Abstract
Whole-transcriptome data were used to study the changes in expression of genes coding proteins involved in the calcium regulation processes in the hippocampus of male mice with symptoms of depression caused by chronic social defeat stress. Cacna1g, Cacnb3, Camk1g, Camk2d, Camk2n2, Caly, Caln1, S100a16, and Slc24a4 genes were upregulated in the hippocampus of depressed mice compared to a control, while C-acna2d1, Cacng5, Grin2a, and Calm2 were downregulated. The greatest number of significant correlations was observed between the expression level of Calm2, which showed the highest transcriptional activity, and other differentially expressed genes. Calcium signaling in the hippocampus was assumed to be disrupted in mice exposed to chronic social defeat stress. The involvement of Calm2, Сamk1g, Camk2d, and Camk2n2 genes in the process is discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Depression is a multifactorial condition caused by an interaction of social, psychological, and physiological factors and is among the most common mental disorders [1]. Molecular mechanisms of depression have been the focus of many studies. Depression-associated genes are sought intensely [2–4]. The multiplicity of related genes agrees with the idea that depression is a complex heterogeneous disorder that involves various neurophysiological processes [5–7].
Distortion of calcium homeostasis in the central nervous system is known to play a role in the mechanisms of the majority of psychoemotional disorders, including depression [8–10] and neurological conditions [11–15]. Calcium ions have been shown to regulate a variety of processes in the normal living cell [16, 17], acting as secondary messengers to trigger important intracellular signaling cascades in response to external stimuli [18] via complexation with the calcium-binding protein calmodulin [19–21]. In nervous tissue, calcium signaling plays a special role in membrane depolarization and synaptic activity [22, 23] to ensure neuronal plasticity and learning- and memory-related processes [24].
The term “calciopathy” has been coined to collectively describe various conditions due to distortion of calcium-related processes. Calcium channelopathies are disorders due to improper work of voltage-gated calcium channels, result from dysfunction of ion channel subunits or their regulatory proteins, and are a major part of calciopathies along with dysfunctions of regulatory pathways and mitochondria [25].
The hippocampus plays an important role in the mechanisms of mental disorders and, in particular, depression [15, 26, 27]. The hippocampus is a central structure of the limbic system and is directly involved in neurogenesis, emotion formation, memory consolidation, and the stress response [28, 29]. In view of this, the objective of this work was to identify the genes that change in expression in the hippocampus under the influence of chronic social stress and code for proteins involved in regulating the calcium processes. We used the data from hippocampal transcriptome sequencing (RNA-seq) in male mice exposed to chronic social stress, which leads to a depression-like condition [30, 31].
We assumed that a study of how and to what extent expression of particular calcium process-related genes changes in the hippocampus would help to better understand the Ca2+-dependent molecular mechanisms of a depression-like condition in mice exposed to chronic social stress.
EXPERIMENTAL
Animals. Experiments were performed in adult male C57BL/6J mice that were 2.5–3 months old and had a body weight of 26–28 g. The mice received enough water and granulated food. The light–dark cycle was 12 : 12 h. All experimental procedures were performed in compliance with international guidelines for animal research (Directive 2010/63/EU of the European Parliament and of the Council on the Protection of Animals Used for Scientific Purposes). The study protocol no. 9 was approved by the Ethics Committee at the Institute of Cytology and Genetics (Minutes no. 613 dated March 24, 2010).
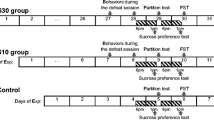
Induction of a depression-like state in male mice. The chronic social defeat stress model was used to simulate a depression-like condition [31, 32]. Mice were placed in pairs into an experimental cage with a transparent perforated divider in the center. The divider allowed the mice to see, hear, and smell each other (sensory contact), but prevented their physical interaction. The divider was removed every day in the afternoon (from 3 to 5 p.m.), leading to agonistic interactions. The first two or three days were used to identify the winners, which demonstrated aggressive behavior daily, and mice always defeated when interacting with the same partner. The defeated males were then daily transferred after the test into another cage with a new aggressive partner separated by the divider. When intense attacks by an aggressive male lasted more than 3 min during an agonistic interaction test, the interaction was terminated by placing the divider back to avoid damage to the partner. The test was continued for up to 10 min otherwise. A depression-like condition developed in the males with repeated social defeat experience after 3 weeks of the experiment [31].
To perform a neurogenomic study, we selected the mice with negative defeat experience and an overt depressive condition, which was accompanied by anxiety, fear, and lower locomotor activity (the mice are hereafter referred to as depressive mice). An alternative group used to study the specific changes included mice that belonged to a contrasting social group, had positive experience of winning between-male confrontations, and demonstrated aggressive behavior in the test (hereafter, aggressive mice).
Mice of the depressive and aggressive test groups were decapitated on the next day after the last confrontation. Males without consistent experience of agonistic interactions were used as a control. The hippocampus was excised according to the Allen Mouse Brain Atlas (http://mouse.brain-map.org/static/atlas) by the same researcher. The specimens were labeled, placed into an RNAlater solution to prevent RNA degradation, and stored at –70°C until sequencing.
Transcriptome analysis. Transcriptome sequencing in hippocampal specimens of male mice was performed by Genoanalytica (Moscow, Russia; http:// genoanalytica.ru/). Three mice were examined in each group. Each specimen was sequenced separately. Total mRNA was isolated with a Dynabeads RNA Purification kit (Ambion, United States), and cDNA libraries were constructed according to the NEBNext protocol for Illumina (NEB, United States). Sequencing of the cDNA libraries was performed on an Illumina Hiseq 1500 platform (Illumina Sequencing, United States). Only annotated genes were included in further analysis. A gene expression level was measured in fragments per kilobase of transcript per million mapped reads (FPKM), using the Cufflinks program.
Gene ontology categories of differentially expressed genes (DEGs) were determined using a bioinformatics web resource. DEGs were identified as genes whose expression levels differed significantly (p < 0.05) between depressive and control mice, aggressive and control mice, and depressive and aggressive mice. The significance of differences was additionally assessed using a correction for multiple comparisons (a q-value was obtained as a p-value corrected by the Benjamini–Hochberg method (FDR). The following web resources were used to evaluate the gene ontology categories for DEG sets:
(1) STRING: Functional Protein Association Networks (http://string-db.org),
(2) GeneCards: The Human Gene Database (https://genecards.org), and
(3) MalaCards: Human Disease Database (https:// malacards.org).
We initially focused on 75 genes whose protein products are involved in various Ca2+-related processes from Ca2+ transport through ion channels to change the intracellular Ca2+ concentration to the induction of Ca2+/calmodulin-dependent signaling cascades via activation of the respective enzymes. The genes are listed in Table S1 (see Supplementary Information at http://www.molecbio.ru/downloads/2023/ 2/supp_Pavlova_rus.pdf).
RESULTS AND DISCUSSION
Differences in DEGs set and the direction and extent of changes in gene expression in the hippocampus were observed between mice with alternative (negative or positive) types of social interaction experience.
Expression of 26 out of the 75 genes in depressive mice significantly differed from that in control and aggressive mice (Table 1). The genes code for functionally different proteins involved in calcium signaling. Eleven genes (Cacna1a, Cacna1b, Cacna1g, C-acna1h, Cacna1i, Cacna2d1, Cacnb1, Cacnb3, Cacng2, Cacng3, and Cacng5) code for proteins of voltage-gated calcium channels; seven genes (Calb1, Calcoco1, Calm2, Caln1, Hpcal4, Ppp3r1, and S100a16) code for calcium-binding proteins; one gene (Caly) codes for the neuron-specific protein calcyon, which is necessary for maximal Ca2+ release; three genes (Camk1g, Camk2d, and Ca-mk2n2) code for Ca2+/calmodulin-dependent protein kinases Iγ and IIϭ and a protein kinase II inhibitor; two genes (Slc24a2 and Slc24a4) code for Na/K/Ca transporters; and two genes (Grin2a and Grin2c) code for subunits 2A and 2C of the NMDA glutamate receptor with a high Ca2+ permeability.
A comparison of gene expression between the depressive and aggressive mice (D vs. A) revealed differences in expression of 24 genes. Of these, 17 genes (Cacna1, Cacna1b, Cacna1g, Cacna1h, Cacna1i, Cacnb1, Cacnb3, Cacng2, Cacng3, Calcoco1, Caln1, Camk1g, Camk2d, Camk2n2, Hpcal4, S100a16, and Grin2c) were expressed to a higher level in the depressive mice compared with the aggressive mice, and the other seven genes (Cacna2d1, Cacng5, Calb1, Calm2, Ppp3r1, Slc24a2, and Grin2a) were expressed to a higher level in the aggressive mice (Table 1).
Expression levels of 13 genes in the depressive mice significantly differed from the levels observed in the control mice (Fig. 1, Table 1). In particular, nine genes (Cacna1g, Cacnb3, Camk1g, Camk2d, Camk2n2, Caly, Caln1, S100a16, and Slc24a4) were upregulated and four genes (Cacna2d1, Cacng5, Grin2a, and Calm2) were downregulated. Two genes (Cacng2 and Cacna1a) were downregulated in the aggressive mice compared with the control mice; genes upregulated in stress were not detected (Fig. 1, Table 1).
A correlation analysis of expression of the 15 DEGs shown in Fig. 1 showed differences in the type and degree of correlations between them (Table 2; more detailed data are summarized in Table S2, see Supplementary Information).
Opposite correlations were observed between the two groups of genes that changed in expression in the depressive mice. The upregulated DEGs (Cacna1g, Cacnb3, Camk1g, Camk2d, Camk2n2, Caly, Caln1, S100a16, and Slc24a4) correlated positively, if at all, with each other and negatively, if at all, with the downregulated DEGs (Cacna2d1, Cacng5, Grin2a, and Calm2). Only positive, if any, correlations were observed between the downregulated DEGs. Expression of Cacng5 did not correlate with expression of the other genes (Table S2, see Supplementary Information at http://www.molecbio.ru/downloads/2023/ 2/supp_Pavlova_rus.pdf).
A positive correlation was found between the two calcium channel DEGs (Cacna1a and Cacng2) that changed in expression in the aggressive mice. The finding indicates that the calcium channel subunits encoded by the genes act in a coordinated manner to regulate the calcium processes in the hippocampus in aggressive mice (Table S2).
Five (Cacna1g, Camk1g, Camk2n2, Caln1, and Calm2) out of the 15 DEGs showed maximal numbers of correlations, including highly significant ones (р < 0.01 or р < 0.001), with each other and the other genes in the alternative behavioral mouse groups (Table 2). Саlm2 had the greatest number of correlations with other genes, and the majority of these correlations were negative (Table 2).
To further study the interactions of the 15 DEGs in regulating the calcium processes, functional associative relationships of the DEG-encoded proteins were analyzed using the STRING database (http://string-db.org). The associative relationships of proteins encoded by 11 out of the 15 DEGs are shown in Fig. 2.
Functional associative relationships of 11 DEG-encoded proteins according to STRING data (http://string-db.org).
The scheme (Fig. 2) indicates that Сalm2, Сamk1g, Camk2d, Сamk2n2, and Grin2a (NMDA glutamate receptor subunit α with a high Ca2+ permeability) may play a key role in the formation of possible pathways regulating the function of calcium channels.
DISCUSSION
Our study showed for the first time that genes for proteins involved in regulating the calcium signaling proteins change in expression in the hippocampus of male mice exposed to chronic social defeat stress. The findings agree with the experimental data that the hippocampus as a central structure of the limbic system of the brain is involved in the response to chronic social stress [28, 29, 33, 34] and the pathogenesis of various mental disorders, including autism, anxiety disorders, schizophrenia, and depression in particular [26, 27, 35–39].
Specific changes in expression of calcium process-associated genes in the hippocampus were found to accompany the response to chronic social stress in mice of both test groups. The changes were greater in depressive mice, which provide a model of human depressive disorders [30, 31]. In total, 13 out of the 15 genes examined changed in expression in the depressive mice (relative to the control). Only two genes were regulated in the aggressive mice, and expression of these genes did not change in the depressive mice (Fig. 1; Table 2, columns D vs. C and A vs. C).
However, 24 genes showed differences in expression when we compared the two mouse groups with alternative types of social behavioral experience (Table 1, column D vs. A). Opposite changes occurring in the groups relative to the control are probably responsible for the greater difference observed in the comparison of the aggressive vs. depressive mice.
It should be noted that expression of other gene groups in the hippocampus has been studied in the same model and that similar results have been obtained. For example, five genes for collagen proteins changed in expression in the hippocampus in aggressive mice, while 15 genes were regulated in depressive mice; only two genes were common in the gene sets of the two mouse groups, and opposite changes in expression were observed for these genes [40]. An analysis of glutamatergic genes has revealed changes in expression of only one gene in aggressive mice and seven genes in depressive mice [41]. The above data and the results of this work indicate that depressive animals are more susceptible to the negative effect of chronic social stress on expression of various genes in the hippocampus.
Membrane voltage-gated calcium channels play the most important role in calcium signaling because Ca2+ ions transferred through these channels affect many intracellular processes, such as electrical impulse transmission in neurons, synaptic transmission, regulation of cell secretion mechanisms, activation of Ca2+-dependent cascades, and gene expression [23, 42–44]. It is thought that voltage-gated channels are inactive and are activated when the membrane is depolarized and the potential shifts to a positive region, thus allowing Ca2+ influx into the cell [45, 46]. Uncontrolled chronic stress has been shown to change the dendrite structure in the hippocampus and to cause a higher calcium loading of cells upon depolarization [28].
Six genes for proteins of voltage-gated channels showed opposite changes in expression in the two test groups (Fig. 1) and correlated both positively and negatively to a various extent with each other and other genes (Table S2, see Supplementary Information at http://www.molecbio.ru/downloads/2023/2/supp_ Pavlova_rus.pdf). For example, Cacna1g expression correlated positively with expression of nine other DEGs and was increased in depressive mice. Сасna2d1, which belongs to the same family, decreased in expression and correlated negatively with five genes. Cacng5 decreased in expression and lacked correlation with other DEGs. The findings did not produce a detailed picture of a concerted regulation of calcium processes by DEGs of voltage-gated channels, but point to their different roles in the response to chronic social defeat stress.
Chronic social defeat stress, which causes a depression-like condition in animals, changed expression of not only the genes for calcium channel subunits, but also of several other genes whose protein products are involved in the regulation of calcium signaling at its various steps. Changes in expression were observed for the genes for calcium-binding proteins, including calmodulin, calcium/calmodulin-dependent protein kinases, a protein kinase inhibitors, transmembrane proteins that regulate Ca2+ influx into the cell, and transporter proteins (Fig. 1, Table 1).
Interesting data were obtained for Саlm2, which codes for the calcium-binding protein calmodulin (a calcium-modulating protein). First, the Calm2 expression level (>800 FPKM) is manifold greater than expression levels of other genes in the hippocampus (Fig. 1). Second, Calm2 showed the greatest number of correlations with other genes: ten positive and two negative correlations (Table 2).
Calmodulin is responsible for signal transmission to Ca2+/calmodulin-dependent protein kinases and plays a key role in the pathogenesis of psychoneurological disorders [47]. Ca2+/calmodulin-dependent protein kinases are especially active in brain tissues, where the enzymes perform many functions, regulating synaptic plasticity, gene expression, and remodeling of the cytoskeleton. A role of Ca2+/calmodulin-dependent protein kinases in human depression and animal depression-like conditions is a matter of discussion [48]. The Ca2+/calmodulin complex has been shown to inactivate the voltage-gated calcium channels when excessive Ca2+ enters the cell, and the term “calmodulation” has been proposed, meaning the regulation of calcium channel activity by calmodulin [49].
Downregulation of Саlm2; upregulation of Camk1g, Camk2d, and Camk2n2, which are functionally associated with Сalm2; and their correlations with other genes indicate that substantial changes in the function of the calcium/calmodulin-dependent protein kinase complex and the respective calcium signaling system in the hippocampus occur in animals in response to chronic social defeat stress.
Based on our findings, the above genes are possible to consider as key regulators of the calcium-dependent signaling system in the hippocampus in chronic social defeat stress. The idea is indirectly supported by the associative relationships of their protein products.
A substantial decrease in Grin2a expression was observed in the depressive mice, supplementing the picture of DEG regulation in the hippocampus (Fig. 1). Grin2a codes for a protein of glutamate-dependent Ca2+-permeable ion channels and plays a key role in the mechanisms of certain types of memory and learning [50]. Grin2a showed a positive correlation with Саlm2 and negative correlations with six other genes, including Camk1g, Camk2d, Camk2n2, and Саln1 for a calmodulin family protein. The finding indicates that activities of Grin2a and the above genes in the hippocampus are coordinated in the depressive mouse group.
Our data suggest that calciopathy, including calcium channelopathy, may develop to distort calcium signaling in the mouse hippocampus in response to chronic social defeat stress.
Genes of the Сасna group attract particular attention now. The genes code for proteins of voltage-gated calcium channels, which are thought to play a crucial universal role in the pathogenesis of many neuropsychological disorders [51–55]. For example, Cacna1b, Cacna1g, Cacna1h, and Cacna1i are associated with autism, and their expression was found to change in depressive mice in our study. The finding agrees with data from other studies that have used the same behavioral model and demonstrated that genes associated with autistic traits are upregulated in depressive mice [41, 56].
The 26 DEGs associated with calcium signaling in mice (Table 1) were compared with their human counterparts with the use of genetic databases (https://www.malacards.org/ and https://www.genecards.org/) and published data [54]. In total, 24 genes, that were found to be differentially expressed in the hippocampus of mice, were associated with neurodegenerative and mental disorders in humans (Table 3). A similarity in DEGs associated with calcium processes between mice and humans make it possible to expect that common mechanisms may be found to underlie various psychoneurological disorders and their risk may be predicted using the genetic markers identified.
CONCLUSIONS
The Calm2, Camk1g, Сamk2d, and Camk2n2 genes, which are differentially expressed in the hippocampus of depressive mice, correlate with DEGs coding for voltage-gated calcium channels and may play a key role in regulating the channel function. Grin2a, which codes for an ionotropic glutamate receptor subunit, may play a substantial role. In total, our findings demonstrate that calcium signaling-related genes change in expression, making it possible to assume that calciopathy, including calcium channelopathy, develops in the hippocampus of mice with a depression-like condition due to negative experience of defeats and chronic social defeat stress.
REFERENCES
Li Z., Ruan M., Chen J., Fang Y. 2021. Major depressive disorder: advances in neuroscience research and translational applications. Neurosci. Bull. 37, 863–880.
Lohoff F.W. 2010. Overview of the genetics of major depressive disorder. Curr. Psychiatry Rep. 12, 539–546.
Sall S.S., Thompson W., Santos A., Dwyer D.S. 2021. Analysis of major depression risk genes reveals evolutionary conservation, shared phenotypes, and extensive genetic interactions. Front. Psychiatry. 12, 698029.
Mariani N., Cattane N., Pariante C., Cattaneo A. 2021. Gene expression studies in depression development and treatment: an overview of the underlying molecular mechanisms and biological processes to identify biomarkers. Translat. Psychiatry. 11, 354.
Stacey D., Cohen-Woods S., Toben C., Arolt V., Dannlowski U., Baune B.T. 2013. Evidence of increased risk for major depressive disorder in individuals homozygous for the high-expressing 5-HTTLPR/rs25531 (LA) allele of the serotonin transporter promoter. Psychiatr. Genet. 23, 222–223.
Fan T., Hu Y., Xin J., Zhao M., Wang J. 2020. Analyzing the genes and pathways related to major depressive disorder via a systems biology approach. Brain Behav. 10, e01502.
Nobis A., Zalewski D., Waszkiewicz N. 2020. Peripheral markers of depression. J. Clin. Med. 9, 3793.
Duman R.S., Voleti B. 2012. Signaling pathways underlying the pathophysiology and treatment of depression: novel mechanisms for rapid-acting agents. Trends Neurosci. 35, 47–56.
Donev R., Alawam K. 2015. Alterations in gene expression in depression: prospects for personalize patient treatment. Adv. Protein Chem. Struct. Biol. 101, 97–124.
Norkeviciene A., Gocentiene R., Sestokaite A., Sabaliauskaite R., Dabkeviciene D., Jarmalaite S., Bulotiene G.A. 2022. Systematic review of candidate genes for major depression. Medicina (Kaunas). 58, 285.
Berridge M.J. 2014. Calcium signaling and psychiatric disease: bipolar disorder and schizophrenia. Cell Tissue Res. 357, 477–492.
Fairless R., Williams S.K., Diem R. 2014. Dysfunction of neuronal calcium signaling in neuroinflammation and neurodegeneration. Cell Tissue Res. 357, 455–462.
Czeredys M. 2020. Dysregulation of neuronal calcium signaling via store-operated channels in Huntington’s disease. Front. Cell Dev. Biol. 8, 611735.
Da Silva P.R., Gonzaga do N.T.K.S, Maia R.E., da Silva B.A. 2022. Ionic channels as potential targets for the treatment of autism spectrum disorder: a review. Curr. Neuropharmacol. 20, 1834–1849.
Xu J., Minobe E., Kameyama M. 2022. Ca2+ dyshomeostasis links risk factors to neurodegeneration in Parkinson’s disease. Front. Cell. Neurosci. 16, 867385.
Schmunk G., Gargus J.J. 2013. Channelopathy pathogenesis in autism spectrum disorders. Front. Genet. 4, 222.
Cortés-Mendoza J., de León-Guerrero S.D., Pedraza-Alva G., Pérez-Martínez L. 2013. Shaping synaptic plasticity: the role of activity mediated epigenetic regulation on gene transcription. Int. J. Dev. Neurosci. 6, 359–369.
Berridge M.J., Lipp P., M.D., Bootman M.D. 2000. The versatility and universality of calcium signaling. Nat. Rev. Mol. Cell Biol. 1, 11–21.
Van Eldik L.J., Watterson D.M. 1998. Calmodulin and calcium signal transduction: an introduction. In Calmodulin and Signal Transduction. Van Eldik L.J., Watterson D.M., Eds. Elsevier: Academic, pp. 1–15.
Brandt P.C., Vanaman T.C. 1998. Calmodulin and ion flux regulation. In Calmodulin and Signal Transduction. Van Eldik L.J., Watterson D.M., Eds. Elsevier: Academic, pp. 397–471.
Zhang M., Abrams C., Wang L., Gizzi A., He L., Lin R., Chen Y., Loll P.J., Pascal J.M., Zhang J.-F. 2012. Structural basis for calmodulin as a dynamic calcium sensor. Structure. 20, 911–923.
Salińska E., Łazarewicz J.W. 2012. Role of calcium in physiology and pathology of neurons. Postepy Biochem. 58, 403–417.
Brini M., Calì T., Ottolini D., Carafoli E. 2014. Neuronal calcium signaling: function and dysfunction. Cell. Mol. Life Sci. 71, 2787–2814.
Napolioni V., Persico A.M., Porcelli V., Palmieri L. 2011. The mitochondrial aspartate/glutamate carrier AGC1 and calcium homeostasis: physiological links and abnormalities in autism. Mol. Neurobiol. 44, 83–92.
Schmunk G., Gargus J.J. 2013. Channelopathy pathogenesis in autism spectrum disorders. Front. Genet. 4, 222.
Savitz J.B., Drevets W.C. 2009. Imaging phenotypes of major depressive disorder: genetic correlates. Neuroscience. 164, 300–330.
Grace A.A. 2016. Dysregulation of the dopamine system in the pathophysiology of schizophrenia and depression. Nat. Rev. Neurosci. 17, 524–532.
Krugers H.J., Lucassen P.J., Karst H., Joëls M. 2010. Chronic stress effects on hippocampal structure and synaptic function: relevance for depression and normalization by anti-glucocorticoid treatment. Front. Synaptic Neurosci. 2, 24.
Lagace D.C., Donovan M.H., DeCarolis N.A., Farnbauch L.A., Malhotra S., Berton O., Nestler E.J., Krishnan V., Eisch A.J. 2010. Adult hippocampal neurogenesis is functionally important for stress-induced social avoidance. Proc. Natl. Acad. Sci. U. S. A. 107, 4436–4441.
Golden S.A., Covington H.E., Berton O., Russo S.J. 2011. A standardized protocol for repeated social defeat stress in mice. Nat. Protoc. 6, 1183–1191.
Kudryavtseva N.N. 2021. Development of mixed anxiety/depression-like state as a consequences of chronic anxiety: review of experimental data. Curr. Topics Behav. Neurosci. Berlin: Springer, 54, 125–152.
Kudryavtseva N.N., Bakshtanovskaya I.V., Koryakina L.A. 1991. Social model of depression in mice of C57BL/6J strain. Pharmacol. Biochem. Behav. 38, 315–320.
Karst H., Joëls M. 2007. Brief RU 38486 treatment normalizes the effects of chronic stress on calcium currents in rat hippocampal CA1 neurons. Neuropsychopharmacology. 32, 1830–1839.
Smagin D.A., Bondar N.P., Kovalenko I.N., K-udryavtseva N.N., Michurina T.V., Enikolopov G., Park J.-H., Peunova N., Glass Z., Sayed K. 2015. Altered hippocampal neurogenesis and amygdalar neuronal activity in adult mice with repeated experience of aggression. Front. Neurosci. 9, 443.
DeLong G.R. 1992. Autism, amnesia, hippocampus and learning. Neurosci. Biobehav. Rev. 16, 63–70.
Irle E., Ruhleder M., Lange C., Seidler-Brandler U., Salzer S., Dechent P., Weniger G., Leibing E., Leichsenring F. 2010. Reduced amygdalar and hippocampal size in adults with generalized social phobia. J. Psychiatry Neurosci. 35, 126–131.
Moon A.L., Haan N., Lawrence S. Wilkinson L.S., Thomas K.L., Hall J. 2018. CACNA1C: Association with psychiatric disorders, behavior and neurogenesis. Schizophrenia Bull. 44, 958–965.
Xu W., Yao X., Zhao F., Zhao H., Cheng Z., Yang W., Cui R., Xu S., Li B. 2020. Changes in hippocampal plasticity in depression and therapeutic approaches influencing these changes. Neural Plasticity. 8861903, 16.
Schwarz K., Moessnang C., Schweiger J.I., Harneit A., Schneider M., Chen J., Cao H., Schwarz E., Witt S.H., Rietschel M., Nöthen M., Degenhardt F., Wackerhagen C., Erk S., Romanczuk-Seiferth N., Walter H., Tost H., Meyer-Lindenberg A. 2022. Ventral striatal-hippocampus coupling during reward processing as a stratification biomarker for psychotic disorders. Biol. Psychiatry. 91, 216–225.
Smagin D.A., Galyamina A.G., Kovalenko I.L., Babenko V.N., Kudryavtseva N.N. 2019. Aberrant expression of collagen gene family in the brain regions of male mice with behavioral psychopathologies induced by chronic agonistic interactions. BioMed. Res. Int. 7276389.
Kovalenko I.L., Galyamina A.G., Smagin D.A., Kudryavtseva N.N. 2020. Co-expression of glutamatergic and autism spectrum genes in the hippocampus of male mice with impaired social behavior. Vavilov. Zh. Genet. Sel. 24, 191–199.
Berridge M.J., Bootman M.D., Roderick H.L. 2003. Calcium: calcium signaling: dynamics, homeostasis and remodeling. Nat. Rev. Mol. Cell Biol. 4, 517–529.
Clapham D.E. 2007. Calcium signaling. Cell. 131, 1047–1058.
Nicholls J.G., Martin A.R., Wallas B.J., Fuchs P.A. 2003. Ot neirona k mozgu. (From Neuron to Brain). Moscow: Editorial URSS.
Dolgacheva L.P., Tuleukhanov S.T., Zinchenko V.P. 2020. Involvement of Ca2+-permeable AMPA receptors in synaptic plasticity. Biol. Membr.: Zh. Membr. Klet. Bio-l. 37, 175–187.
Melьnikov K.N. 2006. Diversity and properties of calcium channels in excitable membranes. Psikhofarmakol. Biol. Narkol. 6, 1139–1155.
Stratton M.M., Chao L.H., Schulman H., Kuriyan J. 2013. Structural studies on the regulation of Ca2+/calmodulin dependent protein kinase II. Curr. Opin. Struct. Biol. 23, 292–301.
Sałaciak K., Koszałka A., Zmudzka E., Pytka K. 2021. The calcium/calmodulin-dependent kinases II and IV as therapeutic targets in neurodegenerative and neuropsychiatric disorders. Int. J. Mol. Sci. 22, 1–32.
Ben-Johny M., Yue D.T. 2014. Calmodulin regulation (calmodulation) of voltage-gated calcium channels. J. Gen. Physiol. 143, 679–692.
Lucia D., Burgess D., Cullen C.L., Dorey E.S., Rawashdeh O., Moritz K.M. 2019. Periconceptional maternal alcohol consumption leads to behavioural changes in adult and aged offspring and alters the expression of hippocampal genes associated with learning and memory and regulators of the epigenome. Behav. Brain Res. 362, 249–257.
Dedic N., Pohlmann M.L., Richter J.S., Mehta D., Czamara D., Metzger M.W., Dine J., Bedenk B.T., Hartmann J., Wagner K.V., Jurik A., Almli L.M., Lori A., Moosmang S., Hofmann F., Wotjak C.T., Rammes G., Eder M., Chen A., Ressler K.J., Wurst W., Schmidt M.V., Binder E.B., Deussing J.M. 2018. Cross-disorder risk gene CACNA1C differentially modulates susceptibility to psychiatric disorders during development and adulthood. Mol. Psychiatry. 23, 533–543.
O′Roak B.J., Vives L., Girirajan S., Karakoc E., Krumm N., Coe B.P., Levy R., Ko A., Lee C., Smith J.D.,Turner E.H., Stanaway I.B., Vernot B., Malig M.,Baker C.,Reilly B., Akey J.M., Borenstein E., Rieder M.J., Nickerson D.A., Bernier R., Shendure J., Eichler E.E. 2012. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature. 485, 246–250.
Li B., Tadross M.R., Tsien R.W. 2016. Sequential ionic and conformational signaling by calcium channels drives neuronal gene expression. Science. 351, 863–867.
Kessi M., Chen B., Peng J., Yan F., Yang L., Yin F. 2021. Calcium channelopathies and intellectual disability: a systematic review. Orphanet. J. Rare. Dis. 16, 219.
Andrade A., Brennecke A., Mallat S., Brown J., Rivadeneira J., Czepiel N., Londrigan L. 2019. Genetic associations between voltage-gated calcium channels and psychiatric disorders. Int. J. Mol. Sci. 20, 3537.
Kudryavtseva N.N., Kovalenko I.L., Smagin D.A., Galyamina A.G., Babenko V.N. 2017. Abnormality of social behavior and dysfunction of autism related gene expression developing under chronic social defeat stress in male mice. Eur. Neuropsychopharmacol. 27, S678.
Funding
This work was supported by State Program 47 “Scientific and Technological Development of the Russian Federation” (2019–2030) (project no. 0134-2019-0002) an agreement with the Institute of Cytology and Genetics SB RAS (project no. FWNR-2022-0019).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interests. The authors declare that they have no conflict of interest.
Statement on the welfare of animals. All experiments with mice were performed in compliance with international guidelines for animal research (Directive 2010/63/EU of the European Parliament and of the Council on the Protection of Animals Used for Scientific Purposes). The study protocol (no. 9) was approved by the Ethics Committee at the Institute of Cytology and Genetics SB RAS (Minutes no. 613 dated March 24, 2010).
Additional information
Translated by T. Tkacheva
Supplementary Information
Rights and permissions
About this article
Cite this article
Pavlova, M.B., Smagin, D.A., Kudryavtseva, N.N. et al. Changes in Expression of Genes Associated with Calcium Processes in the Hippocampus in Mice Exposed to Chronic Social Stress. Mol Biol 57, 356–365 (2023). https://doi.org/10.1134/S0026893323020176
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893323020176