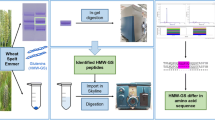
Abstract
This work was devoted to a study of the composition of protein concentrates from amaranth grain (Amaranthus hypochondriacus L.) of variety Voronezh. Amaranth protein concentrates were obtained by alkaline extraction of proteins and neutralization of the solution followed by ultrafiltration, by separating the starch fraction with amylolytic enzymes, and by alkaline extraction of proteins and their precipitation at pH 4.5. Conditions for the extraction of proteins followed by chromatography–mass spectrometric analysis and identification were selected. It was found that proteins from amaranth grain were more effectively extracted with a buffer with urea at protein concentrations of 1.7, 1.9, and 2.9 mg/cm3 in solution, respectively, while a buffer with detergents was more effective for the extraction of low-molecular-weight proteins at protein concentrations of 4.9, 2.9, and 9.0 mg/cm3 in solution, respectively. As a result of HPLC–MS/MS analysis followed by identification and search in the UNIPROT database, it was established that the main protein of amaranth grain is 11S-globulin, which acts as a reserve protein of amaranth seeds. In amaranth concentrates, 14 unique proteins belonging only to A. hipochondriacus L. were identified, and also proteins that did not belong to this species were reliably identified. Based on the results of semiquantitative analysis of the peptide profile of amaranth grain protein concentrates, a high frequency of occurrence of the main 11S-globulin proteins was established in all samples. The frequency of occurrence of other proteins in samples obtained by different methods differed significantly due to the peculiarities of protein isolation from amaranth grain. The results obtained can be used to prepare plant protein concentrates with a given protein composition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Plant proteins are widely used in the food industry and feed production. Amaranth is among promising protein sources as an alternative to proteins of plant and animal origins [1, 2]. This crop attracts attention due to its high level of adaptation, rapid growth of biomass, high protein content, and its balanced amino acid composition [3, 4]. The biological functions of amaranth proteins largely depend on the area of cultivation, climatic conditions, and variety [5]; protein structure [6]; and methods of protein isolation from the plant [7]. In the assessment of the biological properties of protein concentrates from amaranth grains, it is important to determine and identify the proteins included in their composition. It is well known that solubility decreases as a result of physicochemical actions on protein molecules in the course of isolation [8], and this fact complicates protein extraction for further analysis and identification.
The purpose of this work was to select conditions for protein extraction in the production of protein concentrates from amaranth grains of the Voronezh variety and the subsequent chromatography–mass spectrometric analysis and identification of the protein composition.
EXPERIMENTAL
The test material was amaranth grain of the Voronezh variety. Protein concentrates from the amaranth grain were obtained as follows: (1) by alkaline extraction of proteins and neutralization of the solution followed by ultrafiltration on a Vodopad UMTKp-1 unit with a pore size of 0.14 µm; (2) by separation of a starch fraction from the grain using amylolytic enzymes; and (3) by alkaline extraction of proteins and their precipitation at pH 4.5. The protein weight fractions in the obtained concentrates were 73, 55, and 83%, respectively.
Protein concentrates were dissolved in a phosphate buffer solution, and a lysis buffer solution (Buffer 1) (50 mM Tris, pH 8.0, 150 mM NaCl, 0.1% sodium dodecyl sulfate, 0.25% sodium deoxycholate, and 0.5% Nonidet P-40) with a protease inhibitor cocktail (Roche, Switzerland) was added. Then, they were extracted again with a buffer solution containing 8 M urea and 50 mM Tris (Buffer 2) for 30 min at room temperature with constant stirring and centrifuged at 10 000g, +4°C.
Protein concentrations were determined by the BCA method using a bicinchoninic acid kit (Thermo Fisher, the United States). The efficiency of protein extraction was evaluated by polyacrylamide gel electrophoresis (PAGE).
Hydrolysis of proteins in solution was carried out sequentially with enzyme preparations Lys C (Promega, the United States) 1/100 for 4 h and then with trypsin (Promega, the United States) 1/50 for 16 h.
An Agilent 1100 nanoflow chromatograph with a homemade C18-based column and with an electrospray emitter was used for HPLC–MS analysis. The column was prepared immediately before measurements.
Mass spectrometric analysis was carried out on an LTQ FT Ultra tandem mass spectrometer in a two-stage mode of automatic measurement of spectra [9, 10].
The results of HPLC–MS analysis were monitored using the Qual Browser software. Lists of the exact masses of peptides and the masses of their fragments were obtained from mass chromatograms using the Raw2msm application, and they were used to search and identify proteins in the database using the Peaks Studio software (Bioinformatics Solutions Inc., the United States, version 8.5). To identify peptides, the amino acid sequences of the proteins were used according to the Uniprot KB database.
RESULTS AND DISCUSSION
To select an optimal buffer solution for the dissolution and denaturation of proteins, we applied two most commonly used lysis buffers: Buffer 1 (detergent-based) and Buffer 2 (urea-based). For the subsequent mass spectrometric analysis and PAGE electrophoresis, the protein concentrations in samples weighing 10 mg were determined (Table 1).
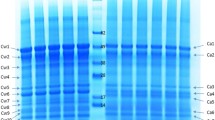
According to the results of polyacrylamide gel electrophoresis (Fig. 1), it was found that proteins were extracted more effectively with Buffer 2 (with urea); however, Buffer 1 (with detergents) was more effective for the extraction of low-molecular-weight proteins. Similarities were noted between samples 1', 2', and 3', and the protein content increased with the weight fraction of protein in the preparations. Because the buffer solutions differed in extraction ability, both fractions were used for further studies.
Electrophoresis of protein concentrates from amaranth grain in 12% polyacrylamide gel: (1) ultrafiltration concentrate (protein weight fraction, 73%), (2) protein concentrate (protein weight fraction, 55%), and (3) protein concentrate (protein weight fraction, 83%). Sequential extracts were applied to paired tracks: Buffer 1 and then Buffer 2 (').
As a result of hydrolysis of the protein content of protein concentrates from amaranth and HPLC–MS/MS analysis followed by identification and search in the UNIPROT database, we identified 188 proteins without taking into account species; Table 2 summarizes the main of these proteins. However, by selecting proteins that belong only to A. hipochondriacus L. and unique proteins that were not assigned to this species but were reliably identified, we obtained a list of identifications of 14 proteins (Table 2). The small number of identifications was due to the low completeness of the database not only for plants of this species but also for other types of amaranth.
As can be seen in Table 2, the main proteins of amaranth grain are 11S-globulin, which belongs to the main storage proteins of seeds, stress-protective heat shock protein (Hsp70), agglutinin, ribosomal protein, lipid transfer protein, and enzyme proteins.
Semiquantitative analysis of the hydrolysates of amaranth protein concentrates allowed us to reveal proteins and corresponding peptides that were most characteristic of samples obtained by different methods (Fig. 2). As can be seen in Fig. 2, the protein compositions of amaranth samples varied significantly. A high frequency of occurrence of the main storage proteins, 11S-globulins, was found in all samples. However, sample 2 differed from sample 3 in the presence of a larger amount of proteins such as heat shock protein (Hsp70) with a stress-protective effect, the enzymes glyceraldehyde-3-phosphate dehydrogenase, 5-methyltetrahydropteroyltriglutamate–homocysteine S-methyltransferase involved in the synthesis of methionine, malate dehydrogenase, enzyme related to type IV glycogenosis, etc. The small number of other identified proteins can be explained by the effect of aggressive media on the structure of proteins in the course of their isolation. A large amount of agglutinin proteins, ribosomal proteins, and nonspecific lipid transfer proteins was observed in the protein concentrate obtained after ultrafiltration (sample 3), while the amount of other proteins was minimal, probably, as a consequence of protein losses upon passage through the membrane. In the amaranth protein concentrate without chemical extraction of proteins (sample 1), a high frequency of occurrence was observed for all identified proteins with the exception of the proteins present in sample 3, which can be explained by the presence of strong bonds of these proteins with biopolymers of plant seeds and the impossibility of their extraction.
CONCLUSIONS
We selected conditions for the extraction of proteins from amaranth grains obtained by different isolation methods. Using chromatography–mass spectrometric analysis and identification of the main proteins, we found that the storage protein 11S-globulin was the main protein of all amaranth concentrates. A high frequency of occurrence of stress-protective heat shock proteins (Hsp70) and enzyme proteins was established in the amaranth protein concentrate obtained by alkaline extraction and protein precipitation at pH 4.5. The composition of the ultrafiltration concentrate was characterized by high concentrations of agglutinin proteins, ribosomal proteins, and nonspecific lipid transport proteins and a minimal amount of enzyme proteins. The largest number of identified proteins was found in the amaranth protein concentrate obtained after enzymatic removal of starch. At the same time, low concentrations of agglutinins, ribosomal proteins, and nonspecific lipid transport proteins were noted. The results obtained allowed us to recommend different methods for obtaining plant protein concentrates with a given protein composition.
REFERENCES
Pavlenkova, S., Shuvaeva, G., Meshcheryakova, O., Miroshnichenko, L., and Korneeva, O., Amaranth, J. Biotechnol., 2017, vol 256, p. S27. https://doi.org/10.1016/J.JBIOTEC.2017.06.642
Chirkova, T.V., Soros. Obrazovat. Zh., 1999, no. 10, p. 22.
Khandaker, L., Masum-Akond, A.S.M.G., Ali, M.B., and Oba, S., Sci. Hortic., 2010, vol. 123, p. 289. https://doi.org/10.1016/j.scienta.2009.09.012
Santos, M.Y., Osuna-Castro, J.A., Borodanenko, A., and Parades-Lopez, O., Food Sci. Technol. Int., 2005, vol. 11, p. 269. https://doi.org/10.1177/1082013205056491
Taipova, R.M. and Kuluev, B.R., Biomika, 2015, vol. 7, no. 4, p. 284.
Avanza, M.V., Puppo, M.C., and Anon, M.C., J. Food Sci., 2005, vol. 70, p. 223. https://doi.org/10.1111/j.1365-2621.2005.tb07139.x
Nasseri, A.T., Rasoul-Amini, S., Morowvat, M.H., and Ghasemi, Y., Am. J. Food Technol., 2011, vol. 6, p. 103. https://doi.org/10.3923/ajft.2011.103.116
Meshcheryakova, O.L., Vasilenko, L.I., Gubin, A.S., Sviridova, T.V., and Korneeva, O.S., Sorbtsionnye Khromatogr. Protsessy, 2022, vol. 22, no. 6, p. 841. https://doi.org/10.17308/sorpchrom.2022.22/10890
Aebersold, R. and Mann, M., Nature, 2003, vol. 422, p. 198. https://doi.org/10.1038/nature01511
Weisser, H., Nahnsen, S., Grossmann, J., Nilse, L., Quandt, A., Brauer, H., Sturm, M., Kenar, E., Kohlbacher, O., Aebersold, R., and Malmström, L., J. Proteome Res., 2013, vol. 12, p. 1628. https://doi.org/10.1021/pr300992u
Funding
This research was supported by the Russian Science Foundation (project no. 22-26-00277).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors of this work declare that they have no conflicts of interest.
Additional information
Translated by V. Makhlyarchuk
Publisher’s Note.
Pleiades Publishing remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Meshcheryakova, O.L., Bugrova, A.E., Kononikhin, A.S. et al. Chromatography–Mass Spectrometry Analysis of Plant Protein Concentrates. J Anal Chem 79, 1318–1321 (2024). https://doi.org/10.1134/S1061934824700679
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1061934824700679