Abstract
This study aimed to evaluate the etiology and pregnancy outcomes of fetuses underwent invasive prenatal diagnosis for fetal growth restriction (FGR) accompanied by structural malformations. Data from 130 pregnancies referred for prenatal diagnosis for FGR accompanied by structural malformations were obtained between July 2011 and July 2023. Traditional karyotyping was conducted for all the subjects. A total of 37 (28.5%) cases of chromosomal abnormalities were detected by karyotyping, including 30 cases of numerical anomalies and seven cases of unbalanced structural anomalies. Trisomy 18 was the most common abnormalities, accounting for 51.4%, significantly higher than any other chromosomal abnormality. The cohort was predominantly comprised of early-onset FGR (88.5%) compared to late-onset FGR (11.5%). The incidences of chromosomal abnormalities in this two groups were 29.6% (34/115) and 20.0% (3/15), respectively (p > 0.05). The majority (74.6%, 97/130) of the cohort were affected by a single system malformation, with chromosomal abnormalities found in 19.6% (19/97) of cases. In pregnancies of structural malformations involving two and multiple systems, the frequencies were 56.5% (13/23), and 50.0% (5/10), respectively. Single nucleotide polymorphism array (SNP array) was performed in parallel for 65 cases, revealing additional 7.7% cases of copy number variants (CNVs) compared to karyotyping. Polymerase chain reaction (PCR) was used for detection of cytomegalovirus (CMV) DNA in 92 cases. All fetuses with FGR associated with two or more system malformations were either terminated or stillborn, irrespective of chromosomal aberrations. Conversely, 71.8% of pregnancies with a single-system malformation and normal genetic testing results resulted in live births. Furthermore, two (2.2%) cases tested positive for CMV DNA, leading to one termination and one case of serious developmental disorder after birth. Our study suggests that structural malformations associated with FGR are more likely to affect a single organ system. When multiple systems are involved, the incidence of chromosomal abnormalities and termination rates are notably high. We advocate for the use of CMA and CMV DNA examinations in FGR cases undergo invasive prenatal diagnosis, as these tests can provide valuable insights for etiological exploration and pregnancy management guidance.
Similar content being viewed by others
Introduction
Fetal growth restriction (FGR) is a condition that fetus does not reach its intrauterine biological potential for growth and development. Traditionally, FGR has been defined as fetuses with an estimated fetal weight (EFW) below the 10th percentile for gestational age, and it could be symmetric or asymmetric. Some FGR symmetric may be actually normal small for gestational age (SGA), and many fetuses diagnosed with FGR exhibit have unexpectedly normal growth afterbirth1,2,3,4. Nonetheless, when FGR is suspected, it is suggestive to conduct an etiology evaluation to assess the prognosis and make informed decisions regarding the pregnancy.
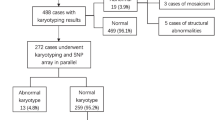
Multiple factors have been implicated in FGR. Maternal factors and uteroplacental factors are primarily related to obstetric management, while Genetic diseases play a significant role in fetal factors, often necessitating invasive prenatal genetic diagnosis. In recent decades, convention karyotyping and chromosomal microarray analysis (CMA) have become widely accepted for the routine genetic diagnosis of FGR, revealing a range of chromosomal and submicroscopic disorders. Survival data for growth-restricted fetuses without structural defect have been well-documented5,6,7. In our previous study related to genetic findings of FGR without structural malformations, karyotyping detected chromosomal abnormalities in 3.9% of cases, while CMA identified an additional 4.2% with clinically significant submicroscopic aberrations8. In clinical practice, the co-occurrence of FGR and structural malformations is a common observation, yet specific estimations of etiology are seldom reported. The current study retrospectively reviews the profiles of malformations, genetic etiology, and pregnancy outcomes in 130 pregnancies affected by both FGR and structural abnormalities. Additionally, considering that congenital cytomegalovirus (CMV) infection is the most relevant infection factors for FGR9,10, the quantitative determination of CMV DNA in 92 cases of prenatal specimens was analyzed. The purpose of this study is to explore the etiology and pregnancy outcomes in pregnancies with FGR complicated by structural malformations. We hope to provide a more comprehensive understanding of the etiology and pregnancy outcomes in cases of FGR complicated by structural malformations. This work builds upon our previous publication on FGR and will aid in guiding clinical consultations.
Materials and methods
Patients and samples
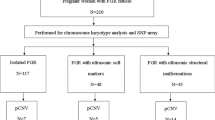
The retrospective data were collected between July 2011 and July 2023, encompassing 130 pregnancies that underwent invasive prenatal diagnosis due to the diagnosis of FGR accompanied by structural malformations. Among these, 92 cases were identified as symmetric FGR, and 38 cases were classified as asymmetric FGR. FGR was diagnosed based via ultrasound when EFW fell below the 10th percentile based on the Hadlock formula. The gestational age at FGR initially diagnosed was 25.8 ± 4.5 weeks, with 88.5% of cases being diagnosed before 32 weeks (early-onset FGR) and 11.5% diagnosed after 32 weeks (late-onset FGR). Detailed anatomical scans were performed when FGR was diagnosed. Structural abnormalities affected various organ systems, including the cardiac, craniocerebral, gastrointestinal, genitourinary, skeletal, and faciocervical systems. Depending on the number of organ systems involved in the structural malformation, they were further classified into groups of single-system, two-system, and multiple-system malformations. The basic characteristics are presented in Table 1.
Our samples included 70 cases of amniotic fluid and 60 cases of umbilical cord blood. Among them, 25 cases of umbilical cord blood was collected during induction of labor.
Methods
All experiments were performed in accordance with relevant guidelines and regulations.
Cytogenetic analysis
Karyotyping was performed on all the subjects. The cytogenetic analysis progress involving cell culture and G-banded karyotyping was performed according to the standard protocols in local laboratory, similar to those described in previous publication11. The karyotype was analyzed at a resolution of 320–500 bands level, using International System international system for human cytogenetic nomenclature 2020 (ISCN 2020)12 for karyotype description. Numerical chromosomal abnormalities and unbalanced structural abnormalities by conventional karyotyping were deemed clinically significant.
SNP array analysis
SNP array analysis has been utilized in our center since late 2016, thus only 65 pregnancies underwent SNP array analysis in parallel. Genomic DNA was extracted from uncultured amniotic fluid and cord blood samples using QIAGEN kit (Qiagen, Germany), following the manufacturer’s instructions. We employed the Affymetrix CytoScan 750 K array (Affymetrix Inc., Santa Clara, CA, USA) for SNP array analysis. This array includes 200,000 probes targeting single nucleotide polymorphisms and 550,000 probes designed to detect copy number variations (CNVs) throughout the entire human genome. As described in our prior publication8, microarray-based CNV analysis was performed using Chromosome Analysis Suite software (ChAS), version 3.1 (Affymetrix, Santa Clara, CA, USA), and genomic imbalances were annotated based on the GRCh37/hg19 Genome Build (July 2013). We maintained a general threshold of significance: gains or losses of ≥ 400 kb and regions of homozygosity (ROH) ≥ 10 Mb. Uniparental disomy (UPD) was identified based on the presence of a region of homozygosity (ROH) encompassing an entire chromosome. A specialized UPD tool was employed for comprehensive genome-wide detection of UPD within child-parent trios to confirm the maternal or paternal origin of UPD.
All identified CNVs were cross-referenced with both our institutional database and national public CNV repositories, including the Database of Genomic Variants (DGV), Database of Chromosome Imbalance and Phenotype in Humans Using Ensemble Resources (DECIPHER), International Standards for Cytogenomic Arrays Consortium, and Online Mendelian Inheritance in Man (OMIM).
The results from SNP array analysis was analyzed following the definition provided by the American College of Medical Genetics (ACMG)13, and were categorized into five levels: pathogenic, benign, likely pathogenic, likely benign, and variants of uncertain significance (VOUS). Clinically significant findings included pathogenic and likely pathogenic variants. Parental SNP array analysis was recommended to ascertain the inheritance of CNVs.
CMV-DNA testing
CMV DNA was extracted from amniotic fluid or cord blood on the Magna Pure LC Instrument (RocheMolecular Biochemicals, Meylan, France) with the Total NA serum-plasma kit (Roche Diagnostic). The viral load was assessed through quantitative polymerase chain reaction (qPCR) analysis. A viral load of ≥ 500copies/ml was considered as a positive result.
Pregnancy outcomes follow up
Information on pregnancy outcomes, including stillbirth, termination of pregnancy (TOP), and live birth, was collected from the hospital’s clinical database or through direct telephone inquiries. The follow-up ages ranged from 2 months to 5 years.
Statistical analysis
The data were analyzed using SPSS software v26.0 (SPSS Inc., Chicago, IL, USA). Statistical comparisons were performed using the chi-square test, the Fisher’s exact test, and p < 0.05 was considered statistically significant.
Ethical approval and consent to participate
The present study was approved by the Protection of Human Ethics Committee of Fujian Provincial Maternity and Children’s Hospital. Written informed consent was obtained from each pregnant woman.
Results
As outlined in Table 2, among the 130 pregnancies associated with structural malformations, a total of 37 (28.5%) chromosomal abnormalities were identified, including 30 numerical aberrations and seven cases of unbalanced structural abnormalities. The most common aberration was trisomy 18, accounting for 51.4% (19/37), followed by trisomy 21 (8.1%) and trisomy 13 (8.1%). The frequencies of chromosomal abnormalities between early-onset FGR and late-onset FGR showed no significant differences (29.6% vs. 20.0%, p > 0.05).
Table 3 displayed the types and chromosomal abnormality frequencies in pregnancies with anomalous FGR. The majority of FGR cases (73.1%) involved structural malformation in a single organ system, followed by two-system involvement (17.7%), and multiple-system involvement pregnancies (9.3%). Their rates of chromosomal abnormalities were 18.6% (18/95), 56.5% (13/23), and 50.0% (6/12), respectively. Cardiac malformations were the most common type, occurring in 93 pregnancies, with 59 of them being affected only by cardiac malformation, showing a chromosomal abnormality rate of 18.6%.
Among the 65 cases underwent both karyotyping and SNP array in parallel, additional 6 cases of submicroscopic aberration were revealed by SNP array compared to karyotyping, with 5 (7.7%) cases being clinically significant. Detailed information is presented in Table 4. Three of these cases were related to known syndrome: 15q24 Microdeletion Syndrome (#613,406), 8q21.11 Microdeletion Syndrome (#614,230), and DiGeorge Syndrome (#188,400/#192,430).
CMV infection was confirmed in 2 (2.1%, 2/92) cases who both exhibited craniocerebral malformation. Fetus 1 manifestied FGR, cerebral cortical dysplasia, Blake’s pouch cyst, posterior fossa abnormalities, and ventriculomegaly. The fetus resulted in live birth, and the child suffered from severe speech, hearing, and motor impairments at 4-years follow-up. Fetus 2 had FGR, severe ventriculomegaly, and broadening cisterna magna, and was finally terminated.
Pregnancy outcomes were available for 126 cases (95.4%). All pregnancies with clinically relevant genetic aberrations were terminated. The outcomes of the remaining 84 cases with normal genetic testing are shown in Table 5. All 13 cases with two and multiple system malformations ended in TOP. In the 71 pregnancies complicated by FGR and a single system malformation, 51 (71.8%) resulted in live births. Normal development was observed in 49 of them. The rest two cases showed abnormal development: one was of CMV infection, as mentioned above (fetus 1); the other one, with physical development delay, was found in case of sever FGR (EFW < 3th percentile) and duodenal stenosis.
Discussion
As expected, fetuses with FGR complicated by structural malformations were at a high risk of chromosomal abnormalities. The incidence of chromosomal abnormalities by karyotyping was 28.5%, much higher than 3.9% in our previous study on FGR without structural malformation8. Trisomy 18 was the most frequently encountered aberration, similar to the finding in the manyreports5,11,14. Additionally, many studies have reported a decrease in the rate of chromosomal abnormalities in isolated FGR fetuses as the gestational age at which FGR is first diagnosed increases, with fewer or no chromosomal abnormalities observed in pregnancies with isolated FGR diagnosed after 32 gestational weeks8,16,17. According to the latest SMFM (Society for Maternal–Fetal Medicine) guideline, late-onset FGR is not considered an indication for invasive diagnosis18. However, our study demonstrated that fetuses with FGR complicated by structural malformations diagnosed before and after 32 weeks showed similarly high detection rates (29.6% and 20.0%) of chromosomal anomalies. Therefore, we suggest that genetic evaluation should be considered when structural malformations is present in late-onset FGR.
We explored the influence of malformations, and found that the frequencies of genetic defects and pregnancy outcomes depended on the number of organ system involved in malformations. In FGR with a single-system malformation, the incidence of microscopic abnormalities (18.9%) was much lower than those involving two or multiple systems malformations (56.5% and 50.0%, respectively). Among pregnancies with follow-up data available, all those with two or more systems malformations were terminated, regardless of the presence of genetic abnormalities, whereas live births were observed in 72.9% of single-system malformation pregnancies. The findings demonstrate that when there are more than two system malformations involved in FGR, the rates of chromosomal abnormalities and pregnancy termination are extremely high. The cardiac system was most frequently involved, affecting up to 62.1% of the single-system malformation group, followed by craniocerebral system (13.7%) and genitourinary system (9.5%). Its rate of chromosome abnormalities was close to those involving other systems. Many scholars are concerned about changes in the structure and function of the cardiac and craniocerebral system of FGR fetuses, as they may be related to short-term and long-term adverse effects on FGR fetuses19,20. In this study, we focused on the genetic etiology, and we found that the rates of chromosomal abnormalities were similar between them.
All live births were from pregnancies involving single systemic malformation. Only two out of 71 survivors exhibited abnormal phenotypes. One of them had complex craniocerebral malformation in prenatal ultrasound. The clinical features of severe speech, hearing, and motor impairments after birth can be largely explained by intrauterine CMV infection in this case. Intrauterine CMV infection often lead to craniocerebral malformation, which was present in both two cases with CMV infection in our study, and can lead to a series of serious developmental disorders after birth9,21. The intrauterine manifestation and postnatal phenotype of this case highlighted the significant harm of intrauterine CMV infection and the potential association of CMV infection with FGR10,22.
An increase in pathogenic CNVs has been recognized to associate with FGR. In our previous study on FGR without structural malformations, SNP array analysis yielded additional 4.2% of clinically relevant aberrations compared with karyotyping8. In current study, FGR pregnancy with structural anomalies showed a slightly higher value of 7.7%. The finding was similar to that reported by Schaeffer et al.23. However, in a recent study by Chen et al.24, the pathogenic CNVs detection rate in FGR with structural anomalies was as high as 33.33%. The significantly varied results may be explained by different sample size and different FGR definition standards. Three known syndromes were involved in our study. Among them, 15q24 Microdeletion Syndrome (#613,406) and 8q21.11 Microdeletion Syndrome (#614,230) were rarely characterized by FGR25. As for 22q11.21 microdeletion, which is responsible for DiGeorge Syndrome or Velocardiofacial Syndrome, FGR was frequently reported in FGR with structural malformations12,26,27,28. In one case (case 1) presenting FGR and ventricular septal defect (VSD), a 2.6 Mb deletion was detected in the region of 3q26.33-3q27.2, involving 30 OMIM genes. This aberration is a rare condition in which all previously reported cases have experienced FGR and some other phenotypes29,30,31,32. In addition, maternal UPD of chromosome 16 was revealed in a fetus with FGR, VSD, aortic stenosis, and hypoplastic or absent left kidney. According to existing database and literatures, UPD (16) has been well believed to be correlated with FGR, mainly due to its potential impact on the function of the placenta33,34,35. This further strengthens the practical value of CMA in the etiological diagnosis of FGR and the associated malformations.
Our study was limited by the small sample size. In addition, not all cases underwent SNP array analysis and CMV DNA testing, which may introduce bias in etiology evaluation of submicroscopic aberration and intrauterine infection. A study with a larger sample size, comprehensive examination, and long-term follow-up is required to accurately assess the etiology and prognosis of fetuses with FGR associated with structural abnormalities.
In conclusion, structural malformations associated with FGR were more likely to involve a single organ system. When more than one system is involved, the incidence of chromosomal abnormalities and pregnancy termination is very high. In cases of invasive prenatal diagnosis, we recommend conducting CMA and CMV DNA examinations for etiological exploration and guidance in pregnancy management.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Selvaratnam, R. J. et al. Does public reporting of the detection of fetal growth restriction improve clinical outcomes: A retrospective cohort study. BJOG Int. J. Obstet. Gynaecol. 127, 581 (2020).
Ego, A., Goffinet, F., Kaminiski, M., Monier, I. & Zeitlin, J. Poor effectiveness of antenatal detection of fetal growth restriction and consequences for obstetric management and neonatal outcomes: A French National Study. Obstet. Gynecol. Surv. 122, 518 (2015).
Selvaratnam, R. J., Wallace, E. M., Hunt, R. W. & Davey, M. Preventing harm: A balance measure for improving the detection of fetal growth restriction. Aust. N. Z. J. Obstet. Gynaecol. 61, 715 (2021).
Lees, C. C. et al. 2 year neurodevelopmental and intermediate perinatal outcomes in infants with very preterm fetal growth restriction (TRUFFLE): A randomised trial. Lancet 385(9983), 2162–2172 (2015).
Meler, E., Sisterna, S. & Borrell, A. Genetic syndromes associated with isolated fetal growth restriction. Prenat. Diagn. 40(4), 432–446. https://doi.org/10.1002/pd.5635 (2020).
Sagi-Dain, L., Peleg, A. & Sagi, S. Risk for chromosomal aberrations in apparently isolated intrauterine growth restriction: A systematic review. Prenat. Diagn. 37(11), 1061–1066. https://doi.org/10.1002/pd.5160 (2017).
Temming, L. A. & Cahill, A. G. Early second-trimester fetal growth restriction and adverse perinatal outcomes reply. Obstet. Gynecol. J. Am. Coll. Obstet. Gynecol. 4, 131 (2018).
Wu, X. et al. Fetal genetic findings by chromosomal microarray analysis and karyotyping for fetal growth restriction without structural malformations at a territory referral center: 10-year experience. BMC Pregnancy Childbirth 23(1), 73. https://doi.org/10.1186/s12884-023-05394-y (2023).
Navti, O. B., Al-Belushi, M. & Konje, J. C. Cytomegalovirus infection in pregnancy—An update. Eur. J. Obstet. Gynecol. Reprod. Biol. 258, 216–222. https://doi.org/10.1016/j.ejogrb.2020.12.006 (2021).
Tanimura, K. et al. Fetal ultrasound and magnetic resonance imaging abnormalities in congenital cytomegalovirus infection associated with and without fetal growth restriction. Diagnostics (Basel) 13(2), 306. https://doi.org/10.3390/diagnostics13020306 (2023).
Li, H., Li, Y., Zhao, R. & Zhang, Y. Cytogenetic analysis of amniotic fluid cells in 4206 cases of high-risk pregnant women. Iran. J. Public Health 48(1), 126–131 (2019).
Schindewolf, E. Expanding the fetal phenotype: Prenatal sonographic findings and perinatal outcomes in a cohort of patients with a confirmed 22q11.2 deletion syndrome. Am. J. Med. Genet. Part A 176(8), 1735–1741 (2018).
Capalbo, A., Hoffmann, E. R., Cimadomo, D., Maria Ubaldi, F. & Rienzi, L. Human female meiosis revised: New insights into the mechanisms of chromosome segregation and aneuploidies from advanced genomics and time-lapse imaging. Hum. Reprod. Updat. 23(6), 706–722. https://doi.org/10.1093/humupd/dmx026 (2017).
Dall’Asta, A. et al. Etiology and perinatal outcome of periviable fetal growth restriction associated with structural or genetic anomaly. Ultrasound Obstet. Gynecol. 55(3), 368–374. https://doi.org/10.1002/uog.20368 (2020).
Maulik, D. Fetal growth restriction: The etiology. Clin. Obstet. Gynecol. 49(2), 228–235. https://doi.org/10.1097/00003081-200606000-00006 (2006).
Drummond, C. L. et al. Fetal karyotyping after 28 weeks of gestation for late ultrasound findings in a low risk population. Prenat. Diagn. 23(13), 1068–1072. https://doi.org/10.1002/pd.715 (2003).
Peng, R., Yang, J., Xie, H. N., Lin, M. F. & Zheng, J. Chromosomal and subchromosomal anomalies associated to small for gestational age fetuses with no additional structural anomalies. Prenat. Diagn. 37(12), 1219–1224. https://doi.org/10.1002/pd.5169 (2017).
Martins, J. G., Biggio, J. R. & Abuhamad, A. Society for maternal-fetal medicine consult series #52: Diagnosis and management of fetal growth restriction: (Replaces Clinical Guideline Number 3, April 2012). Am. J. Obstet. Gynecol. 223(4), B2-b17. https://doi.org/10.1016/j.ajog.2020.05.010 (2020).
Mäkikallio, K. et al. Fetal growth restriction and cardiovascular outcome in early human infancy: A prospective longitudinal study. Heart Vessel. 31(9), 1504–1513. https://doi.org/10.1007/s00380-015-0742-5 (2015).
McLean, G. et al. Three-dimensional ultrasound cranial imaging and early neurodevelopment in preterm growth-restricted infants. J. Paediatr. Child Health 54(4), 420–425. https://doi.org/10.1111/jpc.13808 (2018).
Kagan, K. O. & Hamprecht, K. Cytomegalovirus infection in pregnancy. Birth Defects Res. 296(297), 336–346 (2017).
Tsuge, M. et al. Prospective cohort study of congenital cytomegalovirus infection during pregnancy with fetal growth restriction: Serologic analysis and placental pathology. J. Pediatr. 206, 42-48.e2. https://doi.org/10.1016/j.jpeds.2018.10.003 (2019).
Shaffer, L. G. et al. Detection rates of clinically significant genomic alterations by microarray analysis for specific anomalies detected by ultrasound. Prenat. Diagn. 32(10), 986–995 (2012).
Tzadikevitch Geffen, K. et al. The yield of chromosomal microarray in pregnancies complicated with fetal growth restriction can be predicted according to clinical parameters. Fetal Diagn. Ther. 48(2), 140–148. https://doi.org/10.1159/000511475 (2021).
Liu, Y. & Mapow, B. Coexistence of urogenital malformations in a female fetus with de novo 15q24 microdeletion and a literature review. Mol. Genet. Genomic Med. 8(7), e1265. https://doi.org/10.1002/mgg3.1265 (2020).
Lin, I. et al. Central 22q11.2 deletion (LCR22 B-D) in a fetus with severe fetal growth restriction and a mother with severe systemic lupus erythematosus: Further evidence of CRKL haploinsufficiency in the pathogenesis of 22q11.2 deletion syndrome. Am. J. Med. Genet. A 185(10), 3042–3047. https://doi.org/10.1002/ajmg.a.62346 (2021).
Chen, Y. et al. The genetic etiology diagnosis of fetal growth restriction using single-nucleotide polymorphism-based chromosomal microarray analysis. Front. Pediatr. 9, 743639. https://doi.org/10.3389/fped.2021.743639 (2021).
Srebniak, M. I. et al. Prenatal SNP array testing in 1000 fetuses with ultrasound anomalies: Causative, unexpected and susceptibility CNVs. Eur. J. Hum. Genet. 24(5), 645–651. https://doi.org/10.1038/ejhg.2015.193 (2016).
Bouman, A. et al. An interstitial de-novo microdeletion of 3q26.33q27.3 causing severe intrauterine growth retardation. Clin. Dysmorphol. 24(2), 68–74. https://doi.org/10.1097/MCD.0000000000000075 (2015).
Ounap, K., Pajusalu, S., Zilina, O., Reimand, T. & Zordania, R. An 8.4-Mb 3q26.33–3q28 microdeletion in a patient with blepharophimosis-intellectual disability syndrome and a review of the literature. Clin. Case Rep. 4(8), 824–830. https://doi.org/10.1002/ccr3.632 (2016).
Robilliard, R. & Caylan, M. Infantile presentation of 3q26.33–3q27.2 deletion syndrome. BMJ Case Rep. 13(11), e233215. https://doi.org/10.1136/bcr-2019-233215 (2020).
Dasouki, M., Roberts, J., Santiago, A., Saadi, I. & Hovanes, K. Confirmation and further delineation of the 3q26.33–3q27.2 microdeletion syndrome. Eur. J. Med. Genet. 57(2–3), 76–80. https://doi.org/10.1016/j.ejmg.2013.12.007 (2014).
Eggenhuizen, G. M., Go, A., Koster, M. P. H., Baart, E. B. & Galjaard, R. J. Confined placental mosaicism and the association with pregnancy outcome and fetal growth: A review of the literature. Hum. Reprod. Updat. 27(5), 885–903. https://doi.org/10.1093/humupd/dmab009 (2021).
Grau Madsen, S., Uldbjerg, N., Sunde, L. & Becher, N. Prognosis for pregnancies with trisomy 16 confined to the placenta: A Danish cohort study. Prenat. Diagn. 38(13), 1103–1110. https://doi.org/10.1002/pd.5370 (2018).
Soong, Y. K. et al. Genome-wide detection of uniparental disomy in a fetus with intrauterine growth restriction using genotyping microarrays. Taiwan J. Obstet. Gynecol. 48(2), 152–158. https://doi.org/10.1016/s1028-4559(09)60277-1 (2009).
Funding
The Natural Science Foundation of Fujian Province, China (Grant No. 2021J01413). Key Special Projects of Fujian Provincial Department of Science and Technology (Grant No. 2021YZ034011). Joint Funds for the innovation of science and Technology, Fujian province (Grant No. 2020Y9135).
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, X., He, S., Shen, Q. et al. Etiologic evaluation and pregnancy outcomes of fetal growth restriction (FGR) associated with structural malformations. Sci Rep 14, 9220 (2024). https://doi.org/10.1038/s41598-024-59422-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-59422-8
- Springer Nature Limited