Abstract
Salmonella enterica and Escherichia coli are major food-borne human pathogens, and their genomes are routinely sequenced for clinical surveillance. Computational pipelines designed for analyzing pathogen genomes should both utilize the most current information from annotation databases and increase the coverage of these databases over time. We report the development of the GEA pipeline to analyze large batches of E. coli and S. enterica genomes. The GEA pipeline takes as input paired Illumina raw reads files which are then assembled followed by annotation. Alternatively, assemblies can be provided as input and directly annotated. The pipeline provides predictive genome annotations for E. coli and S. enterica with a focus on the Center for Genomic Epidemiology tools. Annotation results are provided as a tab delimited text file. The GEA pipeline is designed for large-scale E. coli and S. enterica genome assembly and characterization using the Center for Genomic Epidemiology command-line tools and high-performance computing. Large scale annotation is demonstrated by an analysis of more than 14,000 Salmonella genome assemblies. Testing the GEA pipeline on E. coli raw reads demonstrates reproducibility across multiple compute environments and computational usage is optimized on high performance computers.
Similar content being viewed by others
Introduction
Salmonella enterica (hereafter Salmonella) are estimated to cause at least 1 million illnesses in the United States each year1. Escherichia coli are ubiquitous in a wide variety of environments relevant to food safety including food animal gastrointestinal systems, animal production sites, human gastrointestinal systems, meats, and manure impacted soils. A small but clinically important sub-set of E. coli are pathogenic. The ubiquitous nature of E. coli contributes to their relevance beyond food safety. Due to their prominence and small genome size, Salmonella and E. coli are also two of the top organisms with available whole genome sequencing read archives (https://www.ncbi.nlm.nih.gov/sra?term=(%22public%22%5BAccess%5D)%20AND%20%22genomic%22%5BSource%5D) and assemblies (https://www.ncbi.nlm.nih.gov/genome/browse#!/overview/). Such large datasets often rely on high-performance computing to accelerate computational tasks via increased RAM, threads, and parallelization2. Large datasets can benefit from data analysis pipelines, which process many input files with an initial set of user specifications and distill the results to a small number of organized outputs for interpretation3.
Useful pipelines for epidemiologic annotation should take advantage of the most up-to-date reference information available. Actively curated reference databases meet this need by rapidly incorporating newly released genomic data. The interplay between these two dependencies can be thought of as a positive feedback loop wherein 1. Running the pipeline on new strains improves the database coverage and quality by exposing knowledge gaps and 2. The database improvement leads to more accurate search hits when running the pipeline on new strains.
Here, we introduce the GEA pipeline. GEA stands for Gammaproteobacteria Epidemiologic Annotation. Analyses central to the GEA pipeline are those using Center for Genomic Epidemiology (CGE) developed tools4. The databases for these tools to search are updated frequently, facilitating the positive feedback loop between our pipeline and these databases. Several existing pipelines utilize CGE developed tools5,6,7,8,9. But we are unaware of any other published pipelines, which use FimTyper10, MLST11, PlasmidFinder12, ResFinder13, SerotypeFinder14, and VirulenceFinder15 in tandem.
Another important feature of the pipeline is the use of a container. Containers allow for compute mobility16 and provide an increased level of reproducibility17. The container housing the software tools for running the GEA pipeline also takes advantage of the Scientific File System (SCIF)18, providing independent mount points to different apps in the container with incompatible environmental requirements.
Prior versions of the GEA pipeline have been used in published research19,20,21 in food safety, risk assessment, antimicrobial resistance gene transfer, and virulence research at the US Meat Animal Research Center. This demonstrates the utility of the pipeline and the benefit of the distilled annotation summary across hundreds of genomes. This paper describes the methods used for creating the pipeline and provides a kilo-scale demonstration of the pipeline on S. enterica assemblies. The pipeline is available from github.com/Phylloxera/GEA-dev.
Results
Computer resource usage
Table 1 provides the usage summary for testing. The run time varied from 5.04 to 34.56 h (3.15 to 21.6 min-per-E.-coli-library). Assembly accounted for 89–95% of pipeline run time.
Output
The principal output of the GEA pipeline is the tab-delimited file, metadata.txt. The metadata can be opened in Excel. The number of rows corresponds to the number of assemblies annotated and the number of columns is dataset dependent. The annotation quality is expected to improve as the CGE database coverage improves over time. The metadata from the test dataset of 96 E. coli raw read libraries contained 444 columns and the metadata from the demonstration dataset of 14,310 Salmonella assemblies had 597 columns. Table 2 summarizes the components of the two metadata files. Tables 3 and 4 provide single annotation snapshots from VirulenceFinder15 E. coli and ResFinder13 Salmonella respectively, while Table 5 provides a snapshot of the all tools summary from E. coli. Table 5 suggests a relationship between FIM type and Antimicrobial Resistance Gene (ARG) content for the 96 E. coli libraries test dataset. The complete summary output file (GEA_ecoli_test_Ceres_metadata.txt) from USDA Ceres22 is provided in Supplementary Data S1.
Demonstration
The pipeline ran on the 14,310 S. enterica assemblies in ~ 72 h (~ 18 s-per-S.-enterica-assembly) on the USDA/Mississippi State University Atlas cluster22. The complete summary output file is in GEA_senterica_demo_Atlas_metadata.txt in Supplementary Data S1.
Discussion
A diverse set of bioinformatic tools has been developed for phenotypic prediction based on genomic data, especially for human pathogens. These tools often grow out of the requirements sought by a group of researchers. In the case of GEA, these included the desire to assemble and run CGE tools at the command-line on large numbers of strains with a single summary output and to have genome assemblies in a single directory ready for submission to NCBI.
Users have reported different results with the same input data, sometimes with analyses conducted many months apart. This is suspected to be caused by updates in the actively curated CGE databases. This has been confirmed in some instances. In the interest of reproducibility, the user has the option to update their local copy of the CGE databases and use the most up-to-date versions, or to leave the databases static for reproducibility across independent computational runs. Reproducibility has long been an aspiration of scientific analysis, however database dependent analyses may demonstrably benefit from non-reproducibility as database coverage increases with the passage of time.
In our testing phase, we sought to have reproducibility across multiple computational environments and this was largely achieved. The only difference in the outputs across high performance computers was due to the contig names assigned by shovill (https://github.com/tseemann/shovill). In all these cases, the length, coverage, and Pilon23 name were identical whereas shovill assigned contig integers differed by 1. Furthermore, shovill contig names included the date assembled, an additional possible source of discrepancy for runs taking place on different days. E.g., contig00205 len = 509 cov = 31.1 corr = 0 origname = NODE_348_length_509_cov_31.119617_pilon sw = shovill-spades/1.1.0 date = 20231107 on Ceres vs contig00204 len = 509 cov = 31.1 corr = 0 origname = NODE_348_length_509_cov_31.119617_pilon sw = shovill-spades/1.1.0 date = 20231101 on Moose (discrepancies in bold; example from row 15, column 128 of GEA_ecoli_test_Ceres_metadata.txt in Supplementary Data S1). The identical coverage and Pilon designation indicates that, in all cases, these were identical contigs, but that the final integer contig name assignment in shovill may not be deterministic. Additionally, E. coli test data Assembly_bp and Ncontigs statistics were identical across computing environments. The other discrepancy was caused by insufficient memory being available to Skesa24 on the Desktop computer causing two libraries to not assemble resulting in missing Skesa plasmid annotations (rows 33 and 35, columns 21 and 22 of GEA_ecoli_test_Ceres_metadata.txt in Supplementary Data S1). Importantly, these assembly failures were documented by the GEA pipeline log, which alerted the Desktop user. Apart from these two discrepancy sources (shovill final contig naming and Skesa memory requirements not being met), we expect that GEA will provide reproducible results for a given version of the pipeline and databases.
GEA has clear advantages and limitations relative to tools with similar goals. First, long available tools, such as nullarbor (https://github.com/tseemann/nullarbor) and TORMES7 have the distinction of an active user base, citations, and more time under development. Bacannot25 is a newer tool, which is container based like GEA. Software containerization increases reproducibility over OS specific source-compile-install-run and cross-platform package manager methodologies17. RSYD-BASIC26 is also a newer tool which produces a tabular output somewhat like that produced by GEA. The lack of a web server option is a limitation of GEA. However, having a batch command-line implementation of CGE tools was a central functionality driving the development of GEA. GEA also lacks phylogenetic methods, except to the extent that typing predictions are phylogenetically informative. Currently, GEA is only indicated for E. coli and S. enterica. The strongest advantages of GEA are a single dependency (Apptainer27), batch processing in an HPC environment proven by hundreds of successful analyses of illumina raw read libraries21, and the successful demonstration at the kilo-scale of a processing rate of ~ 18-s-per-S.-enterica-assembly as demonstrated in this report.
Shovill and Skesa can both handle low levels of contamination. The pipeline has been tested with a diversity of libraries from three different Illumina sequencing platforms at the USMARC Core Lab, but exhaustive testing on the types and degrees of contamination has not been conducted. Regardless, preprocessing or quality control of libraries should be unnecessary.
The Gammaproteobacteria name derivation of the GEA pipeline implies a much broader set of pathogens than are currently included. One noticeable impact of incorporating new species is a substantially smaller set of annotations for the added species, at least initially. This is because all other species have fewer applicable CGE tools relative to E. coli. This is surmountable in instances where there is a non-CGE tool available for the task (e.g. GEA uses SeqSero228 for serotyping Salmonella since the CGE tool, SerotypeFinder14, does not serotype Salmonella). Other enhancements, which could be incorporated into GEA in the future include long-read sequence inputs, identification of additional genotypes/phenotypes as new CGE tools are released, and downsampling reads to accelerate Skesa assembly as is done with shovill. GEA is now available for other users with institutional HPCs for rapid characterization of large batches of E. coli and Salmonella genomes from diverse sample sources. GEA is available for download from github.com/Phylloxera/GEA-dev.
Methods
Pipeline
The GEA pipeline implements the following steps in sequential order (Fig. 1). GEA processes the user command-line options and inputs. If the inputs are raw reads, the workflow proceeds to Assembly.
-
1.
Assembly is first carried out by Shovill(https://github.com/tseemann/shovill) followed by SKESA24. Skesa is used for identifying complete plasmids and can circularize some novel small plasmids not yet on the plasmidfinder database.
-
2.
Epidemiologic Prediction is conducted on shovill assemblies or user supplied fasta contigs. Local copies of the CGE tool and 5 loci databases (https://github.com/Phylloxera/5loci) are updated by the pipeline unless otherwise specified by the user. The databases are stored locally in the user’s home directory (e.g. $HOME/share/resfinder). BLAST29 is used as the search method for the CGE tools and to query the custom 5 loci databases.
-
a.
PlasmidFinder12 (CGE)
-
b.
MLST11 (CGE) is run. If the user does not specify the taxon, mlst is run against both databases and the species is determined by GEA.
-
c.
Serotyping is conducted using SerotypeFinder14 (CGE—E. coli) or SeqSero228 (Salmonella).
-
d.
Resfinder413 (CGE) is run against resfinder_db and pointfinder_db databases.
-
e.
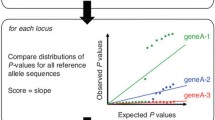
The 5 loci databases are queried with Blast.
-
f.
VirulenceFinder15 (CGE—E. coli)
-
g.
Ezclermont30 (E. coli)
-
h.
FimTyper10 (CGE—E. coli)
-
a.
-
3.
The results are compiled and written to the tab delimited output file, metadata.txt. GC and N50 summary statistics are calculated by stats.sh(https://jgi.doe.gov/data-and-tools/software-tools/bbtools/bb-tools-user-guide/statistics-guide/) during results compilation and additional summary statistics are extracted from the shovill logs.
GEA is written in Bash. The current version has several new features relative to prior development iterations used in previous work.
-
1.
The user can specify the taxon.
-
2.
The user can specify whether to update their local copy of the databases.
-
3.
The input data can be raw gzipped paired reads (fastq) or genome assemblies (fasta).
-
4.
The following new tools have been added: Ezclermont, FimTyper, VirulenceFinder, and the 5 loci databases.
-
5.
The container recipe utilizes SCIF18 for software environment modularity inside the software container.
-
6.
The container, and pipeline are made available via https://github.com/Phylloxera/GEA-dev.
The shovill pipeline utilizes SPAdes31, Velvet32, Lighter33, FLASh34, SAMtools35, BWA-MEM36, KMC37, seqtk(https://github.com/lh3/seqtk), pigz(https://zlib.net/pigz/), Pilon23, Trimmomatic38 and samclip(https://github.com/tseemann/samclip). The container with the needed software was created using Apptainer27.
Testing
GEA was tested on 3 linux high performance computers and a desktop computer with Hyper-V enabled on Windows 10 Professional (Table 1). The data used in testing were a single plate of 96 illumina E. coli raw read libraries from a long-term evolutionary study. Testing utilized the -u F option to query identical versions of the CGE databases and evaluate reproducibility across the compute environments.
Demonstration
To demonstrate the pipeline at kilo-scale, GEA was run on 14,310 Salmonella enterica genome assemblies released during October, 2023. The assemblies were downloaded on November 6, 2023 using datasets(https://www.ncbi.nlm.nih.gov/datasets) with download genome options: taxon 28901,–include genome,–exclude-atypical,–released-after 10/1/2023,–released-before 10/31/2023,–assembly-source GenBank, and–dehydrated. Fasta files were moved to a single folder to be used as input. GEA was run on the Atlas high performance computer system of Mississippi State University and the US Department of Agriculture22 on November 11, 2023, with options: -t senterica, -u F, -r 336:00:00, -m 360G, and -c 48. Initial tests predicted a run time of 2–5 days.
Data availability
Underlying data: The test dataset used during the current study can be made available from the corresponding author on reasonable request. The demonstration dataset used during the current study are publicly available from the National Center for Biotechnology Information. The assemblies can be downloaded using the publicly available software, datasets (https://www.ncbi.nlm.nih.gov/datasets), with download genome options: taxon 28901,–include genome,–exclude-atypical,–released-after 10/1/2023,–released-before 10/31/2023,–assembly-source GenBank, and–dehydrated. Extended data: USDA-NAL: Supplementary Data S1. This project contains the following extended data:–GEA_ecoli_test_Ceres_metadata.txt. GEA Pipeline summary output (metadata.txt) for 96 E. coli Illumina libraries used as test data.–GEA_senterica_demo_Atlas_metadata.txt GEA Pipeline summary output (metadata.txt) for 14,310 S. enterica assemblies used as demonstration data. Extended data are available under the terms of the Creative Commons Attribution 4.0 International license (CC-BY 4.0).
Code availability
The full pipeline (as an apptainer container) and files required for building the pipeline from scratch are available from: [https://github.com/Phylloxera/GEA-dev]. The version used in all analyses herein is 0.0.1-alpha.1. GEA License: [https://github.com/Phylloxera/GEA-dev/blob/main/LICENSE]. The 5 loci BLAST databases are available from: [https://github.com/Phylloxera/5loci]. 5 loci License: [https://github.com/Phylloxera/5loci/blob/main/LICENSE].
References
Scallan, E. et al. Foodborne illness acquired in the United States–major pathogens. Emerg. Infect. Dis. 17, 7–15. https://doi.org/10.3201/eid1701.P11101 (2011).
Fjukstad, B. & Bongo, L. A. A review of scalable bioinformatics pipelines. Data Sci. Eng. 2, 245–251. https://doi.org/10.1007/s41019-017-0047-z (2017).
Leipzig, J. A review of bioinformatic pipeline frameworks. Brief. Bioinform. 18, 530–536. https://doi.org/10.1093/bib/bbw020 (2017).
Center for Genomic Epidemiology Repositories [internet]. [cited 29 November 2023]. https://bitbucket.org/genomicepidemiology/workspace/repositories/
Chukamnerd, A. et al. BacSeq: A user-friendly automated pipeline for whole-genome sequence analysis of bacterial genomes. Microorganisms. 11, 1769. https://doi.org/10.3390/microorganisms11071769 (2023).
Couvin, D., Stattner, E., Segretier, W., Cazenave, D. & Rastogi, N. simpiTB–a pipeline designed to extract meaningful information from whole genome sequencing data of Mycobacterium tuberculosis complex, allows to combine genomic, phylogenetic and clustering analyses in existing SITVIT databases. Infect. Genet. Evol. 113, 105466. https://doi.org/10.1016/j.meegid.2023.105466 (2023).
Quijada, N. M., Rodríguez-Lázaro, D., Eiros, J. M. & Hernández, M. TORMES: An automated pipeline for whole bacterial genome analysis. Bioinformatics 35, 4207–4212. https://doi.org/10.1093/bioinformatics/btz220 (2019).
Thomsen, M. C. F. et al. A bacterial analysis platform: An integrated system for analysing bacterial whole genome sequencing data for clinical diagnostics and surveillance. PloS One 11, e0157718. https://doi.org/10.1371/journal.pone.0157718 (2016).
Xavier, B. B. et al. BacPipe: A rapid, user-friendly whole-genome sequencing pipeline for clinical diagnostic bacteriology. IScience. 23, 100769. https://doi.org/10.1016/j.isci.2019.100769 (2020).
Roer, L. et al. Development of a web tool for Escherichia coli subtyping based on fimH alleles. J. Clin. Microbiol. 55, 2538–2543. https://doi.org/10.1128/jcm.00737-17 (2017).
Larsen, M. V. et al. Multilocus sequence typing of total-genome-sequenced bacteria. J. Clin. Microbiol. 50, 1355–1361. https://doi.org/10.1128/jcm.06094-11 (2012).
Carattoli, A. & Hasman, H. PlasmidFinder and in silico pMLST: Identification and typing of plasmid replicons in whole-genome sequencing (WGS). Horiz. Gene Transf. 285–294. (2020). https://doi.org/10.1007/978-1-4939-9877-7_20
Bortolaia, V. et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemoth. 75, 3491–3500. https://doi.org/10.1093/jac/dkaa345 (2020).
Joensen, K. G., Tetzschner, A. M. M., Iguchi, A., Aarestrup, F. M. & Scheutz, F. Rapid and easy in silico serotyping of Escherichia coli isolates by use of whole-genome sequencing data. J. Clin. Microbiol. 53, 2410–2426. https://doi.org/10.1128/jcm.00008-15 (2015).
Joensen, K. G. et al. Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J. Clin. Microbiol. 52, 1501–1510. https://doi.org/10.1128/jcm.03617-13 (2014).
Kurtzer, G. M., Sochat, V. & Bauer, M. W. Singularity: Scientific containers for mobility of compute. PloS One 12, e0177459. https://doi.org/10.1371/journal.pone.0177459 (2017).
Grüning, B. et al. Practical computational reproducibility in the life sciences. Cell Syst. 6, 631–635. https://doi.org/10.1016/j.cels.2018.03.014 (2018).
Sochat, V. The scientific filesystem. GigaScience giy023, 023. https://doi.org/10.1093/gigascience/giy023 (2018).
Guragain, M., Schmidt, J. W., Kalchayanand, N., Dickey, A. M. & Bosilevac, J. M. Characterization of Escherichia coli harboring colibactin genes (clb) isolated from beef production and processing systems. Sci. Rep. 12, 5305. https://doi.org/10.1038/s41598-022-09274-x (2022).
Guragain, M., Schmidt, J. W., Dickey, A. M. & Bosilevac, J. M. Distribution of extremely heat-resistant Escherichia coli in the beef production and processing continuum. J. Food Protect. 86, 100031. https://doi.org/10.1016/j.jfp.2022.100031 (2023).
Schmidt, J. W. et al. Twenty-four-month longitudinal study suggests little to no horizontal gene transfer in situ between third-generation cephalosporin-resistant Salmonella and third-generation cephalosporin-resistant Escherichia coli in a beef cattle feedyard. J. Food Protect. 85, 323–335. https://doi.org/10.4315/JFP-21-371 (2022).
SCINet Scientific Computing U.S. DEPARTMENT OF AGRICULTURE. [internet]. [cited 29 November 2023]. scinet.usda.gov
Walker, B. J. et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PloS One 9, e112963. https://doi.org/10.1371/journal.pone.0112963 (2014).
Souvorov, A., Agarwala, R. & Lipman, D. J. SKESA: Strategic k-mer extension for scrupulous assemblies. Genome Biol. 19, 153. https://doi.org/10.1186/s13059-018-1540-z (2018).
de Almeida, F. M., de Campos, T. A. & Pappas, G. J. Jr. Scalable and versatile container-based pipelines for de novo genome assembly and bacterial annotation. F1000Research 12, 1205. https://doi.org/10.12688/f1000research.139488.1 (2023).
Steinke, K. et al. RSYD-BASIC: A bioinformatics pipeline for routine sequence analysis and data processing of bacterial isolates for clinical microbiology. Access Microbiol. https://doi.org/10.1099/acmi.0.000646.v2 (2023).
Dykstra, D. Apptainer without setuid. arXiv 2208, 12106. [preprint] (2022). https://doi.org/10.48550/arXiv.2208.12106
Zhang, S. et al. SeqSero2: Rapid and improved Salmonella serotype determination using whole-genome sequencing data. Appl. Environ. Microbiol. 85, e01746-e1819. https://doi.org/10.1128/AEM.01746-19 (2019).
Camacho, C. et al. BLAST+: Architecture and applications. BMC Bioinform. 10, 44935. https://doi.org/10.1186/1471-2105-10-421 (2009).
Waters, N. R., Abram, F., Brennan, F., Holmes, A. & Pritchard, L. Easy phylotyping of Escherichia coli via the EzClermont web app and command-line tool. Access Microbiol. 2, acmi000143. https://doi.org/10.1099/acmi.0.000143 (2020).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477. https://doi.org/10.1089/cmb.2012.0021 (2012).
Zerbino, D. R. & Birney, E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829. https://doi.org/10.1101/gr.074492.107 (2008).
Song, L., Florea, L. & Langmead, B. Lighter: Fast and memory-efficient sequencing error correction without counting. Genome Biol. 15, 44939. https://doi.org/10.1186/s13059-014-0509-9 (2014).
Magoč, T. & Salzberg, S. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963. https://doi.org/10.1093/bioinformatics/btr507 (2011).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008. https://doi.org/10.1093/gigascience/giab008 (2021).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 1303, 3997 [preprint] (2013). https://doi.org/10.48550/arXiv.1303.3997
Kokot, M., Długosz, M. & Deorowicz, S. KMC 3: Counting and manipulating k-mer statistics. Bioinformatics 33, 2759–2761. https://doi.org/10.1093/bioinformatics/btx304 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Acknowledgements
We thank two anonymous reviewers who identified gaps in the manuscript, a bug in the pipeline script, and suggested a beneficial pipeline enhancement that improved the paper. Stephanie Schmidt provided secretarial support. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the USDA. The USDA is an equal opportunity provider and employer.
Funding
Funding for this research was provided by the USDA. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization (J.W.S., M.G.); Data Curation (A.M.D.); Formal Analysis (A.M.D.); Investigation (A.M.D.); Methodology (A.M.D., J.W.S., J.L.B., M.G.); Resources (A.M.D., J.W.S., J.L.B., M.G.); Software (A.M.D.); Visualization (A.M.D.); Writing (A.M.D., J.W.S., M.G.); Editing (J.W.S., J.L.B., M.G.).
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dickey, A.M., Schmidt, J.W., Bono, J.L. et al. The GEA pipeline for characterizing Escherichia coli and Salmonella genomes. Sci Rep 14, 13257 (2024). https://doi.org/10.1038/s41598-024-63832-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-63832-z
- Springer Nature Limited