Abstract
Malaria infection involves an obligatory, yet clinically silent liver stage1,2. Hepatocytes operate in repeating units termed lobules, exhibiting heterogeneous gene expression patterns along the lobule axis3, but the effects of hepatocyte zonation on parasite development at the molecular level remain unknown. Here we combine single-cell RNA sequencing4 and single-molecule transcript imaging5 to characterize the host and parasite temporal expression programmes in a zonally controlled manner for the rodent malaria parasite Plasmodium berghei ANKA. We identify differences in parasite gene expression in distinct zones, including potentially co-adaptive programmes related to iron and fatty acid metabolism. We find that parasites develop more rapidly in the pericentral lobule zones and identify a subpopulation of periportally biased hepatocytes that harbour abortive infections, reduced levels of Plasmodium transcripts and parasitophorous vacuole breakdown. These ‘abortive hepatocytes’, which appear predominantly with high parasite inoculum, upregulate immune recruitment and key signalling programmes. Our study provides a resource for understanding the liver stage of Plasmodium infection at high spatial resolution and highlights the heterogeneous behaviour of both the parasite and the host hepatocyte.
Similar content being viewed by others
Main
Malaria is a mosquito-borne disease caused by Plasmodium spp. parasites. When an infected female Anopheles mosquito bites a mammalian host, it inoculates around 100 sporozoites, a motile form of the parasite capable of target recognition and host interaction1. The sporozoites migrate through the dermis and travel in the bloodstream until they reach the liver through the portal vein2. Upon colonizing a hepatocyte, the parasite encapsulates itself in the plasma membrane, forming a parasitophorous vacuole6, where it rapidly replicates to form a coenocyte of a few thousand nuclei1. At the end of the liver stage of the parasite’s life cycle7 (48–60 h post-infection (hpi)) thousands of individual merozoites are released into the bloodstream, giving rise to the pathological blood stage.
The mammalian liver exhibits spatial heterogeneity. It is composed of repeating anatomical units termed lobules. Each lobule has a diameter of around 0.5 mm in mice and consists of 9 to 12 concentric layers of hepatocytes3. Blood flows from portal nodes through radial sinusoidal channels into draining central veins, creating gradients of oxygen, nutrients and hormones. As a result, hepatocytes in different zones exhibit different gene expression signatures8. Periportal hepatocytes engage in protein secretion, ureagenesis and gluconeogenesis, whereas pericentral hepatocytes specialize in processes such as bile acid production, xenobiotic metabolism and glutamine biosynthesis3. Previous ex vivo studies have suggested that the pace of Plasmodium infection could differ between pericentral and periportal hepatocytes9,10. Studies using bulk RNA measurements have characterized the transcriptomes of host and parasite during the liver stage of infection11,12, and a malaria single-cell atlas has been generated using an ex vivo platform13. However, accounting for the hepatocyte spatial heterogeneity and identifying heterogeneous host and parasite responses requires in vivo single-cell approaches.
Single-cell atlas of liver-stage malaria
To study the Plasmodium liver stage at single-cell resolution, we injected mice with GFP-expressing P. berghei ANKA14 (approximately 106 parasites per injection; Fig. 1a and Methods). We euthanized infected mice at different time points after infection (2, 12, 24, 30 and 36 hpi), and extracted livers for single-cell RNA sequencing (scRNA-seq; Extended Data Fig. 1a–f). We also embedded liver tissue for single-molecule fluorescence in situ hybridization (smFISH) experiments. We sorted GFP+ and GFP− hepatocytes and performed scRNA-seq using the MARS-seq protocol4 (Extended Data Fig. 1g–k). We aligned the reads to both mouse and Plasmodium genomes, obtaining 24,968 hepatocytes of which 4,115 were infected (Fig. 1b,c and Extended Data Fig. 2). Both infected and uninfected hepatocytes exhibited clear zonated expression programmes, as evident from the mutually exclusive expression of the periportally zonated hepatocyte gene Cyp2f2 and the pericentrally zonated gene Cyp2e1 (Fig. 1d). We used a previously established set of hepatocyte landmark genes15, and additionally filtered for those that did not exhibit changes between infected and uninfected hepatocytes to establish a zonation score for each hepatocyte that is correlated with its location along the lobule radial axis (Fig. 1e and Methods). The natural course of malaria infection involves a mosquito bite and inoculation of roughly 100 sporozoites1. To examine whether the inoculum size might affect our results, we complemented our experiments with natural mosquito bite delivery by exposing individual mice to 35 infected mosquitos for 30 min. We further analysed control mice, either non-infected or mock-infected with mosquito saliva (Extended Data Fig. 2a–c and Methods).
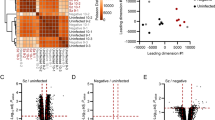
a, Experimental schematic. GFP+ parasites are injected into mice (left) and livers are extracted (middle) at different ensuing time points for scRNA-seq and in situ imaging (right). Middle top, schematic of hepatocyte zonation. CV, central vein; PV, portal vein. b, Uniform manifold approximation and projection for dimension reduction (UMAP) of hepatocytes coloured by time after infection. c, UMAP of hepatocytes coloured by log10 of the sum of Plasmodium reads (ΣPBA). d, UMAP coloured by the expression of the pericentral hepatocyte gene Cyp2e1 and the periportal gene Cyp2f2. Black dots in c,d denote infected hepatocytes. e, Zonation score inferred from the sum of zonated hepatocyte landmark genes. UMAP projections are reconstructed on the basis of the combined mouse and Plasmodium transcriptomes.
Hepatocyte response to malaria infection
Unbiased clustering of the data on the basis of the combined mouse and Plasmodium transcriptomes revealed nine clusters (Extended Data Fig. 2c–g). These included pericentral and periportal uninfected hepatocytes, as well as early and late-infected hepatocytes. The uninfected hepatocytes further clustered according to stress genes, such as Saa1 and Saa2, with mosquito-bitten and non-infected mice exhibiting lower stress responses (Extended Data Fig. 2d,g). A cluster that we termed ‘activated hepatocytes’ was enriched in uninfected hepatocytes at 36 hpi, with strong increases in interferon activity genes such as Ifit1 and Ifit316 (Extended Data Fig. 2c–g). At 2 hpi, both infected and uninfected hepatocytes exhibited a global increase in expression of genes that have previously been shown to be associated with the stress involved in tissue dissociation17, including Cxcl10 and Mt1 (Extended Data Fig. 3a–c). Using smFISH, we demonstrated that expression of these genes indeed does not increase at this stage following infection (Extended Data Fig. 3d–f). Notably, the dissociation-stress signature also appeared at 2 hpi in mosquito-bitten mice, suggesting this programme represents a transient stress stemming from the infection process (Extended Data Fig. 3c). Hepatocytes therefore seem to have a modified cellular state that prompts the induction of stress genes upon tissue dissociation at 2 hpi. Our atlas also included a small cluster consisting of paired hepatocytes and non-parenchymal cells (Extended Data Fig. 2d,e). Each of the clusters included several mice, excluding major batch effects and inter-mouse variability (Extended Data Fig. 2g).
To identify host programmes modified by infection at different time points, we performed differential gene expression analysis between the infected and uninfected hepatocytes. We stratified single hepatocytes by their inferred zone so that comparisons were performed between infected and uninfected hepatocytes that resided at similar lobule coordinates and were sampled at the same time points (Fig. 2a, Extended Data Fig. 4, Supplementary Table 1 and Methods). We found that infected hepatocytes upregulated genes enriched for several programmes, including TNF signalling, interferon-α (IFNα) and IFNγ responses and glutathione metabolism (Fig. 2b). Upregulated genes included Ftl1 and Fth1 (which encode the iron chelator ferritin light and heavy chains, respectively) and Slc40a1 (which encodes the iron export ferroportin-1 transporter) (Fig. 2a). Iron is essential for liver-stage malaria development, and ferritin has been shown to be increased in the serum of infected individuals18. The increased levels of hepatocyte ferritin and ferroportin genes might be an adaptation to deprive the parasite of available iron. Downregulated processes in infected hepatocytes included fatty acid metabolism, bile acid metabolism and complement and coagulation cascades (Fig. 2b). Given the essential role of fatty acids for the parasite development19,20, the reduction in hepatocyte fatty acid biosynthesis genes such as Acly and Fasn (Fig. 2a) might serve to deprive the parasite of these key building blocks. We used smFISH to validate the predicted change in the expression of representative genes in infected hepatocytes (Fig. 2c,d), demonstrating a significant increase in Ftl1 and a decrease in G6pc, Fasn and Apob. We further validated the respective changes in expression in the mosquito-bitten model, using both scRNA-seq (Extended Data Fig. 3h) and smFISH (Extended Data Fig. 3i).
a, MA plot showing the median expression ratio over time and space between infected and uninfected hepatocytes. Cells were binned by time (2, 12, 24, 30 and 36 hpi) and zone (pericentral or periportal), and ratios shown for each gene are the median of the ratios for that gene over all time points. The y-axis shows the log2 of the median ratio per gene, and the x-axis shows the log10 of the gene’s maximum average expression across all time and zone bins. Genes significantly increased or decreased in infected hepatocytes are plotted in red or blue, respectively (false discovery rate (FDR) q-value < 0.01; Methods). Gene symbols are shown for selected genes highlighted with a black circle. b, Gene set enrichment analysis (GSEA) shows an increase in the expression of genes associated with immune and stress pathways alongside a decrease in multiple metabolic pathways in infected hepatocytes. H, Hallmark gene sets; K, KEGG gene sets. c, Quantification of smFISH experiments validates predicted differentially expressed genes in hepatocytes. Two-sided Wilcoxon rank-sum test. Infected: n = 4,109 cells; uninfected: n = 11,739 cells; both from 21 mice sampled at five time points. scRNA-seq Benjamini–Hochberg adjusted P-values: Ftl1: 1.4 × 10−80; G6pc: 1.4 × 10−215; Fasn: 7.5 × 10−58; Apob: 3.4 × 10−172. smFISH Benjamini–Hochberg adjusted P-values: Ftl1: 5.6 × 10−21; G6pc: 2.9 × 10−4; Fasn: 6.4 × 10−3; Apob: 3.0 × 10−4. Bounds of boxes span the interquartile range (IQR), horizontal black lines denote the median, whiskers span 1.5× IQR and black bold circles indicate data points outside this range. d, smFISH images of validated genes at 36 hpi. Phal, phalloidin; PBA-18S, Plasmodium Ch12 18S rRNA (PBANKA_1245821). Blue indicates DAPI signal. Images are representatives of 86 (left) or 85 (right) imaging fields from 3 mice. Scale bars, 10 μm. In all figures, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001; NS, not significant (P ≥ 0.05).
To identify potential zonated patterns in infection rates we analysed the computationally inferred zonation scores of infected hepatocytes (Extended Data Fig. 5a). The single-cell data suggested that pericentral infections were more abundant at all time points; however this differential abundance could stem from differences in the efficiency of single-cell extraction from different lobule zones. To identify zonated features of infection rates in an unbiased manner, we therefore analysed the zonal abundances of infected hepatocytes in situ. We combined smFISH for the periportally zonated albumin-encoding gene Alb8, and established an in situ zonation score on the basis of Alb levels in the hepatocytes that neighbour each infected cell (Extended Data Fig. 5b–d and Methods). We found that infected hepatocytes were not zonated at 2, 15, 24 and 36 hpi but were significantly more abundant in the pericentral zones at 40 hpi (Extended Data Fig. 5d). We further used our scRNA-seq data to demonstrate that the parasite mRNA content was significantly higher in pericentral infections specifically at the later sequenced time points of 30 hpi and 36 hpi (Extended Data Fig. 5e). Parasites infecting pericentral hepatocytes therefore seem to survive and to develop at a higher rate compared with parasites in periportal hepatocytes in the retro-orbital setting with high levels of inoculum.
Periportally enriched abortive infections
We next explored the reasons for the lower abundance and smaller fraction of parasite transcripts in periportal hepatocytes at late time points. We examined the scRNA-seq data of infected hepatocytes at 36 hpi (Fig. 3a). Infected hepatocytes exhibited two distinct clusters based on the host transcriptome. The minor cluster was enriched in periportal hepatocytes (Fig. 3b) and contained cells with significantly lower numbers of parasite reads (Fig. 3c), despite the overall number of counts being the same (log10(sum of counts) = 4.50 ± 0.31 compared with 4.47 ± 0.44 in the major cluster). This minor cluster was enriched in genes related to immune programmes, such as Cxcl10, Nfkbia and Sqstm1, p53 pathway genes such as Mdm2, the transcription factor MYC and its downstream targets, and the Notch downstream target transcription factor HES1 (Fig. 3d–h and Extended Data Fig. 6a–d).
a, Infected hepatocytes exhibit a distinct subcluster at 36 hpi, annotated as ‘Abortive’. b, Zonation scores show that abortive hepatocytes are more periportally zonated compared with productive hepatocytes. c, Abortive hepatocytes harbour a smaller fraction of parasite mRNA at 36 hpi. In b,c, box bounds span IQR, horizontal yellow lines denote the median, whiskers span 1.5× IQR and black bold circles indicate data points outside this range; two-sided Wilcoxon rank-sum test. Cells sampled from five biologically independent mice. d, Volcano plot showing differentially expressed hepatocyte genes between the two clusters. e, UMAP plots of representative genes (bold in d) that are upregulated in abortive hepatocytes. f, GSEA analysis demonstrates increased expression of immune-recruiting pathways, and MYC and p53 pathways. g, Abortive cells harbour a disintegrated vacuole. Images are representatives of 107 (56 productive and 51 abortive) imaging fields from 3 mice. h, SmFISH validation of genes that are increased in expression in abortive hepatocytes. n = 3 biologically independent mice. Two-sided Wilcoxon rank-sum test. Exact P-values: Myc uninfected versus productive: 0.065, productive versus abortive: 4.3 × 10−11, uninfected versus abortive: <2.22 × 10−16; Sqstm1 uninfected versus productive: 0.05, productive versus abortive: 1.1 × 10−7, uninfected versus abortive: 6.8 × 10−13; Mdm2 uninfected versus productive: 0.83, productive versus abortive: 7.2 × 10−8, uninfected versus abortive: <2.22 × 10−16; Cdkn1a uninfected versus productive: 0.17, productive versus abortive: 0.13, uninfected versus abortive: 0.0022. Box bounds span IQR, horizontal red lines denote the median, whiskers span 1.5× IQR, and black bold circles indicate data points outside of this range. i, Abortive cells are distinct from egressing parasites. The dotted white line indicates the region of interest for analysis. Images are representatives of 74 (25 at 48 hpi and 49 at 36 hpi) imaging fields from 6 mice (3 per time point). Scale bars, 10 μm.
Our in situ validations of the signatures of the minor cluster (Fig. 3g,h and Extended Data Fig. 6a–d), identified by their increased levels of marker transcripts (Fig. 3d,e), uncovered distinct morphological features of the minor cluster cells. The parasitophorous vacuole was disintegrated, as evident by the ubiquitous fluorescence of both GFP and parasite 18S rRNA (Fig. 3g). The parasitic nuclei were also dispersed throughout the hepatocyte cytoplasm. Our in situ analysis validated the portal enrichment of these cells (Extended Data Fig. 7a). These events most probably do not represent hepatocytes harbouring productive merozoites, since merozoite formation and release from hepatocytes starts at 50–60 hpi in vitro7. Moreover, blood transfer from liver-stage infected mice gave rise to parasitaemia16 in recipient mice only at 52 hpi and not at 42 hpi. To validate that these hepatocytes constitute a cellular state distinct from merozoite egress, we performed smFISH analysis, probing for the gene encoding the merozoite marker merozoite surface protein 1 (MSP1; PBANKA_0831000) gene. MSP1 expression increased in infected hepatocytes with intact vacuole at 48 hpi and 52 hpi, but was barely expressed in the hepatocytes with vacuole breakdown, either at 36 hpi or 48 hpi (Fig. 3i). Given the pattern of vacuole breakdown and relatively early phase of their appearance, we therefore termed these cells ‘abortive hepatocytes’.
The fractions of hepatocytes with disintegrated vacuole increased over time from 3% at 24 hpi to 17% at 36 hpi and 27% at 40 hpi (Extended Data Fig. 7a). Moreover, abortive hepatocytes also appeared in the mosquito-bitten model (Extended Data Fig. 7b,e). At 36 hpi, the fraction of abortive hepatocytes increased with inoculum size, ranging from 3.5% for the mosquito-bitten mice, to 17% with inoculum containing 2 × 104 parasites and 45% with inoculum containing 106 parasites (Extended Data Fig. 7b). Notably, the pericentral bias of infected hepatocytes increased concordantly with the fraction of abortive hepatocytes (Extended Data Fig. 7c).
Abortive hepatocytes harboured a distinct parasite gene expression signature compared with productive hepatocytes (Supplementary Table 2) that included higher expression of genes encoding the Plasmodium heat shock proteins HSP90, HOP and UIS24 (Extended Data Fig. 7d). We also identified distinctly different mRNA localization patterns for several parasite genes between abortive and non-abortive hepatocytes—the Plasmodium transcripts for LISP1, LISP2 and BIN3 (PBANKA_090330) were localized in distinct foci in productive hepatocytes but were completely interspersed in abortive hepatocytes (Extended Data Fig. 7g). Our analysis therefore highlights a molecular blueprint of periportally biased infected hepatocytes with an abortive phenotype and elevated expression of immune-recruiting programmes. Elimination of these abortive hepatocytes by the immune system could account for the lower abundance of periportally infected hepatocytes at late time points of infection.
Spatio-temporal programmes of the parasite
We next used our single-cell atlas to examine the developmental programmes of the parasite during the liver stage (Fig. 4a,b, Extended Data Figs. 8–10 and Supplementary Table 3). Unbiased clustering of the infected hepatocytes according to the parasite transcriptome revealed clusters that corresponded to the time after infection, as well as abortive hepatocytes (Extended Data Fig. 8). Since parasites develop asynchronously, we applied the Monocle tool21,22 (Fig. 4c,d, Extended Data Fig. 9 and Methods) to assign a pseudotime for each infected hepatocyte. The pseudotime coordinates correlated with the discrete times (Fig. 4d). Notably, pseudotime coordinates were higher in the pericentral hepatocytes at late time points (Fig. 4d), consistent with the higher fraction of Plasmodium transcripts that we observed in this zone (Extended Data Fig. 5e). Our pseudotime analysis enabled characterization of the parasite expression programmes at high temporal resolution (Fig. 4e–h and Supplementary Table 3). Early parasite genes included sporozoite genes13,23,24 (Extended Data Fig. 10a–c), and those encoding RNA polymerases (Extended Data Fig. 10e) and ubiquitin (Fig. 4e), presumably serving to remove the protein content of the preceding sporozoite state. The parasites next sequentially upregulated purine metabolism genes (crucial for the synthesis of mRNA during the ensuing massive parasite proliferation; Fig. 4g), DNA polymerases (Extended Data Fig. 10f) and metabolic programmes for glycolysis, gluconeogenesis and pyruvate metabolism (Fig. 4g and Extended Data Fig. 10i,j). Finally, parasites induced genes associated with biotin metabolism and fatty acid metabolism (Fig. 4h and Extended Data Fig. 10k,l). The late increase in the parasite genes encoding de novo fatty acid biosynthesis coincided with a decline in the earlier expression of the genes encoding UIS4 and UIS3 (Fig. 4b and Extended Data Fig. 10a), which have been suggested to facilitate transport of free fatty acids from the hepatocyte host25. We used smFISH to validate the temporal programmes of selected parasite genes (Fig. 4i–k).
a, UMAP projections of the infected hepatocytes based on the associated Plasmodium transcriptomes, coloured by time after infection. Abortive cells (black outline) are annotated according to the minor cluster in Fig 3a. b, Representative genes colour-coded by stage of infection in which gene expression peaks: early (UIS4; PBANKA_0501200, blue), intermediate–early (GRX1; PBANKA_0404700, yellow), intermediate–late (GS; PBANKA_1111800, orange) and late (LISP2;l PBANKA_1003000, red). c, UMAP projection coloured by pseudotime, calculated with Monocole. d, Pseudotime distributions in cells binned by zone and time after infection. Bounds of boxes span IQR, whiskers span 1.5× IQR, black bold circles are outside of this range, and the dashed red line connects the medians. Two-sided Wilcoxon rank-sum test. e–h, k-means clustering of Plasmodium averaged expression programmes over pseudotime. Titles indicate Plasmodium KEGG pathways enriched in each pseudotime cluster. e, Glycerolipid metabolism, glycerophospholipid metabolism, glyoxylate and dicarboxylate metabolism and ubiquitin genes (asterisk denotes ubiquitin genes, which were the major component of the enriched KEGG arginine biosynthesis pathway) (Supplementary Table 3). f, Porphyrin and chlorophyll metabolism. g, Glycolysis and gluconeogenesis, purine metabolism and pyruvate metabolism. h, Biotin metabolism and fatty acid biosynthesis. Data are mean ± s.e.m. Expression of genes in e–h is normalized to the maximum across time points. i, Pseudotime expression profiles of Plasmodium genes: SAMdMT (PBANKA_1404800 SAM dependent methyltransferase), LCMT (PBANKA_130360 leucine carboxyl methyltransferase) and FABI (PBANKA_122980). Data are mean ± s.e.m., vertical dashed lines are median pseudotime values of the corresponding time after infection. j,k, smFISH validation of genes in i at 24, 40 (n = 2 biologically independent mice) and 36 hpi (n = 3) biologically independent mice. j, Two-sided Wilcoxon rank-sum test. FABI 24 versus 36 hpi: P = 9 × 10−5, 36 versus 40 hpi: P = 0.001; LCMT 24 versus 36 hpi: P = 6.6 × 10−7, 36 versus 40 hpi: P = 0.451; SAMdMT 24 versus 36 hpi: P = 0.71, 36 versus 40 hpi: P = 0.16). Box bounds span IQR, horizontal black lines denote medians, whiskers span 1.5× IQR, and black bold circles show points outside of this range. k, Representative images of all imaging fields analysed in j. Scale bars, 10 μm.
We further identified Plasmodium genes that were differentially expressed between pericentral and periportal hepatocyte host cells (Extended Data Fig. 9i and Supplementary Table 2). To this end, we compared periportal and pericentral hepatocytes at late pseudotime coordinates while ensuring that they were matched in distributions, to account for the more advanced pericentral pseudotime coordinates (Methods). Parasites in periportal hepatocytes exhibited higher levels of GILP—which encodes the glyoxalase I-like protein26—and TLP2—which encodes thioredoxin-like protein 227. Parasites in pericentral hepatocytes exhibited higher levels of GPI16—which encodes glycosylphosphatidylinositol-anchored protein28—as well as several fam-b genes29 (Supplementary Table 2).
Discussion
Here we provide a comprehensive molecular overview of the life cycle of individual parasites and their hepatocyte hosts throughout the Plasmodium liver stage. The lack of zonated abundances at early time points indicates that the parasite does not appear to preferentially colonize hepatocytes at specific zones under a regime of high-inoculum retro-orbital injections. Rather, our results align with a random process of colonization with zone-dependent rates of development and/or survival. The higher pericentral abundance of infections at late time points could be explained either by the lower oxygen tension in the pericentral lobule layers3, which has been shown to promote parasite development ex vivo9, or owing to lower survival of periportal infected hepatocytes. It is also consistent with recent findings of enhanced ex vivo growth of Plasmodium falciparum in pericentral-like hepatocytes10. The liver lobule exhibits immunological zonation30, with a higher periportal abundance of myeloid and lymphoid cells. The decreased frequencies of periportal infections in the high-inoculum setting could be explained by higher elimination rates of abortive hepatocytes, which we found to be more abundant periportally. Indeed, infections at later time points were more pericentrally zonated with higher inoculum sizes, concordantly with the higher abundance of the abortive hepatocytes (Extended Data Fig. 7b,c). Notably, in mosquito-bitten mice, infections at 36 hpi were not significantly zonated. Technical challenges associated the low inoculum size prevent us from determining whether initial infection is zonally random in this more natural scenario.
Abortive hepatocytes exhibit a distinct gene expression signature enriched in immune recruitment genes, such as the IFNγ response and—to a lesser extent—the IFNα response. Type I interferon has been shown to mediate the liver-stage immune response16,31, as well as contribute toimmune mediated pathology32. The abortive cells that we have identified may eventually become immune-infiltrated and eliminated16. It will be informative to apply paired-cell approaches15,33 to explore the interactions between abortive hepatocytes and specific immune cell subsets. Specifically, such studies could resolve whether the abortive state is an intrinsic autonomic response of the hepatocyte to the infection, or rather is induced by interactions with immune cells.
Our scRNA-seq data suggested that abortive hepatocytes strongly increase the activities of key pathways, such as Notch, p53 and MYC. p53 inhibition has been shown to affect liver-stage malaria progression34, and MYC inhibition dampens acute liver failure35. It would be informative to apply our approach of spatially resolved scRNA-seq of infected hepatocytes in mice with drug-induced or genetically induced perturbations of these pathways, to explore the possibility of increasing the rate of the abortive hepatocyte state. Although we observed abortive hepatocytes also in the mosquito-bitten mouse model—which represents a more physiological model of infection compared to inoculation via injection—their frequencies were substantially lower. This implies that abortive hepatocytes may therefore be a rare event in natural settings of infection.
The combined measurements of the parasite and host presented in this study forms a resource for the detailed analysis of the Plasmodiumliver stage, while accounting for the liver’s spatial heterogeneity3. We have identified several potentially adaptive programmes of the host hepatocyte and the parasite, including processes such as iron and fatty acid metabolism, as well as zonal trends in both the host and the parasite. Our atlas can serve as a basis for future exploration of potential vulnerabilities and the identification of targetable host and parasite pathways36.
Methods
Mice and tissues
Experiments were conducted on 6- to 7-week old C57BL/6J female mice. Mice were purchased from the Charles River Breeding Laboratories and were housed in the facilities of the Instituto de Medicina Molecular in Lisbon in a germ-free environment supplied with water and food ad libitum. Mice were kept in 22–24 °C with 45–65% humidity, under 14h/10h light/dark cycle. All in vivo protocols were approved by the internal animal care committee of Instituto de Medicina Molecular and were performed according to national and European regulations. For smFISH, 2 or 3 mice were sampled per time point. For scRNA-seq 4, 5, 4, 3 and 5 mice were sampled at 2, 12, 24, 30 and 36 hpi, respectively, with 4 mice for control (3 non-infected and one mock-infected sampled 2 h after injection). For the mosquito-bitten infection model, scRNA-seq samples were collected from 2 mice at 2 hpi and 3 mice at 36 hpi. Due to technical limitations, mice from time points 24 hpi and 12 hpi were collected ±1–2 h from the designated time point.
Parasite
GFP-expressing P. berghei ANKA (clone 259cl214) was used. Sporozoites were obtained by dissection of the salivary glands of infected female Anopheles stephensi mosquitoes bred and infected at the Instituto de Medicina Molecular. Mice were inoculated using retro-orbital injection. For scRNA-seq, each mouse was injected with 106 sporozoites in 200 µl DMEM. Mock-infected control mouse was injected with filtered mosquito salivary gland debris devoid of the parasite. For smFISH, each mouse was injected with 2 × 104 sporozoites in 200 µl DMEM. In addition, some mice were inoculated via natural exposure to infected mosquitos. Individual mice were exposed to 35 infected mosquitos for 30 min.
Liver dissociation and FACS sorting of hepatocytes
Mice livers were perfused and dissociated into single cells using Liberase Blendzyme 3 recombinant collagenase (Roche Diagnostics) as previously described8. Isolated hepatocytes were sorted on BD FACS Aria IIu using a 130 μm nozzle and 1.5 neutral density filter, running on BD FACSDiva (v6.1.3) (Extended Data Fig. 1). Samples were stained with anti-CD45 (BioLegend 103137, lot B296484, 1:300), anti-CD31 (BioLegend 102509, lot B312608, 1:300) antibodies and DAPI or propidium iodide (PI). Cells were gated to include live cells only (DAPI or PI negative) and exclude doublets (FSC, SSC) and non-parenchymal (CD31− and CD45−). The remaining cells were then gated for infected (GFP+) or uninfected (GFP−) and sorted accordingly. The cells were sorted into 384-well capture plates containing 2 μl lysis solution and barcoded poly(T) reverse transcription primers for MARS-seq4, allowing for both single-cell barcoding and unique molecular identifiers (UMIs) barcoding of mRNA transcripts. Every plate contained uninfected hepatocytes and several rows (3, 5 or 10) of infected hepatocytes. Four wells were left empty on the bottom left corner of each plate (wells O1, O2, P1 and P2) for background control. Sorted plates were spun down, frozen on dry ice and then kept at −80 °C until library preparation.
MARS-seq library preparation and sequencing
Libraries were prepared as previously described4. In brief, mRNA in capture plates were barcoded and reverse transcribed into cDNA then pooled together. The pooled cDNA libraries were then amplified using T7 in vitro transcription and fragmented. Resulting RNA libraries were then tagged with pool-specific barcodes and Illumina sequencing adapters and converted to cDNA again. Pooled libraries were quality controlled at different times of the protocol and prior to sequencing. The final libraries were then pooled together (15–20 at a time) and sequenced using the NextSeq 500/550 kit High Output Kit v2.5 (Illumina 20024906) or NovaSeq 6000 SP Reagent Kit v1.5 (100 cycles, Illumina 20028401). The Illumina output files were converted to fastq format using bcl2fastq (v2.20.0.422) and then aligned to a combined reference genomes of Mus musculus (GRCm38.p6) and P. berghei ANKA (PBANKA01.43) using STAR (v2.7.3a)37 and zUMI (v0.0.6c)38.
scRNA-seq data processing
Data processing (Extended Data Fig. 1g–k) was done in Python (3.7.6; packages include scanpy, numpy, pandas and scipy). For every plate, the background counts per gene were calculated based on the mean expression in the empty wells (defined as wells with less than 1,000 total UMIs). The background was then subtracted from all the wells in the plate. Using the Ensembl database, rows were renamed using ‘gene-id’ indication and pseudogenes were filtered out based on the ‘gene-biotype’ indication. Rows with duplicated gene-ids were merged into a single row. Cells were divided into ‘infected’ and ‘uninfected’ based on sorting scheme per plate. Since sorting/barcoding errors occasionally led to miss-classification of the infected wells, a classifier was further applied based on the logarithm of the ratio between the summed PBA UMIs and the total cellular UMIs (after adding a pseudocount of 1 to both, Extended Data Fig. 1h). For each plate, 100 thresholds for this ratio were scanned, and for each threshold, false positive rates (FPR) and false negative rates (FNR) were computed. FPR was defined as the fraction of cells annotated as uninfected that were above the threshold. FNR was defined as the fraction of cells annotated as infected that were below the threshold. The equal error rate was selected as the threshold for which FPR = FNR. Only cells annotated as infected or uninfected that had a log PBA summed ratio above or below this threshold, respectively, were retained.
Following the filtered annotation, a PBA transcriptomic background was subtracted from all wells in plates that included infected cells. To this end, the annotated uninfected cells were considered as the background. The 99th percentile of every PBA gene in uninfected cells was subtracted from the same gene in all cells. The pooled and background-subtracted dataset was next filtered for high mitochondrial content. The fraction of mouse mitochondrial genes was calculated for every remaining cell, and cells with mitochondrial fraction above the 95th percentile per mouse were removed. Cells were next filtered to retain cells with more than 1,500 and less than 150,000 reads, as well as more than 200 and less than 8,000 genes. Based on the filtered cells, low yielding plates were excluded as follows – For each plate the ratio between the median log summed UMI counts per plate and the median log summed UMI counts per mouse was computed. A z-score was then calculated for this value, and plates with z-score <−1.5 were removed from the dataset (Extended Data Fig. 1k). From this point onwards, all analyses were performed on the reads of protein-coding genes only, excluding mouse mitochondrial genes and major urinary proteins (Mups) known to be highly variable between mice39. In addition, remaining ‘infected’ cells with less than 30 PBA reads were filtered out.
scRNA-seq processed data analysis
The processed UMI table was analysed using Seurat 4.0.140 running on R4.0.2 (packages include dplyr, tidyr, ggpubr, stats, and EnhancedVolcano). Data normalization and scaling followed the suggested default settings for most of the package functions. The data were log-normalized and scaled without regression. Top 2,000 variable genes were selected using the FindVariableFeatures function with the vst method. Principal component analysis was based on these genes, and the first 10 components were used for clustering (0.2 resolution, Louvain algorithm) and dimension reduction (UMAP, using the cosine metric).
Subsets of the main Seurat structure were used to cluster and analyse the 36 hpi infected cells using only mouse transcripts (Fig. 3), and all the infected cells using PBA transcripts (Fig. 4). The subsetted cells were renormalized and scaled based on the respective subset of transcripts. When scaling the 36 hpi subset, the data were batch-corrected (using the regress option in the ScaleData function) based on technical repeats. For each subset, the top 2,000 variable genes were selected, and the first 10 principal components were used for clustering (resolution 0.1 in Fig. 3 and 0.2 in Fig. 4) and UMAP reductions were generated (using the cosine metric in Fig. 3 and the Euclidian metric in Fig. 4). For differential gene expression analysis and pathway enrichment, the raw data was normalized to relative counts per cell in MATLAB R2019a by dividing by the sum of all genes that individually take up less than 0.01 of the cellular summed UMIs when averaged over all cells.
Zonation reconstruction
Single-cell spatial locations along the lobule axis were inferred computationally from the data based on the sum of a panel of landmark genes, as previously described15. However, for this study a smaller subset of landmark genes was used, retaining only genes that showed no significance change in expression between infected and unifected cells. Periportal landmark genes used were: Apof, Apom, Asgr2, Atp5a1, C1s1, C8b, Cpt2, Tkfc, Eef1b2, Fads1, Gc, Hsd17b13, Ifitm3, Igf1, Igfals, Ndufb10, Pigr, S100a1, Serpina1c, Serpina1e, Serpind1, Serpinf1, Uqcrh, Vtn, Arg1 and Cps1. Pericentral landmark genes used were: Alad, Aldh1a1, Nat8f2, Cpox, Cyb5a, Cyp3a11, Lect2, Mgst1, Prodh and Slc16a10.
Pseudotime trajectories
Cell trajectories were calculated using the Monocle 3.1022 package. The normalized data of the infected hepatocyte PBA genes subset was used to find the top 10 principal components using preprocess_cds function (with no additional normalization). Cells were then clustered and UMAP reductions were generated. Branching trajectories were calculated using learn_graph with a minimum branching of length of 10. The single-connection node closest to 2 hpi cell cluster was chosen as a starting node for pseudotime assignment.
Cells were ordered by their pseudotime values and for each gene a moving average of the normalized expression was calculated along this axis (sliding window of 800 cells), generating pseudotime expression matrix for further analysis. When analysing the differences in Plasmodium transcriptomes between hepatocytes at different zones, we compared pericentral and periportal hepatocytes, defined as hepatocytes with a zonation score lower or higher respectively than the 30th percentile over all hepatocytes. Since pseudotime values were higher in pericentral-residing parasites at late time points, we further ensured that the pseudotime distributions were identical. To this end, we compared the periportal infected hepatocytes with pseudotimes above the 80th percentile of that zonal set, with pericentral hepatocytes with pseudotimes sampled to match the same periportal pseudotime distribution, using the inverse transform sampling method41.
Stratified ratio change
To find global markers of Liver-stage infection while excluding spatio-temporal biases, cells were binned based on their metadata into 5 time points (2, 12, 24, 30 and 36 hpi) and 2 zones (pericentral or periportal, according to whether zonation score was lower or higher than the 30th percentile of overall zonation scores). The ratio change of mean normalized expression between infected and uninfected cells was calculated for every mouse gene (Supplementary Table 1) and one-sided Wilcoxon rank-sum tests were used to establish statistical significance of increased or decreased expression in infected cells. Global ratio change was calculated as the median value of the ratio changes per gene in each of the ten bins. Overall significance was calculated using Fisher combined method corrected using Benjamini–Hochberg FDR < 0.01.
Pathways enrichment
For mouse gene set enrichment, genes with mean relative expression greater than 10−5 of summed mouse UMIs were ranked based on their ratio change between cell subsets (infected/uninfected; abortive/productive; pericentral/periportal; and so on). The ranked ratio was the basis for GSEA (v3.0)42. Curated KEGG and Hallmark annotations were used, filtered for minimum 15 genes in set and maximum 500. Default setting of 1,000 permutation was used to establish significance. For abortive cells DGE stricter thresholds were used: relative expression > 10−4, and minimum 30 genes in set.
KEGGREST (v1.28.0) was used in R4.0.2 to assign Plasmodium genes to different Malaria KEGG pathways. Pathways were grouped into four clusters based on their peak expression in pseudotime using k-means clustering in MATLAB (‘Distance’ metric = ‘cosine’).
smFISH
Mice were killed by cervical dislocation and their livers were collected. Tissues were fixed in 4% paraformaldehyde (Santa Cruz Biotechnology sc-281692) for 3 h, incubated overnight with 30% sucrose in 4% paraformaldehyde, embedded in OCT (Tissue-Tek 4583) and stored at −80 °C. Cryosections 8–15 µm thick were used for probe hybridization as previously described43. In brief, the sections were permeabilized in cold 70% ethanol for 2 h, then rehydrated in 2× SSC (Ambion AM9763). Rehydrated tissues were treated with proteinase K (10 µg ml−1 Merck 124568) and then incubated with 5% or 15% formamide (Ambion AM9342) in 2× SSC (5% formamide concentration was used for hybridization of Plasmodium probe libraries with low GC content). Treated sections were then mounted with hybridization buffer (5% or 15% formamide; 10% dextran sulfate (Sigma D8906); 0.02% BSA (Ambion AM2616); 1 mg ml−1 E. coli tRNA (Sigma R1753); 2 mM vanadyl ribonucleoside complex (NEB S1402S); 2× SSC) containing diluted probes and incubated over night at 30 °C. Probe libraries (Supplementary Table 4) were coupled with Cy5 or Alexa594 and diluted to 1:3,000, with the exception of the 18Sp probe that was coupled with Atto 488 and was used in 1:3,000, 1:6,000 and 1:30,000 dilutions for 2 hpi, 15 hpi and 24–52 hpi, respectively. After hybridization, the sections were incubated with 50 ng ml−1 DAPI (Sigma-Aldrich, D9542) in 5% or 15% formamide for 30 min at 30 °C, for nuclear staining and then washed in GLOX buffer (0.01M Tris pH 8.0 (Ambion M9856); 0.4% glucose (Sigma-Aldrich G8270); 2× SSC) until mounting and imaging. Samples that required hepatocyte segmentation, underwent additional staining with 1:500 rhodamine-conjugated phalloidin (Invitrogen R415) in GLOX for 15 min at room temperature. Imaging was performed on Nikon Ti-E inverted fluorescence microscope using the NIS elements software AR 5.11.01. The dot stack images were first filtered with a three-dimensional Laplacian of Gaussian filter of size 15 pixels and standard deviation of 1.5 pixels. Images were used for gene expression validations or in situ zonation analysis.
Single-cell in situ gene expression validations
Single field 100× images were taken of minimum 21 consecutive 0.3 µm z-stacks. ImageM43 was used for segmentation and dot counting. For mouse genes: segmentation was done manually based on phalloidin staining of the hepatocyte borders. Cytoplasms of productive infected cells were segmented to exclude the parasite, while abortive infected cells were segmented as a whole. Nuclei were segmented semi-automatically by the software. Gene transcripts were counted in 10 consecutive z-stacks and divided by the total volume of the segmented cell (excluding nuclei). mRNA concentration of every infected cell was compared to those of 5–10 adjacent uninfected hepatocytes (Wilcoxon rank-sum test, one-sided) and a P-value was obtained. For Plasmodium genes: the parasite vacuole was segmented based on a probe for the parasite chromosome 12 18S rRNA (18Sp, PBANKA_1245821). Gene transcripts were counted in 5–10 consecutive z-stacks and divided by the total volume of the segmented parasite. mRNA concentrations were compared between different time points using Wilcoxon rank-sum test (two-sided).
In situ zonation analysis
Tissue samples were hybridized with Alb mRNA Cy5 and 18Sp Atto488 probes. Scans of the whole tissue sections were imaged at 10× in a single z-stack. Single fields were autofocused and stitched together using NIS elements ‘Scan large image’ feature. Images were manually processed to remove out-of-focus pixels and to identify the positions of infected cells. The processed images were then analysed in MATLAB as follows: A small section of background pixels (usually central/portal vein) were marked and a background threshold established (mean pixel value + 5 × s.d. of the pixel value). A 40 × 40-pixel window was sampled around the centre of every infected cell, then a number of randomly placed 40 × 40-pixel windows was sampled from the image (excluding areas below the background threshold). The median pixel value of the Alb mRNA signal was then calculated for every window, excluding background pixels. Then, for each infected cell, the fraction of random windows with a median value lower than that of the cell was calculated and regarded as an approximate zonation score for every infected cell, 0 being very pericentral and 1 being periportal. The zonation score distributions of consecutive time point were compared using two-sided Wilcoxon rank-sum tests.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Code availability
The code used to process the raw data to a Scanpy/Seurat structure used for analysis is available at https://github.com/AmichayAfriat/SpatioTemporal_malaria_liver_stage_atlas/. Codes for further analysis and data visualization are available at https://doi.org/10.5281/zenodo.7081863.
References
Despommier, D. D. et al. Parasitic Diseases 6th edn (Parasites Without Borders, 2017).
Vaughan, A. M. & Kappe, S. H. I. Malaria parasite liver infection and exoerythrocytic biology. Cold Spring Harb. Perspect. Med. 7, a025486 (2017).
Ben-Moshe, S. & Itzkovitz, S. Spatial heterogeneity in the mammalian liver. Nat. Rev. Gastroenterol. Hepatol. 16, 395–410 (2019).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Bahar Halpern, K. et al. Bursty gene expression in the intact mammalian liver. Mol. Cell 58, 147–156 (2015).
Nyboer, B., Heiss, K., Mueller, A.-K. & Ingmundson, A. The Plasmodium liver-stage parasitophorous vacuole: a front-line of communication between parasite and host. Int. J. Med. Microbiol. 308, 107–117 (2018).
Graewe, S. et al. Hostile takeover by plasmodium: reorganization of parasite and host cell membranes during liver stage egress. PLoS Pathog. 7, e1002224 (2011).
Halpern, K. B. et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature 542, 352–356 (2017).
Ng, S. et al. Hypoxia promotes liver-stage malaria infection in primary human hepatocytes in vitro. Dis. Model. Mech. 7, 215–224 (2014).
Yang, A. S. P. et al. Zonal human hepatocytes are differentially permissive to Plasmodium falciparum malaria parasites. EMBO J. 40, e106583 (2021).
Albuquerque, S. S. et al. Host cell transcriptional profiling during malaria liver stage infection reveals a coordinated and sequential set of biological events. BMC Genomics 10, 270 (2009).
Toro-Moreno, M., Sylvester, K., Srivastava, T., Posfai, D. & Derbyshire, E. R. RNA-seq analysis illuminates the early stages of Plasmodium liver infection. mBio 11, e03234–19 (2020).
Howick, V. M. et al. The malaria cell atlas: single parasite transcriptomes across the complete Plasmodium life cycle. Science 365, eaaw2619 (2019).
Franke-Fayard, B. et al. A Plasmodium berghei reference line that constitutively expresses GFP at a high level throughout the complete life cycle. Mol. Biochem. Parasitol. 137, 23–33 (2004).
Halpern, K. B. et al. Paired-cell sequencing enables spatial gene expression mapping of liver endothelial cells. Nat. Biotechnol. 36, 962–970 (2018).
Liehl, P. et al. Host-cell sensors for Plasmodium activate innate immunity against liver-stage infection. Nat. Med. 20, 47–53 (2014).
van den Brink, S. C. et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat. Methods 14, 935–936 (2017).
Spottiswoode, N., Duffy, P. E. & Drakesmith, H. Iron, anemia and hepcidin in malaria. Front. Pharmacol. 5, 125 (2014).
Yu, M. et al. The fatty acid biosynthesis enzyme FabI plays a key role in the development of liver-stage malarial parasites. Cell Host Microbe 4, 567–578 (2008).
Vaughan, A. M. et al. Type II fatty acid synthesis is essential only for malaria parasite late liver stage development. Cell. Microbiol. 11, 506–520 (2009).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Bogale, H. N. et al. Transcriptional heterogeneity and tightly regulated changes in gene expression during Plasmodium berghei sporozoite development. Proc. Natl Acad. Sci. USA 118, e2023438118 (2021).
Real, E. et al. A single-cell atlas of Plasmodium falciparum transmission through the mosquito. Nat. Commun. 12, 3196 (2021).
Mikolajczak, S. A., Jacobs-Lorena, V., MacKellar, D. C., Camargo, N. & Kappe, S. H. I. L-FABP is a critical host factor for successful malaria liver stage development. Int. J. Parasitol. 37, 483–489 (2007).
Soga, A., Shirozu, T. & Fukumoto, S. Glyoxalase pathway is required for normal liver-stage proliferation of Plasmodium berghei. Biochem. Biophys. Res. Commun. 549, 61–66 (2021).
Kehr, S., Sturm, N., Rahlfs, S., Przyborski, J. M. & Becker, K. Compartmentation of redox metabolism in malaria parasites. PLoS Pathog. 6, e1001242 (2010).
Liu, Q. et al. The glycosylphosphatidylinositol transamidase complex subunit PbGPI16 of Plasmodium berghei is important for inducing experimental cerebral malaria. Infect. Immun. 86, e00929–17 (2018).
Fougère, A. et al. Variant exported blood-stage proteins encoded by Plasmodium multigene families are expressed in liver stages where they are exported into the parasitophorous vacuole. PLoS Pathog. 12, e1005917 (2016).
Gola, A. et al. Commensal-driven immune zonation of the liver promotes host defence. Nature 589, 131–136 (2021).
Miller, J. L., Sack, B. K., Baldwin, M., Vaughan, A. M. & Kappe, S. H. I. Interferon-mediated innate immune responses against malaria parasite liver stages. Cell Rep. 7, 436–447 (2014).
Ribot, J. C. et al. γδ-T cells promote IFN-γ–dependent Plasmodium pathogenesis upon liver-stage infection. Proc. Natl Acad. Sci. USA 116, 9979–9988 (2019).
Giladi, A. et al. Dissecting cellular crosstalk by sequencing physically interacting cells. Nat. Biotechnol. 38, 629–637 (2020).
Kaushansky, A. et al. Suppression of host p53 Is critical for Plasmodium liver-stage infection. Cell Rep. 3, 630–637 (2013).
Kolodziejczyk, A. A. et al. Acute liver failure is regulated by MYC- and microbiome-dependent programs. Nat. Med. 26, 1899–1911 (2020).
Caldelari, R. et al. Transcriptome analysis of Plasmodium berghei during exo-erythrocytic development. Malar. J. 18, 330 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Parekh, S., Ziegenhain, C., Vieth, B., Enard, W. & Hellmann, I. zUMIs—a fast and flexible pipeline to process RNA sequencing data with UMIs. Gigascience 7, giy059 (2018).
Droin, C. et al. Space–time logic of liver gene expression at sub-lobular scale. Nat. Metab. 3, 43–58 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Devroye, L. in Handbooks in Operations Research and Management Science, Vol. 13 (eds Henderson, S. G. & Nelson, B. L.) Ch. 4 (Elsevier, 2006).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Lyubimova, A. et al. Single-molecule mRNA detection and counting in mammalian tissue. Nat. Protoc. 8, 1743–1758 (2013).
Acknowledgements
S.I. is supported by the Wolfson Family Charitable Trust, the Edmond de Rothschild Foundations, the Fannie Sherr Fund, the Dr. Beth Rom-Rymer Stem Cell Research Fund, the Helen and Martin Kimmel Institute for Stem Cell Research, the Minerva Stiftung grant, the Israel Science Foundation grant no. 1486/16, the Broad Institute‐Israel Science Foundation grant no. 2615/18, the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme grant no. 768956, the Chan–Zuckerberg Initiative grant no. CZF2019‐002434, the Bert L. and N. Kuggie Vallee Foundation and the Howard Hughes Medical Institute (HHMI) international research scholar award. L.B. is supported by the European Molecular Biology Organization under EMBO Long-Term Fellowship ALTF 724–2019. This work was also financed by la Caixa Foundation (HR17/52150010) to M.M.M. and Fundação para a Ciência e Tecnologia (LISBOA-01-0145-FEDER-030751 and PTDC/BIA-MOL/30112/2017) to M.M.M. and V.Z.-L.
Author information
Authors and Affiliations
Contributions
S.I. and M.M.M. conceived the study. V.Z.-L., S.M., Â.F.C. and A.L. dissected the mosquitoes, obtained sporozoites and infected the mice. A.A., V.Z.-L., K.B.H., S.M., Â.F.C. and A.L. fixed and embedded liver tissue for imaging, performed liver perfusion, and FACS-sorted single-cell hepatocytes. A.A. and K.B.H. prepared and sequenced the MARS-seq libraries and designed, imaged and quantified smFISH protocols. A.A., L.B. and S.I. wrote the code and performed the data analysis. I.A. contributed to project design. S.I., A.A. and M.M.M. wrote the paper. All of the authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 scRNAseq sampling.
a-f, Representative FACS gating for scRNAseq. (a) FACS gating based on general size to find hepatocytes. (b-c) Gating to discern single-cells from clumps of cells. (d) Negative selection to exclude non-parenchymal cells. e, Gating to include only live hepatocytes. (f) gating for infected and uninfected hepatocytes based on GFP content. g-k, Quality control steps in the scRNAseq process. (g) UMIs (unique molecular identifier) are summed per gene per cell and denoted as either mouse or Plasmodium gene. (h) Quality control for infected tag assigned to cells. Fraction of PBA gene UMIs out of all reads was calculated per cell. Based on the log10 of that value (Y axis) a threshold was established (red line, Methods) to identify erroneous tagging of cells. Infected/Uninfected cells below/above the threshold are discarded. Bounds of boxes span IQR, horizontal black lines denote medians, whiskers span 1.5× IQR. (i) Identifying cells with high mitochondrial gene fraction. Per mouse, the 95th percentile of cells, based on their Mouse mitochondrial gene content (Y axis), was discarded from the dataset. (j) filtering all cells based on their high/low total transcriptome yield. X axis – total number of UMIs; Y axis – Total number of genes with at least 1 UMI assigned; cells colored by infection tag (green – Infected, blue – uninfected). Cells were discarded if they fell below (teal lines) or above (red lines) certain thresholds for each category (dashed line – number of UMIs; dotted line – number of genes). (k), Filtering plates outliers based on Z-score for the ratio between median UMIs per cell in plate compared to all cells from mouse. Per mouse, every cell total UMI count was summed, then the median value of log10 that sum for all cells assigned to plate was divided by the median value of log10 that sum for all cell assigned to mouse (Y axis). Based on that value a Z-score (white text on bars) was calculated per plate. Plates with Z-score lower than −1.5 (red outline) were excluded from further analysis.
Extended Data Fig. 2 Unsupervised clustering of the Plasmodium liver stage atlas.
a, UMAP of hepatocytes clustered based on both host and parasite transcriptome, colored by hours post-infection (hpi) and inoculation method. NI – non infected mouse, MB – mosquito bitten mouse. b, UMAP colored by Infected/Uninfected tag assigned to cells based on sorting. Infected cells are in red. c, UMAP colored by Louvain cluster (Methods). d, Heat map for marker genes per cluster. purple – low expression; yellow – high expression. e, Dotplot of specific markers for dividing (Mki67, Tubb5), immune (Ptprc, and specifically macrophages - Marco) and endothelial (Ptprb) cells, as well as interferon activation (Isg15, Ifit1, Irf7). f, The contribution of different sampled time points to the composition of individual clusters and the complete atlas. g, The contribution of different mice to the composition of individual clusters and the complete atlas.
Extended Data Fig. 3 Additional scRNAseq analysis.
a-g, ScRNAseq of uninfected cells acquired at 2hpi show increase in genes associated with tissue dissociation. a, Volcano plot of differentially-expressed hepatocyte genes between uninfected cells acquired at 2hpi and at every other time-point. Bold genes are highlighted in (d-f). b, UMAP plot colored by log10 summed relative expression of dissociation marker genes (van den Brink et al. 2017)17. c, Violin plot of stress indication in uninfected cells binned by time demonstrates notable increase in dissociation stress at 2hpi, both for injection inoculated and mosquito-bitten mice. Bounds of boxes span IQR, horizontal black lines denote the median, whiskers show data within 1.5*IQR. In order from left-to-right, n cells = 3,205/1,445/1,917/2,308/2,838/1,560/3,116/1,632/2,832 over 3/1/4/5/4/3/5/2/3 biologically independent mice. d-e, UMAP plots colored by normalized expression of representative up-regulated genes in the 2hpi cluster. UMAP projections reconstructed based on the combined mouse and Plasmodium transcriptomes. f, smFISH images of Cxcl10 and Mt1 at 2hpi and 24hpi uninfected hepatocyte show no discernible change in expression. Cxcl10 mRNA in red (positive control signal in non-parenchymal cell indicated by white arrow on left panel), Mt1 mRNA in green, Dapi in blue. Images are representatives of 40 imaging fields (15 for 2hpi and 25 for 24hpi) from 4 mice (2 per time point). g, Volcano plot of differentially-expressed hepatocyte genes between mock infected mouse acquired at 2hpi and hepatocytes from uninfected mice. h-i, Host hepatocyte transcriptome changes are also observed in mosquito-bitten mice. h, scatter plot of the log10 mean expression of genes in non-infected hepatocytes (Y axis) vs. infected hepatocytes (X axis) in the mosquito-bitten mice at 36hpi (n = 3 mice). Genes significantly increased or decreased in Fig. 2a are plotted in red or blue respectively. The names of selected genes from Fig. 2a are higlighted. i, Representative smFISH image of genes at 36hpi in mosquito-bitten mouse show similar changes to those observed in injected mice (Fig. 2d). Phalloidin (Phal) in white, Dapi blue; Ftl1 mRNA in red, Apob mRNA in green. Dashed white line - infected hepatocyte; Dotted yellow line - parasite. Image is representative of five imaging fields over two mice.
Extended Data Fig. 4 Differential gene expression in different spatio-temporal bins.
a, MA plots of the median expression ratio between infected and uninfected hepatocytes, split into 10 spatio-temporal bins. Y axis indicates log2 of the median ratio per gene. X axis indicates log10 of the gene’s max average expression. Genes significantly increased or decreased in infected hepatocytes are plotted in red or blue respectively (FDR q-value < 0.01). The names of selected genes from Fig. 2a marked with black circle are highlighted. b, Gene set enrichment analysis (GSEA) for highly expressed genes ranked based on ratio changes indicated in (a). (H) denote Hallmark gene sets, (K) denote KEGG gene sets. PC - pericentral, PP - periportal.
Extended Data Fig. 5 Pericentral infected hepatocytes become more abundant at late time points.
a, Violin plots of computationally-inferred zonation scores computed on the scRNAseq data. bounds of boxes span IQR, horizontal black lines denote the median, whiskers span 1.5*IQR (Interquartile range), dashed line demonstrates the median change between groups. Significance determined by two-sided Wilcoxon rank-sum test, with p-value adjusted using Benjamini–Hochberg procedure. b, smFISH of Albumin (Alb, white) and parasite Ch12 18S rRNA (18Sp, green). Yellow squares represents the 40x40 pixels window around the infected cells, number indicates median pixel intensity. Red dots represent the centers of 1248 40x40 pixel windows (“neighborhoods”) randomly placed onto the tissue area. “CV” denotes central-vein locations. Red blow up of 100x100 pixels (11x11 micron) around a pericentral infected cell. Letters indicate the infected cells plotted on c, Histogram of median pixel value of random neighborhoods in the image shown in (b), red X represent values of infected neighborhoods labeled correspondingly to the yellow labels in (b). Note that infected cell ‘C’ resides in a more periportal zone (as evident by the higher Alb levels in its neighborhood). d, smFISH quantification demonstrates lower proportions of infected periportal hepatocytes at 40hpi. Shown is the zonation score based on the smFISH images, namely the probability to observe a neighborhood zonation lower than the value observed in a random cell within the image field. Data points per category represent cells imaged from multiple non-consecutive tissue sections, sampled from 2 biologically independent mice per time point (Methods). e, scRNAseq analysis demonstrates that periportal parasites have lower transcript counts. Cells per time-point (Pericentral/periportal): 2hpi = 69/122, 12hpi = 179/715, 24hpi = 760/692, 30hpi = 182/307, 36hpi = 364/719 taken from 4/5/4/3/5 mice respectively. For (d) and (e), bounds of boxes span IQR, horizontal red lines denote the median, whiskers span 1.5*IQR (Interquartile range), black-bold circles indicate datapoints outside of said range. Significance determined by two-sided Wilcoxon rank-sum test, with p-value adjusted using Benjamini–Hochberg procedure.
Extended Data Fig. 6 Abortive hepatocytes show unique transcriptomic markers.
a-d, smFISH validations of host genes up regulated in abortive hepatocytes at 40hpi. Top row – Abortive; Bottom row – Productive. Per pair of images: Left panel - Phalloidin (Phal) in red, parasite Ch12 18S rRNA (18Sp, PBANKA_1245821) in green, Dapi blue; Right panel – Cxcl10/Cdkn1a/Sqstm1/Hes1 mRNA in white. Dashed yellow line: infected hepatocyte. Images are representatives of 35/30/30/49 (Cxcl10/Cdkn1a/Sqstm1/Hes1) imaging fields from 2 mice.
Extended Data Fig. 7 Properties of abortive hepatocytes.
a, Abortive hepatocytes are periportally zonated in-situ at 24hpi and 36hpi. Shown is the zonation score based on the smFISH images, namely the probability to observe a neighborhood zonation lower than the value observed in a random cell within the image field, bounds of boxes span IQR, horizontal black lines denote the median, whiskers span 1.5*IQR (Interquartile range). Significance was calculated using fisher’s method on one-sided Wilcoxon rank-sum test per mouse repeat (n = 2/3/2 for 24hpi/36hpi/40hpi respectevly). b-c, Pericentral bias of infection increase concordantly with the frequency of abortive cells. b, Average frequency of abortive infection at 36hpi in increasing inoculums. Error-bars denotes the standard deviation out of n = 3 biologically independent mice for injection model and n = 2 for mosquito bite model. c, zonation score in-situ similar to (a). d, MA plots of the mean PBA expression ratio between abortive and productive hepatocytes at 36hpi. Y axis indicates log2 of the mean ratio per gene. X axis indicates log10 of the gene’s max expression. PBA genes significantly increased or decreased in infected hepatocytes are plotted in red or blue respectively (FDR q-value < 0.01, ratio > 1.5). e, Abortive hepatocytes exhibit similar transcriptional changes in mosquito-bite model. Left panel - Phalloidin (Phal) in white, parasite Ch12 18S rRNA (18Sp, PBANKA_1245821) in green, Dapi blue; Right panel – Mdm2 mRNA in red, Myc mRNA in green. Dashed yellow line: infected hepatocyte. f, Abortive cells exhibit similar transcriptional changes even at 52hpi. Left panel - Phalloidin (Phal) in white, parasite Ch12 18S rRNA (18Sp, PBANKA_1245821) in green, Dapi blue; Right panel – Mdm2 mRNA in red, Myc mRNA in green. Dashed yellow line: infected hepatocyte. g, Abortive parasites exhibit loss of mRNA localization for some Plasmodium genes.
Extended Data Fig. 8 Unsupervised clustering of the infected hepatocytes based on Plasmodium transcriptome.
a, UMAP colored by hours post-infection (hpi) and inoculation method. b, UMAP colored by Abortive/Productive tag assigned to cells based on clustering analysis in Fig. 3a. c, UMAP colored by Louvain cluster (Methods). d, Heatmap for marker genes per cluster. purple – low expression; yellow – high expression. e, The contribution of different sampled time points to the composition of individual clusters and the complete infected hepatocytes atlas. g, Total Plasmodium (PBA) reads increase with time in productive clusters. Y axis - log10 of the sum of PBA reads. Pairwise significance between clusters was determined by two-sided Wilcoxon rank-sum test (ns:p > 0.05; ****:p < = 0.0001; ELS-MELS:p < 2.22e-16, MELS-MLLS:p < 2.22e-16, MLLS-LLS:p = 0.15, LLS-Abortive:p = 3.9e-7; n cells per cluster from left-to-right: 1,657/1,434/507/319/198). bounds of boxes span IQR, horizontal black lines denote the median, whiskers span 1.5*IQR (Interquartile range), black-bold circles indicate datapoints outside of said range.
Extended Data Fig. 9 Pseudotime and spatial analysis of the infected hepatocytes based on Plasmodium transcriptome.
a-d, Monocle UMAP visualization of the infected hepatocytes superimposed with their pseudotime trajectories. Initial node marked with white dot. a, Cells colored by pseudotime value. b, cells colored by sampled time point (hours post infection, hpi). c, Cells colored by their Abortive/Productive tag assigned based on clustering analysis in Fig. 3a. d, Cells colored by the 3 main branching trajectories. e, main UMAP figure colored by the 3 main branching trajectories. Note that branch B3 is in complete overlap with the “Abortive” cell cluster as shown in Extended Data Fig. 8c, while branch B2 is a subset of cluter “Mid-early LS”. f-g, B2 branch is a batch effect of 2 plates with elevated number of reads. f, Heatmap of marker genes per branch. purple – low expression; yellow – high expression. B2 markers are mainly ribosomal genes. g, log10 of total Plasmodium reads is significantly higher in branch B2 compared to B1 (two-sided Wilcoxon rank-sum test). h, Pseudotime distributions in cells binned by cluster. Pairwise significance between consecutive groups was determined by two-sided Wilcoxon rank-sum test (**:p < = 0.01; ****:p < = 0.0001; adjusted p-values for all comparisions p < 2.22e-16; n cells per cluster from left-to-right: 1,657/1,434/507/319/198). bounds of boxes span IQR, horizontal black lines denote the median, whiskers span 1.5*IQR, black-bold circles indicate datapoints outside of said range. i, MA plot (log ratio – M; to max average - A) showing zonally enriched genes in late pseudotime. Y axis indicates log2 of the mean ratio per gene. X axis indicates log10 of the gene’s max average expression. Genes significantly increased pericentral or periportal hepatocytes are plotted in red or blue respectively (FDR q-value < 0.2, Methods). The names of selected genes marked with black circle are highlighted.
Extended Data Fig. 10 Pseudotemporal expression in different Plasmodium gene sets.
a, Marker genes for salivated sporozoites (taken from Bogale et al23). b, Marker genes for Activating sporozoites (taken from Real et al24). c, Marker genes for sporozoites in the Malaria Cell Atlas (taken from Howick et al13). d, Marker genes for exoerythrocytic forms (EEF) in the Malaria Cell Atlas (taken from Howick et al13). e-f, RNA/DNA polymerases. g, Genes related to Plasmodium KEGG pathway “Glycerolipid metabolism”. h, Genes related to Plasmodium KEGG pathway “Glycerophospholipid metabolism”. i, Genes related to Plasmodium KEGG pathway “Pyruvate metabolism”. j, Genes related to Plasmodium KEGG pathway “Glycolysis / Gluconeogenesis”. k, Genes related to Plasmodium KEGG pathway “Biotin metabolism”. l, Genes related to various Plasmodium KEGG pathways relating to fatty acids (e.g. biosynthesis, elongation, degradation). Pseudotemporal expression Color indicates gene-wise normalization over pseudotime with yellow = 1 = max expression. m, Pie chart showing the distributions of the times post infection for cells included in each of the 8 pseudotime bins in a-l.
Supplementary information
Supplementary Table 1
Stratified MUS expression over time and space. Mouse gene expression of zonally-stratified cells at different time points tagged as uninfected (NI) or infected (INF). Shown are means, SEMs (Units are fraction of cellular UMIs), stratified median ratio per-gene, one-sided Wilcoxon rank-sum test p-value and Benjamini–Hochberg FDR q-value (left: up regulation in INF, right: down regulation).
Supplementary Table 2
DGE. Differential Plasmodium/Mouse gene expression of (1) 36hpi cells tagged as either abortive or productive (2) zonally-stratified infected cells in late pseudotime. Shown are means [Units are fraction of cellular UMIs], ratio between means, two-sided Wilcoxon rank-sum test p-value, and Benjamini–Hochberg FDR q-value.
Supplementary Table 3
Stratified expression over pseudotime. Plasmodium and Mouse gene expression of pseudotemporally-stratified infected cells. Shown are means, SEMs [Units are fraction of cellular UMIs], center of mass (COM) and KEGG pathways that include this gene (for PBANKA).
Supplementary Table 4
Probes used in this study. Sequences of the smFISH probes libraries used in this study.
Supplementary Table 5
Sample statistics. Detailed breakdown of the contribution of different samples to the scRNA-seq dataset.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Afriat, A., Zuzarte-Luís, V., Bahar Halpern, K. et al. A spatiotemporally resolved single-cell atlas of the Plasmodium liver stage. Nature 611, 563–569 (2022). https://doi.org/10.1038/s41586-022-05406-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05406-5
- Springer Nature Limited
This article is cited by
-
Host-pathogen interactions in the Plasmodium-infected mouse liver at spatial and single-cell resolution
Nature Communications (2024)
-
Disentanglement of single-cell data with biolord
Nature Biotechnology (2024)
-
A spatiotemporal atlas of mouse liver homeostasis and regeneration
Nature Genetics (2024)
-
Analysis of codon usage bias of thioredoxin in apicomplexan protozoa
Parasites & Vectors (2023)
-
Liver in infections: a single-cell and spatial transcriptomics perspective
Journal of Biomedical Science (2023)