Abstract
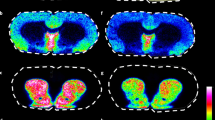
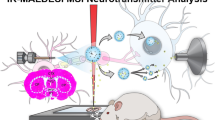
We present a mass spectrometry imaging (MSI) approach for the comprehensive mapping of neurotransmitter networks in specific brain regions. Our fluoromethylpyridinium-based reactive matrices facilitate the covalent charge-tagging of molecules containing phenolic hydroxyl and/or primary or secondary amine groups, including dopaminergic and serotonergic neurotransmitters and their associated metabolites. These matrices improved the matrix-assisted laser desorption/ionization (MALDI)–MSI detection limit toward low-abundance neurotransmitters and facilitated the simultaneous imaging of neurotransmitters in fine structures of the brain at a lateral resolution of 10 µm. We demonstrate strategies for the identification of unknown molecular species using the innate chemoselectivity of the reactive matrices and the unique isotopic pattern of a brominated reactive matrix. We illustrate the capabilities of the developed method on Parkinsonian brain samples from human post-mortem tissue and animal models. The direct imaging of neurotransmitter systems provides a method for exploring how various neurological diseases affect specific brain regions through neurotransmitter modulation.






Similar content being viewed by others
Data availability
The MALDI–MSI datasets that support the findings of this study are available from the corresponding author upon request. NMR data for all compounds is available in the Supplementary Note. MS and MS/MS spectra for derivatized standards are available in the supplementary information. Source data are available online for Fig. 4.
Code availability
Quantitative data analyses were performed using the msIQuant imaging software (v.2.0.1.15). The software is freely available at https://ms-imaging.org/wp/paquan/.
References
Ng, J., Papandreou, A., Heales, S. J. & Kurian, M. A. Monoamine neurotransmitter disorders—clinical advances and future perspectives. Nat. Rev. Neurol. 11, 567–584 (2015).
Falck, B., Hillarp, N. A., Thieme, G. & Torp, A. Fluorescence of catechol amines and related compounds condensed with formaldehyde. J. Histochem. Cytochem. 10, 348–354 (1962).
Jones, B. E. & Beaudet, A. Distribution of acetylcholine and catecholamine neurons in the cat brainstem: a choline acetyltransferase and tyrosine hydroxylase immunohistochemical study. J. Comp. Neurol. 261, 15–32 (1987).
Ametamey, S. M., Honer, M. & Schubiger, P. A. Molecular imaging with PET. Chem. Rev. 108, 1501–1516 (2008).
Caprioli, R. M., Farmer, T. B. & Gile, J. Molecular imaging of biological samples: localization of peptides and proteins using MALDI–TOF MS. Anal. Chem. 69, 4751–4760 (1997).
Norris, J. L. & Caprioli, R. M. Analysis of tissue specimens by matrix-assisted laser desorption/ionization imaging mass spectrometry in biological and clinical research. Chem. Rev. 113, 2309–2342 (2013).
Kompauer, M., Heiles, S. & Spengler, B. Atmospheric pressure MALDI mass spectrometry imaging of tissues and cells at 1.4-μm lateral resolution. Nat. Methods 14, 90–96 (2017).
Passarelli, M. K. et al. The 3D OrbiSIMS-label-free metabolic imaging with subcellular lateral resolution and high mass-resolving power. Nat. Methods 14, 1175–1183 (2017).
Zenobi, R. & Knochenmuss, R. Ion formation in MALDI mass spectrometry. Mass Spectrom. Rev. 17, 337–366 (1998).
Shariatgorji, M. et al. Direct targeted quantitative molecular imaging of neurotransmitters in brain tissue sections. Neuron 84, 697–707 (2014).
Quirke, J. M. E., Adams, C. L. & Vanberkel, G. J. Chemical derivatization for electrospray-ionization mass-spectrometry 1. alkyl-halides, alcohols, phenols, thiols, and amines. Anal. Chem. 66, 1302–1315 (1994).
Bergh, M. S., Bogen, I. L., Lundanes, E. & Oiestad, A. M. L. Validated methods for determination of neurotransmitters and metabolites in rodent brain tissue and extracellular fluid by reversed phase UHPLC–MS/MS. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 1028, 120–129 (2016).
Bjorklund, A. & Dunnett, S. B. Dopamine neuron systems in the brain: an update. Trends Neurosci. 30, 194–202 (2007).
Eisenhofer, G., Kopin, I. J. & Goldstein, D. S. Catecholamine metabolism: a contemporary view with implications for physiology and medicine. Pharm. Rev. 56, 331–349 (2004).
Tank, A. W. & Lee Wong, D. Peripheral and central effects of circulating catecholamines. Compr. Physiol. 5, 1–15 (2015).
Breese, G. R. & Traylor, T. D. Effect of 6-hydroxydopamine on brain norepinephrine and dopamine evidence for selective degeneration of catecholamine neurons. J. Pharm. Exp. Ther. 174, 413–420 (1970).
Herr, N. R. et al. In vivo voltammetry monitoring of electrically evoked extracellular norepinephrine in subregions of the bed nucleus of the stria terminalis. J. Neurophysiol. 107, 1731–1737 (2012).
Chiu, A. S., Godse, D. D. & Warsh, J. J. Determination of brain regional normetanephrine levels by liquid chromatography with electrochemical detection (LC–EC). Prog. Neuropsychopharmacol. 5, 559–563 (1981).
Polotis, M. & Niccolini, F. Serotonin in Parkinson’s disease. Behav. Brain Res. 277, 136–145 (2015).
Albin, R. L. Neurobiology of basal ganglia and Tourette syndrome: striatal and dopamine function. Adv. Neurol. 99, 99–106 (2006).
Kita, H., Chiken, S., Tachibana, Y. & Nambu, A. Serotonin modulates pallidal neuronal activity in the awake monkey. J. Neurosci. 27, 75–83 (2007).
Kveder, S., Iskric, S. & Keglevic, D. 5-Hydroxytryptophol—a metabolite of 5-hydroxytryptamine in rats. Biochem. J. 85, 447 (1962).
Panula, P., Revuelta, A. V., Cheney, D. L., Wu, J. Y. & Costa, E. An immunohistochemical study on the location of GABAergic neurons in rat septum. J. Comp. Neurol. 222, 69–80 (1984).
Yasumi, M., Sato, K., Shimada, S., Nishimura, M. & Tohyama, M. Regional distribution of GABA transporter 1 (GAT1) mRNA in the rat brain: comparison with glutamic acid decarboxylase67 (GAD67) mRNA localization. Brain Res. Mol. Brain Res. 44, 205–218 (1997).
Wang, H., Katz, J., Dagostino, P. & Soghomonian, J. J. Unilateral 6-hydroxydopamine lesion of dopamine neurons and subchronic l-DOPA administration in the adult rat alters the expression of the vesicular GABA transporter in different subsets of striatal neurons and in the substantia nigra, pars reticulata. Neuroscience 145, 727–737 (2007).
Kageyama, T. et al. The 4F2hc/LAT1 complex transports l-DOPA across the blood–brain barrier. Brain Res. 879, 115–121 (2000).
Boado, R. J., Li, J. Y., Nagaya, M., Zhang, C. & Pardridge, W. M. Selective expression of the large neutral amino acid transporter at the blood–brain barrier. Proc. Natl Acad. Sci. USA 96, 12079–12084 (1999).
Murata, M. Pharmacokinetics of l-dopa—special reference to food and aging. J. Neurol. 253, 47–52 (2006).
Meissner, W. et al. Time-course of nigrostriatal degeneration in a progressive MPTP-lesioned macaque model of Parkinson’s disease. Mol. Neurobiol. 28, 209–218 (2003).
Bezard, E., Brotchie, J. M. & Gross, C. E. Pathophysiology of levodopa-induced dyskinesia: potential for new therapies. Nat. Rev. Neurosci. 2, 577–588 (2001).
Bagley, M. C., Ekelof, M., Rock, K., Patisaul, H. & Muddiman, D. C. IR-MALDESI mass spectrometry imaging of underivatized neurotransmitters in brain tissue of rats exposed to tetrabromobisphenol A. Anal. Bioanal. Chem. 410, 7979–7986 (2018).
Fernandes, A. M. et al. Direct visualization of neurotransmitters in rat brain slices by desorption electrospray ionization mass spectrometry imaging (DESI–MS). J. Am. Soc. Mass Spectrom. 27, 1944–1951 (2016).
Esteve, C., Tolner, E. A., Shyti, R., van den Maagdenberg, A. M. & McDonnell, L. A. Mass spectrometry imaging of amino neurotransmitters: a comparison of derivatization methods and application in mouse brain tissue. Metabolomics 12, 30 (2016).
Kaya, I. et al. On-tissue chemical derivatization of catecholamines using 4-(N-Methyl)pyridinium boronic acid for ToF-SIMS and LDI-ToF mass spectrometry imaging. Anal. Chem. 90, 13580–13590 (2018).
Ciliax, B. J. et al. The dopamine transporter: immunochemical characterization and localization in brain. J. Neurosci. 15, 1714–1723 (1995).
Ramsay, W. J. et al. Designed enclosure enables guest binding within the 4200 Å3 cavity of a self-assembled cube. Angew. Chem. Int. Ed. Engl. 54, 5636–5640 (2015).
Zhang, X., Andren, P. E., Chergui, K. & Svenningsson, P. Neurokinin B/NK3 receptors exert feedback inhibition on l-DOPA actions in the 6-OHDA lesion rat model of Parkinson’s disease. Neuropharmacology 54, 1143–1152 (2008).
Fernagut, P. O. et al. Dopamine transporter binding is unaffected by l-DOPA administration in normal and MPTP-treated monkeys. PLoS One 5, e14053 (2010).
Porras, G. et al. PSD-95 expression controls l-DOPA dyskinesia through dopamine D1 receptor trafficking. J. Clin. Invest 122, 3977–3989 (2012).
Santini, E. et al. Distinct changes in cAMP and extracellular signal-regulated protein kinase signalling in l-DOPA-induced dyskinesia. PLoS One 5, e12322 (2010).
Bezard, E., Imbert, C., Deloire, X., Bioulac, B. & Gross, C. E. A chronic MPTP model reproducing the slow evolution of Parkinson’s disease: evolution of motor symptoms in the monkey. Brain Res. 766, 107–112 (1997).
Bezard, E. et al. Relationship between the appearance of symptoms and the level of nigrostriatal degeneration in a progressive 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine-lesioned macaque model of Parkinson’s disease. J. Neurosci. 21, 6853–6861 (2001).
Bezard, E., Ravenscroft, P., Gross, C. E., Crossman, A. R. & Brotchie, J. M. Upregulation of striatal preproenkephalin gene expression occurs before the appearance of parkinsonian signs in 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine monkeys. Neurobiol. Dis. 8, 343–350 (2001).
Ahmed, M. R. et al. Lentiviral overexpression of GRK6 alleviates l-DOPA-induced dyskinesia in experimental Parkinson’s disease. Sci. Transl. Med 2, 28ra28 (2010).
Fasano, S. et al. Inhibition of Ras–guanine nucleotide-releasing factor 1 (Ras–GRF1) signaling in the striatum reverts motor symptoms associated with L-dopa-induced dyskinesia. Proc. Natl Acad. Sci. USA 107, 21824–21829 (2010).
Munoz, A. et al. Combined 5-HT1A and 5-HT1B receptor agonists for the treatment of l-DOPA-induced dyskinesia. Brain 131, 3380–3394 (2008).
Porras, G., Li, Q. & Bezard, E. Modeling Parkinson’s disease in primates: the MPTP model. Cold Spring Harb. Perspect. Med. 2, a009308 (2012).
Kallback, P., Nilsson, A., Shariatgorji, M. & Andren, P. E. msIQuant—quantitation software for mass spectrometry imaging enabling fast access, visualization, and analysis of large data sets. Anal. Chem. 88, 4346–4353 (2016).
Acknowledgements
This work was supported by the Swedish Brain Foundation (grant FO2018-0292 to P.E.A.), the Swedish Foundation for Strategic Research (grant RIF14-0078 to P.E.A), the European FP7 ITN Community’s Seventh Framework Program ARIADME (grant 607517 to P.E.A.), the Swedish Research Council, Medicine and Health (grant 2018–03320 to P.E.A.), the Swedish Research Council, Natural and Engineering Science (grant 2018–05501 to P.E.A. and 2018–05133 to L.R.O.) and the Science for Life Laboratory (to P.E.A.). Confocal microscope images were acquired at the BioVis core facility at Uppsala University with help of the BioVis staff. We thank M. Witt for support using the 12T FTICR MS instrument and S. Dengler for experimental assistance.
Author information
Authors and Affiliations
Contributions
M.S. and A.N. conceived the methodology, designed experiments, acquired, analyzed and interpreted data, and wrote the manuscript; E.F. acquired and analyzed data and wrote part of the manuscript; T.V. acquired and analyzed data; P.K. analyzed data; L.K. and J.S. synthesized the reactive matrices; I.M. and X.Z. performed animal experiments and provided tissue samples; E.B. and P.S. provided tissue samples, supervised animal experiments and edited the manuscript; L.R.O. conceived the methodology, designed experiments, synthesized the reactive matrices, interpreted data and edited the manuscript; P.E.A. conceived the methodology, designed experiments, interpreted data, wrote the manuscript and is principal investigator for the grants that fund this research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nina Vogt and Allison Doerr were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–14 and Supplementary Tables 1–2
Supplementary Video
MALDI-MS imaging of the nigrostriatal dopaminergic pathway. The nigrostriatal pathway is visualized changing the color scaling for ion intensity of the FMP-10 derivatized DA signal (m/z 674.28). MALDI-MSI experiment was replicated four times independently from different animals with similar results.
Source data
Rights and permissions
About this article
Cite this article
Shariatgorji, M., Nilsson, A., Fridjonsdottir, E. et al. Comprehensive mapping of neurotransmitter networks by MALDI–MS imaging. Nat Methods 16, 1021–1028 (2019). https://doi.org/10.1038/s41592-019-0551-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0551-3
- Springer Nature America, Inc.
This article is cited by
-
Spatial multimodal analysis of transcriptomes and metabolomes in tissues
Nature Biotechnology (2024)
-
Novel mimetic tissue standards for precise quantitative mass spectrometry imaging of drug and neurotransmitter concentrations in rat brain tissues
Analytical and Bioanalytical Chemistry (2024)
-
Prosaposin maintains lipid homeostasis in dopamine neurons and counteracts experimental parkinsonism in rodents
Nature Communications (2023)
-
Targeted Energy Metabolomics Combined with Spatial Metabolomics Study on the Efficacy of Guhong Injection Against Cerebral Ischemia Reperfusion
Molecular Neurobiology (2023)
-
Pushing the frontiers: tools for monitoring neurotransmitters and neuromodulators
Nature Reviews Neuroscience (2022)