Abstract
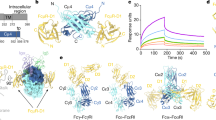
Immunoglobulin Fc receptors are cell surface transmembrane proteins that bind to the Fc constant region of antibodies and play critical roles in regulating immune responses by activation of immune cells, clearance of immune complexes and regulation of antibody production. FcμR is the immunoglobulin M (IgM) antibody isotype-specific Fc receptor involved in the survival and activation of B cells. Here we reveal eight binding sites for the human FcμR immunoglobulin domain on the IgM pentamer by cryogenic electron microscopy. One of the sites overlaps with the binding site for the polymeric immunoglobulin receptor (pIgR), but a different mode of FcμR binding explains its antibody isotype specificity. Variation in FcμR binding sites and their occupancy reflects the asymmetry of the IgM pentameric core and the versatility of FcμR binding. The complex explains engagement with polymeric serum IgM and the monomeric IgM B-cell receptor (BCR).





Similar content being viewed by others
Data availability
The structural data that support the findings of this study have been deposited in the Protein Data Bank and EM Data bank. The 8:1 FcμR/IgM-Fc model displayed in Fig. 1 has entry number EMD-16150 and PDB 8BPE. The IgM-Fc core with one FcμR at subunit Fcμ1 has entry number EMD-16151 and PDB 8BPF. The IgM subunit Fcμ3 with two FcμR has entry number EMD-16152 and PDB 8BPG. Source data are provided with this paper.
References
Kubagawa, H. et al. Identity of the elusive IgM Fc receptor (FcμR) in humans. J. Exp. Med. 206, 2779–2793 (2009).
Akula, S., Mohammadamin, S. & Hellman, L. Fc receptors for immunoglobulins and their appearance during vertebrate evolution. PLoS ONE 9, e96903 (2014).
Kubagawa, H. et al. The long elusive IgM Fc receptor, FcμR. J. Clin. Immunol. 34, 35–45 (2014).
Li, Y. et al. Structural insights into immunoglobulin M. Science 367, 1014–1017 (2020).
Kumar, N., Arthur, C. P., Ciferri, C. & Matsumoto, M. L. Structure of the secretory immunoglobulin A core. Science 367, 1008–1014 (2020).
Kumar, N., Arthur, C. P., Ciferri, C. & Matsumoto, M. L. Structure of the human secretory immunoglobulin M core. Structure 29, 1–8 (2021).
Wang, Y. et al. Structural insights into secretory immunoglobulin A and its interaction with a pneumococcal adhesin. Cell Res. 30, 602–609 (2020).
Shima, H. et al. Identification of TOSO/FAIM3 as an Fc receptor for IgM. Int. Immunol. 22, 149–156 (2009).
Lloyd, K. A., Wang, J., Urban, B. C., Czajkowsky, D. M. & Pleass, R. J. Glycan-independent binding and internalization of human IgM to FCMR, its cognate cellular receptor. Sci. Rep. 7, 42989 (2017).
Nyamboya, R. A., Sutton, B. J. & Calvert, R. A. Mapping of the binding site for FcμR in human IgM-Fc. Biochim. Biophys. Acta Proteins Proteom. 1868, 140266 (2020).
Skopnik, C. M. et al. Identification of amino acid residues in human IgM Fc receptor (FcµR) critical for IgM binding. Front. Immunol. 11, 618327 (2021).
Putnam, F. W., Florent, G., Paul, C., Shinoda, T. & Shimizu, A. Complete amino acid sequence of the mu heavy chain of a human IgM immunoglobulin. Science 182, 287–291 (1973).
Chen, Q., Menon, R., Calder, L. J., Tolar, P. & Rosenthal, P. B. Cryomicroscopy reveals the structural basis for a flexible hinge motion in the immunoglobulin M pentamer. Nat. Commun. 13, 6314 (2022).
Ma, X. et al. Cryo-EM structures of two human B cell receptor isotypes. Science 377, 880–885 (2022).
Su, Q. et al. Cryo-EM structure of the human IgM B cell receptor. Science 377, 875–880 (2022).
Dong, Y. et al. Structural principles of B-cell antigen receptor assembly. Nature 612, 156–161 (2022).
Kubagawa, H. et al. Differences between human and mouse IgM Fc receptor (FcμR). Int. J. Mol. Sci. 22, 7024 (2021).
Kubagawa, H. et al. Functional roles of the IgM Fc receptor in the immune system. Front. Immunol. 10, 945 (2019).
Ouchida, R. et al. FcμR interacts and cooperates with the B cell receptor to promote B cell survival. J. Immunol. 194, 3096–3101 (2015).
Nguyen, T. T. T. et al. The IgM receptor FcμR limits tonic BCR signaling by regulating expression of the IgM BCR. Nat. Immunol. 18, 321–333 (2017).
Vire, B., David, A. & Wiestner, A. TOSO, the FcµR, is highly expressed on chronic lymphocytic leukemia B cells, internalizes upon IgM binding, shuttles to the lysosome, and is downregulated in response to TLR activation. J. Immunol. 187, 4040–4050 (2013).
Stadtmueller, B. M. et al. The structure and dynamics of secretory component and its interactions with polymeric immunoglobulins. eLife 5, e10640 (2016).
Hamburger, A. E., West, A. P. & Bjorkman, P. J. Crystal structure of a polymeric immunoglobulin binding fragment of the human polymeric immunoglobulin receptor. Structure 12, 1925–1935 (2004).
Rochereau, N. et al. Essential role of TOSO/FAIM3 in intestinal IgM reverse transcytosis. Cell Rep. 37, 110006 (2021).
Martin, S. R., Ramos, A. & Masino, L. (2021). Biolayer interferometry: Protein–RNA interactions. In Protein-Ligand Interactions Methods in Molecular Biology, Vol. 2263 (Daviter, T. et al. eds) https://doi.org/10.1007/978-1-0716-1197-5_16 (Humana, 2021).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Du, Z. et al. The trRosetta server for fast and accurate protein structure prediction. Nat. Protoc. 16, 5634–5651 (2021).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr. D D74, 814–840 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Jakobi, A. J., Wilmanns, M. & Sachse, C. Model-based local density sharpening of cryo-EM maps. eLife 6, e27131 (2017).
Burnley, T., Palmer, C. M. & Winn, M. Recent developments in the CCP-EM software suite. Acta Crystallogr. D 73, 469–477 (2017).
Agirre, J. et al. Privateer: software for the conformational validation of carbohydrate structures. Nat. Struct. Mol. Biol. 22, 833–834 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Acknowledgements
We thank A. Nans of the Structural Biology Science Technology Platform for advice on data collection and computing; A. Purkiss and P. Walker of the Structural Biology Science Technology Platform; the Scientific Computing Science Technology Platform for computational support. This work was supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (CC2106 (P.B.R.), CC2006 (P.T.)), the UK Medical Research Council (CC2106 (P.B.R.), CC2006 (P.T.)) and the Wellcome Trust (CC2106 (P.B.R.), CC2006 (P.T.)). For the purpose of Open Access, the authors have applied a CC BY public copyright license to any Author Accepted Manuscript version arising from this submission.
Author information
Authors and Affiliations
Contributions
Q.C., R.P.M. and L.M. performed experiments. Q.C., R.P.M., L.M., P.T. and P.B.R. contributed to experimental design, data analysis and manuscript writing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Brian Sutton and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Editor recognition statement: Primary Handling Editor: Katarzyna Ciazynska, in collaboration with the Nature Structural & Molecular Biology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Binding of Full-length IgM and IgM-Fc core to surface-immobilised FcμR.
(a) Structure schematic for full-length IgM. (b) Structure schematic for proteolysed IgM-Fc core. (c) Binding of full-length (green) and proteolysed IgM-Fc core (blue) to FcμR monitored by Bio-Layer Interferometry (BLI). Representative data sets for each form of IgM are shown. Instrument response values are plotted against IgM concentration. Fitting curves are shown as black lines. The apparent equilibrium dissociation constants (Kd) for full-length IgM and IgM-Fc core are 0.3 ± 0.1 nM and 0.7 ± 0.1 nM respectively. Technical replicates gave values of 0.4 ± 0.1 nM and 0.7 ± 0.1 nM respectively. Raw data for the plot and the technical replicates are provided in the Source Data file.
Extended Data Fig. 2 Single particle analysis of FcμR/IgM-Fc.
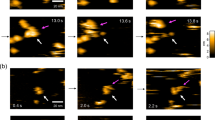
(a) Flow chart of the data processing for both non-tilted and tilted datasets. Steps are conducted in Relion 3.1 (clear box) or Cryosparc 3.2.0 (grey box). (b) A typical micrograph with three particles highlighted in red dotted circles. (c) Typical 2D classes of the complex. (d) 3D auto-refined map. (e) Half-map Fourier shell correlation (FSC) plot showing 3.5 Å global resolution.
Extended Data Fig. 3 Cryo-EM structure of IgM-Fc/FcμR complex.
(a) Particle subset selection by focused 3D classification at the IgM-Fc core and map refinement. (b) Front and side view of the complex model. IgM in grey, FcμR in rainbow colours. (c) Fitting of model and map shown in (a) and (b). (d) Fourier shell correlation (FSC) with 3.6 Å resolution at 0.143 cut-off and map-model FSC plot showing 3.9 Å resolution at 0.5 cut-off calculated with the model shown in (b) calculated in Phenix. (e) Local resolution of the refined map calculated in Cryosparc. (f) Eulerian angle distribution of the particles in the non-uniform refinement. (g) 3DFSC histogram calculated in Cryosparc of the refined map showing anisotropy between 3.2 Å −5.4 Å.
Extended Data Fig. 4 Maps of focused 3D classification at all IgM subunits Fcμ1 to Fcμ5 for quantification of FcμR occupancy at each subunit.
The central EM density is the refined map shown in Extended Data Fig. 3, reconstructed with 516,875 particles. Mask used for focused 3D classification for each subunit (containing Cμ4 dimer, Cμ3 dimer and FcμR) is shown in a specific colour (subunit Fcμ1, red; subunit Fcμ2, orange; subunit Fcμ3, yellow; subunit Fcμ4, green; subunit Fcμ5, pink). The 3D classes show different FcμR binding states (at front, back, both or neither) at each subunit individually. The front and the back of IgM is shown at the top-left of the figure using the same definition described in the main text.
Extended Data Fig. 5 Cryo-EM structure of FcμR/IgM-Fc complex focused on subunit Fcμ1.
(a) Particle subset selection by focused 3D classification at subunit Fcμ1 and map refinement. (b) Front and side view of the complex model. IgM in grey, FcμR in red. (c) Fitting of model and map shown in (a) and (b). (d) Fourier shell correlation (FSC) with 3.5 Å resolution at 0.143 cut-off and map-model FSC plot showing 3.6 Å resolution at 0.5 cut-off calculated in Phenix. (e) Local resolution of the refined map calculated in Cryosparc. (f) Eulerian angle distribution of the particles in the non-uniform refinement. (g) 3DFSC histogram of the refined map calculated in Cryosparc of the refined map showing anisotropy between 3.2 Å −7.6 Å.
Extended Data Fig. 6 Cryo-EM structure of FcμR/IgM-Fc complex focused on subunit Fcμ3.
(a) Particle subset selection by focused 3D classification at subunit Fcμ3 and map refinement. (b) Front and side view of the complex model. IgM in grey, FcμR in yellow. (c) Fitting of model and map shown in (a) and (b). (d) Fourier shell correlation (FSC) with 3.1 Å resolution at 0.143 cut-off and map-model FSC plot showing 3.3 Å resolution at 0.5 cut-off calculated in Phenix. (e) Local resolution of the refined map calculated in Cryosparc. (f) Eulerian angle distribution of the particles in the non-uniform refinement. (g) 3DFSC histogram calculated in Cryosparc of the refined map showing angular resolution distribution from 3.0 Å − 3.8 Å.
Extended Data Fig. 7 Density maps of key regions at FcμR and FcμR/IgM binding interface.
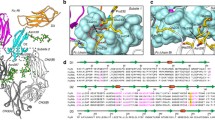
(a–b) The two conserved disulfide bonds in FcμR. (c–g) Densities of the interacting residues on FcμR and Cμ4 domains, corresponding to the interactions shown in Fig. 4b. FcμR in dark yellow, Cμ4-B chain in light grey, and Cμ4-A chain in slate grey. (h) Densities of the residues in CDR3 region of FcμR interacting with the neighbouring Cμ4 domain (in brown), corresponding to Fig. 4c. (i) Densities of the residues in CDR3 regions of FcμR interacting with the tailpiece of Fcμ5 chain (in pink), corresponding to Fig. 4d.
Extended Data Fig. 8 N-linked glycosylation at Asn563 contacting FcμR at subunit Fcμ3.
(a) The tailpiece assembly of the IgM pentamer showing ten N-linked glycosylation sites (purple). (b) The map of subunit Fcμ3 (same map as Extended Data Fig. 6a, before postprocessing, map threshold = 0.2). (c) Zoom-in view of the N-Acetylglucosamine (NAG) molecules (orange) linking from Asn563 (purple) at the tailpiece of Fcμ3B chain to FcμR-3f (yellow). (d) The overall map of FcμR/IgM-Fc (same map as Extended Data Fig. 3a, before postprocessing, map threshold = 0.2). (e) Cross-section of the map in (d) indicated by the red dotted line showing the densities of the two NAG chains (orange arrowheads) extending from Asn563 of the two Fcμ chains (Fcμ3A and Fcμ3B) to the two FcμR molecules at both sides.
Supplementary information
Source data
Source Data Extended Data Fig. 1
Raw data for the BLI experiments.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, Q., Menon, R.P., Masino, L. et al. Structural basis for Fc receptor recognition of immunoglobulin M. Nat Struct Mol Biol 30, 1033–1039 (2023). https://doi.org/10.1038/s41594-023-00985-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-00985-x
- Springer Nature America, Inc.
This article is cited by
-
Secreted IgM modulates IL-10 expression in B cells
Nature Communications (2024)
-
Multi-faceted immunoglobulin M meets its elusive receptor
Nature Structural & Molecular Biology (2023)