Abstract
Technology of production of single-domain antibodies (NANOBODY® molecules, also referred to as nanoantibodies, nAb, or molecules based on other stable protein structures) and their derivatives to solve current problems in biomedicine is becoming increasingly popular. Indeed, the format of one small, highly soluble protein with a stable structure, fully functional in terms of specific recognition, is very convenient as a module for creating multivalent, bi-/oligo-specific genetically engineered targeting molecules and structures. Production of nAb in periplasm of E. coli bacterium is a very convenient and fairly universal way to obtain analytical quantities of nAb for the initial study of the properties of these molecules and selection of the most promising nAb variants. The situation is more complicated with production of bi- and multivalent derivatives of the initially selected nAbs under the same conditions. In this work, extended linker sequences (52 and 86 aa) between the antigen-recognition modules in the cloned expression constructs were developed and applied in order to increase efficiency of production of bispecific nanoantibodies (bsNB) in the periplasm of E. coli bacteria. Three variants of model bsNBs described in this study were produced in the periplasm of bacteria and isolated in soluble form with preservation of functionality of all the protein domains. If earlier our attempts to produce bsNB in the periplasm with traditional linkers no longer than 30 aa were unsuccessful, the extended linkers used here provided a significantly more efficient production of bsNB, comparable in efficiency to the traditional production of original monomeric nAbs. The use of sufficiently long linkers could presumably be useful for increasing efficiency of production of other bsNBs and similar molecules in the periplasm of E. coli bacteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
New drugs and tools based on monoclonal antibodies and their derivatives have revolutionized biomedicine and immunobiotechnology. In recent years, in addition to classical monoclonal antibodies, engineering of the recombinant antibody fragments as well as other antigen-binding molecules has received increasing attention from many researchers. Today, one of the most promising technologies for creation of the targeting genetically engineered molecules and structures that are multivalent and bi-/oligo-specific is production of the recombinant derivatives of single-domain antigen-binding fragments (VHH) of special antibodies (HCAb, heavy-chain only antibodies), which comprise a homodimer of the shortened heavy chain with complete absence of light chains. Such special antibodies are present normally in the blood of members of the Camelidae family and in some cartilaginous fish species in addition to the classical types of immunoglobulins [1, 2]. Camelids naturally produce antibodies in which the target recognition module consists of a single variable domain (VHH), recombinant version of which is also referred to as a single-domain antibody, nanoantibody (nAb) or NANOBODY® molecule (NANOBODY® and NANOBODIES® are registered trademarks of Ablynx N.V., a subsidiary of Sanofi, so we will not use these names further in the article). The main features of nAbs are their small size (10-fold smaller than a classical antibody), high solubility, stability (in a wide range of temperatures and pH), ability to recognize unusual epitopes hidden for classical antibodies, possibility of using a very effective phage display method for selection of optimal nAb variants, as well as ease of their various modifications by genetic engineering methods. Nanoantibodies could be coupled to Fc-domains, other nAbs, peptide tags or toxins and could be chemically conjugated with drugs, radionuclides, photosensitizers, and nanoparticles [3-5]. Efficient methods of genetically engineered modification (formatting) of the initially selected monovalent nAbs have been developed in order to adapt them for specific use. More complex derivative molecules, multivalent, bispecific, and other constructs created on their basis could exhibit significantly higher specificity and binding efficiency, as well as significantly higher biological activity. On the basis of formatted nAbs, new biological tools and materials with improved properties could be developed, for example, to fight viral or other infections [6-10]. Despite the numerous examples of designing and application of bispecific nAbs [11-14], including using bacterial expression systems [15-17], currently there is no universal enough method for their rapid production in soluble and functional form for preliminary testing and selection of the most promising variants. Researchers pay attention to the methods of protein synthesis induction, suitable cell strains, and specific plasmid vectors. In the most recent studies [11, 13] and reviews [15-17] bispecific nAbs (bsNBs) were isolated from the bacterial cell lysate, but under such conditions bsNBs could denature and form undesirable bonds with other proteins present in the sample. At the same time, very few approaches to obtain bispecific nAbs with sufficiently good yield under conditions close to physiological ones have been described. One of the effective options may be isolation of such nAbs from the bacterial periplasm, where, unlike in the cytosol, cysteine bonds that stabilize immunoglobulin structure are properly formed [18].
Design of expression constructs with two or more such domains requires particular attention to the linker sequences (linkers) between the variable antigen-recognizing antibody domains. It has been shown that both length and composition of the linkers could influence the production efficiency, stability, and biological activity of both genetically engineered single-chain antibody [19, 20] and bispecific nAb [21]. The most detailed information on the development of linkers between the antigen-recognizing modules in the course of creation of bispecific nAbs is given in an extensive patent on this topic [22] from the world’s main developer of nAb technology and biomedical products based on it – the Belgian company ABLYNX NV, which recently became part of the global pharmaceutical company Sanofi. The authors of the patent provide the list of a large number of linkers with references that have been described and used up to the end of 2014. In the patent, lengths of the reported linkers range from 1 to 50 amino acid residues (aa).
It should be noted that production of bi- and multivalent nAb derivatives in bacterial periplasm is often a difficult task. In particular, our laboratory has accumulated considerable experience in the production of a variety of monovalent nAbs in bacterial periplasm, but our initial experiments on design and production of bispecific and multivalent nAbs in bacterial periplasm revealed that the procedure using for this purpose the currently most popular method of cloning two copies of VHH separated by a traditional linker sequence such as (GGGGS)3 or a 28 aa linker created on the basis of the hinge region of the non-canonical Camel antibody was unreliable (instability of the constructs, high mutability, very low efficiency of production and isolation of soluble nAbs) [8]. The aim of this study was to test the assumption that it is possible to increase efficiency of bispecific nAb production by significant increase of the linker length using combination of the shorter linker sequences that we have previously used. The main objectives of this work were: (i) to create new constructs for expression of bsNBs with two variants of elongated linkers based on the previously cloned sequences of monovalent nAbs; (ii) to use the obtained constructs to test efficiency of the bsNBs production in bacterial periplasm and their isolation in soluble form, (iii) to analyze functionality of the antigen-binding modules of isolated bsNBs.
MATERIALS AND METHODS
Molecular cloning of expression constructs for production of bispecific nAbs in E. coli periplasm. Cloning of the constructs schematically shown in Fig. 1 (see the Results and Discussion section) based on the previously obtained nAb variants [8, 23-25], which already contain the left N-terminal part of the indicated linkers at the end of the formatted nAb sequence (the hinge region of 28 aa is grayed out in Fig. 1; the hinge region is additionally followed by trimerizing domain (ILZ) in some taken clones) into the phagemid vector pHEN6 was performed using competent E. coli XL-1 Blue cells (Eurogen, Russia), [26], which contain the gene of resistance to ampicillin. PCR cloning was performed in several steps using pairs of synthesized oligonucleotides (table), restriction and ligation reactions similar to those described previously [8]. The final bispecific constructs were incorporated into the pHEN6 phagemid vector at the NcoI and EcoRI sites. These constructs consist of four principle fragments (Fig. 1): NcoI to BamHI (VHH1 and N-terminus of the linker), BamHI to SalI (C-terminus of the linker, PGGGGSGGGGSGGGGLE), SalI to NotI (VHH2), and NotI to EcoRI (HA-tag, His-tag). These fragments were obtained by restriction of PCR products synthesized with the following primer pairs: NcoI-VHH-fd and BamHI-rev (previously obtained nAb clones were used as a template); BamHI-fd and SalI-rev (without DNA template); SalI-fd and NotI-rev (nAb clones); NotI-fd and EcoRI-rev (nAb clones). Correctness of the intermediate constructs was confirmed by PCR and by checking length of the resulting products; sequences of the final constructs were verified by sequencing after cloning was completed. Full information on the sequences of the obtained constructs can be found in our patent application [27].
Primers for PCR cloning
Primer | Sequence (5′-3′) |
|---|---|
NcoI-VHH-fd | CGAGCCGACCATGGCCCTGCAGGTGCAGCTGGTGGAGTCTGG |
BamHI-rev (for linker 1) | ACCTGGATCCCTCGAGCCGCTTTCTGGTTCAGG |
BamHI-rev (for linker 2) | ACCTGGATCCCTCGAGCCGCTACGTTCACCCAC |
BamHI-fd | CGAGGGATCCAGGTGGAGGCGGTTCAGGCGGAGGTGGCA |
Sal-rev | ATGTCCGTCGACTCTAGACCGCCACCGCCGCTGCCACCT |
SalI-fd | GCGGTCGACGGCTCAGGTGCAGCTGGTGGAGT |
NotI-rev | GGACTAGTGCGGCCGCTTGAGGAGACGGTGACCTGGGT |
NotI-fd | GCGAGCGGCCGCTACCCGTACGAC |
EcoRI-rev | GCGGAATTCTATTAGCCGGAAGCGTAGTC |
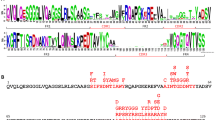
Scheme of the expression construct used for synthesis of bispecific nAbs and sequence of the linkers separating the sequences encoding nAbs. Designations from left to right: lactose promoter site (Plac/operator); ribosome binding site (RBS); translation initiation (arrow); signal peptide for periplasmic localization (pelB); sequences encoding nAbs (VHH1 and VHH2) separated by the linker sequence (amino acid sequences of the used linkers are indicated at the bottom of the figure); HA-tag sequence (YPYDVPDYA) and six histidine residues at the very end. ILZ, trimerizing domain, “isoleucine zipper.”
Production and purification of bispecific nAbs. Expression of nanoantibodies and preparation of periplasmic extract were performed according to the previously described protocol [28] with minor modifications. Overnight bacterial culture of E. coli XL1 Blue containing plasmid DNA with the expression construct was obtained from a freshly grown single colony on a LB-agar supplemented with 1% glucose and 100 μg/ml ampicillin by incubation in 5 ml of LB medium with 0.2% glucose and 70 μg/ml ampicillin overnight on an Excella C24R Shaker Incubator (New Brunswick Scientific, USA) at 180 rpm and 37°C. Overnight culture was diluted 50-100-fold in 300 ml of 2xYT medium containing 0.2% glucose. The bacteria were cultured for 30 min at 180 rpm and 37°C, after which ampicillin was added to concentration 70 μg/ml and incubation was continued. When the bacterial culture reached an optical density of 0.6 at a wavelength of 600 nm, 0.5 mM isopropyl-beta-D-galactopyranoside (IPTG) solution was added, incubation temperature was lowered to 28-30°C, and the cells were incubated for 5 h. The cells were isolated by centrifugation for 10 min at 3000g. Next, osmotic shock procedure was performed to isolate the periplasmic extract. The cell precipitate was suspended in a sucrose-containing buffer (50 mM Tris-HCl, pH 8.0, 0.5 mM EDTA, 20% sucrose, 10 mM imidazole, freshly added 1 mM PMSF) and incubated in an ice bath for 30 min. Aqueous buffered saline solution (10 mM Tris-HCl, pH 8.0, 1 mM MgCl2) was then added, mixed thoroughly and rapidly, and the cell suspension was left on ice for another 30 min, after which the cells were centrifuged for 20 min at 16,000g. The supernatant containing periplasmic extract was taken and NaCl was added to a final concentration of 0.3 M. In the case of isolating a periplasmic extract containing trimerizing bispecific nAbs, it was important to use additional modifications of the procedure, including disruption of the cell wall by adding 5 mg/ml lysozyme to the solution containing sucrose and adding 20 mM imidazole to the subsequent aqueous solution.
The target protein was purified by metal-chelate affinity chromatography on a Ni2+-NTA-agarose using a HIS-Select Nickel Affinity Gel (Sigma-Aldrich, USA). This purification is possible due to the presence of six histidine residues (His-tag) at the C-terminus of formatted nAb. Purification was performed according to the manufacturer’s protocol. The column volume was selected according to the volume of the initial culture and periplasmic extract obtained; on average, a 0.5 ml column was used to purify nAb extracted from 300 ml of culture. The column was equilibrated with a “lysis” buffer (50 mM NaH2PO4 pH 8.0, 0.3 M NaCl, 10 mM imidazole). When purifying presumably trimerizing bispecific antibodies, the amount of imidazole in the buffer was increased to 20 mM. Periplasmic extract was added to the column and protein sorption was performed by “batch”-method, mixing the extract and agarose on a Bio RS-24 mini-rotator (Biosan, Latvia) for 30-40 min. Next, the column was placed in a vertical position, Ni2+-NTA-agarose was allowed to settle, and the extract was passed through it. The column was then washed in three steps using a lysis buffer with a total volume equal to 10 column volumes. Protein elution was performed fractionally by adding one column volume of elution buffer (50 mM NaH2PO4 pH 8.0, 0.3 M NaCl, 250 mM imidazole) three times. Thus, an eluate volume of 1.5 ml was obtained from one 0.5 ml column.
Protein electrophoresis. Quality and quantity of nAb contained in the obtained eluate were determined by analyzing aliquots of purified nAb with electrophoresis in a 5-19% gradient SDS-polyacrylamide gel (SDS-PAGE) performed according to the method of Laemmli [29] under reducing conditions. A MiniProtean II electrophoresis cell (Bio-Rad, USA) was used with a power source Elf-4 (DNA-Technology, Russia). DTT (dithiothreitol) was used as a reducing agent. Before sample application, a buffer (31.25 mM Tris-HCl pH 6.8; 12.5% glycerol; 1% SDS; 0.005% bromophenol blue) was added to the samples, and the samples were heated for 10 min at 97°C. Electrophoresis was performed in a standard SDS buffer (0.2 M glycine, 0.025 M Tris, 0.1% SDS). A Precision Plus Protein Standards protein set (161-0363, Bio-Rad) was used as molecular weight markers. Approximate amount of nAb in the corresponding band on the stained gel was estimated relative to the intensity of the marker bands (for example, in 5 μl of the marker set, intensity of the 50-kDa band corresponds to approximately 375 ng and of the 20-kDa band corresponds to 75 ng of protein).
Enzyme-linked immunosorbent assay (ELISA) to test functionality of both nAb modules in composition of the resulting dual-module bispecific nAb. The following antigens were used: recombinant human ICAM-1 (R&D Systems, USA), recombinant birch pollen allergen Bet v 1 (Biomay AG, Austria), recombinant hemagglutinin (HA-H5) of influenza virus H5N2 (Sino Biotechnological Inc., China), and purified human IgA (Imtek, Russia). An antigen solution at concentration of 2 µg/ml in PBS (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 2 mM KH2PO4, pH 7.4) was used to immobilize an antigen overnight at 4°C in the wells of a 96-well flat-bottomed plate (MaxiSorp, Nuns). Control wells did not contain antigen and were further processed in parallel with the rest of experimental wells. The wells were washed with PBS three times, then blocked in a 1% solution of bovine serum albumin (BSA) in PBS for one hour and washed twice with PBS. The tested nAbs in a 0.1% BSA solution in 1 × PBS were applied to the wells at different concentrations, mostly in the range of 0.4-1 μg/ml, and incubated for 1 h with occasional stirring. The wells were washed thoroughly (5 times in PBS) and horseradish peroxidase-conjugated secondary mouse anti-HA tag antibodies (Sigma-Aldrich) diluted 1 : 2000 in a 0.1% BSA/PBS were added. After 1 h, the wells were again washed thoroughly and then a 1-Step Ultra TMB indicator mix (Thermo Scientific, USA) was added. 2 M sulfuric acid was used to stop the reaction. Optical density was measured at a wavelength of 450 nm with a flatbed photometer. The data were obtained in three repeats.
RESULTS AND DISCUSSION
For many years, our laboratory has been conducting research related to the production of new single-domain antibodies (nAbs) for a wide range of target antigens and aimed at solving urgent problems in biomedicine. At present, for a number of ongoing projects of the laboratory, creation of bispecific nAbs on the basis of obtained nAbs has been required. For this purpose, a bacterial expression system (E. coli) is preferable, since it is in the periplasm of bacteria that we routinely produce nAbs and their derivatives. Previously, our laboratory has already conducted experiments on production of two nAbs in a single reading frame with coding sequences separated by a linker. However, this revealed problems of instability of the resulting constructs in bacteria (unusually high mutagenesis was detected by sequencing and PCR analysis of different resulting clones) and very low yield of recombinant protein with two nAb modules (below the level of background bacterial proteins). These problems emerged both when using the traditional linker (Gly4-Ser)×3 and the linker based on the long hinge site of camelid specific antibodies, consisting of 28 aa. In this work, previous experience was taken into account and an even longer combined linker between the sequences encoding nAb was proposed. Figure 1 shows the scheme of the structure of expression construct based on the plasmid vector pHEN6 [26] for inducible expression of bispecific nAb in the periplasm of E. coli, as well as two variants of the developed linkers (to the best of our knowledge such elongated linkers have not been suggested before).
The linker 1 (52 aa) consists of a long hinge region of camelid specific antibodies (28 aa) connected to the conventional Gly4-Ser-Gly4-Ser-Gly4 linker (as in phagemid pIT2) with several additional amino acid residues, and in the linker 2 (86 aa), an additional sequence of the recombinant trimerizing “isoleucine zipper” domain, ILZ, is incorporated between the two indicated parts of the linker 1 [8]. It has been previously shown that nAbs to which the ILZ domain has been added are indeed efficiently trimerized in the bacterial periplasm immediately after their synthesis [8].
In this short communication, examples and primary characteristics of the first bsNBs obtained using these two linker variants are presented.
Preparation and analysis of bispecific nAbs with linker 1. In collaboration with our Austrian partners, we have recently succeeded in obtaining nAb (aBet) to the major birch pollen allergen (Bet v 1) [24], as well as nAb (aICAM1) to the major epithelial surface receptor protein ICAM1 [23]. One of the further goals of this work was to obtain the bsNBs that bind to ICAM1 on the surface of human upper respiratory tract epithelial cells and can then also bind the pollen allergen coming from an external environment, blocking it and thus preventing an allergic reaction. In this study, we obtained two bsNB variants containing both nAbs, aBet-L1-aICAM1 and aICAM1-L1-aBet, but in different positions relative to the linker. Interestingly, the first bsNB variant (in three independent repeats) was produced markedly and reproducibly better under the same conditions compared to the second variant (lanes 3 and 4, respectively, in Fig. 2a). The level of production for the aBet-L1-aICAM1 bsNTs was quite comparable to the average level of production for the original single-module nAbs (approximately 2 mg from 1 liter of culture). The detectable bsNBs moved in electrophoresis gel in full agreement with the expected size of 36 kDa. At the concentration of 0.4 μg/ml, both bsNBs specifically recognized each of the immobilized antigens, Bet v 1 and ICAM1 (Fig. 2b). Thus, these preliminary data indicate that it is feasible to produce the specified bsNBs in soluble and active form in the bacterial periplasm. Relative location of these two different nAb sequences in the construct may be important for the level of production.
Results of SDS-PAGE and ELISA for bispecific aBet-L1-aICAM1 and aICAM1-L1-aBet nanoantibodies. a) Electrophoretic separation of proteins from periplasmic extracts in a 5-19% gradient SDS-polyacrylamide gel (1, 2) and affinity-purified bsNBs (3, 4). The arrow shows position of the bispecific nanoantibodies (bsNBs). The marker (M) is shown in the leftmost lane and sizes of the protein marker bands are indicated (in kilodaltons, kDa). b) Immunoassay of binding of the obtained bsNBs at concentration of 0.4 μg/ml to the immobilized recombinant antigens (shown at the bottom). Mean values of three measurements are presented with the area of data scatter.
Preparation and analysis of bispecific nAb with linker 2. In our other current project, the task is to develop means to block/detain pathogenic respiratory viruses in the upper respiratory tract of humans based on bispecific nAbs. To do this, we have obtained and continue to obtain both nAbs that specifically bind conservative surface epitopes of the current pathogenic viruses (primarily influenza viruses and coronaviruses), and nAbs to major components of mucus or, in particular, to major immunoglobulin (IgA) of mucous tissues. By binding to the viral particle, IgA molecules cause opsonization of the pathogen and give the immune system a signal to destroy it. It was previously shown that it is preferable to use trimerized nAb to bind and block the virus [8]. In order to study how the addition of ILZ to the linker affects production of bsNB, in this work we tested the second version of the linker (linker 2, Fig. 1). This time, two other previously obtained nAbs were used. One, aHN13, binds hemagglutinin (HA) of the avian influenza virus (H5N2) [8], and the other (E7) binds the human IgA [25]. The expected size of the monomeric version of this bsNB is approximately 39 kDa, which is in good agreement with the mobility of the protein detected by electrophoresis under reducing conditions (Fig. 3a), under which the putative nascent trimer breaks down into monomers.
Results of SDS-PAGE and ELISA for bsNB aHA-L2-E7 (aIgA), also denoted (HN-E7)×3. a) Electrophoretic separation of the affinity-purified bsNB in a 5-19% gradient SDS-polyacrylamide gel: initial preparation (1), after filtering through a filter with a nominal molecular weight cutoff of 100 kDa (2), presumably, trimerized bsNB retained over a filter with molecular weight cutoff of 100 kDa purified from small impurities (3). The arrow shows position of monomeric bsNB. The marker (M) in the rightmost lane is labeled and sizes of the marker bands are indicated (in kilodaltons, kDa). b) Immunoassay of binding of the obtained bsNB (HN-E7)×3 as well as control monovalent nAbs (E7 – against IgA and HN13 – against HA-H5 hemagglutinin of avian influenza virus) at concentration of 1 μg/ml with immobilized recombinant antigens (IgA and HA-H5). Control wells were antigen-free (no AG). Mean values of three measurements are presented with the area of data scatter.
In the case of trimerization, the size of bsNB should increase to about 117 kDa. In this short report, we have not yet set the task of thoroughly studying trimerization of the resulting bsNB, but two facts could be pointed out as indirect indications of trimerization of the produced protein. First, during ultrafiltration using a centrifuge concentrator with a nominal molecular weight cutoff of 100 kDa (Vivaspin Turbo 4, Sartorius), this bsNB behaves as a high molecular weight protein and is almost completely concentrated above the filter, whereas other protein molecules contaminating the initial preparation of bsNB and having a size (in the electrophoregram) of up to 75 kDa pass through the filter noticeably, in contrast to the nAT moving in denaturing electrophoresis as a 39 kDa monomeric protein. Another indication are difficulties in isolation of this bsNB in the composition of periplasmic extract. Based on the assumption of its three-dimensional structure, we specifically selected conditions for more efficient extraction of this bsNB from bacterial periplasm. We added a cell wall disruption procedure with lysozyme and increased ionic strength of the elution solution (details in the methods description). Production rate for the aHA-L2-E7 bsNB using the modified extraction conditions was also comparable to the average production rate of the original single-module nAb (approximately 1 mg from 1 liter of culture). The resulting bsNB had the required functionality and recognized both target proteins in ELISA (Fig. 3b). The monomeric nAbs taken as positive controls, anti-IgA (E7) [25] and nAb against the HA-H5 hemagglutinin of avian influenza virus (HN13) [8], gave a slightly stronger signal upon binding. This may reflect the twofold molar excess of these monospecific nAbs compared to the proportion of E7 and HN13 in the bispecific nAb (at the same concentration of all three nAb variants), and may also indicate possible steric limitations when the presumably trimerized bsNB are used in this assay. Note that for comparison this bsNB was also prepared with the linker 1 (aHA-L1-E7), without the trimerizing domain. The yield of bsNB was 2-fold higher in this case. All three variants of the described bsNBs could not be produced in the detectable amounts at all, when shorter conventional linkers are used.
Thus, we have demonstrated that the produced bispecific nAbs contain all defined domains in a functional state: they are released from periplasm, His-tag is functional for the protein isolation with the help of affinity metal-chelate chromatography, and both antigen-recognizing modules of nAbs and HA-tag are functional in immunoassays. Longer linker sequences apparently contribute to the weakening of undesirable interactions (potential recombinations) between the conserved GC-rich core sites in the consecutive VHH sequences in the case of heterologous bacterial expression system, which is reflected in the stability of constructs (producers) and in the increased production of these bispecific nAbs. We assume that our work is the first one considering the use of such elongated linkers for the purpose of increasing efficiency of production of bispecific nanobodies in periplasm. The obtained results definitely are encouraging for further developments of the similar variants of nAb derivatives for a wide range of current biomedical problems.
Patent application for the invention has been filed to protect priority right of the developments described in this article [27].
Abbreviations
- bsNB:
-
bispecific nanoantibody
- ILZ:
-
trimerizing domain
- nAb:
-
nanoantibody
- PBS:
-
phosphate-buffered saline
References
Hamers-Casterman, C., Atarhouch, T., Muyldermans, S., Robinson, G., Hamers, C., Songa, E. B., Bendahman, N., and Hamers, R. (1993) Naturally occurring antibodies devoid of light chains, Nature, 363, 446-448, https://doi.org/10.1038/363446a0.
Flajnik, M. F., and Kasahara, M. (2010) Origin and evolution of the adaptive immune system: genetic events and selective pressures, Nat. Rev. Genet., 11, 47-59, https://doi.org/10.1038/nrg2703.
Bannas, P., Hambach, J., and Koch-Nolte, F. (2017) Nanobodies and nanobody-based human heavy chain antibodies as antitumor therapeutics, Front Immunol., 8, 1603, https://doi.org/10.3389/fimmu.2017.01603.
Jovčevska, I., and Muyldermans, S. (2020) The therapeutic potential of nanobodies, BioDrugs, 34, 11-26, https://doi.org/10.1007/s40259-019-00392-z.
Tillib, S. B. (2020) Prospective applications of single-domain antibodies in biomedicine [in Russian], Mol. Biol. (Mosk.), 54, 362-373.
Stone, E., Hirama, T., Tanha, J., Tong-Sevinc, H., Li, S., MacKenzie, C. R., and Zhang, J. (2007) The assembly of single domain antibodies into bispecific decavalent molecules, J. Immunol Methods, 318, 88-94, https://doi.org/10.1016/j.jim.2006.10.006.
Hultberg, A., Temperton, N. J., Rosseels, V., Koenders, M., Gonzalez-Pajuelo, M., Schepens, B., Ibañez, L. I., Vanlandschoot, P., Schillemans, J., Saunders, M., Weiss, R. A., Saelens, X., Melero, J. A., Verrips, C. T., Van Gucht, S., and de Haard, H. J. (2011) Llama-derived single domain antibodies to build multivalent, superpotent and broadened neutralizing anti-viral molecules, PLoS One, 6, e17665, https://doi.org/10.1371/journal.pone.0017665.
Tillib, S., Ivanova, T. I., Vasilev, L. A., Rutovskaya, M. V., Saakyan, S. A., Gribova, I. Y., Tutykhina, I. L., Sedova, E. S., Lysenko, A. A., Shmarov, M. M., Logunov, D. Y., Naroditsky, B. S., and Gintsburg, A. L. (2013) Formatted single-domain antibodies can protect mice against infection with influenza virus (H5N2), Antiviral Res., 97, 245-254, https://doi.org/10.1016/j.antiviral.2012.12.014.
Huet, H. A., Growney, J. D., Johnson, J. A., Li, J., Bilic, S., Ostrom, L., Zafari, M., Kowal, C., Yang, G., Royo, A., et al. (2014) Multivalent nanobodies targeting death receptor 5 elicit superior tumor cell killing through efficient caspase induction, mAbs, 6, 1560-1570, https://doi.org/10.4161/19420862.2014.975099.
Laursen, N. S., Friesen, R. H. E., Zhu, X., Jongeneelen, M., Blokland, S., Vermond, J., van Eijgen, A., Tang, C., van Diepen, H., Obmolova, G., van der Neut Kolfschoten, M., Zuijdgeest, D., Straetemans, R., Hoffman, R. M. B., Nieusma, T., Pallesen, J., Turner, H. L., Bernard, S. M., Ward, A. B., Luo, J., Poon, L. L. M., Tretiakova, A. P., Wilson, J. M., Limberis, M. P., Vogels, R., Brandenburg, B., Kolkman, J. A., and Wilson, I. A. (2018) Universal protection against influenza infection by a multidomain antibody to influenza hemagglutinin, Science, 362, 598-602, https://doi.org/10.1126/science.aaq0620.
Efimov, G. A., Kruglov, A. A., Khlopchatnikova, Z. V., Rozov, F. N., Mokhonov, V. V., Rose-John, S., Scheller, J., Gordon, S., Stacey, M., Drutskaya, M. S., Tillib, S. V., and Nedospasov, S. A. (2016) Cell-type-restricted anti-cytokine therapy: TNF inhibition from one pathogenic source, Proc. Natl. Acad. Sci. USA, 113, 3006-3011, https://doi.org/10.1073/pnas.1520175113.
Hanke, L., Das, H., Sheward, D. J., Perez Vidakovics, L., Urgard, E., Moliner-Morro, A., Kim, C., Karl, V., Pankow, A., Smith, N. L., Porebski, B., Fernandez-Capetillo, O., Sezgin, E., Pedersen, G. K., Coquet, J. M., Hällberg, B. M., Murrell, B., and McInerney, G. M. (2022) A bispecific monomeric nanobody induces spike trimer dimers and neutralizes SARS-CoV-2 in vivo, Nat. Commun., 13, 155, https://doi.org/10.1038/s41467-021-27610-z.
Liu, Y., Ao, K., Bao, F., Cheng, Y., Hao, Y., Zhang, H., Fu, S., Xu, J., and Wu, Q. (2022) Development of a bispecific nanobody targeting CD20 on B-cell lymphoma cells and CD3 on T cells, Vaccines (Basel), 10, 1335, https://doi.org/10.3390/vaccines10081335.
Ma, H., Zhang, X., Zeng, W., Zhou, J., Chi, X., Chen, S., Zheng, P., Wang, M., Wu, Y., Zhao, D., Gong, F., Lin, H., Sun, H., Yu, C., Shi, Z., Hu, X., Zhang, H., Jin, T., and Chiu, S. A. (2022) A bispecific nanobody dimer broadly neutralizes SARS-CoV-1 & 2 variants of concern and offers substantial protection against Omicron via low-dose intranasal administration, Cell Discov., 8, 132, https://doi.org/10.1038/s41421-022-00497-w.
De Marco, A. (2015) Recombinant antibody production evolves into multiple options aimed at yielding reagents suitable for application-specific needs, Microb. Cell Factories, 14, 125, https://doi.org/10.1186/s12934-015-0320-7.
Sandomenico, A., Sivaccumar, J. P., and Ruvo, M. (2020) Evolution of Escherichia coli expression system in producing antibody recombinant fragments, Int. J. Mol. Sci., 21, 6324, https://doi.org/10.3390/ijms21176324.
Huleani, S., Roberts, M. R., Beales, L., and Papaioannou, E. H. (2022) Escherichia coli as an antibody expression host for the production of diagnostic proteins: significance and expression, Crit. Rev. Biotechnol., 42, 756-753, https://doi.org/10.1080/07388551.2021.1967871.
Skerra, A., and Plückthun, A. (1988) Assembly of a functional immunoglobulin Fv fragment in Escherichia coli, Science, 240, 1038-1041, https://doi.org/10.1126/science.3285470.
Le Gall, F., Reusch, U., Little, M., and Kipriyanov, S. M. (2004) Effect of linker sequences between the antibody variable domains on the formation, stability and biological activity of a bispecific tandem diabody, Protein Eng. Des. Sel., 17, 357-366, https://doi.org/10.1093/protein/gzh039.
Wang, Q., Chen, Y., Park, J., Liu, X., Hu, Y., Wang, T., McFarland, K., and Betenbaugh, M. J. (2019) Design and production of bispecific antibodies, Antibodies (Basel), 8, 43, https://doi.org/10.3390/antib8030043.
Huang, C., Huang, J., Zhu, S., Tang, T., Chen, Y., and Qian, F. (2023) Multivalent nanobodies with rationally optimized linker and valency for intravitreal VEGF neutralization, Chem. Eng. Sci., 270, 118521, https://doi.org/10.1016/j.ces.2023.118521.
Roobrouck, A., and Stortelers, C. (2015) Bispecific nanobodies. Applicant – ABLYNX NV (Belgium). WIPO/PCT patent publication number WO2015044386 A1. Publication date April 2, 2015.
Zettl, I., Ivanova, T., Zghaebi, M., Rutovskaya, M. V., Ellinger, I., Goryainova, O., Kollárová, J., Villazala-Merino, S., Lupinek, C., Weichwald, C., Drescher, A., Eckl-Dorna, J., Tillib, S. V., and Flicker, S. (2022) Generation of high affinity ICAM-1-specific nanobodies and evaluation of their suitability for allergy treatment, Front. Immunol., 13, 1022418, https://doi.org/10.3389/fimmu.2022.1022418.
Zettl, I., Ivanova, T., Strobl, M. R., Weichwald, C., Goryainova, O., Khan, E., Rutovskaya, M. V., Focke-Tejkl, M., Drescher, A., Bohle, B., Flicker, S., and Tillib, S. V. (2022) Isolation of nanobodies with potential to reduce patients’ IgE binding to Bet v 1, Allergy, 77, 1751-1760, https://doi.org/10.1111/all.15191.
Goryainova, O. S., Ivanova, T. I., Rutovskaya, M. V., and Tillib, S. V. (2017) A method for the parallel and sequential generation of single-domain antibodies for the proteomic analysis of human blood plasma, Mol. Biol. (Mosk.)., 51, 985-996, https://doi.org/10.7868/S0026898417060106.
Conrath, K. E., Lauwereys, M., Galleni, M., Matagne, A., Frère, J. M., Kinne, J., Wyns, L., and Muyldermans, S. (2001) Beta-lactamase in-hibitors derived from single-domain antibody fragments elicited in the Camelidae, Antimicrob. Agents Chemother., 45, 2807-2812, https://doi.org/10.1128/AAC.45.10.2807-2812.2001.
Tillib, S. V., and Goryaynova, O. S. (2024) Bispecific nanobodies with extended linker sequences between antigen-recognition modules for production in the periplasm of E. coli, Russian Federation Patent Application No. 2024101830, FIPS, Moscow.
Baral, T. N., and Arbabi-Ghahroudi, M. (2012) Expression of single-domain antibodies in bacterial systems, Methods Mol Biol., 911, 257-275, https://doi.org/10.1007/978-1-61779-968-6_16.
Laemmli, U. K. (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4, Nature, 227, 680-685, https://doi.org/10.1038/227680a0.
Funding
This work was financially supported by the Ministry of Science and Higher Education of the Russian Federation (Agreement No. 075-15-2021-1086, contract No. RF–193021X0015).
Author information
Authors and Affiliations
Contributions
S.V.T. concept and management of work, performing cloning, and further functional verification of nAb in enzyme immunoassay; O.S.G. and S.V.T. experiments on production and isolation of nAbs; S.V.T. and O.S.G. writing the article.
Corresponding author
Ethics declarations
This work does not contain any studies involving human and animal subjects performed by any of the authors. The authors of this work declare that they have no conflicts of interest.
Additional information
Publisher’s Note. Pleiades Publishing remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tillib, S.V., Goryainova, O.S. Extending Linker Sequences between Antigen-Recognition Modules Provides More Effective Production of Bispecific Nanoantibodies in the Periplasma of E. coli. Biochemistry Moscow 89, 933–941 (2024). https://doi.org/10.1134/S0006297924050134
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297924050134