Abstract
The loglinear pattern of respiratory scaling has been studied for over a century, while an increasing number of non-loglinear patterns have been found in the plant kingdom. Several previous studies had attempted to reconcile conflicting patterns from the aspects of statistical approaches and developmental stages of the organisms. However, the underlying enzymatic mechanism was largely ignored. Here, we propose an enzyme-driven law of photosynthetic scaling and test it in typical crop seedlings under different water conditions. The results showed that the key enzyme activity, the relative photosynthetic assimilation and the relative growth rate were all constrained by the available water, and the relationship between these biological traits and the available water supported our predictions. The enzyme-driven law appears to be more suitable to explain the curvature of photosynthetic scaling than the well-established power law, since it provides insight into the biochemical origin of photosynthetic assimilation.
Similar content being viewed by others
Introduction
Photosynthesis is the fundamental metabolism for most of world’s ecosystems1. The scaling model of the power law has been widely employed to describe net primary production scaling2,3,4,5 in log–log space. Some researchers have applied log-transformed power law to dark respiration rates in trees6,7,8 and shrubs9. A few authors have used photosynthetic rates as metabolic indicators in ocean and plants10,11, but the subjects are marine systems and saplings, respectively. However, the relationship between photosynthetic rate and biomass in crop plants has not been studied under the light of power law.
Photosynthetic assimilation and respiration are both important metabolic processes, and they reverse each other. However, metabolic scaling of basic respiration is a longstanding problem12,13, and towards its resolution, a series of arguments about the power law and the ‘true’ value of scaling exponents have been suggested14,15,16,17. It has theoretically been shown that the scaling exponents should range between 0.31 and 1.00 in aquatic invertebrates and algae18. The models based on polynomial equation have been used to describe and quantify the double-log linear deviation in basic metabolic scaling19,20. However, there are still contentious in the origin of photosynthetic assimilation and the methods used to estimate parameters21.
Many log–log relationships of metabolic scaling have been shown to be nonlinear and cannot be represented by a simple power function20,21,22. Transformations of logarithms are referred to as patterns of “non-loglinear allometry”23 or patterns of “nonlinear allometry”24, which can be represented by adding other terms based on power laws25,26,27 or energy dynamics13,28. Models can explain the variation in scaling exponents in terms of resource allocation (Dynamic Energy Budget Model)29, metabolic level30 and thermodynamics processes31. A sigmoidal model predicted by the plant adaptive strategies hypothesis in NPP scaling1. However, although photosynthesis and respiration are a series of biochemical reactions, chemical kinetics is rarely used to explain the challenges of metabolic scaling. We have developed an enzyme-driven law (EDL), which was successfully applied to quantify the curvature and exponent variations of respiration scaling in microbes32. For terrestrial plants, photosynthesis differs significantly from respiration in microbes. Thus, a modified law is required to describe the photosynthetic scaling by incorporating biochemical mechanisms.
In this study, a new enzyme-driven law for photosynthetic scaling was derived from the hypothesis and both the relative rates of photosynthetic assimilation and growth were constrained by the key enzymatic activities. The law was tested by the data of photosynthesis and growth under different water conditions in represented crop seedlings (rice and maize33,34) to explain the biochemical origin of curvature in photosynthetic scaling.
Results
Key enzymatic constraint in relative photosynthetic assimilation and growth
Our experiment supported the basic hypothesis, both relative photosynthetic assimilation and growth were constrained by the activity of key enzymes (Eq. 1). The key enzyme activity (RuBPcase in rice, PEPcase in maize) increased linearly with relative photosynthetic assimilation (Fig. 1A), and relative change in body mass (Fig. 1B). The photosynthetic key enzyme drove the assimilation and growth respectively in both rice (C3) and maize (C4) with different parameters (Fig. 1).
The water potential dependence of relative photosynthetic assimilation and relative growth
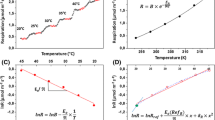
The available water was the limited substrate constrained by the key enzyme in C3 (rice) and C4 (maize) plants under the experiment conditions. Key enzyme activities varied with available water in accordance with the Michaelis–Menten equation (Fig. 2), which meant that available water could be treated as a limiting substrate for the key pathway of photosynthesis at least for seeding stage. The K value of available water in RuBPcase (0.6) was bigger than that in PEPCase (0.04), which meant PEPCase was more sensitive to available water; the water potential of rice was lower than that in maize when the key enzyme activity was zero (Fig. 2).
The activity of the two key enzymes changed with a gradient of available water nonlinearly. PEPCase activity in maize (solid circle) correspond to right vertical axis, and RuBPCase activity in rice (open triangle) correspond to left vertical axis. The total sample number of each treatment is fifteen.
As an important part of plants, water is directly involved in important metabolic processes in plants, one of the raw materials for plant photosynthesis and a medium for many important biochemical reactions. Meanwhile, water stress is one of the main reasons for crop yield reduction under drought conditions. Water potential could be steadily controlled relatively and affected photosynthesis and growth synchronously.
The pattern of water potential dependence appeared in relative photosynthetic assimilation and relative growth at the individual level (Fig. 3). The trends of relative photosynthetic assimilation and relative growth versus water potential were consistent in both rice and maize. The Kq and Km in rice (0.32, 0.28) were bigger than that in maize (0.07, 0.18), which meant maize was more sensitive to available water; The water potential in rice was lower than that in maize when the key enzyme activity was zero (Fig. 3). Equations 2 and 3 were checked and the results were in accordance with our predictions.
The interdependence between relative photosynthetic assimilation (lnQ) and growth (lnM)
The interdependent prediction (Eq. 4) was supported by the relationship between relative photosynthetic assimilation (lnQ) and growth (lnM) (Fig. 4). Meanwhile, the curvatures of the scaling between lnQ and lnM were different in rice and maize, because different species had different Kq and Km (Eqs. 2, 3, Fig. 3). The effect of body size on relative rate of metabolism followed our assumptions at individual level (Fig. 4). The AIC values of traditional law were higher than that of the enzyme-driven law in rice and maize, respectively (Table 1). While the R2 value of enzyme driven law were also higher than that of traditional law in rice and maize, which matched and verified with AIC value. It meant that enzyme-driven law was better than traditional law.
Discussion
The enzyme-driven law had been tested gradually from enzymatic (Eq. 1, Fig. 1), resource response (Eqs. 2, 3, Figs. 2, 3) to photosynthetic scaling (Eq. 4, Fig. 4). The AIC value indicated the enzyme-driven law (Eq. 4) was better than the traditional power law (although the traditional power equation could also be derived from the enzyme-driven hypothesis under special conditions (Eq. 5). The enzymatic dynamics had been used to describe the scaling of the metabolic balance in the micro algae of oceans10,11, in which the effect of body size on the metabolism had not been considered. However, the remarkable36 effects of body size on the metabolism35 could not be ignored. It is widely acceptable that both the metabolism and growth are a series of enzymatic process, which are constrained by the key enzymatic activities37, so it is reasonable to deduce the general law of metabolic scaling. The enzyme-driven law may be the origin of various metabolic scaling.
The AIC comparison shown that enzyme-driven law was better (Table 1) than the power law for the photosynthetic scaling. As these two equations could be deduced from enzyme-driven mechanism (Eq. 5), the enzyme-driven law might be more general than the traditional power law in photosynthetic scaling. Instead, previous studies on the curvilinear scaling were based on traditional power law with incorporation of other terms that derived from statistics25,26,27 or the energy dynamics13,28. The constraint factor of limited resource(s) have been considered in metabolic level boundary hypothesis, which was used to explain the variations of scaling exponent against metabolic level30. Moreover, the similar nonlinear pattern of metabolic scaling had also been found in teleost fish38. Meanwhile, a model of growth scaling was used to describe the growth of many diverse species proposed with parameter-less curve39. In addition, the polynomial equation of energy dynamics had been used to describe the nonlinear curve of active metabolic scaling during ontogeny in birds and mammals28. Instead, our equation (Eq. 4) could be used to describe the active and maintenance metabolic scaling, either linear or nonlinear pattern of scaling at log-scale. Overall, two kinds of linear and nonlinear pattern of photosynthetic scaling could be described by enzyme-driven law (Eqs. 4, 5), which might promote the quantitative integration between biochemical mechanism and ecological scaling.
Material and methods
Material and treatments
Maize (Zea mays L. Zhefengnuo NO.3) and rice (Oryza sativa L. Nipponbare) are representative species of the C4/C3 photosynthetic pathways33,34 and therefore were used in this study. Seedlings of two leaves were cultivated in black plastic buckets filled with Hoagland solution (pH 6.0) under a series of water potentials, which were adjusted to using polyethyleneglycol 6,000 (PEG-6000). Before the stage of two leaves, photosynthetic photon flux density (PPFD) and temperature were controlled in a stepwise diurnal sequence, with daily PPFD of 300 μmol m−2 s−1, and a photoperiod of 16 h. Temperature varied between 25 ℃/daily and 23 ℃/might.
Methods of measurements
A portable open system (LI-6400; Li-Cor, Inc., Lincoln, NE, USA), equipped with a CO2 mixer, 30 mm × 20 mm chamber and red-blue LED light source (6400-2B) was used for the measurement of photosynthetic rate. Sample CO2, block temperature and photosynthetic active radiation (PAR) were set at 400 μmol mol−1, 36 ℃ and 1,200 μmol m−2 s−1 respectively. Dry weights of plants were obtained by oven drying at 105 °C for 30 min and then at 65 °C for 48 h.
The fluid enzymes were extracted by the modified li-ren’s method42. 0.2 g of leaf was put into a precooled mortar, then 100 mmol/ L of precooled Tris–HCl buffer (contained 7 mmol/L β-Mercaptoethanol, 1 mmol/ L EDTA, 5% glycerol and 1% PVP, pH 8.2); This sample was then centrifuged 4 ℃ for 20 min at 15,000 r/m. Then, we used supernatant fluid to measure the activity of RuBPCase and PEPCase with spectrophotometry43.
Models
According to metabolism is a series of enzymatic processes37, we hypothesized that the relative change in photosynthetic assimilation (dQ/Q) was constrained by the change in key enzymatic activity (dvq) at individual level, including the effects of body size8,40. Hence, the relationship between photosynthetic assimilation rate (Q) and the key enzymatic activities (v) was obtained:
The relationship between v and substrate concentration S was assumed to abide by the Michaelis–Menten equation, therefore the relationship between Q and S can be described as follows:
where lnQm is the maximum metabolic rate when S approaches saturation, Kq is a half-saturation constant (i.e., the substrate concentration at which the rate of substrate conversion is equal to lnQm/2), R represent resource. S = R − R0, lnQ0 = lnQ when S equals zero.
We could hypothesize that the relative change in growth (\(\frac{dm}{m}\)) was also constrained by change in the key enzyme activity (dvg)41, which included the effects of body size on the growth8,40. The substrate-dependent equation of logarithmic body mass can be obtained by the integration of \(\frac{dm}{m} \le dv_{g}\):
where lnMm is the maximum body mass when S approaches saturation. lnM = lnM0 when S = 0, Km is the half-saturation constant.
The enzyme-driven law of photosynthetic scaling were obtained by integrating Eqs. 2 and 3:
- 1.
The interdependent relationship between lnQ and lnM was obtained When Kq ≠ Km:
$$ \ln Q \le \frac{{\ln Q_{Mm} \times (\ln M - \ln M_{0} )}}{{K_{qm} + (\ln M - \ln M_{0} )}} + \ln Q_{0} $$(4)where \(\ln Q_{Mm} = \frac{{K_{m} \times \ln Q_{m} }}{{K_{m} - K_{q} }}\), \(K_{qm} = \frac{{K_{q} \times \ln M_{m} }}{{K_{m} - K_{q} }}\), it displayed that lnQ varied a saturation curve with lnM
- 2.
The log-transformed power equation was obtained, that is, lnQ increases linearly with lnM when Kq = Km:
$$ \ln Q \le \frac{{\ln Q_{m} }}{{\ln M_{m} }}\ln M + \ln Q_{0} - \frac{{\ln Q_{m} }}{{\ln M_{m} }}\ln M_{0} $$(5)where lnQm/lnMm is the exponent in the original power equation. Equation 5 shows that the famous traditional power law is a special form of the enzyme dynamics. Equations 4 and 5 both could be called as the enzyme-driven law of metabolic scaling.
Statistical analysis
All the mathematical regressions and statistical analysis were performed in Origin 8.5. The methods were compared by the Akaike's information criterion (AIC)39. The model with the lowest AIC was regarded as the best representation of a curve44.
References
Jenkins, D. G. & Pierce, S. General allometric scaling of net primary production agrees with plant adaptive strategy theory and has tipping points. J. Ecol. 105, 1094–1104 (2017).
Kerkhoff, A. J. & Enquist, B. J. Ecosystem allometry: the scaling of nutrient stocks and primary productivity across plant communities. Ecol. Lett. 9, 419–427 (2006).
Coomes, D. A., Lines, E. R. & Allen, R. B. Moving on from metabolic scaling theory: hierarchical models of tree growth and asymmetric competition for light. J. Ecol. 99, 748–756 (2011).
Michaletz, S. T., Cheng, D., Kerkhoff, A. J. & Enquist, B. J. Convergence of terrestrial plant production across global climate gradients. Nature 512, 39–43 (2014).
Hatton, I. A. et al. The predator-prey power law: biomass scaling across terrestrial and aquatic biomes. Science 349, 1070 (2015).
Peng, Y., Niklas, K. J., Reich, P. B. & Sun, S. Ontogenetic shift in the scaling of dark respiration with whole-plant mass in seven shrub species. Funct. Ecol. 24, 502–512 (2010).
Reich, P. B., Tjoelker, M. G., Machado, J. & Oleksyn, J. Universal scaling of respiratory metabolism, size and nitrogen in plants. Nature 439, 457–461 (2006).
Cheng, D. L., Li, T., Zhong, Q. L. & Wang, G. X. Scaling relationship between tree respiration rates and biomass. Bio Lett. 6, 715–717 (2010).
Xu, K. et al. Indirect effects of water availability in driving and predicting productivity in the Gobi desert. Sci. Total Environ. 133952 (2019).
Lopez-Urrutia, A., San Martin, E., Harris, R. P. & Irigoien, X. Scaling the metabolic balance of the oceans. Proc. Natl. Acad. Sci. USA 103, 8739–8744 (2006).
Chen, X. & Li, B. Testing the allometric scaling relationships with seedlings of two tree species. Acta Oecol. 24, 125–129 (2003).
Brown, J. H., Gillooly, J. F., Allen, A. P., Savage, V. M. & West, G. B. Toward a metabolic theory of ecology. Ecology 85, 1771–1789 (2004).
Norin, T. & Gamperl, A. K. Metabolic scaling of individuals vs. populations: evidence for variation in scaling exponents at different hierarchical levels. Funct. Ecol. 32, 379–388 (2018).
Li, H. T., Han, X. G. & Wu, J. G. Lack of evidence for 3/4 scaling of metabolism in terrestrial plants. J. Integr. Plant Biol. 47, 1173–1183 (2005).
Hoppeler, H. Scaling functions to body size: theories and facts. J. Exp. Biol. 208, 1573–1574 (2005).
West, G. B., Brown, J. H. & Enquist, B. J. A general model for the origin of allometric scaling laws in biology. Science 276, 122–126 (1997).
Hui, D. & Jackson, R. B. Uncertainty in allometric exponent estimation: a case study in scaling metabolic rate with body mass. J. Theor. Biol. 249, 168–177 (2007).
Patterson, M. R. A mass-transfer explanation of metabolic scaling relations in some aquatic invertebrates and algae. Science 255, 1421–1423 (1992).
Dodds, P. S., Rothman, D. H. & Weitz, J. S. Re-examination of the ’ “3/4-law” of metabolism. J. Theor. Biol. 209, 9–27 (2001).
Kozlowski, J. & Konarzewski, M. West, Brown and Enquist’s model of allometric scaling again: the same questions remain. Funct. Ecol. 19, 739–743 (2005).
Packard, G. C. & Birchard, G. F. Traditional allometric analysis fails to provide a valid predictive model for mammalian metabolic rates. J. Exp. Biol. 211, 3581–3587 (2008).
Glazier, D. S. Beyond the “3/4 power-law”: Variation in the intra- and interspecific scaling of metabolic rate in animals. Biol. Rev. 80, 611–662 (2005).
Strauss, R. E. & Huxley, J. S. The study of allometry since Huxley. In: Problems of Relative Growth (Johns Hopkins University Press, Baltimore, 1993).
Knell, R. J. On the analysis of non-linear allometries. Ecol. Entomol. 34, 1–11 (2009).
Stumpf, M. P. H. & Porter, M. A. Critical truths about power laws. Science 335, 665–666 (2012).
Kolokotrones, T., Savage, V., Deeds, E. J. & Fontana, W. Curvature in metabolic scaling. Nature 464, 753–756 (2010).
Packard, G. C. Why allometric variation in mammalian metabolism is curvilinear on the logarithmic scale. J. Exp. Zoo A Ecol. Integr. Physiol. 327, 537–541 (2017).
Hou, C. et al. Energy uptake and allocation during ontogeny. Science 322, 736–739 (2008).
Ledder, G. The basic dynamic energy budget model and some implications. Lett. Biomath. 1, 221–233 (2014).
Glazier, D. S. A unifying explanation for diverse metabolic scaling in animals and plants. Biol. Rev. 85, 111–138 (2010).
Ballesteros, F. J. et al. On the thermodynamic origin of metabolic scaling. Sci. Rep. 8, 1448 (2018).
Li, L. & Wang, G. Enzymatic origin and various curvatures of metabolic scaling in microbes. Sci. Rep. 9 (2019).
Imaizumi, N., Usuda, H., Nakamoto, H. & Ishihara, K. Changes in the rate of photosynthesis during grain filling and the enzymatic activities associated with the photosynthetic carbon metabolism in rice panicles. Plant Cell Physiol. 31, 835–843 (1990).
Langdale, J. A. & Nelson, T. Spatial regulation of photosynthetic development in C4 plants. Trends Genet. 7, 191–196 (1991).
Deng, J. et al. Insights into plant size-density relationships from models and agricultural crops. Proc. Natl. Acad. Sci. 109, 8600–8605 (2012).
Deng, J. et al. Plant mass-density relationship along a moisture gradient in north-west China. J. Ecol. 94, 953–958 (2006).
Webb, J. L. Enzyme and Metabolic Inhibitors (Academic Press, New York, 1966).
Killen, S. S., Atkinson, D. & Glazier, D. S. The intraspecific scaling of metabolic rate with body mass in fishes depends on lifestyle and temperature. Ecol. Lett. 13, 184–193 (2010).
Akaike, H. A new look at the statistical model identification. IEEE Trans. Autom. Control 19, 716–723 (1974).
Mori, S. et al. Mixed-power scaling of whole-plant respiration from seedlings to giant trees. Proc. Natl. Acad. Sci. 107, 1447–1451 (2010).
Scott, M., Gunderson, C. W., Mateescu, E. M., Zhang, Z. & Hwa, T. Interdependence of cell growth and gene expression: origins and consequences. Science 330, 1099–1102 (2010).
Li, L. R. W. W. the regulation of ribulose-1, 5-biosphosphate carboxylase activation in alealfa leaves. Acta Phytophysiol. Sin. 33–39 (1986).
Yonghua D., J. S. G. L. Effect of ABA and 6-BA on activity of key enzymes in carbon assimilation in maize seedlings under water stress. Plant Nutr. Fert. Sci. 182–188 (1997).
Yamaoka, K., Nakagawa, T. & Uno, T. Application of Akaike’s information criterion (AIC) in the evaluation of linear pharmacokinetic equations. J. Pharmacokinet. Biopharm. 6, 165–175 (1978).
Acknowledgements
We thank Dr. Kun Jiang for his critical reading of this manuscript. This work was financed by the Natural Science Fund of China (31330010).
Author information
Authors and Affiliations
Contributions
Z.W. and G.W. conceived the ideas and designed methodology. Z.W., L.H., K.X., H.H., M.L. and S.L. preformed the experiments and collected the data. Z.W. and G.W. coordinated data analysis and interpreted results. Z.W., K.X., Y.L., A.K. and G.W. wrote and revised the main manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Z., He, L., Xu, K. et al. An non-loglinear enzyme-driven law of photosynthetic scaling in two representative crop seedlings under different water conditions. Sci Rep 10, 12720 (2020). https://doi.org/10.1038/s41598-020-69702-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-69702-8
- Springer Nature Limited