Abstract
Recovery of cardiac function is the holy grail of heart failure therapy yet is infrequently observed and remains poorly understood. In this study, we performed single-nucleus RNA sequencing from patients with heart failure who recovered left ventricular systolic function after left ventricular assist device implantation, patients who did not recover and non-diseased donors. We identified cell-specific transcriptional signatures of recovery, most prominently in macrophages and fibroblasts. Within these cell types, inflammatory signatures were negative predictors of recovery, and downregulation of RUNX1 was associated with recovery. In silico perturbation of RUNX1 in macrophages and fibroblasts recapitulated the transcriptional state of recovery. Cardiac recovery mediated by BET inhibition in mice led to decreased macrophage and fibroblast Runx1 expression and diminished chromatin accessibility within a Runx1 intronic peak and acquisition of human recovery signatures. These findings suggest that cardiac recovery is a unique biological state and identify RUNX1 as a possible therapeutic target to facilitate cardiac recovery.






Similar content being viewed by others
Data availability
Raw sequencing files and processed normalized data can be found on the Gene Expression Omnibus (GSE226314). Donors were used from published data (GSE183852). All other data supporting the findings in this study are included in the main article and associated files. Source data are provided with this paper.
Code availability
All scripts used for analysis in this manuscript can be found on GitHub (https://github.com/jamrute/2023_Amrute_et_al_NatureCVR_CardiacRecovery).
References
Roger, V. L. Epidemiology of heart failure. Circ. Res. 113, 646 (2013).
Jessup, M. & Brozena, S. Heart failure. N. Engl. J. Med. 348, 2007–2018 (2003).
Topkara, V. K. et al. Myocardial recovery in patients receiving contemporary left ventricular assist devices: results from the Interagency Registry for Mechanically Assisted Circulatory Support (INTERMACS). Circ. Heart Fail. 9, e003157 (2016).
Burkhoff, D., Topkara, V. K., Sayer, G. & Uriel, N. Reverse remodeling with left ventricular assist devices. Circ. Res. 128, 1594–1612 (2021).
Selzman, C. H. et al. Bridge to removal: a paradigm shift for left ventricular assist device therapy. Ann. Thorac. Surg. 99, 360 (2015).
Givertz, M. M. & Mann, D. L. Epidemiology and natural history of recovery of left ventricular function in recent onset dilated cardiomyopathies. Curr. Heart Fail. Rep. 10, 321–330 (2013).
Kanwar, M. K. et al. Clinical myocardial recovery in advanced heart failure with long term left ventricular assist device support. J. Heart Lung Transplant. 41, 1324–1334 (2022).
Shepherd, C. W. & While, A. E. Cardiac rehabilitation and quality of life: a systematic review. Int. J. Nurs. Stud. 49, 755–771 (2012).
Tseliou, E. et al. Biology of myocardial recovery in advanced heart failure with long-term mechanical support. J. Heart Lung Transplant. 41, 1309–1323 (2022).
McGuire, A. L. et al. The road ahead in genetics and genomics. Nat. Rev. Genet. 21, 581–596 (2020).
Eraslan, G. et al. Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function. Science 376, eabl4290 (2022).
Litviňuková, M. et al. Cells of the adult human heart. Nature 588, 466–472 (2020).
Koenig, A. L. et al. Single-cell transcriptomics reveals cell-type-specific diversification in human heart failure. Nat. Cardiovasc. Res. 1, 263–280 (2022).
Chaffin, M. et al. Single-nucleus profiling of human dilated and hypertrophic cardiomyopathy. Nature 608, 174–180 (2022).
Tucker, N. R. et al. Transcriptional and cellular diversity of the human heart. Circulation 142, 466–482 (2020).
Nicin, L. et al. A human cell atlas of the pressure-induced hypertrophic heart. Nat. Cardiovasc. Res. 1, 174–185 (2022).
Amrute, J. M. et al. Targeting the immune-fibrosis axis in myocardial infarction and heart failure. Preprint at https://www.biorxiv.org/content/10.1101/2022.10.17.512579v1 (2022).
Hall, J. L. et al. Molecular signature of recovery following combination left ventricular assist device (LVAD) support and pharmacologic therapy. Eur. Heart J. 28, 613–627 (2007).
Yang, K. C. et al. Deep RNA sequencing reveals dynamic regulation of myocardial noncoding RNAs in failing human heart and remodeling with mechanical circulatory support. Circulation 129, 1009–1021 (2014).
Drakos, S. G. et al. Distinct transcriptomic and proteomic profile specifies patients who have heart failure with potential of myocardial recovery on mechanical unloading and circulatory support. Circulation 147, 409–424 (2022).
Kamimoto, K. et al. Dissecting cell identity via network inference and in silico gene perturbation. Nature 614, 742–751 (2023).
Alexanian, M. et al. A transcriptional switch governs fibroblast activation in heart disease. Nature 595, 438–443 (2021).
Gupta, D. K. et al. Assessment of myocardial viability and left ventricular function in patients supported by a left ventricular assist device. J. Heart Lung Transplant. 33, 372–381 (2014).
Burke, M. A. & Givertz, M. M. Assessment and management of heart failure after left ventricular assist device implantation. Circulation 129, 1161–1166 (2014).
Bajpai, G. et al. Tissue resident CCR2− and CCR2+ cardiac macrophages differentially orchestrate monocyte recruitment and fate specification following myocardial injury. Circ. Res. 124, 263–278 (2019).
Zaman, R. et al. Selective loss of resident macrophage-derived insulin-like growth factor-1 abolishes adaptive cardiac growth to stress. Immunity 54, 2057–2071 (2021).
Dick, S. A. et al. Self-renewing resident cardiac macrophages limit adverse remodeling following myocardial infarction. Nat. Immunol. 20, 29–39 (2019).
Garcia-Alonso, L., Holland, C. H., Ibrahim, M. M., Turei, D. & Saez-Rodriguez, J. Benchmark and integration of resources for the estimation of human transcription factor activities. Genome Res. 29, 1363–1375 (2019).
Asleh, R., Amir, O. & Kushwaha, S. S. Dynamics of myocardial fibrosis after left ventricular assist device implantation: should speeding up the scar have us scared stiff? Eur. J. Heart Fail. 23, 335–338 (2021).
Wilcox, J. E. et al. ‘Targeting the heart’ in heart failure: myocardial recovery in heart failure with reduced ejection fraction. JACC Heart Fail. 3, 661–669 (2015).
Stratton, M. S. et al. Dynamic chromatin targeting of BRD4 stimulates cardiac fibroblast activation. Circ. Res. 125, 662 (2019).
Aghajanian, H. et al. Targeting cardiac fibrosis with engineered T cells. Nature 573, 430–433 (2019).
Bocchi, V. D. et al. The coding and long noncoding single-cell atlas of the developing human fetal striatum. Science 372, eabf5759 (2021).
Rose, E. A. et al. Long-term use of a left ventricular assist device for end-stage heart failure. N. Engl. J. Med. 345, 1435–1443 (2001).
Miller, L., Birks, E., Guglin, M., Lamba, H. & Frazier, O. H. Use of ventricular assist devices and heart transplantation for advanced heart failure. Circ. Res. 124, 1658–1678 (2019).
Dharmavaram, N. et al. National trends in heart donor usage rates: are we efficiently transplanting more hearts? J. Am. Heart Assoc. 10, 19655 (2021).
Bowen, R. E. S., Graetz, T. J., Emmert, D. A. & Avidan, M. S. Statistics of heart failure and mechanical circulatory support in 2020. Ann. Transl. Med. 8, 827 (2020).
Drakos, S. G. et al. Bridge to recovery: understanding the disconnect between clinical and biological outcomes. Circulation 126, 230 (2012).
Lenneman, A. J. & Birks, E. J. Treatment strategies for myocardial recovery in heart failure. Curr. Treat. Options Cardiovasc. Med. 16, 287 (2014).
Halliday, B. P. et al. Withdrawal of pharmacological treatment for heart failure in patients with recovered dilated cardiomyopathy (TRED-HF): an open-label, pilot, randomised trial. Lancet 393, 61–73 (2019).
Bottle, A., Faitna, P., Aylin, P. P. & Cowie, M. R. Original research: five-year outcomes following left ventricular assist device implantation in England. Open Heart 8, 1658 (2021).
Wang, L. et al. Single-cell reconstruction of the adult human heart during heart failure and recovery reveals the cellular landscape underlying cardiac function. Nat. Cell Biol. 22, 108–119 (2020).
Birks, E. J. et al. Prospective multicenter study of myocardial recovery using left ventricular assist devices (RESTAGE-HF [remission from stage D heart failure]): medium-term and primary end point results. Circulation 142, 2016–2028 (2020).
Zhang, J. & Narula, J. Molecular biology of myocardial recovery. Surg. Clin. North Am. 84, 223–242 (2004).
Klotz, S., Jan Danser, A. H. & Burkhoff, D. Impact of left ventricular assist device (LVAD) support on the cardiac reverse remodeling process. Prog. Biophys. Mol. Biol. 97, 479–496 (2008).
Wohlschlaeger, J. et al. Reverse remodeling following insertion of left ventricular assist devices (LVAD): a review of the morphological and molecular changes. Cardiovasc. Res. 68, 376–386 (2005).
Weinheimer, C. J. et al. Load-dependent changes in left ventricular structure and function in a pathophysiologically relevant murine model of reversible heart failure. Circ. Heart Fail. 11, e004351 (2018).
Chaudhary, K. W. et al. Altered myocardial Ca2+. cycling after left ventricular assist device support in the failing human heart. J. Am. Coll. Cardiol. 44, 837–845 (2004).
Ambardekar, A. V. et al. Incomplete recovery of myocyte contractile function despite improvement of myocardial architecture with left ventricular assist device support. Circ. Heart Fail. 4, 425–432 (2011).
Bajpai, G. et al. The human heart contains distinct macrophage subsets with divergent origins and functions. Nat. Med. 24, 1234–1245 (2018).
Epelman, S., Lavine, K. J. & Randolph, G. J. Origin and functions of tissue macrophages. Immunity 41, 21–35 (2014).
Lavine, K. J. et al. Distinct macrophage lineages contribute to disparate patterns of cardiac recovery and remodeling in the neonatal and adult heart. Proc. Natl Acad. Sci. USA 111, 16029–16034 (2014).
Wong, N. R. et al. Resident cardiac macrophages mediate adaptive myocardial remodeling. Immunity 54, 2072–2088 (2021).
Kuppe, C. et al. Decoding myofibroblast origins in human kidney fibrosis. Nature 589, 281–286 (2021).
Khalil, H. et al. Fibroblast-specific TGF-β–Smad2/3 signaling underlies cardiac fibrosis. J. Clin. Invest. 127, 3770–3783 (2017).
Medzhitov, R. Origin and physiological roles of inflammation. Nature 454, 428–435 (2008).
Tzahor, E. & Dimmeler, S. A coalition to heal—the impact of the cardiac microenvironment. Science 377, eabm4443 (2022).
Sood, R., Kamikubo, Y. & Liu, P. Role of RUNX1 in hematological malignancies. Blood 129, 2070–2082 (2017).
Chen, M. J., Yokomizo, T., Zeigler, B. M., Dzierzak, E. & Speck, N. A. Runx1 is required for the endothelial to haematopoietic cell transition but not thereafter. Nature 457, 887–891 (2009).
Koth, J. et al. Runx1 promotes scar deposition and inhibits myocardial proliferation and survival during zebrafish heart regeneration. Development 147, dev186569 (2020).
Hu, B. et al. Origin and function of activated fibroblast states during zebrafish heart regeneration. Nat. Genet. 54, 1227–1237 (2022).
Amrute, J. M. et al. Cell specific peripheral immune responses predict survival in critical COVID-19 patients. Nat. Commun. 13, 882 (2022)
Kong, Y. & Yu, T. A deep neural network model using random forest to extract feature representation for gene expression data classification. Sci. Rep. 8, 16477 (2018).
Khouri-Farah, N., Guo, Q., Morgan, K., Shin, J. & Li, J. Y. H. Integrated single-cell transcriptomic and epigenetic study of cell state transition and lineage commitment in embryonic mouse cerebellum. Sci. Adv. 8, eabl9156 (2022).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wang, F. et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 14, 22–29 (2012).
Setty, M. et al. Characterization of cell fate probabilities in single-cell data with Palantir. Nat Biotechnol. 37, 451–460 (2019).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation (Camb). 2, 100141 (2021).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Acknowledgements
K.L. is supported by the Washington University in St. Louis Rheumatic Diseases Research Resource-Based Center Grant (National Institutes of Health (NIH) P30AR073752, NIH R01 HL138466, R01 HL139714, R01 HL151078, R01 HL161185 and R35 HL161185); the Leducq Foundation Network (20CVD02); the Burroughs Welcome Fund (1014782); the Children’s Discovery Institute of Washington University and St. Louis Children’s Hospital (CH-II-2015-462, CH-II-2017-628 and PM-LI-2019-829); the Foundation of Barnes-Jewish Hospital (8038-88); and generous gifts from Washington University School of Medicine. S.D. is supported by the American Heart Association Heart Failure Strategically Focused Research Network (grant 16SFRN29020000); National Heart, Lung, and Blood Institute (NHLBI) RO1 grant HL135121, NHLBI RO1 grant HL132067, NHLBI R01 grant HL156667 and NHLBI R01 grant HL151924; Merit Review Award I01 CX002291, US Department of Veterans Affairs; and Nora Eccles Treadwell Foundation grants. J.M.A. is supported by an American Heart Association Predoctoral Fellowship (826325) and the Washington University in St. Louis School of Medicine Medical Scientist Training Program. P.M. is supported by an American Heart Association Postdoctoral Fellowship (916955). T.S. is supported by an American Heart Association Postdoctoral Fellowship (23POST1019351). Figures 1a, 2d and 6c,j were created with BioRender. Histology was performed in the Digestive Diseases Research Core Centers Advanced Imaging and Tissue Analysis Core, supported by grant P30 DK52574. Imaging was performed in the Washington University Center for Cellular Imaging, which is funded, in part, by the Children’s Discovery Institute of Washington University and St. Louis Children’s Hospital (CDI-CORE-2015-505 and CDI-CORE-2019-813) and the Foundation for Barnes-Jewish Hospital (3770). We thank the Genome Technology Access Center at the McDonnell Genome Institute at Washington University School of Medicine for help with genomic analysis. The center is partially supported by National Cancer Institute Cancer Center Support Grant P30 CA91842 to the Siteman Cancer Center. This publication is solely the responsibility of the authors and does not necessarily represent the official views of the National Center for Research Resources or the NIH. The authors are grateful to the donor families for their generosity, and DonorConnect (https://www.donorconnect.life/), Salt Lake City, Utah, for facilitating the work of our research team members acquiring myocardial tissue in the operating rooms of several hospitals in Utah and several other states. The authors are grateful to the University of Utah cardiothoracic surgery team for the invaluable help acquiring the myocardial tissue from chronic heart failure patients.
Author information
Authors and Affiliations
Contributions
S.D. contributed to LVAD sample acquisition and clinical phenotyping. P.M., L.L., A.B. and A.K. isolated nuclei for snRNA-seq. J.A. performed all computational analysis. J.M.A. and K.K. performed GRN analysis and in silico transcription factor perturbation analysis. J.A., L.L., P.M., L.S., D.S., F.K., T.S.S. and B.K. performed RNA in situ hybridization and immunohistochemistry experiments and analyzed and processed images. J.A., F.L., R.K., S.M., D.M., S.D. and K.L. assisted in the interpretation of the data. J.A. made all figures, and J.A. and K.L. drafted the manuscript. K.L. is responsible for all aspects of this manuscript, including experimental design, data analysis and manuscript production. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
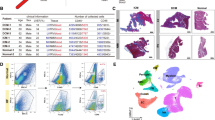
Extended Data Fig. 1 Quality control metrics.
nCount_RNA, nFeature_RNA, percent.mt, and scrublet doublet score split by (A) condition and (B) cell type.
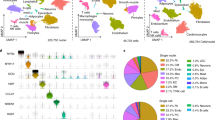
Extended Data Fig. 2 Global clustering.
(A) Heatmap of top10 marker gens for each cell type identified via DE analysis. (B) DotPlot for cell type gene set scores from (A) where the x-axis is cell type gene signature and y-axis is the cluster. (C) Gene set z-scores for top gene markers for each cell type plotted in the UMAP embedding. (D) Cell type composition for each of the patient samples.
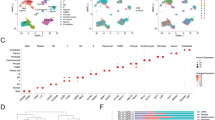
Extended Data Fig. 3 Pseudobulk DE analysis to unravel cardiac recovery.
(A) Pseudobulk DE analysis in each cell type in 3 comparison groups: pre-LVAD HF vs donor, RR-post vs donor, and RR-post vs pre-LVAD HF. Red dots indicate statistically significant genes (adjusted p-value < 0.05). (B) Total number of statistically significant (adjusted p-value < 0.05 and log2FC > 0.58) per cell type in comparisons from (A). (C) Number of overlapping genes in five major cell populations which are up and down in the comparisons from (A). Red number is the number of cardiac recovery genes. P-values calculated using Wald test adjusted for multiple corrections.
Extended Data Fig. 4 Cardiac recovery overlap amongst cell types.
UpSet plot showing overlap in cardiac recovery genes from (Fig. 2) in five major cell populations which are (A) up and (B) down in cardiac recovery.
Extended Data Fig. 5 Cell-specific pseudobulk analysis.
Pseudobulk PCA analysis in each cell type colored by five conditions (donor, U-pre, U-post, RR-pre, and RR-post).
Extended Data Fig. 6 ABRA expression enriched in unloaded group.
(A) DotPlot of cardiomyocyte specific recovery up- and down signature grouped by CM cell states. (B) Density plot of ABRA expression in UMAP embedding. (C) DotPlot of ABRA expression in cardiomyocytes grouped by condition. (D) RNAscope images of ABRA in 5 conditions and scale bar is 100 um. (E) RNAscope images quantified across an array of patients. N = 37 biologically independent samples and p-values calculated using Wald test adjusted for multiple corrections; donor vs U-pre (*P = 0.023), U-pre vs RR-pre (***P < 0.0001), U-pre vs RR-post (***P = 0.0003), U-post vs RR-pre (***P = 0.0007), U-post vs RR-post (*P = 0.0188), and RR-pre vs RR-post (***P = 0.0007).
Extended Data Fig. 7 Macrophage diversity in recovery.
(A) Gene set z-scores for top gene markers for each cell state plotted in the UMAP embedding. (B) Enrich GO using compareclusters from cluster Prolifer across macrophage cell states. P-value calculated using Fisher exact test. (C) WikiPathways enriched in cardiac recovery. P-value calculated using Fisher exact test. (D) Paired comparison of Mac 2 cluster composition at patient level split by U and RR group from biologically independent samples. (E) DoRothEA TF enrichment analysis in U-post and RR-post zoomed in on some key differentially enriched TFs. (F) Overlap between Runx1 target genes and DE genes between U-pre and RR-pre with heatmap of respective genes split by condition.
Extended Data Fig. 8 Fibroblast diversity in recovery.
(A) Gene set z-scores for top gene markers for each cell state plotted in the UMAP embedding. (B) DotPlot for cell type gene set scores from (A) where the x-axis is cell type gene signature and y-axis is the cluster. (C) Enrich GO using compareclusters from cluster profiler across fibroblast cell states. P-value calculated using Fisher exact test.
Extended Data Fig. 9 CellOracle simulation in myeloid cells in TAC.
(A) Myeloid cell states, (B) Marker genes for cell states, (C) Cell state composition and cell density plots in TAC and TAC + JQ1, and (D) Cell oracle Runx1 KO perturbation score with vector field.
Extended Data Fig. 10 CellOracle simulation in fibroblasts in TAC.
(A) Fibroblast cell states, (B) Marker genes for cell states, (C) Cell state composition and cell density plots in TAC and TAC + JQ1, and (D) Cell oracle Runx1 KO perturbation score with vector field.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Supplementary Tables
Supplementary Tables 1–22.
Source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Fig. 5
Statistical source data
Source Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 7
Statistical source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Amrute, J.M., Lai, L., Ma, P. et al. Defining cardiac functional recovery in end-stage heart failure at single-cell resolution. Nat Cardiovasc Res 2, 399–416 (2023). https://doi.org/10.1038/s44161-023-00260-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44161-023-00260-8
- Springer Nature Limited
This article is cited by
-
Macrophages in cardiovascular diseases: molecular mechanisms and therapeutic targets
Signal Transduction and Targeted Therapy (2024)
-
Pharmacological inhibition of RUNX1 reduces infarct size after acute myocardial infarction in rats and underlying mechanism revealed by proteomics implicates repressed cathepsin levels
Functional & Integrative Genomics (2024)
-
Myocardial Recovery and Relapse in Heart Failure With Improved Ejection Fraction
Current Treatment Options in Cardiovascular Medicine (2024)