Abstract
Deoxyribonucleic acid in eukaryotes wraps around the histone octamer to form nucleosomes1, the fundamental unit of chromatin. The N termini of histone H4 interact with nearby nucleosomes and play an important role in the formation of high-order chromatin structure and heterochromatin silencing2,3,4. NuA4 in yeast and its homologue Tip60 complex in mammalian cells are the key enzymes that catalyse H4 acetylation, which in turn regulates chromatin packaging and function in transcription activation and DNA repair5,6,7,8,9,10. Here we report the cryo-electron microscopy structure of NuA4 from Saccharomyces cerevisiae bound to the nucleosome. NuA4 comprises two major modules: the catalytic histone acetyltransferase (HAT) module and the transcription activator-binding (TRA) module. The nucleosome is mainly bound by the HAT module and is positioned close to a polybasic surface of the TRA module, which is important for the optimal activity of NuA4. The nucleosomal linker DNA carrying the upstream activation sequence is oriented towards the conserved, transcription activator-binding surface of the Tra1 subunit, which suggests a potential mechanism of NuA4 to act as a transcription co-activator. The HAT module recognizes the disk face of the nucleosome through the H2A–H2B acidic patch and nucleosomal DNA, projecting the catalytic pocket of Esa1 to the N-terminal tail of H4 and supporting its function in selective acetylation of H4. Together, our findings illustrate how NuA4 is assembled and provide mechanistic insights into nucleosome recognition and transcription co-activation by a HAT.
Similar content being viewed by others
Main
NuA4 contains 13 subunits, of which Esa1 is the catalytic subunit for the acetylation reaction and Tra1 is the scaffold subunit for binding to a variety of transcription activators or transcription factors to recruit NuA4 for targeted gene activation11,12,13. Esa1 interacts with Epl1, Yng2 and Eaf6 to form a Piccolo subcomplex14, which achieves selective H4 acetylation (H4-Ac) in the context of the nucleosome through a position-based mechanism15. Tra1 is the common subunit shared by another transcription co-activator HAT complex, SAGA, which catalyses H3 acetylation on nucleosomes12,13. Despite its fundamental importance, currently available structures of the NuA4 complex are at very low resolution16,17,18 and it remains unclear how HAT complexes interact with the nucleosome substrate15,19.
Structural determination of the complex
To elucidate the mechanism of NuA4, we determined the cryo-electron microscopy (cryo-EM) structure of NuA4 bound to the nucleosome. Endogenous NuA4 complex was purified from yeast cells (Extended Data Fig. 1a), mixed with nucleosome core particles (NCPs) and subjected to single-particle cryo-EM analyses (Extended Data Fig. 2a). A low-resolution structure was achieved, which nevertheless showed the molecular envelope of the large subunit Tra1 and a nucleosome-like density nearby (Extended Data Fig. 2b). To reduce the flexibility of the complex, the upstream activation sequence (UAS) for Gal4 binding was engineered into the linker DNA sequence of the nucleosome (UAS-20N20) and the transcription activator Gal4-VP16 was added to the NuA4–NCP complex. It is interesting to note that the UAS, when sited at different positions, promoted the HAT activity of NuA4 (Extended Data Fig. 1b), suggesting plasticity of the complex. In the presence of Gal4-VP16 the structure was determined at an overall resolution of around 8.7 Å, and showed the envelope of the nucleosome and an unknown density on the disk face (Extended Data Fig. 2c,d). Focused refinements did not resolve the unknown density.
Considering that the bisubstrate inhibitor, carboxymethyl coenzyme A (CMC), stably binds to the Esa1 catalytic pocket20,21 and that the chromo domain of Eaf3 recognizes methylated H3K36 (refs. 22,23), we modified the histones at H4K16 to contain CMC, and at H3 to contain trimethylated Lys36, to further limit the flexibility of the complex (Extended Data Fig. 3). Under these conditions, the structure was determined at an overall resolution similar to that given above, and focused refinements led to resolutions of 3.1 and 3.4 Å, respectively, at the TRA and HAT modules bound on top of the nucleosome, with local resolution of 2.7–2.8 Å (Fig. 1, Extended Data Figs. 4 and 5, Extended Data Table 1 and Supplementary Video 1). Whereas the linker DNA in the high-resolution map was barely detected because of the focused classification and refinement, the overall map of the holo-complex covered a linker DNA of approximately 25 base pairs (bp) and a local map (at a resolution of about 6.7 Å) covered a longer linker DNA (around 30 bp) and helped to orientate the UAS-20N20 nucleosome (Fig. 1a and Extended Data Fig. 4b). The orientation of the nucleosome was also supported by the high-resolution maps of the central base pairs (Extended Data Fig. 5b). The subunit interactions were confirmed by cross-linking mass spectrometry (CL–MS; Extended Data Fig. 1c,d).
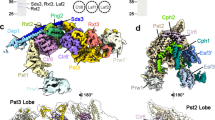
a, Three views of the high-resolution composite maps of the NuA4–NCP complex. The low-resolution molecular envelopes of the all-overall complex (top) and UAS linker DNA (bottom) are shown. b, Three views of the ribbon model of the NuA4–NCP complex. The 25-bp linker DNA is shown, with the last 5 bp of DNA coloured as UAS. c, Model of the assembly and mechanism of nucleosome recognition of NuA4. The transcription factor (TF) is schematically illustrated interacting with the ABS and recognizing the UAS. d, Domain organization of NuA4 subunits and histones. Proteins are colour coded, with those not structurally resolved in white. CC, coiled coil; PBL, polybasic loop.
Overall structure of NuA4
NuA4 is organized into two major modules: the catalytic HAT module and the transcription activator-binding TRA module (Fig. 1a). The HAT module of NuA4 is essentially the core Piccolo subcomplex14, comprising the catalytic subunit Esa1, the N-terminal EPc-A fragment of Epl1 and the PHD domain containing Yng2 and Eaf6 (Fig. 1b–d). The TRA module includes the largest subunit Tra1, Actin, Arp4, Eaf2, Eaf1 and the EPc-B domain of Epl1. Actin, Arp4 and the HSA domain of Eaf1 assemble into a submodule (referred to as the ARP submodule) and bind to the head of Tra1 as described previously17. The two major modules are linked through a disordered connecting loop between the EPc-A and -B domains of Epl1, consistent with the association of Piccolo with the remainder of NuA4 through the Epl1 C-terminal domain14. Eaf3, Eaf5 and Eaf7 formed a subcomplex24, but their structure could not be clearly resolved. Eaf3 extensively cross-linked to the connecting loop of Epl1 (Extended Data Fig. 1c), suggesting that the bridge density connecting the HAT and TRA modules resulted from the Eaf3/5/7 subcomplex (Fig. 1a). Multi-body refinement analysis indicated different modes of relative movement of the two major modules of NuA4 around a polybasic surface (PBS) formed by the Epl1 and Eaf2 subunits of the TRA module (Supplementary Video 2), explaining the plasticity of the complex and the difficulty in resolving a high-resolution structure.
Activator-binding surface of Tra1
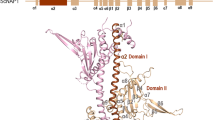
The structure of the complex provides the framework for NuA4 to function as a transcription co-activator. Tra1 comprises over 3,000 residues, binds to various transcription factors and recruits NuA4 for targeted gene expression13. The linker DNA carrying the UAS packs against the edge of the TRA module and projects towards the Finger and FAT domains of Tra1 (Fig. 2a). Intriguingly, the exposed surface of Tra1 facing the UAS is highly conserved from yeast to human, suggesting functional importance, whereas the surface at the opposite side is more variable (Fig. 2b). We propose that the conserved surface of Tra1 serves as the activator-binding surface (ABS). In support of this notion, His400, which is required for Gal4 binding, maps to the conserved surface of the Finger domain25. Furthermore, binding sites Gcn4 and Rap1 (ref. 26) and residues 88–165 and 319–399 also map to the nearby region. The VP16 binding site is less well defined and was proposed to involve the Tra1 C-terminal domain27. The structure of VP16 is not currently resolved. Consistent with the proximity of linker DNA to the FAT domain of Tra1 (Fig. 2a), Gal4-VP16 was cross-linked to multiple residues (Lys2808, Lys2984 and Lys3161) of the FAT domain of Tra1 (Extended Data Fig. 1c and Supplementary Table 1). The conserved surface extends along the Finger domain to the FAT domain, spans a large distance of over 100 Å and potentially interacts with different transcription factors that, independently and/or cooperatively, recognize the adjacent linker DNA (UAS) of the nucleosome to achieve targeted recruitment of NuA4 for gene activation.
a, Ribbon model of the TRA module with Tra1 domains Ring, Finger, FAT, Kinase and FATC coloured differently. The other subunits are coloured as in Fig. 1. The boxed region is enlarged for analysis in d. b, Surface conservation of Tra1. Increasing conservation scores of residues are shown in the red–white–blue spectrum39. Boxed regions are enlarged for analysis in g,h. c, Structural comparison of NuA4 and SAGA (coloured yellow, PDB 6T9I)19. The structures of Tra1 are aligned. The position of the HAT module of SAGA is schematically illustrated and the boxed region (right) is enlarged for analysis. d, Structure of the PBS region of the TRA module. e, Growth phenotype of WT and three mutant yeast cells under the indicated conditions. KAc, potassium acetate. Three independent sets of experiments were performed, with a representative example shown. f, Relative HAT activity of purified WT and DM mutant NuA4 complex. The activity of the WT sample was normalized to 1. Five technical replicates were performed. The activity of an independent batch of samples is shown in Extended Data Fig. 1e. For gel source data, see Supplementary Fig. 1. g, Structure of the ARP submodule. The boxed region is enlarged (right) for analysis. h, Interactions between the Act1–Arp4 dimer with Eaf1, Eaf2 and Epl1.
Tra1 is also responsible for transcription activator binding in the SAGA complex12,13. The Eaf1 binding surface of Tra1 in NuA4 overlaps with the Spt20-Taf12 binding surface in the SAGA complex19 (Fig. 2c), explaining the mutual exclusive assembly of Tra1 into NuA4 and SAGA28. The HAT module in SAGA is oriented relative to Tra1 in a manner analogous to that of NuA4, suggesting that the ABS of Tra1 identified in NuA4 also provides the mechanism of nucleosome recruitment for SAGA.
Polybasic surface of Epl1 and Eaf2
The PBS of the TRA module is composed of polybasic loops of Eaf2 and Epl1 (Fig. 2d) and is positioned adjacent to the linker DNA at the nucleosome edge (Fig. 1b). Lys311, Lys314 and Lys316 of Eaf2 are highly conserved (Extended Data Fig. 6) and exposed to the nucleosomal linker DNA. Likewise, Lys491 and Lys496 of Epl1 are exposed and polybasic sequences at the equivalent positions are found in the homologues (Extended Data Fig. 7). There is a gap between the PBS and linker DNA, suggesting long-range charge–charge interactions. Given the flexibility between TRA and HAT modules, it is very probable that the PBS interacts with varied positions of the DNA, in line with HAT activation by the UAS located at different positions of the linker DNA (Extended Data Fig. 1b).
To examine the importance of the PBS, we replaced wild-type (WT) Epl1 with the polybasic loop mutant (K491E/K496E, Epl1-KE for simplicity). Similarly, we introduced the mutant Eaf2 (K311E/K314E/K316E, Eaf2-KE) and a combination of the Epl1 and Eaf2 mutations in the double mutant (DM). Whereas mutant cells showed normal growth comparable to WT cells in yeast peptone dextrose (YPD) medium, the Epl1-KE mutant showed a growth defect using alternative carbon sources such as glycerol and acetate other than dextrose (Fig. 2e). The Eaf2-KE mutant did not show any obvious defect, and the DM mutant showed a slow growth phenotype similar to the Epl1-KE mutant. These data indicate that the in vivo function of NuA4 depends on the PBS, the polybasic loop of Epl1 in particular, which is closer to the linker DNA (Fig. 2d).
Consistent with the data for cells and their structure, DM mutations did not perturb the integrity of the complex (Extended Data Fig. 1e) but the purified enzyme showed a reduced HAT activity relative to the WT NuA4 complex in vitro (Fig. 2f and Extended Data Fig. 1e). Together, these data indicate that interaction of the nucleosomal linker DNA with the PBS is important for the optimal activity of NuA4.
The polybasic loops of Eaf2 and Epl1 bind to the SANT domain of Eaf1 (Fig. 2d), which was previously assigned to a position close to the ARP submodule based on low-resolution mapping17. However, our high-resolution map indicates that the Eaf1 SANT domain interacts with the FAT domain of Tra1 (Extended Data Fig. 5a), consistent with cross-linking analysis (Extended Data Fig. 1c) and previous studies showing that this domain is required for Tra1 association29. The C-terminal domain of Eaf2 following the polybasic loop is disordered. Given that the YEATS domain-containing subunit Yaf9 cross-linked to the Eaf2 C-terminal domain30 (Extended Data Fig. 1c), the structure suggests that Yaf9 is tethered close to the nucleosome around the PBS of the TRA module, consistent with its role in binding to acetylated H3 tails through the YEATS domain31.
Structure of the ARP submodule
The ARP submodule binds to the head of Tra1 through a helical base formed by Eaf1, Eaf2 and Epl1 (Fig. 2b,g). This structure explains the multiple roles of Eaf1 in the NuA4 assembly18,29,32. The SANT domain of Eaf1 binds to the edge of Tra1 (Fig. 2d) and the HSA helix interacts with the Act1–Arp4 dimer17 (Fig. 2h). The α2–α4 helices of Eaf1 (residues 285–340; Extended Data Fig. 8) bundle with α7–α8 of Eaf2 and α13 of Epl1, forming the helical base that binds to the surfaces of the FAT, kinase and FATC domains of Tra1 (Fig. 2g). This structure is consistent with the importance of a fragment of Eaf1 before the HSA helix for Tra1 binding18. The structure is also consistent with previous studies showing that deletion of the FATC domain (residues 3,711–3,724) of Tra1 causes postactivator-recruitment defects in transcription26, which probably perturbs the binding of the ARP submodule.
Actin and Arp4 in NuA4, previously proposed to function in substrate recruitment17,33, locate at a position proximal to, but not directly contacting, the nucleosome (Fig. 1b), suggesting that their function in substrate recruitment is probably achieved in an indirect manner. The Act1–Arp4 heterodimer in NuA4 is stabilized by a series of previously unrecognized interactions (Fig. 2h). A highly conserved β-hairpin (β6–β7) of Epl1 interacts with the HSA helix and the Act1–Arp4 heterodimer (Extended Data Figs. 5a and 7). Likewise, an N-terminal β-hairpin (β2–β3) of Eaf2 interacts with Actin and Arp4 and may play a role analogous to that of the β-hairpin of Rtt102 in stabilization of Arp7–Arp9 in the SWI/SNF complex34,35. Moreover, Arp4 packs against the SANT domain of Eaf2, the density of which was assigned to the Eaf1 SANT domain17. The high-resolution map clearly supports the sequence identity of Eaf2 (Extended Data Fig. 5a). Our findings show the extensive contacts between Eaf2 and the Act1–Arp4 dimer, which provide the structural basis for them to form a stable submodule shared by the NuA4 and SWR1 complexes11.
Nucleosome recognition of the HAT module
The HAT module of NuA4 is organized into two parts: the catalytic core formed by Esa1 and the EPc-I domain of Epl1 and the helical bundle formed by the EPc-II domain of Epl1, Eaf6 and Yng2 (Fig. 3a). This structure largely maintains the structure of the isolated Piccolo subcomplex15, with the β-hairpin of Esa1 and the helical bundle tilting upward to avoid clashing with the bound nucleosome (Extended Data Fig. 9a). Notably, similar to Piccolo15, the HAT module of the holoenzyme binds to the nucleosome disk face with the catalytic pocket of Esa1 oriented towards the H4 N terminus, which shows notable EM density extending from the nucleosome into the catalytic centre (Extended Data Fig. 5b). The CMC-modified H4K16 binds to the acetyl-CoA-binding pocket of Esa1 at a position similar to that observed previously15,21 (Extended Data Fig. 9b,c). The positioning of the catalytic pocket above the H4 tail is not an artefact caused by the H4 modification, because a similar spatial arrangement was observed for the unmodified nucleosome (Extended Data Fig. 2d). Therefore, the modular nature of NuA4 ensures that the holoenzyme retains the position-based mechanism of the catalytic Piccolo subcomplex for H4 recognition15.
a, Two views of the structure of the HAT module bound to the disk face of the nucleosome. The boxed regions are enlarged for analysis in b,d. b, Binding of the DFL to the nucleosome DNA. c, Local EM density of arginine anchors. d, Interaction of arginine anchors of NuA4 with the H2A–H2B acidic patch. e, Relative HAT activity of NuA4 compared with H4 of the nucleosome in the absence and presence of LANA and arginine anchor (Epl1 RA) peptides. Six technical replicates were performed. Con., control. For gel source data, see Supplementary Fig. 1.
The HAT module binds to the nucleosome disk face through two elements of Epl1: the double-function loop (DFL)15 and the arginine anchors (Fig. 3a). The DFL connects the helical bundle with the catalytic core, contains multiple, positively charged residues including Arg321, Lys322, Arg324 and Arg325 and is in close proximity to the minor grove of the DNA at super-helical location 1.5 of the nucleosome (Fig. 3b and Extended Data Fig. 5b). In support of the functional importance of the DFL, these residues are highly conserved across evolution (Extended Data Fig. 7) and the mutations of Lys322 and Arg322 diminish the HAT activity of Piccolo towards the nucleosome in vitro, resulting in sensitivity to DNA damage in vivo15.
A basic region of the EPc-N domain of Epl1 binds to the nucleosome36. The high-resolution map resolves two arginine anchor residues, Arg56 and Arg58, of the EPc-N domain, which interact with the acidic patch of H2A–H2B (Fig. 3c,d). Consistent with the structure, Lys59 of Epl1 was cross-linked to Lys95 of H2A (Extended Data Fig. 1c and Supplementary Table 1). The importance of acidic patch recognition was demonstrated by the addition of WT LANA peptide to compete with NuA4 for binding to the acidic patch37. WT LANA peptide markedly reduced the HAT activity of NuA4 whereas the binding-defective mutant peptide did not perturb it (Fig. 3e). Likewise, addition of the Epl1 RA peptide to HAT assays inhibited NuA4 whereas addition of the R56E or R58E mutant peptides largely abolished inhibition. Together, these data indicate that the DFL and arginine anchors of NuA4 recognize two epitopes on the disk face of the nucleosome and that this recognition directs the catalytic pocket of Esa1 towards the H4 N-terminal tail, providing the structural basis for the H4 preference of NuA4.
In summary, our findings provide an integrated model of nucleosome recognition by NuA4 through the coordination of multiple elements, including the ABS and PBS of the TRA module and the DFL and arginine anchors of the HAT module (Fig. 1c). PBS and ABS interact with the linker DNA, directly and indirectly, and recruit the nucleosome to the edge of the TRA module such that the closely linked HAT module binds to the disk face of the nucleosome to yield selective acetylation of the H4 tails. Moreover, the tethering histone-tail-binding domains, such as the PHD domain of Yng2 and the YEATS domain of Yaf9, are spatially arranged around the nucleosome, allowing NuA4 to read histone modification signals31,38. The relative orientation of the two major modules of NuA4 shows a large degree of plasticity, which may nevertheless provide the adaptability required in response to different transcription activators and chromatin cues. All subunits, except Eaf5 of yeast NuA4, are conserved in the Tip60 complex in human cells, suggesting that our structure provides a good model for human Tip60.
Methods
Preparation and purification of native NuA4 complex
The yeast strain BJ2168 (a gift from the laboratory of S.-C. Cheng) used in this study carries a tandem His-TEV-protein A tag at the C terminus of Eaf1 subunit. The homology recombination cassettes were amplified by PCR from the modified plasmid pFA6a-TAP-KanMX6. PCR products were transformed into Saccharomyces cerevisiae cells by the lithium acetate method and selected on G418-YPD solid medium40. Integration of the cassettes was confirmed by PCR and immunoblot.
To prepare NuA4, yeast cells were cultured in YPD medium at 30 °C to optical density OD600 = 4. Cells were pelleted, washed and resuspended in lysis buffer (50 mM HEPES, 500 mM KCl, 2 mM MgCl2, 20% glycerol and protease inhibitor cocktail, pH 7.6). The cell suspension was dripped into liquid nitrogen and then ground to powder using a chilled pestle. The powder was thawed at room temperature and centrifuged first at 16,000g for 1 h and the supernatant then centrifuged at 150,000g for 1 h. The supernatant was loaded onto a HisTrap column (GE Healthcare), washed by lysis buffer with 20 mM imidazole and eluted with 300 mM imidazole. The elution was incubated with 1 ml of IgG Sepharose 6 Fast Flow resin (GE Healthcare) overnight and then cleaved by TEV protease at 18 °C for 1 h in a buffer containing 20 mM HEPES, 150 mM KCl, 2 mM MgCl2, 0.1% NP40 and 10% glycerol, pH 7.6. The eluted protein was concentrated and applied to a glycerol gradient centrifugation. Fractions containing NuA4 were pooled and concentrated to approximately 300 nM.
To prepare samples for EM analysis, NuA4 was mixed with 600 nM Gal4-VP16. The mixture was dialysed to the binding buffer containing 20 mM HEPES, 50 mM KCl, 2 mM MgCl2 and 10% glycerol, pH 7.6 for 6 h. Next, 40N20 NCP (900 nM) was added with incubation on ice for 1 h. The mixture was cross-linked and purified using the Grafix method41. The glycerol gradient was prepared using a light buffer containing 10% (v/v) glycerol, 20 mM HEPES, 50 mM KCl and 2 mM MgCl2, pH 7.6, and a heavy buffer containing 40% (v/v) glycerol, 20 mM HEPES, 50 mM KCl, 2 mM MgCl2 and 0.2% glutaraldehyde, pH 7.6. The sample was centrifuged at 38,000 r.p.m. for 16 h at 4 °C using a Beckman SW41 Ti rotor. Fractions containing NuA4-Gal4-VP16-NCP were confirmed by 4.5% native polyacrylamide gel electrophoresis (PAGE) and stained by SYBR gold. Peak fractions were pooled and dialysed in the EM buffer containing 20 mM HEPES, 50 mM KCl, 2 mM MgCl2 and 0.005% NP40, pH 7.6 for 4 h to remove glycerol, and concentrated for EM analysis.
Purification of Gal4-VP16 fusion protein
The Gal4 DNA binding domain (residues 1–147) was cloned from yeast genomic DNA; VP16 activator domain (residues 413–490) was amplified by primer walking and the two fragments were integrated to a pET28b vector by seamless cloning. Escherichia coli Rosetta (DE3) was used for protein expression. Briefly, cells were grown at 37 °C to OD600 = 0.6 and induced by the addition of 0.5 mM isopropyl-ß-d-thiogalactopyranoside. Cells were further cultured for 3 h at 37 °C and harvested. Cells were lysed by sonication in lysis buffer (50 mM Tris, 500 mM NaCl, 10% glycerol, 1 mM phenylmethanesulfonyl fluoride and 2 mM benzamidine, pH 8.0) and the soluble extract was loaded onto Ni2+-NTA-agarose resin. The eluted sample was diluted to 50 mM NaCl, loaded onto a resource 15S column and resolved with a linear gradient of 0.05–1.0 M NaCl. Peak fractions were pooled, concentrated and applied to a Superdex-200 column.
Preparation of histone H3 modified with methyl-lysine analogue
To generate the K36-trimethylated histone H3 variant, DMs K36C and C110A were introduced into the H3 sequence by site-directed mutagenesis. The tri-methyl-aminoethyl group was introduced to the thiol group of K36C as previously described42. The final product was confirmed by quadrupole time-of-flight MS.
Preparation of histone H4 modified with CMC
The chemically synthesized H4 peptide (residues 11–22) modified with CMC at K16 was purchased from Scilight-Peptide. H4 (residues R23C–102) was amplified with PCR using the WT template and integrated to a pET3c vector by seamless cloning. Expression and purification of the truncated histone were performed similarly as for WT, with some modifications. Protein was expressed in DE3 cells. The histone was dialysed once against 5 mM dithiothreitol (DTT) solution and at least twice against 0.05% TFA solution after the ion exchange purification step. Removal of the N-terminal methionine was confirmed by matrix-assisted laser desorption/ionization–ToF MS.
The final product, histone H4 modified with CMC, was assembled by native chemical ligation as previously described43,44 and confirmed by quadrupole time-of-flight MS. First, H4 (residues 11–22) K16CMC (1.0 equiv.) was dissolved in aqueous phosphate (0.2 M) buffer containing 6 M guanidinium chloride. A 1 M NaNO2 aqueous solution (5 equiv.) was added dropwise and stirred for 20 min at −12 °C to produce a peptide acyl azide. MPAA (4-mercaptophenylacetic acid, 50 equiv.) was added and the pH value adjusted to 6.3. The reaction was then removed to room temperature for 5 min. H4 (residues R23C–102) (1.2 equiv.) was added and the reaction buffer adjusted to pH 6.5 with 2 M NaOH to initiate native chemical ligation. The reaction mixture was stirred at 18 °C overnight and reduced by tris(2-carboxyethyl)phosphine (50 mM, pH 7.0). The final reaction mixture was purified using a resource 15S column. Peak fractions were pooled, lyophilized and stored at −80 °C.
Preparation of nucleosome
Wild-type Xenopus laevis histones H2A, H2B, H3 and H4 were expressed and purified as previously described45. For octamer preparation, histones H2A, H2B, H3 (or H3K36me3) and H4 (or H4K16-CMC) were mixed at a molar ratio of 1:1:1:1 and dialysed three times against the refolding buffer (20 mM Tris, 2 M NaCl and 10 mM DTT, pH 7.5). The sample was concentrated and applied to a Superdex 200 size-exclusion column. Peak fractions were pooled, concentrated and frozen in liquid nitrogen.
The 40N20 (UAS-20N20) DNA fragment (TCCGGAGGACTGTCCTCCGGGGACCCTATACGCGGCCGCCATCGAGAATCCCGGTGCCGAGGCCGCTCAATTGGTCGTAGACAGCTCTAGCACCGCTTAAACGCACGTACGCGCTGTCCCCCGCGTTTTAACCGCCAAGGGGATTACTCCCTAGTCTCCAGGCACGTGTCAGATATATACATCCGATAGCTTGTCGAGAAGTACTAG), with the Gal4 DNA binding sequence in bold and the 601 sequence underlined, was prepared by PCR using the Widom 601 sequence as template. The 5′ variant linker DNA sequences used as follows:
UAS-0N20 (TCCGGAGGACTGTCCTCCGG)
UAS-10N20 (TCCGGAGGACTGTCCTCCGGTCGTGCCTGT)
con-20N20 (GAGTTCATCCCTTATGTGATGGACCCTATACGCGGCCGCC)
The PCR product was purified by anion-exchange chromatography and concentrated. Nucleosome reconstitution was performed as previously described45.
Electron microscopy
The NuA4 complex was first studied by negative-stain EM. Briefly, 4 μl of sample was loaded onto a glow-discharged copper grid coated with thin carbon film. The sample was incubated for 1 min and grids were blotted and immersed in uranyl acetate (2%, w/v) for negative staining.
For cryo-EM analysis, 4 μl of sample was applied to Quantifoil R2/1 Au 200 mesh grids, which were glow-discharged for 25 s and subsequently incubated for 60 s before blotting and vitrification by plunging into liquid ethane with a Vitrobot (FEI) operated at 8 °C and 100% humidity.
Negative-stain data collection and image processing
In negative-stain data collection, NuA4–NCP samples were observed using a 200 kV Tecnai F20 microscope (FEI) equipped with a Gatan Ultrascan 4000 camera at a magnification of ×62,000, corresponding to a pixel size of 1.35 Å on the images. Defocus ranging from −1.0 to −2.5 μm and a total dose of about 40 e–/Å2 were used; defocus parameters were estimated by CTFFIND3 (ref. 46). A total of 10,354 particles were picked using the sub-routine e2boxer.py in the EMAN2 suit47. Two-dimensional (2D) classification was carried out in Relion 1.4 to remove 'bad' particles48. A sphere generated by SPIDER was used as the initial model for the first round of three-dimensional (3D) classification49. After several rounds of 3D classification, 8,739 particles were selected and subjected to final 3D reconstruction.
Cryo-EM data collection and image processing
The dataset of NuA4-Gal4-VP16-NCP (H4 modified) was acquired on a Thermo Fisher Scientific Krios G3i operated at 300 kV and equipped with a K3 direct electron detector and GIF Quantum energy filter (Gatan). Video stacks were automatically recorded using AutoEMation2 in super-resolution mode50 at a magnification of ×81,000 for a final pixel size of 0.54125 Å per pixel in super-resolution mode, with defocus values ranging from −1.3 to −1.8 μm. The total electron dose was 50 e–/Å2 fractionated in 32 frames (exposure time, 2.56 s).
A total of 11,000 dose-fractionated image stacks were aligned using MotionCor2 with twofold binning51, resulting in a size of 1.0825 Å per pixel. Contrast transfer function (CTF) parameters were estimated using CTFFIND4 (ref. 52). Particle picking and 2D/3D classification were carried out in Relion 3.0 (ref. 48). Particles were extracted using a box size of 376 × 376 pixels, and normalized. Reference-free 2D classification and global 3D classification with a NuA4–NCP map, derived from the NuA4-Gal4-VP16-40N20 dataset and low passed to 40 Å as the initial model, were performed to screen for 'good' particles in the dataset. All good particles with fourfold binning were used for 3D refinement with local search, resulting in a map at 8.8 Å. Furthermore, these particles were re-extracted without binning and used for global 3D refinement, resulting in a map at 4.0 Å. To improve mapping of the TRA module, focused 3D classification without image alignment was performed using a soft mask. The class that showed optimal density for the TRA module was subjected to another round of focused 3D refinement, resulting in a resolution of 3.0 Å. Focused refinement further improved resolution to 2.7 Å for the ARP submodule and around the Head region of Tra1. To improve the bottom part of Tra1, a soft mask was applied to this region and subsequent 3D classification without image alignment led to an intact TRA module; 3D refinement with a soft mask on the TRA module led to a map at 3.1 Å.
For reconstruction of the NCP-HAT region, the map of the NCP-HAT module low passed to 40 Å was used as reference to obtain an improved initial reference. All classes containing intact NCP-HAT density were re-extracted without binning and used for global refinement, resulting in a map at 7.7 Å resolution. Focused 3D classification without image alignment, and focused refinement, led to resolution at 3.4 Å. For the NCP-arginine anchor region, all classes showing clear features were combined. Focused 3D classification without image alignment and following 3D refinement led to a resolution at 2.8 Å. For reconstruction of the HAT-NCP module with long linker DNA, all classes containing intact HAT-NCP density were re-extracted with twofold binning and used for multiple rounds of 3D classification and 3D refinement with signal subtraction, which led to resolution at 6.7 Å.
The different modes of relative movement of the TRA and HAT modules were carried out by multi-body refinement53, showing rotational motion of the TRA module bound to the nucleosome. The first three eigenvectors shown in Supplementary Video 2 explain 40.4% of variance in the data, with 15.2% for component 1, 12.7% for component 2 and 12.5% for component 3. These represent the principal motions in particles.
The datasets of NuA4–NCP and NuA4-Gal4-VP16-NCP were acquired on an FEI Titan Krios operating at 300 kV with nominal magnification of ×22,500. Images were recorded on a Gatan K2 Summit detector (Gatan company) in super-resolution mode, and binned to a pixel size of 1.306 Å. UCSFImage4 was used for data collection, with defocus values ranging from −1.4 to −1.8 μm54. Each micrograph was dose-fractionated to 32 frames with 0.25 s exposure time in each frame and a total electron dose of 50 e−/Å2. Motion correction, CTF estimation, particle picking and reference-free 2D classification were performed as described above.
For the NuA4–NCP sample, good particles were processed with global 3D classification using RELION 3.0 and a negative-stain reconstruction of NuA4–NCP as the first reference to obtain an improved initial reference. After several rounds of 3D classification and refinement, only a low-resolution map of the intact complex was obtained (Extended Data Fig. 2b).
For the NuA4-Gal4-VP16-NCP sample, all good particles collected from 2D classification were used for global 3D classification. After several rounds of 3D classification, the class containing intact NuA4–NCP density was used for 3D refinement and led to a map with resolution at 8.8 Å. All classes containing the intact TRA module were combined and subjected to 3D refinement, leading to a map with resolution at 3.6 Å (Extended Data Fig. 2d), but the nucleosome-bound HAT did not result in a high-resolution map.
Model building
The model of NuA4 was built by fitting the available structures of Tra1, the catalytic Piccolo subcomplex, the Act1-Arp4-HSAEaf1 subcomplex and Esa1-bound CMC15,17,21,55. The remainder of the molecules, including Eaf2, Eaf1 and the EPc-B domain of Epl1, were manually built based on high-resolution maps in Coot. The structures were refined using Phenix with secondary structure constrains56. The asymmetric cryo-EM density of the linker DNA, which contains DNA of over 25 bp at one side, unambiguously orientates the 40N20 nucleosome into the map. The orientation of the nucleosome is also supported by the high-resolution map of the nucleosome (at a resolution of 2.8 Å), the four central base pairs of which are shown in Extended Data Fig. 5b.
Cross-linking and MS
Sample preparation was the same as for cryo-EM analysis. The mixture was dialysed to binding buffer for 4 h and cross-linked by 3 mM BS3 for 30 min at room temperature. The reaction was quenched with 50 mM Tris, pH 7.5. The cross-linked sample was applied to a 10–40% glycerol gradient as described above. Protein-containing fractions were pooled for MS57. The data were analysed with pLink2 (v.2.3.9) and xiVIEW58 and visualized with chimeraX59.
Yeast genetics
To generate the Eaf2-KE (K311E/K314E/K316E) mutant strain, the C terminus of Eaf2 (residues 311–476) was first replaced by URA3 and yeast cells selected using SC/-Ura medium. URA3 was then replaced by the C terminus of Eaf2 with the desired mutations, and cells were selected using 5-FOA medium (YPD + 5-FOA). The Epl1-KE (K491E/K496E) mutant strain was generated as described above, except that the C terminus of Epl1 (residues 491–832) was replaced by URA3. The DM strain harbouring mutations Eaf2 and Epl1 was generated by introducing the C terminus of Eaf2 mutations into the Epl1-KE mutant strain, as described above. Yeast strain BY4741 (MATα leu2 ura3 his3 met15) was used and transformed with the lithium acetate method. All mutants were confirmed by PCR and DNA sequencing.
For yeast spot sensitivity assays, yeast strains were grown overnight at 30 °C in YPD, diluted to OD600 = 0.1 and grown for a further 6 h in YPD. Fivefold serial dilutions of cultures were then spotted onto different media and grown at 30 °C for 3 days. To examine the utilization of various carbon sources, cells were grown on plates containing 2% glucose (YPD), 2% potassium acetate and 3% glycerol.
HAT assays
In the HAT assay with competitor peptide, reactions were performed in MES buffer (25 mM MES, 180 mM KCl, 2 mM MgCl2, 0.1 mg ml–1 bovine serum albumin (BSA), 50 μM acetyl-CoA, 0.5 μM NCP and 500 μM each competitor peptide, pH 6.5). Peptide sequences used in this assay were as follows: LANA (MAPPGMRLRSGRSTGAPLTRGS), R9E (MAPPGMRLESGRSTGAPLTRGS), Epl1 RA (ASAGSSNSRFRHRKISVKQHL), R56E (ASAGSSNSRFEHRKISVKQHL), R58E (ASAGSSNSRFRHEKISVKQHL). The reactions were initiated by the addition of 10 nM NuA4. After incubation on ice for 25 min, reactions were terminated by the addition of SDS loading buffer and heated at 100 °C for 1 min. Reaction mixtures were resolved with gradient SDS–PAGE, followed by immunoblotting analysis using the primary antibody against pan-acetylated lysine (Cell Signaling Technology, no. 9681S, 1:1,000 dilution) and secondary antibody horseradish peroxidase goat anti-mouse IgG (H+L, ABclonal, no. AS003, 1:5,000 dilution). After exposure, signals were quantified by ImageJ.
In the HAT assay with Gal4-VP16, NuA4 and Gal4-VP16 were premixed at a molar ratio of 1:1. Reactions were performed in buffer (20 mM HEPES, 50 mM KCl, 2 mM MgCl2, 0.1 mg ml–1 BSA, 50 μM acetyl-CoA, 0.1 μM NCP and 1 μg of salmon sperm DNA, pH 7.6). Reactions were initiated by the addition of 0.07 μM NuA4-Gal4-VP16 mixture or NuA4 alone at room temperature for 40 min. Subsequent processes were carried out as described above.
In the HAT assay with WT and DM NuA4, reactions were performed in a buffer containing 20 mM HEPES, 50 mM KCl, 2 mM MgCl2, 0.1 mg ml–1 BSA, 0.5 μM acetyl-CoA and 0.5 μM NCP, pH 7.6. Reactions were initiated by the addition of 10 nM WT or DM NuA4 and kept on ice for 10 min. Subsequent processes were performed as described above.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
References
Luger, K., Mader, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389, 251–260 (1997).
Kayne, P. S. et al. Extremely conserved histone H4 N terminus is dispensable for growth but essential for repressing the silent mating loci in yeast. Cell 55, 27–39 (1988).
Dorigo, B., Schalch, T., Bystricky, K. & Richmond, T. J. Chromatin fiber folding: requirement for the histone H4 N-terminal tail. J. Mol. Biol. 327, 85–96 (2003).
Shogren-Knaak, M. et al. Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 311, 844–847 (2006).
Allard, S. et al. NuA4, an essential transcription adaptor/histone H4 acetyltransferase complex containing Esa1p and the ATM-related cofactor Tra1p. EMBO J. 18, 5108–5119 (1999).
Bird, A. W. et al. Acetylation of histone H4 by Esa1 is required for DNA double-strand break repair. Nature 419, 411–415 (2002).
Babiarz, J. E., Halley, J. E. & Rine, J. Telomeric heterochromatin boundaries require NuA4-dependent acetylation of histone variant H2A.Z in Saccharomyces cerevisiae. Genes Dev. 20, 700–710 (2006).
Durant, M. & Pugh, B. F. NuA4-directed chromatin transactions throughout the Saccharomyces cerevisiae genome. Mol. Cell. Biol. 27, 5327–5335 (2007).
Bruzzone, M. J., Grunberg, S., Kubik, S., Zentner, G. E. & Shore, D. Distinct patterns of histone acetyltransferase and mediator deployment at yeast protein-coding genes. Genes Dev. 32, 1252–1265 (2018).
Dhar, S., Gursoy-Yuzugullu, O., Parasuram, R. & Price, B. D. The tale of a tail: histone H4 acetylation and the repair of DNA breaks. Philos. Trans. R. Soc. Lond. B Biol. Sci. 372, 20160284 (2017).
Doyon, Y. & Cote, J. The highly conserved and multifunctional NuA4 HAT complex. Curr. Opin. Genet. Dev. 14, 147–154 (2004).
Lee, K. K. & Workman, J. L. Histone acetyltransferase complexes: one size doesn't fit all. Nat. Rev. Mol. Cell. Biol. 8, 284–295 (2007).
Cheung, A. C. M. & Diaz-Santin, L. M. Share and share alike: the role of Tra1 from the SAGA and NuA4 coactivator complexes. Transcription 10, 37–43 (2019).
Boudreault, A. A. et al. Yeast enhancer of polycomb defines global Esa1-dependent acetylation of chromatin. Genes Dev. 17, 1415–1428 (2003).
Xu, P. et al. The NuA4 core complex acetylates nucleosomal histone H4 through a double recognition mechanism. Mol. Cell 63, 965–975 (2016).
Chittuluru, J. R. et al. Structure and nucleosome interaction of the yeast NuA4 and Piccolo-NuA4 histone acetyltransferase complexes. Nat. Struct. Mol. Biol. 18, 1196–1203 (2011).
Wang, X., Ahmad, S., Zhang, Z., Cote, J. & Cai, G. Architecture of the Saccharomyces cerevisiae NuA4/TIP60 complex. Nat. Commun. 9, 1147 (2018).
Setiaputra, D. et al. Molecular architecture of the essential yeast histone acetyltransferase complex NuA4 redefines its multimodularity. Mol. Cell. Biol. 38, e00570-17 (2018).
Wang, H. et al. Structure of the transcription coactivator SAGA. Nature 577, 717–720 (2020).
Wu, J., Xie, N., Wu, Z., Zhang, Y. & Zheng, Y. G. Bisubstrate inhibitors of the MYST HATs Esa1 and Tip60. Bioorg. Med. Chem. 17, 1381–1386 (2009).
Yuan, H. et al. MYST protein acetyltransferase activity requires active site lysine autoacetylation. EMBO J. 31, 58–70 (2012).
Sun, B. et al. Molecular basis of the interaction of Saccharomyces cerevisiae Eaf3 chromo domain with methylated H3K36. J. Biol. Chem. 283, 36504–36512 (2008).
Xu, C., Cui, G., Botuyan, M. V. & Mer, G. Structural basis for the recognition of methylated histone H3K36 by the Eaf3 subunit of histone deacetylase complex Rpd3S. Structure 16, 1740–1750 (2008).
Rossetto, D. et al. Eaf5/7/3 form a functionally independent NuA4 submodule linked to RNA polymerase II-coupled nucleosome recycling. EMBO J. 33, 1397–1415 (2014).
Lin, L., Chamberlain, L., Zhu, L. J. & Green, M. R. Analysis of Gal4-directed transcription activation using Tra1 mutants selectively defective for interaction with Gal4. Proc. Natl Acad. Sci. USA 109, 1997–2002 (2012).
Knutson, B. A. & Hahn, S. Domains of Tra1 important for activator recruitment and transcription coactivator functions of SAGA and NuA4 complexes. Mol. Cell. Biol. 31, 818–831 (2011).
Brown, C. E. et al. Recruitment of HAT complexes by direct activator interactions with the ATM-related Tra1 subunit. Science 292, 2333–2337 (2001).
Elias-Villalobos, A., Toullec, D., Faux, C., Seveno, M. & Helmlinger, D. Chaperone-mediated ordered assembly of the SAGA and NuA4 transcription co-activator complexes in yeast. Nat. Commun. 10, 5237 (2019).
Auger, A. et al. Eaf1 is the platform for NuA4 molecular assembly that evolutionarily links chromatin acetylation to ATP-dependent exchange of histone H2A variants. Mol. Cell. Biol. 28, 2257–2270 (2008).
Bittner, C. B., Zeisig, D. T., Zeisig, B. B. & Slany, R. K. Direct physical and functional interaction of the NuA4 complex components Yaf9p and Swc4p. Eukaryot. Cell 3, 976–983 (2004).
Klein, B. J. et al. Yaf9 subunit of the NuA4 and SWR1 complexes targets histone H3K27ac through its YEATS domain. Nucleic Acids Res. 46, 421–430 (2018).
Mitchell, L. et al. Functional dissection of the NuA4 histone acetyltransferase reveals its role as a genetic hub and that Eaf1 is essential for complex integrity. Mol. Cell. Biol. 28, 2244–2256 (2008).
Downs, J. A. et al. Binding of chromatin-modifying activities to phosphorylated histone H2A at DNA damage sites. Mol. Cell 16, 979–990 (2004).
Turegun, B., Kast, D. J. & Dominguez, R. Subunit Rtt102 controls the conformation of the Arp7/9 heterodimer and its interactions with nucleotide and the catalytic subunit of SWI/SNF remodelers. J. Biol. Chem. 288, 35758–35768 (2013).
Schubert, H. L. et al. Structure of an actin-related subcomplex of the SWI/SNF chromatin remodeler. Proc. Natl Acad. Sci. USA 110, 3345–3350 (2013).
Huang, J. & Tan, S. Piccolo NuA4-catalyzed acetylation of nucleosomal histones: critical roles of an Esa1 Tudor/chromo barrel loop and an Epl1 enhancer of polycomb A (EPcA) basic region. Mol. Cell. Biol. 33, 159–169 (2013).
Barbera, A. J. et al. The nucleosomal surface as a docking station for Kaposi's sarcoma herpesvirus LANA. Science 311, 856–861 (2006).
Steunou, A. L. et al. Combined action of histone reader modules regulates NuA4 local acetyltransferase function but not its recruitment on the genome. Mol. Cell. Biol. 36, 2768–2781 (2016).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–350 (2016).
Gietz, R. D. & Schiestl, R. H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 31–34 (2007).
Stark, H. GraFix: stabilization of fragile macromolecular complexes for single particle cryo-EM. Methods Enzymol. 481, 109–126 (2010).
Simon, M. D. et al. The site-specific installation of methyl-lysine analogs into recombinant histones. Cell 128, 1003–1012 (2007).
Dawson, P. E., Muir, T. W., Clark-Lewis, I. & Kent, S. B. Synthesis of proteins by native chemical ligation. Science 266, 776–779 (1994).
Li, Y. T. et al. A semisynthetic Atg3 reveals that acetylation promotes Atg3 membrane binding and Atg8 lipidation. Nat. Commun. 8, 14846 (2017).
Dyer, P. N. et al. Reconstitution of nucleosome core particles from recombinant histones and DNA. Methods Enzymol. 375, 23–44 (2004).
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Ludtke, S. J. 3-D structures of macromolecules using single-particle analysis in EMAN. Methods Mol. Biol. 673, 157–173 (2010).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Nakane, T., Kimanius, D., Lindahl, E. & Scheres, S. H. Characterisation of molecular motions in cryo-EM single-particle data by multi-body refinement in RELION. eLife 7, e36861 (2018).
Li, X., Zheng, S., Agard, D. A. & Cheng, Y. Asynchronous data acquisition and on-the-fly analysis of dose fractionated cryoEM images by UCSFImage. J. Struct. Biol. 192, 174–178 (2015).
Diaz-Santin, L. M., Lukoyanova, N., Aciyan, E. & Cheung, A. C. Cryo-EM structure of the SAGA and NuA4 coactivator subunit Tra1 at 3.7 angstrom resolution. eLife 6, e28384 (2017).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Ye, Y. et al. Structure of the RSC complex bound to the nucleosome. Science 366, 838–843 (2019).
Graham, M., Combe, C. W., Kolbowski, L. & Rappsilber, J. xiView: a common platform for the downstream analysis of crosslinking mass spectrometry data. Preprint at bioRxiv https://doi.org/10.1101/561829 (2019).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Acknowledgements
We thank S.-C. Cheng (Institute of Molecular Biology, Academia Sinica) for providing yeast strain BJ2168 and the Tsinghua University Branch of the China National Center for Protein Sciences (Beijing) for use of the cryo-EM facility. This work was supported by the National Key Research and Development Program (no. 2019YFA0508902 to Z.C), the National Natural Science Foundation of China (nos. 32130016 and 31825016 to Z.C), Beijing Frontier Research Center for Biological Structure, Beijing Advanced Innovation Center for Structural Biology and Tsinghua-Peking Joint Center for Life Sciences.
Author information
Authors and Affiliations
Contributions
K.Q. prepared samples and performed biochemical analysis. K.Q. and K.C. built the atomic model. K.C., K.Q., H.W. and X.L. performed EM analysis. Z.C. wrote the manuscript with help from all authors. Z.C. directed and supervised all of the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Biochemistry of NuA4 and the crosslinking data detected by CL-MS.
(a) Preparation of endogenous NuA4 complex from S.cerevisiae. SDS-PAGE gels were showed with Coomassie stain. More sample of the WT complex was loaded to show the weak bands on the right panel. Over 10 batches of the samples were prepared, and the representative two are shown here. (b) Relative HAT activity of NuA4 towards the nucleosomes in the absence and presence of Gal4-VP16. The Gal4 binding site (UAS) was placed at different positions of the nucleosomes (UAS-0N20, UAS-10N20 and UAS-20N20), and the unrelated sequence was used as a control (con-20N20). Acetylation of H4 was detected by Western blotting. A presentative gel was shown on the top, and quantification at the bottom. The activities were normalized to the ones in the absence of Gal4-VP16. Two technical replicates were performed. (c) Overview of the cross-linking data. Circular plot of high confidence lysine-lysine inter-subunit crosslinks obtained by mass spectrometry for the NuA4-NCP complex. (d) Validated cross-links mapped onto the NuA4-NCP structure. Blue lines, the intra-chain cross-links with cross-linked sites within the 30 Å distance permitted by BS3; green lines, the inter-chain cross-links, whereas red lines depict cross-link over more than 30 Å. Bottom graph displays the distribution of cross-link distances. A vertical line indicates a cutoff at 30 Å, the distance considered reasonable for BS3 crosslinks. (e) Relative HAT activity of an independent batch of WT and DM mutant NuA4 different from that of Fig. 2f. Four technical replicates were performed.
Extended Data Fig. 2 Cryo-EM analysis of the NuA4-NCP complex in the absence and presence of Gal4-VP16.
(a) Representative negative stain images (top panel), cryo-EM images (middle panel), and 2D classification (bottom panel) of NuA4+NCP. (b) Flowcharts of the cryo-EM data processing for the dataset of NuA4+NCP. (c) Representative negative stain images (top panel), cryo-EM images (middle panel), and 2D classification (bottom panel) of NuA4+NCP+Gal4-VP16. (d) Flowcharts of the cryo-EM data processing for the dataset of NuA4+NCP+Gal4-VP16.
Extended Data Fig. 3 Chemistry and Mass-spectrometry analyses of the modified histones.
(a) Schematic diagram of the chemical semi-synthesis of histone H4 with CMC modification at Lys16. (b) Mass-spectrometry identification of the CMC-modified H4 (the expected mass is 11092.57 Da). (c) Mass-spectrometry identification of the H3KC36me3-modified H3 (the expected mass is 15299.76 Da).
Extended Data Fig. 4 Cryo-EM analysis of the NuA4-NCP complex in the presence of Gal4-VP16 and histone-modified nucleosome.
(a) Representative negative stain images (left panel), cryo-EM images (middle panel), and 2D classification (right panel) of NuA4+Gal4-VP16 bound to the H4- and H3-modified nucleosome. (b) Flowcharts of the cryo-EM data processing for the dataset of NuA4+Gal4-VP16 bound to the H4- and H3-modified nucleosome. (c-d) Angular distributions of cryo-EM particles in the final round of refinement of the masked dataset and the estimation of the local resolutions of the TRA module (c) and the nucleosome bound HAT module (d). (e) Gold standard Fourier shell correlation (FSC) curves, showing the overall nominal resolutions of 6.7 Å, 3.4 Å, 3.1 Å, 2.8 Å, 2.7 Å and 2.7 Å for the NCP-HAT with longer linker DNA, the nucleosome bound HAT module, TRA module, the NCP bound with arginine anchors (RAs), ARP submodule, head of Tra1 bound with Eaf1/Eaf2/Epl1, respectively.
Extended Data Fig. 5 Local density maps of the NuA4-nucleosome complex.
(a) Local cryo-EM maps of the TRA module. PBL, polybasic loop. (b) Local cryo-EM maps of the HAT module bound to the nucleosome. EM densities of four consecutive base pairs at the central position of the nucleosome are showed on the left.
Extended Data Fig. 6 Multiple sequence alignments of Eaf2-like proteins.
The conserved residues are indicated by “*”. The secondary structural assignments are based on the structure of Eaf2.
Extended Data Fig. 7 Multiple sequence alignments of Epl1-like proteins.
The conserved residues are indicated by “*”. The secondary structural assignments are based on the structure of Epl1.
Extended Data Fig. 8 Multiple sequence alignments of Eaf1-like proteins.
The conserved residues are indicated by “*”. The secondary structural assignments are based on the structure of Eaf1.
Extended Data Fig. 9 Additional structural analysis of the HAT module.
(a) Comparison of the structures of the HAT module bound to the nucleosome (color coded) and the nucleosome-free Piccolo subcomplex (colored grey, PBD code 5J9W)15. The structure of the Esa1 subunit is aligned. (b) Structural alignment of CMC-binding pocket in NuA4 (color coded) and in Esa1 (colored grey, PBD code 3TO6)21. (c) Structural comparison of CMC-binding in NuA4 (color coded) and AcCoA in Piccolo (colored grey, PBD code 5J9W)15.
Supplementary information
Supplementary Fig. 1
Uncropped gels.
Supplementary Table 1
Data of CL–MS.
Supplementary Video 1
Cryo-EM density and model fitting of NuA4 bound to the nucleosome.
Supplementary Video 2
Multi-body refinement analysis of the NuA4–nucleosome complex.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qu, K., Chen, K., Wang, H. et al. Structure of the NuA4 acetyltransferase complex bound to the nucleosome. Nature 610, 569–574 (2022). https://doi.org/10.1038/s41586-022-05303-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05303-x
- Springer Nature Limited
This article is cited by
-
Structure of the human TIP60 complex
Nature Communications (2024)
-
The NuA4 histone acetyltransferase: variations on a theme of SAGA
Nature Structural & Molecular Biology (2023)
-
Paralog-specific recognition
Nature Chemical Biology (2023)
-
MOF-mediated histone H4 Lysine 16 acetylation governs mitochondrial and ciliary functions by controlling gene promoters
Nature Communications (2023)