analyzed soil profiles to the sea.
An important ecological characteristic of any community is the abundance of taxa, or α-diversity of the community. For the obtained data on the representation of OTUs at the level of similarity of 97%, the Shannon and Pielou indices were calculated (Table 2). The highest indicators of taxonomic richness were found in the moist and relatively rich in organic carbon and nitrogen horizons of the surface moss, where the value of the Shannon index was 8.62 (LG-15). Somewhat lower values of the Shannon index were in the moss (LG-12) and mineral horizons (LG-14 and LG-16). The minimum values of the Shannon index and minimum number of OTUs were noted in the LG-13 mineral horizon: 1.85 and 32, respectively. The number of detected OTUs depended on the same factors as the Shannon index: the highest values were found in the wetter horizons. The values of the Pielou evenness index attest to the approximately the same and high degree of evenness of the microbial communities in the studied soils.
The data obtained by analyzing the sequencing results show a high degree of diversity of prokaryotic communities in the studied soils, which is consistent with the literature data [15]. It is possible that soil moisture is an important factor affecting the taxonomic diversity of the community. There is a positive correlation of the content of the Proteobacteria phylum in the community with an increase in the soil water content, which is also consistent with the data of other researchers [40].
CONCLUSIONS
A molecular biological study of soil microbiomes in the northern part of the NZ archipelago has been performed for the first time.
The microbiome of the studied territories is dominated by bacteria of the Proteobacteria, Actinobacteria, and Acidobacteria phyla, as well as by archaea of the Euryarchaeota phylum. The number of copies of ribosomal genes of all microorganisms decreases, and the taxonomic diversity of prokaryotes increases down the soil profiles in the north of NZ.
According to the data obtained by quantitative real-time PCR, the microbiome of the studied soils is dominated by bacteria, not archaea or fungi. This contradicts the results obtained earlier by luminescent microscopy [9], according to which fungi prevail in the soils of the northern part of NZ. Apparently, this discrepancy is explained by the predominance of dormant forms (spores, conidia, etc.), the content of genetic material in which is lower than in mycelial cells. In order to confirm or refute the fact of the dominance of one of the groups of microorganisms in the studied objects, additional studies by other methods are necessary (for example, by the method of substrate-induced respiration with specific inhibition of fungi by antibiotics/bacteria or biomarker method for determining the content of fatty acids of phospholipids).
Despite the harsh climatic conditions of the Arctic, the NZ soils perform the ecological function of preserving prokaryotic diversity, which is expressed both in the presence of dominant taxa and in a wide variety of candidate phyla, some of which are associated with marine habitats.
REFERENCES
E. V. Blagodatskaya, M. V. Semenov, and A. V. Yakushev, Activity and Biomass of Soil Microorganisms in a Changing Environment (KMK Press, Moscow, 2016) [in Russian].
M. V. Korneykova, D. A. Nikitin, A. V. Dolgikh, and A. S. Soshina, “Soil mycobiota of the Apatity city, Murmansk region,” Mikologiya Fitopatol. 54 (4), 264–277 (2020). https://doi.org/10.31857/S0026364820040078
M. V. Korneykova and D. A. Nikitin, “Qualitative and quantitative characteristics of the soil microbiome in the impact zone of the Kandalaksha aluminum smelter,” Eurasian Soil Sci. 54 (6), 897–906 (2021). https://doi.org/10.1134/S1064229321060089
L. V. Lysak, I. A. Maksimova, D. A. Nikitin, A. E. Ivanova, A. G. Kudinova, V. S. Soina, and O. E. Marfenina, “Microbial communities of soils in East Antarctica,” Moscow Univ. Bull. Ser. 16: Biol. 73 (3), 132–140 (2018).
O. E. Marfenina, D. A. Nikitin, and A. E. Ivanova, “The structure of fungal biomass and diversity of cultivated micromycetes in Antarctic soils (Progress and Russkaya stations),” Eurasian Soil Sci. 49 (8), 934–941. https://doi.org/10.1134/S106422931608007X
D. A. Nikitin, O. E. Marfenina, and I. A. Maksimova, “The use of the succession approach in studying the species composition of microscopic fungi and the content of fungal biomass in Antarctic soils,” Mikologiya Fitopatol. 51 (4), 211–219 (2017).
D. A. Nikitin, M. V. Semenov, A. A. Semikolennykh, I. A. Maksimova, A. V. Kachalkin, and A. E. Ivanova, “Fungal biomass and species diversity of the cultivated mycobiota of soils and substrates of Northbrook, Franz Josef Land,” Mikologiya Fitopatol. 53 (4), 210–222 (2019). https://doi.org/10.1134/S002636481904010X
D. A. Nikitin, E. A. Ivanova, A. D. Zhelezova, M. V. Semenov, R. G. Gadzhiumarov, A. K. Tkhakakhova, T. I. Chernov, N. A. Ksenofontova, and O. V. Kutovaya, “Assessment of the impact of no-till technology and plowing on the microbiome of southern agrochernozems,” Eurasian Soil Sci. 53 (12), 1782–1793 (2020). https://doi.org/10.1134/S106422932012008X
D. A. Nikitin, L. V. Lysak, D. V. Badmadashiev, S. S. Kholod, N. S. Mergelov, A. V. Dolgikh, and S. V. Goryachkin, “Biological activity of soils in the north of the Novaya Zemlya Archipelago: effect of the largest glacial sheet in Russia,” Eurasian Soil Sci. 54 (10), 1496–1516. https://doi.org/10.1134/S1064229321100082
D. A. Nikitin, L. V. Lysak, O. V. Kutovaya, and T. A. Gracheva, “Ecological-trophic structure and taxonomic characteristics of the communities of soil microorganisms in the northern part of the Novaya Zemlya Archipelago,” Eurasian Soil Sci. 54 (11), 1689-1704 (2021). https://doi.org/10.1134/S1064229321110107
D. A. Nikitin and M. V. Semenov, “Characterization of Franz Josef Land Soil Mycobiota by Microbiological Plating and Real-time PCR,” Microbiology (Moscow) 91 (1), 56–66 (2022). https://doi.org/10.1134/S002626172201009X
M. V. Semenov, “Metabarcoding and metagenomics in soil ecology research,” Zhurn. Obshch. Biol. 80 (6), 403–417 (2019). https://doi.org/10.1134/S004445961906006X
M. V. Semenov, N. A. Manucharova, and A. L. Stepanov, “Biomass and taxonomic structure of microbial communities in soils of the right-bank basin of the Oka River,” Eurasian Soil Sci. 52 (8), 971–981. https://doi.org/10.1134/S106422931908012X
P. Baldrian, “The known and the unknown in soil microbial ecology,” FEMS Microbiol. Ecol. 95 (2), fiz005 (2019). https://doi.org/10.1093/femsec/fiz005
A. A. Belov, V. S. Cheptsov, N. A. Manucharova, and Z. S. Ezhelev, “Bacterial communities of Novaya Zemlya Archipelago ice and permafrost,” Geosciences 10 (2), 67 (2020).https://doi.org/10.3390/geosciences10020067
J. E. Box, W. T. Colgan, T. R. Christensen, N. M. Schmidt, M. Lund, F. J. W. Parmentier, R. Brown, et al., “Key indicators of Arctic climate change: 1971–2017,” Environ. Res. Lett. 4 (4), 045010 (2019). https://doi.org/10.1088/1748-9326/aafc1b
J. G. Caporaso, J. Kuczynski, J. Stombaugh, K. Bittinger, F. D. Bushman, E. K. Costello, N. Fierer, et al., “QIIME allows analysis of high-throughput community sequencing data,” Nat. Methods 7 (5), 335–336 (2010). https://doi.org/10.1038/nmeth.f.303
M. Dopson, C. Baker-Austin, A. Hind, J. P. Bowman, P. L. Bond, “Characterization of Ferroplasma isolates and Ferroplasma acidarmanus sp. nov., extreme acidophiles from acid mine drainage and industrial bioleaching environments,” Appl. Environ. Microbiol. 70 (4), 2079–2088 (2004).
P. A. Figueroa-Gonzalez, T. L. Bornemann, P. S. Adam, J. Plewka, F. Revesz, C. Hagen, A. Tancsics, J. Probst, “Saccharibacteria as organic carbon sinks in hydrocarbon-fueled communities,” Front. Microbiol. (2020). https://doi.org/10.3389/fmicb.2020.587782
C. G. Flocco, W. P. Mac Cormack, and K. Smalla, “Antarctic soil microbial communities in a changing environment: their contributions to the sustainability of Antarctic ecosystems and the bioremediation of anthropogenic pollution,” in The Ecological Role of Microorganisms in the Antarctic Environment (Springer, Cham., 2019), pp. 133–161.https://doi.org/10.1007/978-3-030-02786-5_7
R. A. Garrett and H. P. Klenk, Archaea: Evolution, Physiology, and Molecular Biology (John Wiley & Sons, 2008).
F. O. Glöckner, P. Yilmaz, C. Quast, J. Gerken, A. Beccati, A. Ciuprina, G. Brunsa, P. Yarzac, J. Pepliesc, R. Westram, and W. Ludwig, “25 Years of serving the community with ribosomal RNA gene reference databases and tools,” J. Biotechnol. 261, 169–176 (2017). https://doi.org/10.1007/978-3-030-02786-5_7
C. A. Guerra, A. Heintz-Buschart, J. Sikorski, A. Chatzinotas, N. Guerrero-Ramirez, S. Cesarz, L. Beaumelle, et al., “Blind spots in global soil biodiversity and ecosystem function research,” Nature communic. 11 (1), 1–13 (2020). https://doi.org/10.1038/s41467-020-17688-2
C. M. Hansel, S. Fendorf, P. M. Jardine, and C. A. Francis, “Changes in bacterial and archaeal community structure and functional diversity along a geochemically variable soil profile,” Appl. Environ. Microbiol. 74, 1620–1633 (2008). https://doi.org/10.1128/AEM.01787-07
R. Jacoby, M. Peukert, A. Succurro, A. Koprivova, and S. Kopriva, “The role of soil microorganisms in plant mineral nutrition-current knowledge and future directions,” Frontiers Plant Sci. 8, 1617 (2017). https://doi.org/10.3389/fpls.2017.01617
P. H. Janssen, “Identifying the dominant soil bacterial taxa in libraries of 16S RRNA and 16S RRNA genes,” Appl. Environ. Microbiol. 72 (3), 1719–1728 (2006). https://doi.org/10.1128/AEM.72.3.1719-1728.2006
H. M. Kim, J. Y. Jung, E. Yergeau, C. Y. Hwang, L. Hinzman, S. Nam, S. G. Hong, O. Kim, J. Chun, and Y. K. Lee, “Bacterial community structure and soil properties of a subarctic tundra soil in council, Alaska,” FEMS Microbiol. Ecol. 89 (2), 465–475 (2014). https://doi.org/10.1111/1574-6941.12362
K. D. Kits, C. J. Sedlacek, E. V. Lebedeva, P. Han, A. Bulaev, P. Pjevac, A. Daebeler, et al., “Kinetic analysis of a complete nitrifier reveals an oligotrophic lifestyle,” Nature 549, 269–272 (2017). https://doi.org/10.1038/nature23679
L. A. Malard and D. A. Pearce, “Microbial diversity and biogeography in Arctic Soils,” Environ. Microbiol. Rep 10 (6), 611–625 (2018). https://doi.org/10.1111/1758-2229.12680
M. Podar, C. B. Abulencia, M. Walcher, D. Hutchison, K. Zengler, J. A. Garcia, T. Holland, D. Cotton, L. Hauser, and M. Keller, “Targeted access to the genomes of low-abundance organisms in complex microbial communities,” Appl. Environ. Microbiol. 10 (73), 3205–3214 (2007). https://doi.org/10.1128/AEM.02985-06
G. Pold, J. P. Schimel, and S. A. Sistla, “Soil bacterial communities vary more by season than with over two decades of experimental warming in Arctic tussock tundra,” Elementa: Science of the Anthropocene 9 (1) (2021). https://doi.org/10.1525/elementa.2021.00116
E. Post, R. B. Alley, T. R. Christensen, M. Macias-Fauria, B. C. Forbes, M. N. Gooseff et al., “Virginia and Muyin Wang. The polar regions in a 2°C warmer world,” Sci. Adv. 5 (12), 12 (2019). https://doi.org/10.1126/sciadv.aaw9883
E. Pruesse, C. Quast, K. Knittel, B. M. Fuchs, W. Ludwig, J. Peplies, and F. O. Glöckner, “SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB,” Nucleic Acid Res. 35 (21), 7188–7196 (2007). https://doi.org/10.1093/nar/gkm864
C. Rinke, P. Schwientek, A. Sczyrba, N. N. Ivanova, I. J. Anderson, J.-F. Cheng, A. Darling, et al., “Insights into the phylogeny and coding potential of microbial dark matter,” Nature 499 (7459), 431–437 (2013). https://doi.org/10.1038/nature12352
C. M. Sieber, A. J. Probst, A. Sharrar, B. C. Thomas, M. Hess, S. G. Tringe, and J. F. Banfield, “Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy,” Nature Microbiol. 3 (7), 836 (2018).
J. S. Singh and V. K. Gupta, “Soil microbial biomass: a key soil driver in management of ecosystem functioning,” Sci. Total Environ. 634, 497–500 (2018).
A. Spang, R. Hatzenpichler, C. Brochier-Armanet, and T. Rattei, “Distinct Gene Set in Two Different Lineages of Ammonia-Oxidizing Archaea Supports the Phylum Thaumarchaeota,” Trends in Microbiology 18 (8), 331–340 (2010). https://doi.org/10.1016/j.tim.2010.06.003
A. D. Steen, A. Crits-Christoph, P. Carini, K. M. DeAngelis, N. Fierer, K. G. Lloyd, and J. C. Thrash, “High proportions of bacteria and archaea across most biomes remain uncultured,” The ISME J. (2019). https://doi.org/10.1038/s41396-019-0484-y
B. M. Tripathi, H. M. Kim, J. Y. Jung, S. Nam, H. T. Ju, M. Kim, and Y. K. Lee, “Distinct taxonomic and functional profiles of the microbiome associated with different soil horizons of a moist tussock tundra in Alaska,” Frontiers in Microbiology 10, 1442 (2019). https://doi.org/10.3389/fmicb.2019.01442
E. Yergeau, K. K. Newsham, D. A. Pearce, and G. A. Kowalchuk, “Patterns of bacterial diversity across a range of Antarctic terrestrial habitats,” Environ. Microbiol. 9, 2670–2682 (2007). https://doi.org/10.1111/j.1462-2920.2007.01379.x
W. Zhang, P. A. Miller, C. Jansson, P. Samuelsson, J. Mao, B. Smith, “Self-amplifying feedbacks accelerate greening and warming of the Arctic,” Geophys. Rev. Lett. 45 (14), 7102–7111 (2018).
A. Zhelezova, T. Chernov, A. Tkhakakhova, N. Xenofontova, M. Semenov, O. Kutovaya, “Prokaryotic community shifts during soil formation on sands in the tundra zone,” Plos One 14 (4), e0206777 (2019). https://doi.org/10.1371/journal.pone.0206777
ACKNOWLEDGMENTS
The authors thank the Arctic Floating University project of the Lomonosov Northern Arctic Federal University and personally K.S. Zaikov for organizing field work on Novaya Zemlya. The authors thank the staff of the Department of Geography and Evolution of Soils of the Institute of Geography of the Russian Academy of Sciences and personally S.V. Goryachkin for their help in determining the taxonomic position of the studied soils.
Funding
This study was supported by the Russian Foundation for Basic Research, project no. 20-04-00328.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of interest.
Additional information
Translated by D. Konyushkov
SUPPLEMENTARY MATERIALS
Table 1. Soil properties in the northern part of Novaya Zemlya
Supplementary Information
Rights and permissions
About this article
Cite this article
Nikitin, D.A., Lysak, L.V. & Badmadashiev, D.V. Molecular Biological Characteristics of Soil Microbiome in the Northern Part of the Novaya Zemlya Archipelago. Eurasian Soil Sc. 55, 1106–1115 (2022). https://doi.org/10.1134/S1064229322080130
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1064229322080130