Abstract
A global increase in the populations of drug resistant bacteria exerts negative effects on animal production and human health. Our study has been focused on the assessment of resistance determinants in relation to phenotypic resistance of the 74 commensal E. coli isolates present in different ecological environments. The samples were collected from poultry litter, feces, and neck skin. Among the microorganisms isolated from the poultry litter (group A), the highest resistance was noted against AMP and DOX (100%). In the E. coli extracts from the cloacal swabs (group B), the highest resistance was observed against AMP (100%) and CIP (92%). The meat samples (group C) were characterized by resistance to AMP (100%) and STX (94.7%). Genes encoding resistance to β-lactams (blaTEM, blaCTX-M), fluoroquinolones (qnrA, qnrB, qnrS), aminoglycosides (strA-strB, aphA1, aac(3)-II), sulfonamides (sul1, sul2, sul3), trimethoprim (dfr1, dfr5, dfr7/17) and tetracyclines (tetA, tetB) were detected in the studied bacterial isolates. The presence of class 1 and 2 integrons was confirmed in 75% of the MDR E. coli isolates (plasmid DNA), of which 60% contained class 1 integrons, 15% contained class 2 integrons, and 11.7% carried integrons of both classes. Thus, it may be concluded that integrons are the common mediators of antimicrobial resistance among commensal multidrug resistant Escherichia coli at important stages of poultry production.
Similar content being viewed by others
Introduction
The increasing resistance to commonly applied antimicrobial agents is being reflected by growing multiple drug resistance (MDR) in bacteria and is becoming a growing threat to public health. The use of antimicrobial agents in animal husbandry has been linked to the development and spread of the resistant bacteria1. Resistant bacteria can be transferred for example from poultry products to humans via consuming or handling meat contaminated with pathogens2. However, the resistance of commensal bacteria is equally important as they constitute a reservoir and vector of resistance determinants in the environment3.
Exposure to antimicrobials of different classes can lead to cross-resistance and the selection of antibiotic resistance genes (ARGs) that may spread laterally on mobile genetic elements (MGEs) via horizontal gene transfer (HGT)4. HGT is a phenomenon in which genes are transferred between organisms of either the same or different species which often remain in a close ecological relationship5.
It has been shown that conjugation is one of the key mechanisms responsible for the spread of the ARGs6. One of the most efficient mechanism of acquiring ARGs is facilitated by integrons—a site-specific recombination systems capable of recruiting open reading frames (ORF) in the form of mobile gene cassettes 7. Integrons are divided into two distinct subsets, mobile integrons (MIs)—linked to mobile DNA elements, which are primarily involved in the spread of ARGs, and chromosomal integrons (CIs). MIs are associated with conjugation plasmids or transposons (the integron itself is not mobile), which allows the spread and exchange of the resistance genes between individual strains and bacterial species8. MIs may be divided into five classes, which are involved in the propagation of antibiotic-resistance genes9. These classes are divided based on the sequence of the encoded integrase genes, which share 40–58% sequence homology. The first three classes of integrons are involved in the acquisition of the MDR phenotype. Class 1 integrons are mostly found in clinical and animal production isolates, and most of the known antibiotic-resistance gene cassettes belong to this group. Class 1 is associated with functional and nonfunctional transposons derived from Tn402 that can be embedded in larger transposons, such as Tn2110. Class 2 integrons are associated with Tn7 derivatives, and class 3 integrons are thought to be located in a transposon inserted in plasmids11,12. The integron types may be identified based on the detection of the specific integrase gene sequence—Int1, Int2 and Int3 respectively13.
Class 1 and 2 integrons are frequently detected and well characterized, mostly among bacteria belonging to the Enterobacteriaceae family, including E. coli14. The majority of E. coli strains are commensals inhabiting the intestinal tract of humans and warm-blooded animals and rarely causes diseases15. The adaptation ability of these microorganisms into various niches in host organism is determined by its extremely plastic genome. Another important driving force for the evolution of the E. coli genome is the mobile gene pool replaced by the HGT16.
Studies of microorganisms found in the breeding environment of broiler chickens (including E. coli), such as feces and poultry meat provide valuable information about the reservoir of bacterial genes17. While assessing the risks arising from the possible transfer of resistant bacteria within the poultry production chain, it seems important to know the diversity and the prevalence of genetic determinants of antibiotic resistance among the commensal E. coli strains.
In the recent years, a number of studies have been carried out aiming to identify the presence and the structure of the integrons, the type of the resistance cassettes and the relationship between the occurrence of integrons and MDR, in commensal and pathogenic E. coli isolated from human and animal samples18,19,20. A link between the use of antibiotics in animal production and antimicrobial resistance of human pathogens (within which food is one of the possible vectors) was reported in several studies21,22,23. Nevertheless, little is known about the distribution of integrons in E. coli isolated from commercial broiler meat in Poland. Therefore, the purpose of this study was to determine antimicrobial resistance profiles, distribution of class 1 and 2 integrons and integron-associated gene cassettes in commensal strains isolated from poultry litter, broiler chicken feces and meat in western Poland.
Results
Antimicrobial resistance phenotypes of E. coli isolates
We have analyzed the incidence of multidrug resistance gene sequences and the prevalence of class 1 and 2 integrons within 74 commensal E. coli isolates, obtained from poultry litter (group A, n = 23), swabs from broiler chicken cloaca (group B, n = 26) and poultry meat (group C, n = 25). 60 (81.1%) of them exhibited a multiresistant phenotype (resistance to at least three different antimicrobial agent families). In the first step of study, the phenotypic resistance of the E. coli isolates to six antibiotics and chemotherapeutics was assessed.
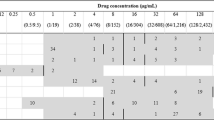
Out of 23 E. coli isolates obtained from poultry litter, 16 showed multidrug resistance. The highest resistance was recorded for AMP (100%), DOX (100%), CIP and STX (81.3%), and AMC (75%); the lowest for GEN (12.5%). Out of 26 isolates obtained from chicken cloaca, 25 exhibited MDR. Among the examined MDR isolates, the highest percentage of resistance was observed for the following antibiotics: AMP (100%), CIP (92%), AMC, DOX (88%), STX (84%) and the lowest for GEN (36%). All E. coli isolates obtained from the cloaca of chickens, showed phenotypic resistance to the antibiotics classes with which the broilers were treated on farms. Among E. coli isolates obtained from meat, 19 of them showed MDR. In this group, the highest resistance was observed for AMP (100%). Lower resistance was noted for: STX (94.7%), CIP (78.9%), DOX (68.4%). Resistance to AMC and GEN was noted in 42.1% and 36.8% of E. coli isolates respectively. Figure 1., shows the percentage of phenotypic resistance to 6 antibiotics among MDR E. coli isolates obtained from poultry litter, cloacal swabs and poultry meat (group A, B and C). Antimicrobial susceptibility tests showed that 11.7% of all MDR isolates of E. coli, were resistant to three antibiotics, 38.3% to four, and 35% showed resistance to five and 15% to six antibiotics (Table 1). Among all E. coli isolates positive for integron sequences, the most common drug resistance profile was that of the resistance to: AMP, AMC, CIP, STX, DOX. In the case of 2 isolates with a pair of integrons, E. coli isolates that showed resistance to 6 antimicrobial agents were reported. Their resistance profile was the same: AMP, AMC, CIP, GEN, STX, DOX.
The percentage of phenotypic resistance to 6 antibiotics among the MDR E. coli isolates obtained from poultry litter (Group A), cloacal swabs (Group B) and poultry meat (Group C): AMP—Ampicillin; AMC—Amoxycyline with Clavulanic Acid; CIP—Ciprofloxacin; GEN -Gentamicine; STX—Sulfamethoxazole with Trimetoprime; DOX—Doxycycline.
Identification and characteristics of resistance genes.
Resistance to AMP and AMC encoded by the narrow spectrum beta lactamase resistance gene (blaTEM) was found in genomic DNA of all E. coli isolates from groups A and B, and in 63,2% of the poultry meat (group C). In case of other genes encoding beta-lactamase resistance, the occurrence of blaCTX-M gene was noted in one colon isolate. BlaSHV gene was detected in one isolate from the poultry meat swabs. Among the MDR isolates showing the ciprofloxacin (CIP) resistance phenotype, a total of 7 qnrA genes, 10 qnrB genes and 6 qnrS genes were reported in genomic DNA. 18 MDR isolates were gentamicin-resistant, and the strA-strB, aphA1, and aac(3)-II genes, giving resistance to aminoglycosides, were present in: 13 isolates of E. coli obtained from litter, 19 isolates from cloacal swabs and 5 isolates from poultry meat. 13 of the 16 tested E. coli MDR isolates showed a sulphonamides resistance phenotype which was encoded by: sul1 (50%), sul2 (18.8%) and sul3 (25%). In the case of 21 MDR isolates obtained from cloaca the sul1 and sul3 genes were recorded in 28% and the sul2 gene in 44% of the cases. In this group, the presence of pairs of sul genes was noted in 5 isolates, in the su1—sul2 combination. In the MDR group of meat isolates, 18 E. coli isolates contained the following genes: sul1 (31.6%), sul2 (26.3%) and sul3 (10.5%) and in one case a pair of genes (sul1 and sul2) were noted. In the case of E. coli isolates recovered from the litter, one of them confirmed 3 genes (sul1, sul2, sul3) determining resistance to sulfonamides. Sulfonamide-resistant isolates of E. coli showed in most cases compatible resistance to trimethoprim resistance genes (dfr1, dfr5, dfr7/17) in 84.6% cases in group A, 71.4% in group B and 38.9% in group C. The tet genes (tetA, tetB) giving resistance to doxycycline were found in 75 and 37.5% isolates from poultry liter, 52 and 36% isolates from cloacal swab, and 36.8 and 10.5% isolates from meat. The tetC gene was not found in any of the studied groups. The pairs of tetA and tetB genes were found in groups A and B in 4 and 3 cases, respectively. The prevalence of the resistance genes among multidrug resistant E. coli isolates obtained from poultry litter (Group A), cloacal swabs (Group B) and poultry meat (Group C) is shown in Table 2 and Fig. 2.
Detection and characterization of integrons
Our study reports a high incidence of integron-bearing E. coli isolates. More than half of the multidrug resistant isolates contained integrons. The presence of class 1 and 2 integrons was confirmed in 45 out of 60 MDR E. coli isolates (75%). Structure of 1 class integron, identified by the presence of the IntI1 gene was detected in 36 (60%) multiresistant isolates in plasmid DNA. Most of this gene was recorded in the DNA of bacteria isolated from cloaca—15 E. coli isolates, and next from litter and meat—11 and 10, isolates, respectively. The frequency of the integrons of class 1 was not significantly different across sampling locations (P > 0.133). In turn, class 2 integrons, identified by the presence of the intI2 gene, were found in a much smaller number of cases, only in cloacal and meat isolates (4 and 5 cases). The frequency of the integtons of class 2 differed significantly (P ≤ 0.05) between these two locations. In the samples in which both classes of the integrons were detected (group B and C), the class 1 integrons were significantly more frequent than class 2 integrons (B: P ≤ 0.01, and C: P ≤ 0.05).
Gene cassette analysis of class 1 and 2 integron genes
Table 3, 4 and 5 shows the phenotypes, antibiotic resistance, and prevalence of class 1 and 2 integrons and their resistance gene cassettes in MDR E. coli isolates obtained from all groups. E. coli isolates containing integrons were grown from poultry litter (group A). 11 out of 16 MDR isolates, contained class 1 integrons only. The variable region of class 1 integrons most frequently contained the aadA1 gene cassette in seven cases. The dfrA1-aadA1 cassette series was present in three cases (18.8%) and dfrA17-aadA5 in one strain. In the class 2 integron variable region, no gene cassettes were found in any case (Table 3).
In case of the multidrug-resistant E. coli isolates obtained from cloacal swabs from broiler chickens (group B), 15 isolates contained class 1 integrons and 4 isolates contained class 2 integrons (Table 4). The results of DNA sequencing of the inserted gene cassettes allowed to identify 11 class 1 integrons containing 5 different gene cassettes: aadA1 (36.4%), dfrA5 (18.2%) and arrays of cassettes: dfrA1-aadA1 (36.4%) and dfrA17-aadA5 (9.1%) in five separate isolates. Genes contained in the four class 2 integron cassettes contained gene cassette arrays: dfrA1-sat2-aadA1.Two isolates containing class 1 and 2 integrons contained cassette arrays: aadA1; dfrA1-sat2-aadA1 and dfrA5; dfrA1-sat2-aadA1. Four (26.7%) class 1 positive integron isolates had no gene cassettes in their variable part.
In the group of MDR E. coli isolated from the poultry meat (group C), genes contained in the class 1 integron variable region were detected in 9 cases (Table 5). VR class 1 integrons showed less variation compared to group B isolates and usually contained 2 gene cassette arrays: aadA1 (22.2%) and dfrA1-aadA1 (77.8%). The variable region of 5 class 2 integrons contained in only two cases a set of cassettes: dfrA1-sat2-aadA1. In one strain, a pair of class 1 and 2 integrons was detected as well as their variable parts were empty.
Discussion
It is estimated that in most developed nations, livestock use 50–80% of antibiotics produced24. Commonly used groups of veterinary medicinal products in the EU are tetracyclines (32%), penicillins (26%), sulfonamides (12%), macrolides (7%), polymyxins (5%) and aminoglycosides (5%)25,26.
In Poland, 829 tones of antibiotics are used annually, of which as much as 578 tones in the agricultural industry27,28. The most commonly used groups of substances in years 2014–2016 were tetracyclines (42.34–49.07%), penicillins (18.98–23.40%) and macrolides (11.69–13.22%)26.
In our study, 74 unrelated commensal isolates of Escherichia coli originated from poultry litter, cloacal swabs and poultry meat were phenotypically and genotypically tested for the antimicrobial resistance and the presence of integrons as factor, for the development of antibiotic resistance and the emergence of MDR strains. Among them, 60 isolates (81.1%) were multiresistant (resistant to a minimum of three classes of antibiotics). We have found the highest level of antimicrobial resistance (96.2%) in the E. coli isolates obtained from broiler intestinal swabs (group B).
The high incidence of multidrug resistance in our study, particularly regarding isolates obtained from feces and meat, is extremely significant and should be regarded as a serious health risk due to the fact, that multidrug resistant isolates may have a chance of contaminating food products, and consequently being transferred to humans.
The percentage of resistance to some antimicrobial agents (Ampicillin, Doxycycline, Trimetophrim, Sulfamethoxazole, and Ciprofloxacin) in all studied groups was particularly high (100–68.4%), which indicates that the commensal E. coli isolates may be a reservoir of resistance to antibiotics and chemotherapeutics. Our data largely overlaps with the data made available by the European Food Safety Authority and the European Centre for Disease Prevention and Control, on the resistance profile of the commensal E. coli isolates obtained from slaughterhouse broilers, collected between 2009 and 2014 in Poland28. It confirms high resistance of E. coli isolates to nalidixic acid ciprofloxacin, and ampicillin (70–90%) and a limited resistance to tetracyclines, sulfonamides and streptomycin28,29.
The blaTEM gene encoding β lactamase, which gives resistance to penicillins and cephalosporins of the first generation, has been detected in all multidrug-resistant isolates obtained from litter and feces. The dominance of the TEM gene over the CTX gene in fecal poultry isolates was also noted in the Nigerian30 study but was not as pronounced as in our experiment (63%-TEM and, 35% CTX-M). Although E. coli isolates with enzymes belonging to the CTX-M family in ESBL-positive bacteria are currently predominant in the world31 in our study the blaCTX-M gene was reported only in one bacterial strain (4%) from cloacal swabs. Similar results were obtained in China, where blaCTX-M was detected in 1.6% of the isolates coming from the meat32.
Quinolone resistance is a current worldwide problem in human and veterinary medicine33. Quinolone resistance can encoded in bacterial chromosome or be present in plasmids. Plasmid-mediated quinolone resistance (PMQR) promote the spread of the multi-drug resistance phenotype. For example, qnr genes present on MDR plasmids are often found with genes encoding β-lactamases34. Of all Qnr determinants present in the our study, the qnrB gene was found most frequently. Similar results were published in other studies33,35. The occurrence of PMQR is also associated with resistance to other groups of antibiotics. The mechanism responsible for this phenomenon is related to the presence of the aminoglycoside acetyltransferase enzyme—AAC(6')-Ib-cr, which modifies both the molecular structure of some fluoroquinolones and aminoglycosides or the oqxAB gene encoding an MDR-type efflux pump contributing to increased resistance to quinolones and chloramphenicol, trimethoprim and quinolones36,37. In Seo and Lee33 study, 10 PMQR-positive E. coli were isolated from chicken meat, and these isolates also showed higher resistance rates to several antimicrobial agents when compared to PMQR-negative E. coli. This is consistent with previous studies showing that the PMQR genes increase resistance to other antimicrobials and cause MDR to drugs such as: aminoglycosides, β-lactams, chloramphenicol, sulfonamides, tetracyclines and trimethoprim38.
Tetracyclines are commonly used to treat bacterial infections in livestock, including poultry in many countries39,40,41. Due to the numerous advantages of tetracyclines, such as their widespread availability, low cost, and several side effects, the use of such antibiotics to treat animal and human infections has been increasing in recent years42. The chickens are treated with tetracycline orally and their metabolites (up to 90%) are excreted in the feces43 on manure44. It is noteworthy that in our study the highest resistance to doxycycline was recorded among E. coli isolates derived from poultry litter, which is a mixture of poultry manure, litter, feathers, feed, and spilled drinking water that accumulates during breeding45. However, the proportion of E. coli isolates with resistance to tetracyclines was lower than the proportion of E. coli isolates resistant to beta lactam antibiotics. Similar results were obtained in study by Islam et al.46 in which MDR isolates were the most resistant to tetracyclines (96.6%) and penicillins (100%). In addition to the antibiotic residues, manure also contains MDR bacteria and resistance genes, which can be transmitted to humans through direct contact between poultry and humans or indirectly via the food chain47. The results of genotyping showed that similarly to other published data, the resistance of commensal E. coli to tetracyclines was induced by the presence of tetA and tetB genes48. The highest content of tet genes in poultry litter isolates confirms the thesis put forward by Furtula et al.44, that the breeding environment significantly contributes to the spread of the resistance, via the transmission of the resistance genes.
Resistance to sulfonamides in Gram-negative bacteria generally results from the presence of the genes sul1, sul2, and/or sul3. Among them, the sul2 gene is the most widely distributed sul gene in porcine, avian, or human E. coli isolates, and it plays an important role in sulfonamide resistance49. In our research, the prevalence of sul2 genes was highest in isolates from cloaca swabs (44%) and was similar to the results of other studies50,51. Interestingly, in only one strain obtained from litter, all tested sul genes were determined. The frequency of sul genes detection in our experiment corresponded to other studies50,51. The selective pressure exerted by sulfonamides in the poultry industry appears to be high, which may favor the maintenance of acquired sul genes among bacteria52.
In our study, the total prevalence of integrons (75%) in MDR isolates was higher than the prevalence of integrons detected in other poultry production prevalence studies53,54. We also found a clear predominance of class 1 integrons in relation to class 2 integrons, which is consistent with previous studies that also showed the highest prevalence of this class in poultry isolates55,56.
Most of the integrons detected in our study contained gene cassettes encoding resistance to trimethoprim (dfrA gene type) and streptomycin/spectinomycin belonging to aminoglycosides (aadA gene type), and the most frequently detected sequence of cassettes was dfrA1-aadA1. The persistence of these genes, which have been reported worldwide in isolates from different sources, may be related to the widespread use of streptomycin/spectinomycin, trimethoprim, sulfonamides and other antibiotics in food producing animals. Although the afore-mentioned aminoglycosides are not used therapeutically in animals in Poland, the presence of aadA genes may be a form of genetic memory, in case of re-exposure of the microorganism to this group of antibiotics57.
The analysis of the variable part of integrons in our experiments indicated the presence of one to three gene cassettes. In group B, we noted the highest number of integrons among all tested groups and a greater variety of gene cassettes within the integron variable part. The higher prevalence of class 1 integrons among the E. coli isolates obtained from the cloaca may be caused by the development and spread of the resistance genes, due to the misuse or abuse of antibiotics in the poultry production1.
Of all MDR E. coli isolates that had integrons, 8 isolates (13.3%) did not contain any of the evaluated gene cassettes. The situation regarding the so-called "empty integrons" has already been described by Fonseca et al.58, where it was indicated that these bacteria could rapidly develop into MDR in the future. However, it cannot be ruled out that integrons may have previously removed cassettes of resistance genes acquired by cutting them out for unknown reasons59.
The data obtained in this study highlights the importance of commensal E. coli in the spread of resistance genes at different stages of poultry production. We confirmed that more than half of the multidrug-resistant isolates (75%) contained integrons. Furthermore, we showed that antibiotic resistance can also occur on non-integron structures. Therefore, there is a need for further detailed genetic studies on the evolution of isolates present in poultry to uncover the underlying mechanisms the acquisition of resistance by these microorganisms and to analyze the implications for humans. Such data may be used to determine the dynamics of resistance development and strategies to counteract antibiotic resistance among zoonotic microorganisms transmitted through food of animal origin at all stages of the food chain, from farm to table.
Materials and methods
E. coli isolates
A total number of 74 E. coli isolates was collected from three areas of poultry production: litter swabs from chicken houses (n = 23), cloacal swabs from chicken (n = 26), chicken meat from slaughterhouses (n = 25). All samples were collected between November 2019 and March 2020.
Litter samples from 23 different chicken houses were acquired in accordance with the boot swabs sampling procedure guidelines of the national control program for Salmonella serotypes in poultry flocks in line with the guidelines of the current EU law60. Samples were collected from 4-week-old chicks (average weight 1.6 kg) 2 weeks before slaughter. The samples of cloacal swabs were collected using swabs (NRS II Transwab swabs with 10 Buffered Peptone Water, Medical Wire & Equipment, Corsham, United Kingdom) in a poultry slaughterhouse. Chickens were raised on 25 unrelated farms located in Greater Poland Voivodship. Birds that were sent to slaughter at 6 weeks of age, weighed an average of 3 kg and belonged to the Ross 308 breed. The chicken meat samples were obtained from neck skin. These samples were delivered to the laboratory for testing, as part Salmonella monitoring program, from 5 different poultry slaughterhouses located in the Greater Poland Voivodeship, from different periods of production.
Information regarding the antibiotics used in the above chickens (name of the antibiotic, withdrawal periods) was included in the food chain documentation. In the treatment of poultry, the most frequently used antibiotics were: Amoxicillin, Enrofloxacin, Doxycycline, Sulfamethoxazole / Trimethoprim.
The samples were placed in buffered peptone water (BioMerieux, Marcy l'Etoile, France) and incubated at 35 °C (± 1 °C) for 18 h (± 2 h) under aerobic conditions. Next, the material was plated on MacConkey agar medium (OXOID, Basingstoke, United Kingdom) and incubated for 24 ± 2 h under aerobic conditions at 37 °C ± 1 °C. Colonies with the typical E. coli phenotype were selected (one per sample) and verified by the MALDI-TOF method (Bruker, Bremen, Germany). The score ranged from 2.015 to 2.152.
Antimicrobial susceptibility testing
Antibiotic susceptibility tests of all 74 E. coli isolates were performed following the standard agar disk diffusion method, according to the CLSI (Clinical and Laboratory Standards Institute-2012) using commercially available antimicrobial disks containing (OXOID, Basingstoke, United Kingdom): penicillins (Ampicillin—AMP 10 μg and Amoxycyline with Clavulanic acid—AMC 20/10 μg), fluoroquinolones (Ciprofloxacin—CIP 5 μg), aminoglycosides (Gentamicine—GEN 10 μg), sulfonamides (Sulfamethoxazole with Trimetoprime—STX 25 μg), tetracyclines (Doxycycline—DOX 30 μg).
The following media were used for the tests: Mueller Hinton Broth (Thermo Fisher Scientific, Waltham, Massachusetts, USA), Mueller–Hinton agar (OXOID, Basingstoke, United Kingdom). The bacterial colonies were classified as sensitive, intermediate, or resistant according to the standardized CLSI guidelines (VET 01S—Performance standards for antimicrobial disk and dilution susceptibility tests for bacteria isolated from animals) and E. coli ATCC 25922 strain was used as control. The isolates were collected and stored at -80℃ for further analyses on Viabank medium (OXOID, Basingstoke, United Kingdom).
Detection of integrons
Class 1 and Class 2 integrons were detected based on the presence of gene sequences characteristic for integrase 1 (IntI1) and integrase 2 (IntI2) respectively. Selected regions were amplified by qualitative PCR carried out on plasmid DNA extracted from the three studied groups (poultry litter, feces and carcasses).
Plasmid DNA from the bacterial samples was extracted with the Gene Matrix Plasmid Miniprep DNA Purification Kit (E3500 EurX, Gdansk, Poland) according to the manufacturer’s protocol. PCR amplification was done in a 10µL mixture containing: 1µL DNA template, 0.3µL of primers (0.3 μM), 2µL of 5× HOT FIREPol® Blend Master Mix kit (04-25-00S25, SolisBiodyne, Tartu, Estonia) and molecular biology graded water (nuclease free, W4502 Merck, Darmstadt, Germany). The thermal cycling conditions included: preincubation at 95 °C (15 min), followed by 38 cycles of denaturation at 95 °C (20 s), annealing at 61 °C (45 s), extension at 72 °C (60 s) and final extension at 72 °C (5 min). Primer pair sequences are listed in Table 6. The specificity of the PCR reaction (product length–base pair, bp) was verified by electrophoresis on a 1.5% agarose gel.
Sequencing of the variable regions of Integron 1 and Integron 2
In the bacterial isolates containing the IntI1 and the IntI2 genes, the variable regions (VRs) of both of the studied integrons were sequenced in order to reveal the specific DNA sequence (plausibly bearing the multidrug resistance genes) within the integron structure. The regions were amplified with a set of specific primers: Integron CL (for integron 1) and Integron CL JJ (for integron 2), sequences are listed in Table 6. The PCR reaction conditions, and primer concentrations were the same as described for IntI1 and IntI2 amplification, with an annealing temperature of 61 °C. The amplified gene cassettes of similar PCR product length (base pair, bp) were sequenced by the Sanger method. Prior to sequencing, the PCR products were purified with FastAP and ExoI enzymes (EF0654 and EN0581, Thermo Fisher Scientific, Waltham, Massachusetts, USA), amplified With the BigDye™ terminator v3.1 Cycle Sequencing Kit (4,337,458, Life Technologies, Carlsbad, California, USA) and purified on Sephadex G50 (G5050, Sigma, St. Louis, Missouri, USA) by filtration. Sequencing of the cassette arrays was done with Applied Biosystems ABI 3130xl 16-capillary array genetic analyzer (Applied Biosystems, Waltham, Massachusetts, USA). Data was analysed with the Seqman software.
Detection of antimicrobial resistance genes outside the integron cassettes
This experiment focused on the detection of 19 antimicrobial resistance genes in the studied genomic samples. Genomic DNA was extracted with Extractme DNA Bacteria Kit (EM02, Blirt, Gdansk, Poland) according to the manufacturer’s protocol. The PCR reaction conditions were the same as described for the detection of introns. Primer sequences are listed in Table 6. We tested for the presence of 4 genes associated with resistance to b-lactam antibiotics—ampicillin and amoxicillin (blaSHV, blaTEM, blaCTX-M, blaOXA)61, ciprofloxacin-resistance (qnrA, qnrB, qnrS)62, streptomycin-resistance (strA-strB)63, kanamycin-resistance (aphA1)63, gentamicin-resistance (aac(3)-II)64, sulphonamides-resistance (sul1, sul2, sul3)63, trimethoprim-resistantance (dfr1, dfr5, dfr7/17)65 and tetracycline-resistance (tetA, tetB, tetC)63. The analyses were done with the set of primers listed in Table 6 and the PCR reaction conditions were as described for the detection of integrons.
Statistical analysis
The significance of the differences between presence and absence of integrons of class 1 and 2, and the predominance of class 1 integrons in relation to class 2 integrons were tested within each group (A, B, and C) using Pearson's Chi-squared Test. Relation between the presence of integrons and the phenotype of multi antibiotic resistance was tested with the use of Poisson Regression analyses where the dependent variable was the number of antibiotics, resistance, and the independent variable was the integron’s presence. These analyses were performed separately for integron class 1 and 2 and for each group (A, B, and C). All of the statistical analyses were performed with the use of the R environment66.
References
Agyare, C., Boamah, V. E., Zumbi, C. N. & Osei, F. B. Antibiotic Use in Poultry Production and Its Effects on Bacterial Resistance. https://doi.org/10.5772/intechopen.79371 (2018).
van den Bogaard, A. E. & Stobberingh, E. E. Epidemiology of resistance to antibiotics. Links between animals and humans. Int. J. Antimicrob. Agents 14, 327–335. https://doi.org/10.1016/s0924-8579(00)00145-x (2000).
Racewicz, P., Majewski, M., Madeja, Z. E., Lukomska, A. & Kubiak, M. Role of integrons in the proliferation of multiple drug resistance in selected bacteria occurring in poultry production. Br Poult Sci 61, 122–131. https://doi.org/10.1080/00071668.2019.1697426 (2020).
Skarżyńska, M., Zając, M. & Wasyl, D. Antibiotics and bacteria: Mechanisms of action and resistance strategies. Postępy Mikrobiologii - Advancements of Microbiology 59, 49–62. https://doi.org/10.21307/pm-2020.59.1.005 (2020).
Kelly, B. G., Vespermann, A. & Bolton, D. J. The role of horizontal gene transfer in the evolution of selected foodborne bacterial pathogens. Food Chem. Toxicol. 47, 951–968. https://doi.org/10.1016/j.fct.2008.02.006 (2009).
Singer, R. S. & Williams-Nguyen, J. Human health impacts of antibiotic use in agriculture: A push for improved causal inference. Curr. Opin. Microbiol. 19, 1–8. https://doi.org/10.1016/j.mib.2014.05.014 (2014).
Stokes, H. W. & Hall, R. M. A novel family of potentially mobile DNA elements encoding site-specific gene-integration functions: integrons. Mol. Microbiol. 3, 1669–1683. https://doi.org/10.1111/j.1365-2958.1989.tb00153.x (1989).
Boucher, Y. et al. Recovery and evolutionary analysis of complete integron gene cassette arrays from Vibrio. BMC Evolu. Biol. https://doi.org/10.1186/1471-2148-6-3 (2006).
Ghazalibina, M. et al. Prevalence of integrons and antibiotic resistance pattern in Acinetobacter baumannii isolated from clinical samples of Iranian patients: A systematic review and meta-analysis. Ethiop. J. Health Sci. 29, 639–648. https://doi.org/10.4314/ejhs.v29i5.15 (2019).
Cambray, G., Guerout, A. M. & Mazel, D. Integrons. Annu. Rev. Genet. 44, 141–166. https://doi.org/10.1146/annurev-genet-102209-163504 (2010).
Barlow, R. S., Pemberton, J. M., Desmarchelier, P. M. & Gobius, K. S. Isolation and characterization of integron-containing bacteria without antibiotic selection. Antimicrob. Agents Chemother. 48, 838–842. https://doi.org/10.1128/aac.48.3.838-842.2004 (2004).
Shibata, N. et al. PCR typing of genetic determinants for metallo-beta-lactamases and integrases carried by gram-negative bacteria isolated in Japan, with focus on the class 3 integron. J. Clin. Microbiol. 41, 5407–5413. https://doi.org/10.1128/jcm.41.12.5407-5413.2003 (2003).
Kargar, M., Mohammadalipour, Z., Doosti, A., Lorzadeh, S. & Japoni-Nejad, A. High prevalence of class 1 to 3 integrons among multidrug-resistant diarrheagenic Escherichia coli in Southwest of Iran. Osong. Public Health Res. Perspect. 5, 193–198. https://doi.org/10.1016/j.phrp.2014.06.003 (2014).
Han, N., Sheng, D. & Xu, H. Role of Escherichia coli strain subgroups, integrons, and integron-associated gene cassettes in dissemination of antimicrobial resistance in aquatic environments of Jinan, China. Water Sci. Technol. 66, 2385–2392. https://doi.org/10.2166/wst.2012.473 (2012).
Ramos, S. et al. Escherichia coli as commensal and pathogenic bacteria among food-producing animals: Health Implications of Extended Spectrum beta-lactamase (ESBL) Production. Animals (Basel). https://doi.org/10.3390/ani10122239 (2020).
Mazurek, J., Bok, E., Stosik, M. & Baldy-Chudzik, K. Antimicrobial resistance in commensal Escherichia coli from pigs during metaphylactic trimethoprim and sulfamethoxazole treatment and in the post-exposure period. Int. J. Environ. Res. Public Health 12, 2150–2163. https://doi.org/10.3390/ijerph120202150 (2015).
Garcia-Aljaro, C., Moreno, E., Andreu, A., Prats, G. & Blanch, A. R. Phylogroups, virulence determinants and antimicrobial resistance in stx(2) gene-carrying Escherichia coli isolated from aquatic environments. Res. Microbiol. 160, 585–591. https://doi.org/10.1016/j.resmic.2009.08.004 (2009).
Drugdova, Z. & Kmet, V. Prevalence of beta-lactam and fluoroquinolone resistance, and virulence factors in Escherichia coli isolated from chickens in Slovakia. Biologia 68, 11–17. https://doi.org/10.2478/s11756-012-0142-6 (2013).
Kaushik, M., Kumar, S., Kapoor, R. K., Virdi, J. S. & Gulati, P. Integrons in Enterobacteriaceae: Diversity, distribution and epidemiology. Int. J. Antimicrob. Agents 51, 167–176. https://doi.org/10.1016/j.ijantimicag.2017.10.004 (2018).
Seo, K. W. & Lee, Y. J. Prevalence and characterization of beta-lactamases genes and class 1 integrons in multidrug-resistant Escherichia coli isolates from chicken meat in Korea. Microb. Drug Resist. https://doi.org/10.1089/mdr.2018.0019 (2018).
Stine, O. C. et al. Widespread distribution of tetracycline resistance genes in a confined animal feeding facility. Int. J. Antimicrob. Agents 29, 348–352. https://doi.org/10.1016/j.ijantimicag.2006.11.015 (2007).
Carattoli, A. Animal reservoirs for extended spectrum beta-lactamase producers. Clin. Microbiol. Infect. 14, 117–123. https://doi.org/10.1111/j.1469-0691.2007.01851.x (2008).
Silbergeld, E. K., Graham, J. & Price, L. B. Industrial food animal production, antimicrobial resistance, and human health. Annu. Rev. Public Health 29, 151–169. https://doi.org/10.1146/annurev.publhealth.29.020907.090904 (2008).
Cully, M. Public health: The politics of antibiotics. Nature 509, S16-17. https://doi.org/10.1038/509S16a (2014).
EMA. Sales of veterinary antimicrobial agents in 30 European countries in 2016. (European Surveillance of Veterinary Antimicrobial Consumption, EMA/275982/2018, 2018).
Majewski, M., Anusz, K., Belkot, Z., Racewicz, P. & Lukomska, A. Impact of residues of veterinary medicinal products in food of animal origin on public health safety in Poland in the years 2003–2017. Med. Weter. 76, 416–422. https://doi.org/10.21521/mw.6412 (2020).
ECDC, EFSA & EMA. Second joint report on the integrated analysis of the consumption of antimicrobial agents and occurrence of antimicrobial resistance in bacteria from humans and food-producing animals. https://doi.org/10.2903/j.efsa.2017.4872 (2017).
EFSA. European Union summary report on antimicrobial resistance in zoonotic and indicator bacteria from humans, animals and food in 2015. Efsa J. https://doi.org/10.2903/j.efsa.2017.4694 (2017).
Roth, N. et al. The application of antibiotics in broiler production and the resulting antibiotic resistance in Escherichia coli: A global overview. Poult. Sci. 98, 1791–1804. https://doi.org/10.3382/ps/pey539 (2019).
Akinbami, O. R., Olofinsae, S. & Ayeni, F. A. Prevalence of extended spectrum beta lactamase and plasmid mediated quinolone resistant genes in strains of Klebsiella pneumonia, Morganella morganii, Leclercia adecarboxylata and Citrobacter freundii isolated from poultry in South Western Nigeria. PeerJ https://doi.org/10.7717/peerj.5053 (2018).
Saliu, E. M., Vahjen, W. & Zentek, J. Types and prevalence of extended-spectrum beta-lactamase producing Enterobacteriaceae in poultry. Anim. Health Res. Rev. 18, 46–57. https://doi.org/10.1017/s1466252317000020 (2017).
Li, L. L., Ye, L., Kromann, S. & Meng, H. C. Occurrence of extended-spectrum beta-lactamases, plasmid-mediated quinolone resistance, and disinfectant resistance genes in Escherichia coli isolated from ready-to-eat meat products. Foodborne Pathog. Dis. 14, 109–115. https://doi.org/10.1089/fpd.2016.2191 (2017).
Seo, K. W. & Lee, Y. J. Prevalence and characterization of plasmid mediated quinolone resistance genes and class 1 integrons among multidrug-resistant Escherichia coli isolates from chicken meat. J. Appl. Poultry Res. 28, 761–770. https://doi.org/10.3382/japr/pfz016 (2019).
Piekarska, K. et al. Prevalence of qnr genes in clinical Enterobacteriaceae non-susceptible to fluoroquinolone in Poland. Med. Dosw. Mikrobiol. 64, 211–219 (2012).
Kindle, P. et al. Phenotypic and genotypic characteristics of Escherichia coli with non-susceptibility to quinolones isolated from environmental samples on pig farms. Porcine Health Manag. 5, 9. https://doi.org/10.1186/s40813-019-0116-y (2019).
Robicsek, A. et al. Fluoroquinolone-modifying enzyme: A new adaptation of a common aminoglycoside acetyltransferase. Nat. Med. 12, 83–88. https://doi.org/10.1038/nm1347 (2006).
Hansen, L. H., Jensen, L. B., Sorensen, H. I. & Sorensen, S. J. Substrate specificity of the OqxAB multidrug resistance pump in Escherichia coli and selected enteric bacteria. J. Antimicrob. Chemother. 60, 145–147. https://doi.org/10.1093/jac/dkm167 (2007).
Amin, M., Dibachi, S., Shahin, M. Prevalence of class 1 integrons and plasmid-mediated qnr-genes among Enterobacter isolates obtained from hospitalized patients in Ahvaz, Iran. Infezioni Medicina, 351–357 (2017).
Sarmah, A. K., Meyer, M. T. & Boxall, A. B. A. A global perspective on the use, sales, exposure pathways, occurrence, fate and effects of veterinary antibiotics (VAs) in the environment. Chemosphere 65, 725–759. https://doi.org/10.1016/j.chemosphere.2006.03.026 (2006).
Daghrir, R. & Drogui, P. Tetracycline antibiotics in the environment: A review. Environ. Chem. Lett. 11, 209–227. https://doi.org/10.1007/s10311-013-0404-8 (2013).
Borghi, A. A. & Palma, M. A. Tetracycline: Production, waste treatment and environmental impact assessment. Braz. J. Pharm. Sci. 50, 25–40. https://doi.org/10.1590/s1984-82502011000100003 (2014).
Jahantigh, M., Samadi, K., Dizaji, R. E. & Salari, S. Antimicrobial resistance and prevalence of tetracycline resistance genes in Escherichia coli isolated from lesions of colibacillosis in broiler chickens in Sistan, Iran. BMC Vet. Res. https://doi.org/10.1186/s12917-020-02488-z (2020).
Kietzmann, M. & Baumer, W. Oral medication via feed and water - pharmacological aspects. Deutsche Tierarztliche Wochenschrift 116, 204–208. https://doi.org/10.2376/0341-6593-116-204 (2009).
Furtula, V. et al. Veterinary pharmaceuticals and antibiotic resistance of Escherichia coli isolates in poultry litter from commercial farms and controlled feeding trials. Poult. Sci. 89, 180–188. https://doi.org/10.3382/ps.2009-00198 (2010).
Font-Palma, C. Characterisation, kinetics and modelling of gasification of poultry manure and litter: An overview. Energy Convers. Manage. 53, 92–98. https://doi.org/10.1016/j.enconman.2011.08.017 (2012).
Islam M.J., S. S., Das K.K., Sharmin N. and Hasan M.N. Isolation of Plasmid- Mediated Multidrug Resistant Escherichia coli from Poultry. Int. J. Sustain. Crop Prod. 3, 46–50 (2008).
Marshall, B. M. & Levy, S. B. Food animals and antimicrobials: Impacts on human health. Clin. Microbiol. Rev. 24, 718. https://doi.org/10.1128/cmr.00002-11 (2011).
Van, T. T. H., Chin, J., Chapman, T., Tran, L. T. & Coloe, P. J. Safety of raw meat and shellfish in Vietnam: An analysis of Escherichia coli isolations for antibiotic resistance and virulence genes. Int. J. Food Microbiol. 124, 217–223. https://doi.org/10.1016/j.ijfoodmicro.2008.03.029 (2008).
Blahna, M. T. et al. The role of horizontal gene transfer in the spread of trimethoprim-sulfamethoxazole resistance among uropathogenic Escherichia coli in Europe and Canada. J. Antimicrob. Chemother. 57, 666–672. https://doi.org/10.1093/jac/dkl020 (2006).
Jouini, A. et al. Detection of multiple-antimicrobial resistance and characterization of the implicated genes in Escherichia coli isolates from foods of animal origin in Tunis. J. Food Prot. 72, 1082–1088. https://doi.org/10.4315/0362-028x-72.5.1082 (2009).
Vinue, L. et al. Genetic environment of sul genes and characterisation of integrons in Escherichia coli isolates of blood origin in a Spanish hospital. Int. J. Antimicrob. Agents 35, 492–496. https://doi.org/10.1016/j.ijantimicag.2010.01.012 (2010).
Soufi, L. et al. Escherichia coli of poultry food origin as reservoir of sulphonamide resistance genes and integrons. Int. J. Food Microbiol. 144, 497–502. https://doi.org/10.1016/j.ijfoodmicro.2010.11.008 (2011).
Adelowo, O. O., Fagade, O. E. & Agerso, Y. Antibiotic resistance and resistance genes in Escherichia coli from poultry farms, southwest Nigeria. J. Infect. Dev. Ctries. 8, 1103–1112. https://doi.org/10.3855/jidc.4222 (2014).
Cavicchio, L. et al. Class 1 and class 2 integrons in avian pathogenic Escherichia coli from poultry in Italy. Poult. Sci. 94, 1202–1208. https://doi.org/10.3382/ps/pev095 (2015).
Kang, H. Y. et al. Characterization of antimicrobial resistance and class 1 integrons found in Escherichia coli isolates from humans and animals in Korea. J. Antimicrob.. Chemother. 55, 639–644. https://doi.org/10.1093/jac/dki076 (2005).
Dakic, I., Petrikkos, G., Dimitrijevic, V. & Charvalos, E. Multidrug resistance and integrons in Escherichia coli isolated from chicken in greece. Acta Vet.-Beogr. 61, 575–584. https://doi.org/10.2298/avb1106575d (2011).
White, P. A., McIver, C. J. & Rawlinson, W. D. Integrons and gene cassettes in the enterobacteriaceae. Antimicrob. Agents Chemother. 45, 2658–2661. https://doi.org/10.1128/aac.45.9.2658-2661.2001 (2001).
Fonseca, E. L., Vieira, V. V., Cipriano, R. & Vicente, A. C. P. Class 1 integrons in Pseudomonas aeruginosa isolates from clinical settings in Amazon region, Brazil. FEMS Immunol. Med. Microbiol. 44, 303–309. https://doi.org/10.1016/j.femsim.2005.01.004 (2005).
Dessie, H. K., Bae, D. H. & Lee, Y. J. Characterization of integrons and their cassettes in Escherichia coli and Salmonella isolates from poultry in Korea. Poult. Sci. 92, 3036–3043. https://doi.org/10.3382/ps.2013-03312 (2013).
EU. Commission Regulation (EU) No 200/2012 of 8 March 2012 concerning a Union target for the reduction of Salmonella enteritidis and Salmonella typhimurium in flocks of broilers, as provided for in Regulation (EC) No 2160/2003 of the European Parliament and of the Council Text with EEA relevance. In: Regulation, C. (Ed.) https://data.europa.eu/eli/reg/2012/200/oj (2012).
Fang, H., Ataker, F., Hedin, G. & Dornbusch, K. Molecular epidemiology of extended-spectrum beta-lactamases among Escherichia coli isolates collected in a Swedish hospital and its associated health care facilities from 2001 to 2006. J. Clin. Microbiol. 46, 707–712. https://doi.org/10.1128/jcm.01943-07 (2008).
Marti, E. & Balcazar, J. L. Real-time PCR assays for quantification of qnr genes in environmental water samples and chicken feces. Appl. Environ. Microbiol. 79, 1743–1745. https://doi.org/10.1128/aem.03409-12 (2013).
Kozak, G. K., Boerlin, P., Janecko, N., Reid-Smith, R. J. & Jardine, C. Antimicrobial resistance in Escherichia coli isolates from swine and wild small mammals in the proximity of swine farms and in Natural Environments in Ontario, Canada. Appl. Environ. Microbiol. 75, 559–566. https://doi.org/10.1128/aem.01821-08 (2009).
Hu, X. et al. A high throughput multiplex PCR assay for simultaneous detection of seven aminoglycoside-resistance genes in Enterobacteriaceae. BMC Microbiol. 13, 58. https://doi.org/10.1186/1471-2180-13-58 (2013).
Grape, M., Motakefi, A., Pavuluri, S. & Kahlmeter, G. Standard and real-time multiplex PCR methods for detection of trimethoprim resistance dfr genes in large collections of bacteria. Clin. Microbiol. Infect. 13, 1112–1118. https://doi.org/10.1111/j.1469-0691.2007.01807.x (2007).
R Core Team. A language and environment for statistical computing. Foundation for Statistical Computing, Vienna, Austria (2021).
Levesque, C., Piche, L., Larose, C. & Roy, P. H. PCR mapping of integrons reveals several novel combinations of resistance genes. Antimicrob. Agents Chemother. 39, 185–191. https://doi.org/10.1128/AAC.39.1.185 (1995).
White, D. G. et al. The isolation of antibiotic-resistant salmonella from retail ground meats. N. Engl. J. Med. 345, 1147–1154. https://doi.org/10.1056/NEJMoa010315 (2001).
Funding
The Publication was co-financed within the framework of the Polish Ministry of Science and Higher Education’s program: “Regional Initiative Excellence” in the years 2019–2022 (No. 005/RID/2018/19) “financing amount 12 000 000,00 PLN”.
Author information
Authors and Affiliations
Contributions
P.R., M.M. and Z.M. designed experiments and prepared the manuscript. J.W., D.W., M.K., H.B. performed experimental work. P.R. and Z.M. assembled and analyzed the genomic data. M.P. carried out the statistical analyses. S.N. supervised the work progress and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Racewicz, P., Majewski, M., Biesiada, H. et al. Prevalence and characterisation of antimicrobial resistance genes and class 1 and 2 integrons in multiresistant Escherichia coli isolated from poultry production. Sci Rep 12, 6062 (2022). https://doi.org/10.1038/s41598-022-09996-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-09996-y
- Springer Nature Limited
This article is cited by
-
Antibiotic Resistance and Virulence Gene Patterns Associated with Multi Drug Resistant Avian Pathogenic Escherichia coli (APEC) Isolated from Broiler Chickens in India
Indian Journal of Microbiology (2024)
-
Global trends in antimicrobial resistance on organic and conventional farms
Scientific Reports (2023)
-
The pan-genome of the emerging multidrug-resistant pathogen Corynebacterium striatum
Functional & Integrative Genomics (2023)
-
Molecular characterization of multi drug resistant Escherichia coli isolates at a tertiary hospital in Abuja, Nigeria
Scientific Reports (2022)