Abstract
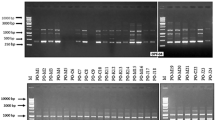
The genetic diversity of the rice blast pathogen, Pyricularia oryzae, was analysed in rice-growing provinces of Argentina. A total of 161 isolates of the fungus was collected from 15 rice cultivars at nine locations during 2000-05 and characterised using Pot2-DNA fingerprinting. Based on DNA analysis (isolates with ≥70% band similarity), five lineages were identified and designated A, B, C,Dand E, with 11, 22, 4, 1 and 4 haplotypes identified, respectively. The predominant lineage, B, representing 38% of the collected isolates, was recovered from four cultivars in five locations. In contrast to lineages A and B, which did not contain a dominant haplotype, a single haplotype predominated in lineages C and E. Isolates representing all haplotypes were examined for virulence on a set of differential rice cultivars, near-isogenic lines and commercial cultivars commonly grown in Argentina, revealing 41 pathotypes and 24 international races. There was no significant association between DNA fingerprint similarities and pathotypes. Overall, these data indicated that populations of P. oryzae in Argentina are genetically simple and predominantly clonal yet have a high pathotype diversity.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Babujee L, Gnanamanickam SS (2000) Molecular tools for characterization of rice blast pathogen (Magnaporthe grisea) population and molecular marker-assisted breeding for disease resistance. Current Science 78, 248–257.
Chen D, Zeigler RS, Leung H, Nelson RJ (1995) Population structure of Pyricularia grisea at two screening sites in the Philippines. Phytopathology 85, 1011–1020. doi: 10.1094/Phyto-85-1011
Consolo VF, Cordo CA, Salerno GL (2005) Mating-type distribution and fertility status in Magnaporthe grisea populations from Argentina. Mycopathologia 160, 285–290. doi: 10.1007/s11046-005-4333-3
Cordo CA, Linquidst JC, Marassi J (1980) Identification of races of Pyricularia oryzae in Argentina. International Rice Research Newsletter 5, 5–6.
Correa-Victoria F, Zeigler R (1995) Stability of partial and complete resistance in rice to Pyricularia grisea under rainfed upland conditions in eastern Colombia. Phytotpathology 85, 977–982. doi: 10.1094/Phyto-85-977
Correa-Victoria F, Levy M, Zeigler R (1994) Relationship between virulence spectrum and genetic families in the rice blast fungus. In ‘Proceedings of the international symposium of rice blast disease’. (Eds SA Leong, RS Zeigler, PS Teng) pp. 211–230. (CAB International: Wallingford, UK)
Couch BC, Kohn LM (2002) A multilocus gene genealogy concordant with host preference indicates segregation of a new species, Magnaporthe oryzae from M. grisea. Mycologia 94, 683–693. doi: 10.2307/3761719
Dobinson KF, Harris RE, Hamer JE (1993) Grasshopper, a long terminal repeat (LTR) retroelement in the phytopathogenic fungus Magnaporthe grisea. Molecular Plant-Microbe Interactions 6, 114–126.
Don LD, Tosa Y, Nakashashiki H, Mayama S (1999) Population structure of the rice blast pathogen in Vietnam. Annals of the Phytopathological Society of Japan 65, 475–479.
Farman ML, Taura S, Leong SA (1996) The Magnaporthe grisea DNA fingerprinting probe MGR586 contains the 30 end of an inverted repeat transposon. Molecular & General Genetics 251, 675–681.
Fern’andez DV (2005) Perfil descriptivo de la cadena del arroz. Subsecretar’ia Agricultura Ganader’ia y Pesca de Argentina. Available at http://www. sagpya.mecon.gov.ar/new/0-0/programas/dma/publicaciones/ perspectivas/Perfiles%20descriptivos/Cadena%20de%20arroz.pdf [Verified 6 March 2008]
Fernandez Valliela MV (1978) ‘Introducci’on a la Fitopatolog’ia.’ 3rd edn. (Colecci’on Cientif’ica INTA: Buenos Aires)
George MLC, Nelson RJ, Zeigler RS, Leung H (1998) Rapid population analysis of Magnaporthe grisea by using rep-PCR and endogenous repetitive DNA sequences. Phytopathology 88, 223–229. doi: 10.1094/ PHYTO.1998.88.3.223
Giarrocco LE, Marassi MA, Salerno GL (2007) Assessment of the genetic diversity in Argentine rice cultivars with SSR markers. Crop Science 47, 851–858.
Gibbons JW, Gonzales D, Delgado D (2000) Use of lineage exclusion in a multi-objective rice breeding program. In ‘Advances in rice blast research’. (Eds D Tharreau, MH Lebrun, NJ Talbot, L Notteghem) pp. 146–153. (Kluwer Academic Publishers: Dordrecht, The Netherlands)
Hamer JE, Farrall L, Orbach MJ, Valent B, Chumley FG (1989) Host species-specific conservation of a family of repeated DNA sequences in the genome of a fungal plant pathogen. Proceedings of the National Academy of Sciences of the United States of America 86, 9981–9985. doi: 10.1073/pnas.86.24.9981
Han SS, Ra DS, Nelson RJ (1993) Comparisons of phylogenetic trees and pathotypes of Pyricularia oryzae in Korea. Journal of Agricultural Science 35, 315–323.
Hittalmani S, Parco A, Mew TV, Zeigler RS, Huang N (2000) Fine mapping and DNA marker-assisted pyramiding of the three major genes for blast resistance in rice. Theoretical and Applied Genetics 100, 1121–1128. doi: 10.1007/s001220051395
International Rice Research Institute (1980) ‘Standard evaluation system for rice. International Rice Testing Program.’ 3rd edn. (International Rice Research Institute: Los Baños, The Philippines)
Javan-Nikkhah M, Hedjaroude GA, Sharifi-Tehrani A, Okhovvat SM (2003) Study on pathogenic diversity in population of Magnaporthe grisea, the rice blast fungus in Guilan province, Iran. Iranian Journal of Agricultural Science 34, 647–658.
Javan-Nikkhah M, Mc Donald B, Banke S, Ali-Hedjaroude G (2004) Genetic structure of Iranian Pyricularia grisea populations based in rep-PCR fingerprinting. European Journal of Plant Pathology 110, 909–919. doi: 10.1007/s10658-004-5570-x
Kachroo P, Leong SA, Chatoo BB (1994) Pot2, an inverted repeat transposon from the rice blast fungus Magnaporthe grisea. Molecular & General Genetics 245, 339–348. doi: 10.1007/BF00290114
Kachroo P, Leong SA, Chattoo BB (1995) Mg-SINE: a short interspersed nuclear element from the rice blast fungus, Magnaporthe grisea. Proceedings of the National Academy of Sciences of the United States of America 92, 11125–11129. doi: 10.1073/pnas.92.24.11125
Levy M, Romao J, Marchetti MA, Hamer JE (1991) DNA fingerprinting with a dispersed repeated sequence resolves pathotype diversity in the rice blast fungus. The Plant Cell 3, 95–102. doi: 10.2307/3869203
Levy M, Correa-Victoria FJ, Zeigler RS, Xu S, Hamer JE (1993) Genetic diversity of the rice blast fungus in a disease nursery in Colombia. Phytopathology 83, 1427–1433. doi: 10.1094/Phyto-83-1427
Ling KC, Ou SH (1969) Standarization of the international race numbers of Pyricularia oryzae. Phytopathology 59, 339–342.
Mantel NA (1967) The detection of disease clustering and a generalized regression approach. Cancer Research 27, 209–220.
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Research 8, 4321–4325. doi: 10.1093/nar/ 8.19.4321
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proceedings of the National Academy of Sciences of the United States of America 76, 5269–5273. doi: 10.1073/pnas.76.10.5269
Ninh Thuan NT, Bigirimana J, Roumen E, Van Der Straeten D, Höfte M (2006) Molecular and pathotype analysis of the rice blast fungus in North Vietnam. European Journal of Plant Pathology 114, 381–396. doi: 10.1007/s10658-006-0002-8
Ou H (1980) Pathogen variability and host resistance in rice blast disease. Annual Review of Phytopathology 18, 167–187. doi: 10.1146/annurev. py.18.090180.001123
Ou H (1985) ‘Rice diseases.’ 2nd edn.(Commonwealth Mycological Institute: Kew, UK)
Park SY, Milgroom MG, Han SS, Kang S, Lee YH (2003) Diversity of pathotypes and DNA fingerprint haplotypes in populations of Magnaporthe grisea in Korea over two decades. Phytopathology 93, 1378–1385. doi: 10.1094/PHYTO.2003.93.11.1378
Piotti E, Rigano MM, Rodino D, Rodolfi M, Castiglione A, Picco M, Sala F (2005) Genetic structure of Pyricularia grisea (Cooke) Sacc. Isolates from Italian paddy fields. Journal of Phytopathology 153, 80–86. doi: 10.1111/j.1439-0434.2005.00932.x
Rohlf FJ (1998) ‘NTSYS — PC version 2.0. Numerical taxonomy and multivariate analysis system.’ (Exeter Software: Setauket, NY)
Roumen E, Levy M, Notteghem JL (1997) Characterization of the European pathogen population Magnaporthe grisea by DNA fingerprinting and pathotype analysis. European Journal of Plant Pathology 103, 363–371. doi: 10.1023/A:1008697728788
Sneath PHA, Sokal RR (1973) ‘Numerical taxonomy. The principles and practice of numerical classification.’ (W. H. Freeman: San Francisco, CA)
Sone T, Suto M, Tomita F (1993) Host species-specific repetitive DNA sequence in the genome of Magnaporthe grisea the rice blast fungus. Bioscience, Biotechnology, and Biochemistry 57, 1228–1230.
Swofford DL (2000) ‘PAUP 4.0. Phylogenetic analysis using parsimony and other methods.’ (Sinauer Associates: Sunderland, MA)
Tuite J (1969) ‘Plant pathological methods. Fungi and bacteria.’ (Burgess Publishing: Minneapolis, MN)
Valent B, Chumley FG (1991) Molecular genetic analysis of the rice blast fungus, Magnaporthe grisea. Annual Review of Phytopathology 29, 443–467. doi: 10.1146/annurev.py.29.090191.002303
Valent B, Crawford MS, Weaver CG, Chumley FG (1986) Genetic studies of fertility and pathogenicity in Magnaporthe grisea (Pyricularia grisea). Iowa State Journal of Research 60, 569–594.
Xia JQ, Correll JC, Lee FN, Marchetti MA, Rhodes DD (1993) DNA fingerprinting to examine microgeographic variation in the Magnaporthe grisea (Pyricularia grisea) population in two rice fields in Arkansas. Phytopathology 83, 1029–1035. doi: 10.1094/ Phyto-83-1029
Xia JQ, Correll JC, Lee FN, Ross WJ, Rhoads DD (2000) Regional population diversity of Pyricularia grisea in Arkansas and the influence of the host selection. Plant Disease 84, 877–884. doi: 10.1094/ PDIS.2000.84.8.877
Zeigler RS (1998) Recombination in Magnaporthe grisea. Annual Review of Phytopathology 36, 249–276. doi: 10.1146/annurev.phyto.36.1.249
Zeigler RS, Thome J, Nelson RJ, Levy M, Correa-Victoria F (1994) Lineage exclusion: a proposal for linking blast populations analysis to resistance breeding. In ‘Rice blast disease’. (Eds RS Zeigler, SA Leong, PS Teng) pp. 267–292. (CABI/IRRI: Wallingford, UK)
Zeigler RS, Cuoc LX, Scout RP, Bernardo MA, Chen DH, Valent B, Nelson RJ (1995) The relationship between lineage and virulence in Pyricularia grisea in the Philippines. Phytopathology 85, 443–451. doi: 10.1094/Phyto-85-443
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Consolo, V.F., Cordo, C.A. & Salerno, G.L. DNA fingerprint and pathotype diversity of Pyricularia oryzae populations from Argentina. Australasian Plant Pathology 37, 357–364 (2008). https://doi.org/10.1071/AP08010
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1071/AP08010