Abstract
Skin color is highly variable in Africans, yet little is known about the underlying molecular mechanism. Here we applied massively parallel reporter assays to screen 1,157 candidate variants influencing skin pigmentation in Africans and identified 165 single-nucleotide polymorphisms showing differential regulatory activities between alleles. We combine Hi-C, genome editing and melanin assays to identify regulatory elements for MFSD12, HMG20B, OCA2, MITF, LEF1, TRPS1, BLOC1S6 and CYB561A3 that impact melanin levels in vitro and modulate human skin color. We found that independent mutations in an OCA2 enhancer contribute to the evolution of human skin color diversity and detect signals of local adaptation at enhancers of MITF, LEF1 and TRPS1, which may contribute to the light skin color of Khoesan-speaking populations from Southern Africa. Additionally, we identified CYB561A3 as a novel pigmentation regulator that impacts genes involved in oxidative phosphorylation and melanogenesis. These results provide insights into the mechanisms underlying human skin color diversity and adaptive evolution.







Similar content being viewed by others
Data availability
The epigenomic data, Hi-C and HiChIP data were uploaded to UCSC browser and are available at https://genome.ucsc.edu/s/fengyq/Tishkoff_Lab%2Dhg38%2DMPRA%2DHiC_Pigmentation. All RNA-seq and epigenomic data generated in this study are available at GEO GSE240717. Genotype data for GWAS are in dbGaP phs001396.vl.pl. Source data are provided with this paper.
Code availability
Public software and packages were used following the developer’s manuals. The custom code used for data analysis has been deposited at GitHub (https://github.com/fengyq/nature_gentics_codes) and Zenodo101 (https://zenodo.org/records/10198223).
References
Jablonski, N. G. & Chaplin, G. Colloquium paper: human skin pigmentation as an adaptation to UV radiation. Proc. Natl Acad. Sci. USA 107, 8962–8968 (2010).
Barsh, G. S. What controls variation in human skin color? PLoS Biol. 1, E27 (2003).
Beleza, S. et al. Genetic architecture of skin and eye color in an African–European admixed population. PLoS Genet. 9, e1003372 (2013).
Liu, F. et al. Genetics of skin color variation in Europeans: genome-wide association studies with functional follow-up. Hum. Genet. 134, 823–835 (2015).
Martin, A. R. et al. An unexpectedly complex architecture for skin pigmentation in Africans. Cell 171, 1340–1353 (2017).
Galván-Femenía, I. et al. Multitrait genome association analysis identifies new susceptibility genes for human anthropometric variation in the GCAT cohort. J. Med. Genet. 55, 765–778 (2018).
Neale lab UK-Biobank GWAS result. Neale Lab http://www.nealelab.is/uk-biobank/ (2018).
Adhikari, K. et al. A GWAS in Latin Americans highlights the convergent evolution of lighter skin pigmentation in Eurasia. Nat. Commun. 10, 358 (2019).
Lona-Durazo, F. et al. Meta-analysis of GWA studies provides new insights on the genetic architecture of skin pigmentation in recently admixed populations. BMC Genet. 20, 59 (2019).
Jiang, L., Zheng, Z., Fang, H. & Yang, J. A generalized linear mixed model association tool for biobank-scale data. Nat. Genet. 53, 1616–1621 (2021).
Batai, K. et al. Genetic loci associated with skin pigmentation in African Americans and their effects on vitamin D deficiency. PLoS Genet. 17, e1009319 (2021).
Pairo-Castineira, E. et al. Expanded analysis of pigmentation genetics in UK Biobank. Preprint at bioRxiv https://doi.org/10.1101/2022.01.30.478418 (2022).
Crawford, N. G. et al. Loci associated with skin pigmentation identified in African populations. Science 358, eaan8433 (2017).
Miller, C. T. et al. cis-Regulatory changes in KIT ligand expression and parallel evolution of pigmentation in sticklebacks and humans. Cell 131, 1179–1189 (2007).
Tsetskhladze, Z. R. et al. Functional assessment of human coding mutations affecting skin pigmentation using zebrafish. PLoS ONE 7, e47398 (2012).
Visser, M., Kayser, M. & Palstra, R.-J. HERC2 rs12913832 modulates human pigmentation by attenuating chromatin-loop formation between a long-range enhancer and the OCA2 promoter. Genome Res. 22, 446–455 (2012).
Praetorius, C. et al. A polymorphism in IRF4 affects human pigmentation through a tyrosinase-dependent MITF/TFAP2A pathway. Cell 155, 1022–1033 (2013).
Fan, S. et al. Whole-genome sequencing reveals a complex African population demographic history and signatures of local adaptation. Cell 186, 923–939.e14 (2023).
Gordon, M. G. et al. lentiMPRA and MPRAflow for high-throughput functional characterization of gene regulatory elements. Nat. Protoc. 15, 2387–2412 (2020).
Akey, J. M. et al. Tracking footprints of artificial selection in the dog genome. Proc. Natl Acad. Sci. USA 107, 1160–1165 (2010).
Myint, L., Avramopoulos, D. G., Goff, L. A. & Hansen, K. D. Linear models enable powerful differential activity analysis in massively parallel reporter assays. BMC Genomics 20, 209 (2019).
Adelmann, C. H. et al. MFSD12 mediates the import of cysteine into melanosomes and lysosomes. Nature 588, 699–704 (2020).
Luecke, S. et al. The aryl hydrocarbon receptor (AHR), a novel regulator of human melanogenesis. Pigment Cell Melanoma Res. 23, 828–833 (2010).
Kayser, M. et al. Three genome-wide association studies and a linkage analysis identify HERC2 as a human iris color gene. Am. J. Hum. Genet. 82, 411–423 (2008).
Lona-Durazo, F. et al. A large Canadian cohort provides insights into the genetic architecture of human hair colour. Commun. Biol. 4, 1253 (2021).
Simcoe, M. et al. Genome-wide association study in almost 195,000 individuals identifies 50 previously unidentified genetic loci for eye color. Sci. Adv. 7, eabd1239 (2021).
Liang, Z. et al. BL-Hi-C is an efficient and sensitive approach for capturing structural and regulatory chromatin interactions. Nat. Commun. 8, 1622 (2017).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Bhattacharyya, S., Chandra, V., Vijayanand, P. & Ay, F. Identification of significant chromatin contacts from HiChIP data by FitHiChIP. Nat. Commun. 10, 4221 (2019).
Ochoa, D. et al. Open Targets Platform: supporting systematic drug-target identification and prioritisation. Nucleic Acids Res. 49, D1302–D1310 (2021).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Albers, P. K. & McVean, G. Dating genomic variants and shared ancestry in population-scale sequencing data. PLoS Biol. 18, e3000586 (2020).
Levy, C., Khaled, M. & Fisher, D. E. MITF: master regulator of melanocyte development and melanoma oncogene. Trends Mol. Med. 12, 406–414 (2006).
Klein, J. C. et al. A systematic evaluation of the design and context dependencies of massively parallel reporter assays. Nat. Methods 17, 1083–1091 (2020).
Tan, B. et al. FOXP3 over-expression inhibits melanoma tumorigenesis via effects on proliferation and apoptosis. Oncotarget 5, 264–276 (2014).
Cao, Y. et al. Accurate loop calling for 3D genomic data with cLoops. Bioinformatics 36, 666–675 (2020).
Takeda, K. et al. Induction of melanocyte-specific microphthalmia-associated transcription factor by Wnt-3a. J. Biol. Chem. 275, 14013–14016 (2000).
Bondurand, N. et al. Interaction among SOX10, PAX3 and MITF, three genes altered in Waardenburg syndrome. Hum. Mol. Genet. 9, 1907–1917 (2000).
Kichaev, G. et al. Leveraging polygenic functional enrichment to improve GWAS power. Am. J. Hum. Genet. 104, 65–75 (2019).
Morgan, M. D. et al. Genome-wide study of hair colour in UK Biobank explains most of the SNP heritability. Nat. Commun. 9, 5271 (2018).
Visconti, A. et al. Genome-wide association study in 176,678 Europeans reveals genetic loci for tanning response to sun exposure. Nat. Commun. 9, 1684 (2018).
Larimore, J. et al. Mutations in the BLOC-1 subunits dysbindin and muted generate divergent and dosage-dependent phenotypes. J. Biol. Chem. 289, 14291–14300 (2014).
Saito, H. et al. Melanocyte-specific microphthalmia-associated transcription factor isoform activates its own gene promoter through physical interaction with lymphoid-enhancing factor 1. J. Biol. Chem. 277, 28787–28794 (2002).
Wang, X. et al. LEF-1 regulates tyrosinase gene transcription in vitro. PLoS ONE 10, e0143142 (2015).
Ishitani, T. et al. The TAK1–NLK–MAPK-related pathway antagonizes signalling between β-catenin and transcription factor TCF. Nature 399, 798–802 (1999).
Ishitani, T., Ninomiya-Tsuji, J. & Matsumoto, K. Regulation of lymphoid enhancer factor 1/T-cell factor by mitogen-activated protein kinase-related Nemo-like kinase-dependent phosphorylation in Wnt/β-catenin signaling. Mol. Cell. Biol. 23, 1379–1389 (2003).
Gai, Z., Gui, T. & Muragaki, Y. The function of TRPS1 in the development and differentiation of bone, kidney, and hair follicles. Histol. Histopathol. 26, 915–921 (2011).
Swoboda, A. et al. STAT3 promotes melanoma metastasis by CEBP-induced repression of the MITF pathway. Oncogene 40, 1091–1105 (2021).
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016).
Sitaram, A. & Marks, M. S. Mechanisms of protein delivery to melanosomes in pigment cells. Physiology 27, 85–99 (2012).
Wang, Z. et al. CYB561A3 is the key lysosomal iron reductase required for Burkitt B-cell growth and survival. Blood 138, 2216–2230 (2021).
Karlsson, M. et al. A single-cell type transcriptomics map of human tissues. Sci. Adv. 7, eabh2169 (2021).
Lee, J. H. et al. Evolutionarily assembled cis-regulatory module at a human ciliopathy locus. Science 335, 966–969 (2012).
Lamason, R. L. et al. SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science 310, 1782–1786 (2005).
Lavado, A., Olivares, C., García-Borrón, J. C. & Montoliu, L. Molecular basis of the extreme dilution mottled mouse mutation: a combination of coding and noncoding genomic alterations. J. Biol. Chem. 280, 4817–4824 (2005).
Seruggia, D., Fernández, A., Cantero, M., Pelczar, P. & Montoliu, L. Functional validation of mouse tyrosinase non-coding regulatory DNA elements by CRISPR–Cas9-mediated mutagenesis. Nucleic Acids Res. 43, 4855–4867 (2015).
Ambrosio, A. L., Boyle, J. A., Aradi, A. E., Christian, K. A. & Di Pietro, S. M. TPC2 controls pigmentation by regulating melanosome pH and size. Proc. Natl Acad. Sci. USA 113, 5622–5627 (2016).
Ploper, D. et al. MITF drives endolysosomal biogenesis and potentiates Wnt signaling in melanoma cells. Proc. Natl Acad. Sci. USA 112, E420–E429 (2015).
Zhang, Y. et al. Lef1 contributes to the differentiation of bulge stem cells by nuclear translocation and cross-talk with the Notch signaling pathway. Int. J. Med. Sci. 10, 738–746 (2013).
Fantauzzo, K. A., Kurban, M., Levy, B. & Christiano, A. M. Trps1 and its target gene Sox9 regulate epithelial proliferation in the developing hair follicle and are associated with hypertrichosis. PLoS Genet. 8, e1003002 (2012).
Fantauzzo, K. A. & Christiano, A. M. Trps1 activates a network of secreted Wnt inhibitors and transcription factors crucial to vibrissa follicle morphogenesis. Development 139, 203–214 (2012).
Yamada, T. et al. Wnt/β-catenin and kit signaling sequentially regulate melanocyte stem cell differentiation in UVB-induced epidermal pigmentation. J. Invest. Dermatol. 133, 2753–2762 (2013).
Andl, T., Reddy, S. T., Gaddapara, T. & Millar, S. E. WNT signals are required for the initiation of hair follicle development. Dev. Cell 2, 643–653 (2002).
Tobias, P. V. & Biesele, M. The Bushmen: San Hunters and Herders of Southern Africa (Human & Rousseau, 1978).
Feng, Y., McQuillan, M. A. & Tishkoff, S. A. Evolutionary genetics of skin pigmentation in African populations. Hum. Mol. Genet. 30, R88–R97 (2021).
Rawofi, L. et al. Genome-wide association study of pigmentary traits (skin and iris color) in individuals of East Asian ancestry. PeerJ 5, e3951 (2017).
Stokowski, R. P. et al. A genomewide association study of skin pigmentation in a South Asian population. Am. J. Hum. Genet. 81, 1119–1132 (2007).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Kang, H. M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Kang, H. M. et al. Efficient and parallelizable association container toolbox, EPACTS v3.3.0. EPACTS http://genome.sph.umich.edu/wiki/EPACTS (2013).
Bhatia, G., Patterson, N., Sankararaman, S. & Price, A. L. Estimating and interpreting FST: the impact of rare variants. Genome Res. 23, 1514–1521 (2013).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Barrett, T. et al. NCBI GEO: mining millions of expression profiles—database and tools. Nucleic Acids Res. 33, D562–D566 (2005).
Phenotype: pigmentation phenotype. International Mouse Phenotyping Consortium https://www.mousephenotype.org/data/phenotypes/MP:0001186 (2023)
Dickinson, M. E. et al. High-throughput discovery of novel developmental phenotypes. Nature 537, 508–514 (2016).
Baxter, L. L., Watkins-Chow, D. E., Pavan, W. J. & Loftus, S. K. A curated gene list for expanding the horizons of pigmentation biology. Pigment Cell Melanoma Res. 32, 348–358 (2019).
Uhlen, M. et al. A pathology atlas of the human cancer transcriptome. Science 357, eaan2507 (2017).
Custom Alt-R™ CRISPR–Cas9 guide RNA. Integrated DNA Technologies https://www.idtdna.com/site/order/designtool/index/CRISPR_CUSTOM (2023).
RNA sequencing frequently asked questions. GENEWIZ https://web.genewiz.com/rna-seq-faq (2023).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
FastQC. GitHub https://github.com/s-andrews/FastQC (2020)
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res. 4, 1521 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Kucukural, A., Yukselen, O., Ozata, D. M., Moore, M. J. & Garber, M. DEBrowser: interactive differential expression analysis and visualization tool for count data. BMC Genomics 20, 6 (2019).
Blighe, K., Rana, S., Lewis, M. EnhancedVolcano: publication-ready volcano plots with enhanced colouring and labeling. R package version 1.14.0. EnhancedVolcano https://github.com/kevinblighe/EnhancedVolcano (2023).
Luo, W. & Brouwer, C. Pathview: an R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics 29, 1830–1831 (2013).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Li, H. et al. The sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Meers, M. P., Tenenbaum, D. & Henikoff, S. Peak calling by Sparse Enrichment Analysis for CUT&RUN chromatin profiling. Epigenet. Chromatin 12, 42 (2019).
Coetzee, S. G., Coetzee, G. A. & Hazelett, D. J. motifbreakR: an R/Bioconductor package for predicting variant effects at transcription factor binding sites. Bioinformatics 31, 3847–3849 (2015).
Kulakovskiy, I. V. et al. HOCOMOCO: towards a complete collection of transcription factor binding models for human and mouse via large-scale ChIP–seq analysis. Nucleic Acids Res. 46, D252–D259 (2018).
An, L. et al. OnTAD: hierarchical domain structure reveals the divergence of activity among TADs and boundaries. Genome Biol. 20, 282 (2019).
Feng, Y. Codes for skin pigmentation paper. Zenodo https://doi.org/10.5281/zenodo.10198223 (2023).
Shin, J. H., Blay, S., Graham, J. & McNeney, B. LDheatmap: an R function for graphical display of pairwise linkage disequilibria between single nucleotide polymorphisms. J. Stat. Softw. 16, 1–9 (2006).
Liu, T. et al. Cistrome: an integrative platform for transcriptional regulation studies. Genome Biol. 12, R83 (2011).
Acknowledgements
This research was supported by the following grants: NIH grants R35 GM134957-01, 3UM1HG009408-02S1, 1R01GM113657-01 and 5R01AR076241-02. We thank the Skin Biology and Disease Resource-based Center (SBDRC, NIH P30-AR069589) at the University of Pennsylvania for funding and providing human primary melanocytes. The sequencing of MPRA was carried out by the DNA Technologies and Expression Analysis Core at the University of California Davis Genome Center, supported by the NIH Shared Instrumentation Grant 1S10OD010786-01. We thank E. Burton for assistance on part of the plasmid cloning. We thank Z. (J.) Zhou from the Department of Genetics at the University of Pennsylvania for sharing their tissue culture room. We thank J. Phillips-Cremins from the Department of Genetics at the University of Pennsylvania for constructive suggestions on Hi-C. We thank H. Wong and H. Wu at the University of Pennsylvania for sharing their experimental equipment. We thank the African participants for their contributions to this study.
Author information
Authors and Affiliations
Contributions
Y.F. and S.A.T. designed the study and wrote the original draft. Y.F. performed the Hi-C, H3K27ac HiChIP, CRISPR, RNA-seq, ATAC-seq and CUT&RUN experiments and related data analysis. Y.F. and F.I. conducted the MPRA under supervision of N.A. Y.F. and C.Z. analyzed the MPRA data. S.F. and M.E.B.H played a role in quality control and analysis of WGS and SNP array data. Y.F. and S.F. conducted the GWAS and Di analysis. Y.F. and N.X. performed CRISPR editing and related assays in MNT-1. S.A.T., T.N., S.W.M., G.G.M., A.K.N., C.F. and G.B. played a role in collecting data from Africa. J.S. and E.O. performed the CYB561A3 immunofluorescence imaging and analyses. E.O. and M.S.M. provided resources and additional experimental insights. All authors assisted with manuscript review and editing. S.A.T. supervised the project.
Corresponding author
Ethics declarations
Competing interests
N.A. is an equity holder of Encoded Therapeutics, a gene regulation therapeutics company and is a cofounder and scientific advisor of Regel Therapeutics and Neomer Diagnostics. The remaining authors declare no competing financial interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quality statistics of the MPRA experiments.
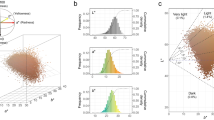
(a) Statistics for FLASH-merged reads in the association library. The plot shows that 46.1% are 200 bp fragments as designed. (b) Statistics of BWA-mapped reads in the association library. The plot shows that 44.1% are 200 bp fragments as designed. (c) Statistics of barcode types per oligo in the association library. On average, each oligo is linked with 126 different barcodes. (d) Statistics of barcode types per oligo in reference (n = 1102), alternative (n = 1103), negative control (n = 153), and positive control (n = 30) oligos. Data is from the association library. (e) Statistics of barcode counts per oligo in reference (n = 1102), alternative (n = 1103), negative control (n = 153), and positive control (n = 30) oligos. Data is from the association library. (f) Barcode types for reference and alternative alleles are comparable. Pearson’s r = 0.91, p < 2 × 10−16. (g) Principal component analysis of DNA and RNA libraries from MNT-1 and WM88 cells. Three replicates. (h) Summary of enhancer activities estimated by MPRA. Enhancer activities were defined as the barcode counts per million in the RNA library divided by the barcode counts per million in the DNA library. Alt: oligos containing alternative alleles (n = 1103). Ref: oligos containing reference alleles (n = 1102). Negative, negative control oligos (n = 148). Positive, positive control oligos (n = 30). For boxplots, central lines are median, with boxes extending from the 25th to the 75th percentiles. Whiskers further extend by ±1.5 times the interquartile range from the limits of each box.
Extended Data Fig. 2 MPRA identifies six allelic skewed variants near MFSD12.
(a) Plot showing allelic skewed variants in regulatory regions near MFSD12. Blue tracks indicate DNase-Seq, ATAC-Seq, and ChIP-Seq from melanocytes; orange tracks indicate ChIP-Seq from melanoma (501-mel) cells; green tracks indicate DNase-Seq from ENCODE cell lines. E1-E4, enhancers. P, promoter. (b-g) Relative enhancer activities of the two alleles at rs142317543, rs6510759, rs734454, rs10416746, rs6510760, rs7246261 estimated by MPRA (n = 3). For b, c, d, f, g, p-values were estimated with a random effects model for mpralm and paired t-tests with multiple testing adjustments; e was without multiple testing adjustments. (h-k) Relative enhancer activities estimated by LRA. Two-tailed paired t-tests (For LRA in MNT1, n = 6. For LRA in WM88, 2 h n = 8; others n = 9). Data were presented as mean ± SEM. ns p > 0.05. (l) rs6510760 and rs7246261 disrupt the binding motifs of AHR and TFAP2, respectively. Predicted by ‘MotifBreakR’98. (m) The LD pattern of candidate functional variants near MFSD12. LD was calculated using the 180 G18 data by the LDheatmap102 package. For boxplots, central lines are median, with boxes extending from the 25th to the 75th percentiles. Whiskers further extend by ±1.5 times the interquartile range from the limits of each box.
Extended Data Fig. 3 The enhancer E4 interacts with the promoter of MFSD12 and affects the expression of MFSD12.
(a, b) Chromatin interactions near MFSD12 identified by Hi-C and H3K27ac HiChIP with Hae3 digestion. The upper matrix is from MNT-1 Hi-C data, and the lower matrix is from MNT-1 H3K27ac HiChIP data. TADs were called by onTAD100 and colored by nested TAD levels. The solid arch was a loop defined using FitHiChIP29 software, the dashed arch was a potential loop based on the observed interaction matrix. The interaction matrix between MFSD12 and HMG20B was highlighted with orange angles. The DNase track of melanocytes was downloaded from ENCODE68. rs6510760 and rs657246261 in E4 were colored in red. The plotted region is chr19:3519998-3589998 (hg19). (c) Schematic showing the location of the two sgRNAs targeting the enhancer E4 of MFSD12. (d) PCR results showing efficient knockout of the enhancer by the two sgRNAs. Three independent experiments. (e) qPCR showed that CRISPRi of E4 reduces the gene expression of MFSD12 and HMG20B in MNT-1 cells. Two-sided Dunnett’s test with adjustments for multiple comparisons (n = 3). (f) CRISPRi of E4 slightly increases melanin levels in MNT-1 cells. Two-tailed unpaired t-tests (n = 19). (g) qPCR showed that CRISPR knockout of E4 decreases the gene expression of MFSD12 and HMG20B in MNT-1 cells. Two-tailed unpaired t-tests without multiple testing adjustments (n = 6). Data are presented as mean ± SEM.
Extended Data Fig. 4 Identification of functional variants associated with skin pigmentation near OCA2.
(a) SNP rs6497271 is in a melanocyte-specific enhancer. Blue tracks indicate DNase-Seq, ATAC-Seq, and ChIP-Seq data from melanocytes; orange tracks indicate ChIP-Seq data from melanoma (501-mel) cells; green tracks indicate DNase-Seq data from ENCODE cell lines. E1-E4, enhancers. The plotted region is chr15: 28,335,146-28,385,146 (hg19). (b) MPRA and LRA reveals that rs4778242 significantly affects the enhancer activity of E1 in MNT-1 and WM88 cells. MPRA (n = 3), LRA (n = 9). (c) MPRA showed that rs6497271 affects the enhancer activity of E2 in MNT-1 and WM88 cells (n = 3). (d) MPRA shows that rs7495989 affects the enhancer activity of E3 in MNT-1 and WM88 cells (n = 3). (e) MPRA and LRA reveals that rs4778141 affects the enhancer activity of E4 in MNT-1 and WM88 cells. MPRA (n = 3), LRA (MNT-1, n = 9; WM88, n = 6). (f) rs6497271 overlaps transcription factor binding sites. Left panel shows rs6497271 disrupts the binding motif of LEF1 and SOX10. Right panel shows that rs6497271 overlaps ChIP-seq peaks from Cistrome database103. LRA data are presented as mean ± SEM, tested with two-tailed paired t-tests. MPRA p-values are estimated with a random effects model for mpralm and paired t-tests with multiple testing adjustments. For MPRA boxplots, central lines are median, with boxes extending from the 25th to the 75th percentiles. Whiskers further extend by ±1.5 times the interquartile range from the limits of each box.
Extended Data Fig. 5 Identification of functional variants near MITF related to skin pigmentation in the San.
(a) A Plot showing functional Di-SNP rs111969762 is in a melanocyte-specific regulatory region. Blue tracks indicate DNase-Seq, ATAC-Seq, and ChIP-Seq from melanocytes; orange tracks indicate ChIP-Seq from melanoma (501-mel) cells; green tracks indicate DNase-Seq from ENCODE cell lines. E1-E2, enhancers. (b) MPRA showed that rs111969762 affects enhancer activity in WM88 cells (n = 3). (c) MPRA shows that rs7430957 impacts enhancer activity in WM88 cells (n = 3). (d) LRA shows that rs7430957 does not significantly alter the activity of the E2 enhancer near MITF. P values were estimated by two-tailed paired t-tests, MNT-1(n = 6), WM88 (n = 11). Data were presented as mean ± SEM. ns p > 0.05. (e) rs111969762 overlaps transcription factor binding sites. Left panel showed rs6497271 disrupts the binding motif of FOXP3. Right panel showed that rs111969762 overlaps ChIP-seq peaks from the Cistrome database103. MPRA p-values were estimated with a random effects model for mpralm and paired t-tests with multiple testing adjustments. For MPRA boxplots, central lines are median, with boxes extending from the 25th to the 75th percentiles. Whiskers further extend by ±1.5 times the interquartile range from the limits of each box.
Extended Data Fig. 6 Functional testing of Di-SNPs near LEF1.
(a) MFVs and regulatory elements near LEF1. rs17038630 and rs11939273 are Di-SNPs from the San population. (b) Plot showing allelic skews at rs17038630 in MNT-1 and WM88 cells estimated by MPRA (n = 3). (c) rs17038630 overlaps SOX10 and LEF1 binding sites. Left panel shows that rs17038630 disrupts the binding motif of SOX10 and LEF1. Right panel shows that rs11939273 overlaps ChIP-seq peaks from the Cistrome database103. (d) MPRA and LRA results showing allelic skews at Di-SNP rs11939273 in MNT-1 and WM88 cells, the allele frequency data was from the 180 G18 and 1000 G31 dataset. MPRA (n = 3), LRA (MNT-1, n = 6; WM88, n = 9). LRA data are presented as mean ± SEM, tested with two-tailed paired t-tests. (e) CRISPR-KO of the enhancer E1 of LEF1 does not affect LEF1 expression and melanin levels in MNT-1 cells. Left panel shows genotyping results of CRISPR-KO of the enhancer E1 of LEF1, three independent experiments. Middle panel shows the RT-qPCR results of CRISPR-KO of the enhancer E1 of LEF1 (n = 9). Right panel shows the melanin levels of CRISPR-KO of the enhancer E1 of LEF1 (n = 9). Two-tailed unpaired t-tests. For MPRA boxplots in b and d, central lines are median, with boxes extending from the 25th to the 75th percentiles. Whiskers further extend by ±1.5 times the interquartile range from the limits of each box. MPRA p-values were estimated with a random effects model for mpralm and paired t-tests with multiple testing adjustments.
Extended Data Fig. 7 MPRA and LRA identified three functional Di-SNPs near NLK.
(a) MFVs and regulatory elements near NLK. rs75827647, rs10468581 and rs113940275 are Di-SNPs from the San population. (b) LRA and MPRA results showing allelic skews at rs75827647 in MNT-1 and WM88 cells. MPRA (n = 3), LRA (MNT-1, n = 6; WM88, n = 9). (c) LRA and MPRA results showing allelic skews at rs10468581 in MNT-1 and WM88 cells. MPRA (n = 3), LRA (MNT-1, n = 6; WM88, n = 9). (d) LRA and MPRA results showing allelic skews at rs113940275 in MNT-1 and WM88 cells. MPRA (n = 3), LRA (MNT-1, n = 6; WM88, n = 11). From b to d, the barplots are results of LRA, two-tailed paired t-tests without adjustments for multiple comparisons; data were presented as mean ± SEM. ns p > 0.05. The boxplots are results from MPRA, p-values were estimated with a random effects model for mpralm and paired t-tests with multiple testing adjustments. The right panels are allele frequency maps constructed using the 180G18 and 1000 G31 dataset. For boxplots, central lines are median, with boxes extending from the 25th to the 75th percentiles. Whiskers further extend by ±1.5 times the interquartile range from the limits of each box.
Extended Data Fig. 8 Functional testing of Di-SNPs near TRPS1.
(a) SNP rs11985280 overlaps a regulatory element of TPRS1. Blue tracks show ATAC-Seq, and ChIP-Seq data from melanocytes; orange tracks indicate ChIP-Seq data from melanoma (501-mel) cells, green tracks indicate ATAC-Seq and DNase-Seq data from ENCODE cell lines. (b) MPRA results showing allelic skews at rs11985280 in MNT-1 and WM88 cells (n = 3). p-values were estimated with a random effects model for mpralm and paired t-tests with multiple testing adjustments. For boxplots, central lines are median, with boxes extending from the 25th to the 75th percentiles. Whiskers further extend by ±1.5 times the interquartile range from the limits of each box. (c) rs11985280 disrupts the binding motif of CEBPA and CEBPB. Right panel shows that rs11985280 overlaps ChIP-seq peaks from the Cistrome database103. (d) CRISPR-KO of the enhancer E1 of TRPS1 affects TRPS1 expression but not melanin levels in MNT-1 cells. Left panel shows genotyping results of CRISPR-KO of the enhancer E1 of TRPS1, three independent experiments. Middle panel shows the RT-qPCR results of CRISPR-KO of the enhancer E1 of TRPS1 (n = 9). Right panel shows the melanin levels of CRISPR-KO of the enhancer E1 of TRPS1 in MNT-1 cells (n = 9). Two-tailed unpaired t-tests. Data are presented as mean ± SEM and p values are listed above the bars.
Extended Data Fig. 9 Identification of functional regulatory variants near the BLOC1S6 locus.
(a). rs72713175 overlaps a regulatory region in melanocytes. Green tracks indicate ATAC-seq for MNT-1 and WM88 cells; blue tracks indicate ATAC-seq and ChIP-Seq from NHM; orange tracks indicate CUT&RUN from MNT-1 cells. (b) MPRA results showing allelic skews at rs11985280 in WM88 cells but not in MNT-1 cells (n = 3). P values were estimated with a random effects model for mpralm and paired t-tests without multiple testing adjustments. For boxplots, central lines are median, with boxes extending from the 25th to the 75th percentiles. Whiskers further extend by ±1.5 times the interquartile range from the limits of each box. (c) Allele frequencies at rs72713175 in global populations, data were from the 180 G18 and 1000 G31 datasets. (d) LRA results showing that rs72713175 did not affect enhancer activity in WM88 and MNT-1 cells. Two-tailed paired t-tests (n = 6). (e) CRISPRi of the enhancer containing rs72713175 significantly reduced the expression of BLOC1S6 (control, n = 8; others, n = 6; Two-sided Dunnett’s test with adjustments for multiple comparisons). (f) CRISPRi of the enhancer containing rs72713175 significantly reduced melanin levels in MNT-1 cells (control, n = 18; others, n = 9, Two-sided Dunnett’s test with adjustments for multiple comparisons). Data are presented as mean ± SEM and p values are listed above the bars.
Extended Data Fig. 10 Identification of functional regulatory variants near the DDB1 locus.
(a) Plots showing allelic skewed variants in regulatory elements near the DDB1 locus. rs7948623 overlaps an open chromatin region in melanocytes and many other cell types. rs2277285 and rs2943806 are located within CTCF binding sites and TAD boundaries. Blue tracks indicate DNase-seq, ATAC-seq, and ChIP-Seq data from melanocytes; orange tracks indicate ChIP-Seq data from melanoma (501-mel) cells; gray tracks indicate CTCF ChIP-Seq data from three cell lines; and green tracks indicate DNase-Seq data from ENCODE68. (b-d) Allelic skews at rs7948623, rs2277285 and rs2943806 as estimated by MPRA (n = 3). P values were estimated with a random effects model for mpralm and paired t-tests without multiple testing adjustments. For boxplots, central lines are median, with boxes extending from the 25th to the 75th percentiles. Whiskers further extend by ±1.5 times the interquartile range from the limits of each box. (e) rs7948623 disrupts a MITF binding motif and overlaps ChIP-seq peaks from the Cistrome database103. (f, g) LD pattern between the MFVs at the DDB1 locus. LD was calculated using the 180G18 dataset.
Supplementary information
Supplementary Information
Supplementary Notes 1–5 and Figs. 1–30.
Supplementary Tables
Supplementary Table 1. GWAS statistics of significant GWAS-All SNPs. Supplementary Table 2. GWAS statistics of significant GWAS-Bots SNPs. Supplementary Table 3. MPRA oligo sequences. Supplementary Table 4. MPRA results. Supplementary Table 5. LRA results. Supplementary Table 6. SNP–gene pairs determined by Hi-C and H3K27ac HiChIP. Supplementary Table 7. Allele frequencies of MPRA significant SNPs in global populations. Supplementary Table 8. Primer sequences.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feng, Y., Xie, N., Inoue, F. et al. Integrative functional genomic analyses identify genetic variants influencing skin pigmentation in Africans. Nat Genet 56, 258–272 (2024). https://doi.org/10.1038/s41588-023-01626-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01626-1
- Springer Nature America, Inc.