Abstract
Plant traits determine how individual plants cope with heterogeneous environments. Despite large variability in individual traits, trait coordination and trade-offs1,2 result in some trait combinations being much more widespread than others, as revealed in the global spectrum of plant form and function (GSPFF3) and the root economics space (RES4) for aboveground and fine-root traits, respectively. Here we combine the traits that define both functional spaces. Our analysis confirms the major trends of the GSPFF and shows that the RES captures additional information. The four dimensions needed to explain the non-redundant information in the dataset can be summarized in an aboveground and a fine-root plane, corresponding to the GSPFF and the RES, respectively. Both planes display high levels of species aggregation, but the differentiation among growth forms, families and biomes is lower on the fine-root plane, which does not include any size-related trait, than on the aboveground plane. As a result, many species with similar fine-root syndromes display contrasting aboveground traits. This highlights the importance of including belowground organs to the GSPFF when exploring the interplay between different natural selection pressures and whole-plant trait integration.
Similar content being viewed by others
Main
The GSPFF3 showed that most of the variability in six key aboveground traits that are essential for plant growth, survival and reproduction can be summarized by one plane defined by two axes: one reflecting the size of the plant and its aboveground organs, and the other representing the leaf economics spectrum5. Within this space, species are aggregated in two functional hotspots: within the trait space of herbaceous plants and trees, respectively3. Owing to the recent development of open-access fine-root trait datasets6,7,8, it is now possible to empirically ask how these traits relate to the GSPFF.
Fine roots (either considered as less than 2 mm in diameter or as roots of orders 1–3)4 are responsible for acquiring essential soil resources, mediating biogeochemical cycling and considerably contributing to stabilizing organic matter in soils9,10. Two alternative hypotheses have been formulated regarding the organization of species with respect to their fine-root traits. First, since fine roots can be considered the belowground equivalent of leaves because their main role is acquiring resources11, the plant economics spectrum hypothesis postulates that plant species are organized along a single dimension of the root trait spectrum, aligned with the leaf spectrum12,13,14. Accordingly, there should be a high degree of covariation between leaf and root traits. However, recent analyses have described a two-dimensional RES4 with limited correspondence with the leaf economics spectrum15,16,17. One of the RES dimensions, defined by root nitrogen and root tissue density, reflects a root tissue conservation gradient that represents a trade-off between fast and slow return on investment4, and can be considered analogous to the leaf economics spectrum. The other dimension represents a gradient of plant–fungal interactions in roots, defined by a trade-off between root diameter and specific root length4,18, with higher degrees of interaction expected for thicker roots. This dimension seems to have no equivalent aboveground.
We performed a joint correlative analysis of the coordination and trade-offs of the aboveground and fine-root traits of vascular plants using a global dataset that combines the data that formed the main basis of the GSPFF3, with fine-root trait data used to describe the RES4. We obtained information for aboveground traits for 39,260 plant species from the TRY database6. Specifically, this included a set of fundamental functional traits: plant height, leaf area, specific leaf area, leaf nitrogen concentration, specific stem density and seed mass, which are associated with different key aspects of plant ecology3 (see Methods; Extended Data Table 1). Fine-root trait information was obtained from the Global Root Trait (GRooT) database8 for 2,050 species. We considered four fine-root traits that have been previously used to characterize the RES4,18,19: root diameter specific root length, root nitrogen concentration and root tissue density (Extended Data Table 1; Extended Data Fig. 1). Both databases had 1,719 species in common, of which 301 (belonging to 73 families; Extended Data Fig. 2) had complete empirical information for all selected traits.
To assess the main trends of trait variation, we characterized the trait space using a principal component analysis (PCA) based on all the aboveground and fine-root traits of the subset of species with complete empirical information (301 species). The first four principal components of the PCA accounted for 76% of the total variation. We applied varimax rotation to these axes to obtain a four-dimensional space where traits are best related to axes (Extended Data Tables 2, 3). The first two components (C1 and C2) were mostly related to aboveground traits (Fig. 1a), and the other two components (C3 and C4) were related to fine-root traits (Fig. 1b). The C1–C2 and C3–C4 planes, therefore, corresponded closely to the two-dimensional spaces that have been reported in the GSPFF3 for aboveground traits and the RES4 for fine-root traits (see Supplementary Note 1 and Extended Data Fig. 3 for a detailed description of the patterns). Hereafter, we refer to the C1–C2 plane as the ‘aboveground plane’ and to the C3–C4 plane as the ‘fine-root plane’. Existing evidence for a whole-plant economic spectrum has produced mixed results, with some studies describing a tight correspondence between leaf and fine-root traits12,14 and others indicating loose, complex or no relationship17,19,20,21. If there was a strong correspondence between aboveground and fine-root traits, the number of dimensions necessary to explain variation after combining traits should be smaller than the sum of the individual dimensionalities of the GSPFF and the RES.
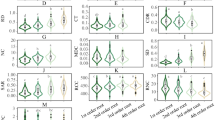
Probabilistic species distributions in the space defined by a PCA followed by varimax rotation based on aboveground and fine-root traits of species with complete trait information (n = 301). Compared with the unrotated PCA, the varimax rotation retrieves the fundamental structure of trait variation in the dataset in a more interpretable and consistent way (less dependent on the subset of species considered; see Methods). a, First and second components (and proportion of variance explained). b, Third and fourth components. The colour gradient depicts different densities of species in the space (the red areas are more densely populated). The arrow length is proportional to the loadings of the considered traits (see Extended Data Table 2). The aboveground traits (represented by green arrows) are specific leaf area (sla), leaf nitrogen concentration (ln), leaf area (la), seed mass (sm), plant height (ph) and specific stem density (ssd). The fine-root traits (represented by brown arrows) are specific root length (SRL), root diameter (D), root tissue density (RTD) and root nitrogen concentration (N). The thick contour lines indicate the 0.5 and 0.99 quantiles, and the thinner lines indicate 0.6, 0.7, 0.8 and 0.9 quantiles.
The fact that four dimensions are required to adequately capture non-redundant trait variation, as found in the present study, suggests that aboveground and fine-root trait syndromes involve traits that are not analogous to each other (Extended Data Fig. 4). As a consequence, combining aboveground and fine-root traits can reveal patterns among species that are not immediately evident when the GSPFF or the RES are examined separately. For example, eastern hemlock (Tsuga canadensis) and black beech (Nothofagus solandri) are two tree species that are very close on the aboveground plane but are far apart on the fine-root plane, because black beech has higher specific root length and root tissue density values. These traits have been associated with the ability of the Nothofagaceae family to dominate in unfertile soils in New Zealand15. The opposite pattern is very common; for example, while Scots pine (Pinus sylvestris), common sunflower (Helianthus annuus) and common persimmon (Diospyros virginiana) are strongly separated from each other on the aboveground plane, these three species occupy similar areas on the fine-root plane (for an examination of species position in the trait space, see https://globaltrait.shinyapps.io/GlobalTraits; Supplementary Application 1).
There are vastly different levels of aggregation of species in different areas of the two planes. Using the same 301 species, we built a null model considering multivariate normal distributions with the same means and covariance matrix as the observed data22, and compared the amount of space occupied by different quantiles of the distributions. We found that the distribution of species on both the aboveground and the fine-root planes is more clumped than expected, with any given observed quantile occupying on average 29% and 14% less trait space than the same quantiles of the aboveground and fine-root null models, respectively (Extended Data Fig. 5). However, the aggregation of species aboveground is mainly around two functional hotspots, whereas there is only one hotspot on the fine-root plane. The two aboveground hotspots, which are far from the centre of the spectrum (leading to higher functional divergence than in the null model; observed functional divergence = 0.58 versus null model functional divergence = 0.38, P = 0.002, n = 500), are associated with a bimodal distribution in size-related traits (C1, described by plant height, seed mass and specific stem density), corresponding to herbaceous species and angiosperm trees3. By contrast, on the fine-root plane, species are concentrated around a central hotspot (leading to low functional divergence: observed functional divergence = 0.34 versus null model functional divergence = 0.38, P = 0.016, n = 500), whereas the surrounding areas are sparsely occupied. Overall, these results are consistent with the idea that a small number of trait syndromes are extremely prevalent, whereas many others, while viable, are rare3,22. This difference may be related to the lack of sufficient data on root traits that scale with plant size (for example, rooting depth or root system size)23 that might contribute to variation along a size-dependent belowground dimension analogous to C1.
Next, we explored how these patterns of occupation of the functional space differed between herbaceous and woody species, and also among families and biomes. For this, we used a larger dataset, including 1,218 species (from 127 families) with empirical information for at least three aboveground and two fine-root traits (Extended Data Table 1, Extended Data Fig. 6). We used a phylogenetically informed imputation procedure22 to complete trait information for those species (Supplementary Methods 1) and estimated the functional space following the same procedure as with the complete dataset.
We examined the patterns of functional differentiation among groups of species, finding that fine-root traits were more similar than aboveground traits, when comparing herbaceous and woody species, different families or biomes. This agrees with the observation of a single functional hotspot on the fine-root plane but two hotspots aboveground, meaning that species with similar fine-root syndromes can display contrasting aboveground traits. Accordingly, the overlap between the distributions of herbaceous and woody species was more than four times higher on the fine-root plane than aboveground (dissimilarity fine-root plane = 18.7%, dissimilarity aboveground = 81.6%). Woodiness explained 36.5% of the variance of the position of species on the aboveground plane versus a mere 0.4% on the fine-root plane (similar to what was found in previous explorations of the RES4), which means that the suites of fine-root traits of herbaceous and woody species are virtually indistinguishable, in contrast to previous results based on a less diverse set of herbaceous species18.
The dissimilarity between pairs of families was also generally higher aboveground than on the fine-root plane (mean ± s.d.: dissimilarity aboveground = 76.2% ± 19.5, dissimilarity fine-root plane = 58.6% ± 12.0). For example, Fagaceae and Pinaceae overlapped only 4% on the aboveground plane, but 62% on the fine-root plane, indicating that the large differences among the aboveground traits of angiosperms and gymnosperms3 are not mirrored at the level of fine-root traits. This was verified by PERMANOVA analyses, where differences among families explained almost twice as much variation aboveground (60.8%) as on the fine-root plane (32.3%). However, 21% of the family pairs presented higher dissimilarities on the fine-root plane than on the aboveground plane. These cases mostly belonged to pairs of families including almost exclusively herbaceous species (for example, Poaceae versus Asteraceae) or pairs of almost exclusively woody families (Fagaceae versus Lauraceae), confirming that differences in traits aboveground are largely driven by plant size.
Although biomes explained less variation on the aboveground plane (PERMANOVA: 13.4%) than families, dissimilarities between pairs of biomes were always larger aboveground than on the fine-root plane (dissimilarity aboveground = 42.8% ± 18.1, dissimilarity fine-root plane = 21.6% ± 6.8). Furthermore, biomes had virtually no explanatory power for fine roots (PERMANOVA: 1.3%), which is in agreement with previous observations at the biome level4 and with the notion that the proportion of total variation that can be found in local communities is larger for fine-root than for aboveground traits24. For example, while the two biomes that were most different on the aboveground plane (temperate grassland/desert versus tropical rainforest) overlapped only 27%, the two most different biomes on the fine-root plane (subtropical desert versus temperate rainforest) still highly overlapped (62%).
We further asked whether the species composing each group were aggregated or dispersed, by examining the patterns of species redundancy25 within herbaceous and woody species, families and biomes. Groups with high redundancy are composed of species with similar trait values. We found that the average redundancy of all groups considering the four-dimensional space was always higher than on the aboveground plane, which was, at the same time, higher than on the fine-root plane (Extended Data Table 4). This result demonstrates that groups tend to partition the total and aboveground trait space (that is, the amount of variation among groups is relatively larger) and share the fine-root trait space. Herbaceous species were more redundant than woody species in all considered aspects of the trait space. This is in agreement with previous descriptions of woody species encompassing a larger amount of the fine-root trait space than herbaceous species4 and with the notion that the hotspot for woody species occupies a particular area of an otherwise very wide functional distribution of woody species3. Altogether, the suites of traits of herbaceous species are more restricted than those of woody species. In addition, herbaceous species follow, on average, a fast return on investment strategy26, which might reduce the potential for fine-root trait variation. Redundancy was higher on the aboveground plane than on the fine-root plane for all families (t20 = −11.13, P < 0.001; Extended Data Table 4), showing that confamilial species are generally more similar in their aboveground traits than in their fine-root traits. The distribution of species within biomes was not as constrained as within families, but redundancy in biomes aboveground also tended to be higher than on the fine-root plane (t7 = −2.03, P = 0.082), which is in agreement with observations showing that individual biomes, and even local sites, contain a large proportion of the global variation in both aboveground and fine-root traits4,5,27,28.
The fact that combining the GSPFF and the RES does not fundamentally change the organization of the two-dimensional spectra that have been described when considering aboveground3 and fine-root4 traits separately, along with the large differences between the dissimilarity and redundancy patterns observed, cast doubts about the generality of the high coordination between fine-root and leaf economic traits12,13,14. However, more-detailed analyses revealed evidence of coordination between aboveground and fine-root trait syndromes. For example, dissimilar families aboveground tended to be also dissimilar in their fine-root traits (Fig. 2a). Dissimilarities among biomes on both planes were positively correlated (Fig. 2b), which is probably associated with the fact that biomes with similar climates occupy similar areas of the functional space, on both the aboveground and the fine-root planes (Fig. 2c, d). Furthermore, more-redundant families were redundant in all considered aspects of the functional space (the four-dimensional space and the aboveground and fine-root planes), suggesting that families are organized along a gradient of whole-plant functional specialization, ranging from families with many species displaying very similar trait syndromes (for example, Poaceae and Pinaceae) to more functionally diverse families (for example, Polygonaceae and Rubiaceae; Fig. 3a). By contrast, we observed a negative relationship between redundancy patterns on the aboveground and fine-root planes in biomes (Fig. 3b), which stems from different associations between redundancy on each of the planes and net primary productivity (NPP). On the one hand, biomes with higher aboveground productivity hosted more redundant species on the aboveground plane (Fig. 3c). On the other hand, redundancy on the fine-root plane was negatively associated with belowground NPP (Fig. 3d). These results suggest that, while low biomass aboveground leads to lower dominance and higher trait diversity aboveground, low water availability and high seasonality reduce the diversity of belowground traits in biomes18. The contrasting relationships with NPP resulted in a lack of relationship between the redundancy of species in the four-dimensional space and the total NPP of the biome (that is, aggregating aboveground and belowground NPP; Fig. 3e).
a, b, The relationship between the dissimilarity on the aboveground and fine-root planes between pairs of: families (n = 210 pairs, 21 families; r = 0.19, P = 0.037, Mantel test with 999 repetitions) (a) and biomes (n = 28 pairs, 8 biomes; r = 0.73, P = 0.001, Mantel test with 999 repetitions) (b). c, d, Climate dissimilarity is positively correlated with the dissimilarities between pairs of biomes on the aboveground plane (n = 28 pairs, 8 biomes; r = 0.76, P = 0.001, Mantel test with 999 repetitions) (c) and the fine-root trait plane (n = 28 pairs, 8 biomes; r = 0.54, P = 0.036, Mantel test with 999 repetitions) (d). The thick lines show the fits of major axis regressions.
a, b, The relationship between aboveground and fine-root redundancy for major taxonomic families (n = 21 families; r = 0.78, P < 0.001, major axis regression) (a) and biomes (n = 8 biomes; r = −0.67, P = 0.029, major axis regression) (b). c–e, Within biomes, the degree of redundancy on the aboveground plane is positively associated with aboveground productivity (n = 8 biomes; F1,6 = 10.63, P = 0.017, linear regression) (c), whereas redundancy on the fine-root plane is negatively related to belowground productivity (n = 8 biomes; F1,6 = 9.25, P = 0.023, linear regression) (d), in contrast to a lack of relationship (indicated by the dashed line) between redundancy in the four-dimensional space and total productivity (n = 8 biomes; F1,6 = 1.16, P = 0.323, linear regression) (e).
Combining the most comprehensive trait databases available, we found that incorporating fine-root traits into the GSPFF3 enriches it with non-redundant information and exposes a general pattern of higher functional trait differentiation for aboveground traits than for fine-root traits. Our results confirm the paramount importance of the size and leaf tissue-quality dimensions aboveground, and the acquisition versus conservation and the plant–fungal interaction trade-offs between fine-root traits4,17,18 belowground. However, our results do not confirm the strong covariation between leaf and fine-root traits predicted by the plant economics spectrum hypothesis12,13,14. In addition, we found that plants differentiate preferentially in the aboveground rather than in the fine-root part of this functional trait space. This result seems to be directly associated with the very different nature of organ function and resource acquisition in the soil environment. Greater differentiation in aboveground trait syndromes than in fine roots may have emerged, for example, from less-stable environmental conditions29, and stronger and more heterogeneous effects of fire and large herbivores30 aboveground. Nonetheless, despite this apparent decoupling of the variation of aboveground and fine-root strategies, we also found evidence of whole-plant coordination, indicated by the correlated patterns of dissimilarity and redundancy among families and biomes.
Understanding the covariation of aboveground and belowground traits will illuminate how evolution has shaped plant strategies to cope with biotic and abiotic environments. Such understanding will require considering that fine-root traits are by no means the only relevant belowground traits, and that the belowground trait space is likely to have more dimensions beyond those described here, partially independent or associated with those aboveground. For example, previous assessments of biomass allocation patterns23,31 have shown that aboveground and belowground biomass are positively correlated, so that the size of the belowground system is likely to be associated with the first, size-related, component. This suggests that this dimension could be interpreted in terms of the size of the whole plant, and thus the two functional hotspots observed in the aboveground space might also be observed in the belowground space. However, not enough empirical data are currently available to adequately test these ideas. Future inclusion of underrepresented clades and other belowground traits, such as mycorrhizal traits18,32, root architectural or clonal traits17,33, may further modify the belowground trait dimensionality and improve our understanding of the full form and function of vascular plants.
Methods
Data collection and processing
Aboveground traits
We selected six aboveground traits previously shown to capture the GSPFF3: plant height (ph; measured in m), leaf area (la; measured in mm2), specific leaf area (sla; measured in mm2/mg; the inverse of leaf mass per area, used by those authors), leaf nitrogen concentration (ln; measured in mg/g), specific stem density (ssd; measured in g/m3) and seed mass (sm; measured in mg). We used publicly available data for these traits from the latest version of the TRY Plant Trait Database6 (version 5.0; https://www.try-db.org/TryWeb/Home.php, accessed April 2019). Altogether, the TRY dataset included over 955,000 trait measurements for 44,431 vascular plant taxa. We removed observations marked as juveniles or seedlings and those done in non-natural conditions (for example, growth chamber, greenhouse, field experiment, herbarium) whenever this information was available. In the analyses, each taxon was represented by an average trait value (excluding outliers with more than 3 s.d.) that was calculated first within individuals (if multiple measurements were taken from a single individual), then within datasets (if multiple individuals were measured in the same location), and finally within species (if multiple individuals were measured in various locations).
Plant height data included 143,429 measurements of adult plant vegetative height for 20,009 taxa. In most datasets, this was represented as observed height or average of measurements. In some cases, plant height was represented as the maximum observation (8,327 records). Specific stem density data included 26,216 measurements for 8,727 taxa. As this trait is usually measured for woody species, we estimated specific stem density for herbaceous plants using leaf dry mass content information (123,470 measurements for 5,684 taxa), following the procedures described in ref. 3. Leaf area data included 111,855 measurements for 13,808 taxa. Different datasets in TRY reported various measurements of leaf area (for example, leaf or leaflet or unknown, petiole included or excluded or unknown). To maximize our data coverage, we included each observation for which any measurement type was available. If different leaf area measurements were available for an individual observation, we included whole-leaf measurements (48% of records). If such data was not available, we included measurements where it was unknown if a leaf or leaflet was measured (12% of records) or where a leaflet was measured (40% of records including both simple and compound leaves). Similarly, if observations reported different petiole measurements, we included those where petiole was included (33% of records). For 50% of records, it was unknown if petiole was included or excluded, and for 17% of records, petiole was excluded. Specific leaf area data included 203,896 measurements for 14,222 taxa. Similarly to leaf area data, we included measurements with petiole (50% of records). For observations where such data was not available, we included measurements where it was unknown if petiole was included or excluded (47% of records) or where petiole was excluded (3% of records). Data for leaf nitrogen concentration included 86,211 measurements for 10,458 taxa. Data for seed mass included 183,170 measurements for 25,831 taxa. Our final aboveground plant trait dataset included 878,247 observations for 39,260 vascular plant species.
Fine-root traits
We collected data from the GRooT database8 and selected four fine-root traits: root diameter (D; measured in mm), specific root length (SRL; measured in m/g), root tissue density (RTD; measured in g/cm3) and root nitrogen concentration (N; measured in mg/g). These traits are deemed to be relevant for root economics4,9,11,16,17,18,19,34. We followed the steps for data curation and preparation included in ref. 4. Specifically, we removed data from dead roots and excluded ferns (Polypodiopsida) and lycopods (Lycopodiopsida) due to their particular root morphology. We only selected data for roots that were either classified as ‘fine roots’ by the original authors, or were defined as roots of orders 1–3 (we kept the minimum order in the cases in which data for more than one order was available in the dataset), or had a diameter smaller than 2 mm (keeping the roots of minimum diameter in case a range of diameters was provided from the same study). We excluded roots with root tissue density > 1. Unlike the TRY database, the GRooT database does not provide information on whether the root data were measured on a seedling or a mature plant. However, since fine roots are relatively young regardless of plant age35, it is unlikely that the fine-root trait data are strongly affected by plant age. Finally, we accounted for the study design (67% of measurements were performed on plants in natural conditions, 32% in pots, less than 0.1% in hydroponic experiments and 0.5% in unspecified conditions) and for the original publications of the trait measurements (to account for other study-specific factors) by making a linear mixed model for each trait. In these models, the log10-transformed and scaled (to 0 mean and 1 s.d.) trait values were used as the response variable, study design as a fixed factor and publication as a random factor4. We then used the residuals of these models as values for each trait. Finally, we removed outliers (with trait values more than 3 s.d. from the species average). To explore the potential effect of measurements not made in natural conditions, we repeated the whole procedure considering only field data, and found that the average species root trait values were highly correlated with the data in which pot data were also included (Pearson’s correlation > 0.97 for all traits). Furthermore, the relationships among different traits were not affected by the exclusion of pot data (Extended Data Table 2c). Therefore, to keep as many species as possible in subsequent analyses, we decided to use the data including measurements taken in different conditions.
Root diameter data included 10,251 observations for 1,592 taxa, specific root length data had 9,966 individual measurements for 1,736 species, root tissue density included 7,662 observations for 1,390 species and root nitrogen concentration data included 5,219 observations for 1,253 species. In total, our belowground dataset included 33,098 observations for 2,050 species.
Woodiness data was mainly extracted from the GRooT database8. For 77 species with missing woodiness values, we used other published databases on woodiness36 or growth form37,38,39. Following the growth form data, trees, shrubs and subshrubs were classified as woody, whereas herbs and graminoids were assigned as non-woody. If a single database or different sources reported different values for a species, we assigned them to both groups (woody/non-woody). For 26 species, we searched for woodiness info from online specimen photos or descriptions.
Imputation of missing traits
Taxonomies from the used trait sources (TRY and GRooT) were standardized to ‘The Plant List’ taxonomy40 using the R package Taxonstand41. We combined the aboveground and fine-root trait information, resulting in a dataset that included 1,719 species with at least one aboveground and one fine-root trait measured (‘full dataset’). Neither the aboveground nor the fine-root traits were complete (see Supplementary Table 1), with only 301 species having complete empirical trait measurements for the ten traits (‘complete dataset’). We completed trait information by performing a trait-imputation procedure for the ten traits at the same time using the missForest R package42,43. Before the imputation process, all traits were log10-transformed, centred and scaled. We incorporated the evolutionary relationships between species in the imputation process by including the first ten phylogenetic eigenvectors in the matrix to be imputed, as recommended in ref. 44. The phylogeny was obtained using the R package V.Phylomaker45, with the GBOTB phylogeny46 as the backbone. To reduce the uncertainty about the imputed trait values, after the imputation procedure, we only retained those species in which empirical trait measurements were available for at least 50% of the aboveground traits (that is, at least three aboveground traits) and at least 50% of the fine-root traits (that is, at least two fine-root traits). Finally, we estimated the reliability of the imputation procedure (Supplementary Methods 1). This procedure resulted in a final dataset that included 1,218 species from 127 families in which 84.6% of trait information (10,305 records) came from empirical measurements and 15.4% of data (1,875 records) were imputed (‘imputed dataset’; Extended Data Table 1).
Ascribing species to biomes
We defined biomes by their vegetation type, according to Whittaker’s definition47, in the basis of the average temperature and precipitation, using the R package plotbiomes48 (adapted from ref. 49.). We downloaded all of the records belonging to vascular plants (phylum: Tracheophyta) with coordinates from GBIF50 and then selected the records belonging to the 1,218 species of the imputed dataset. We filtered out the records with clearly false locality coordinates (for example, equal latitude and longitude, both latitude and longitude equal to zero, and longitude/latitude outside possible ranges) and the records from living specimens (that is, from zoos and botanical gardens), conserved specimens (that is, museums) and unknown sources. This resulted in a final dataset of 1,131 species. The number of geolocalized species and the number of records by species varied between 1 and 906,097 (1st quartile = 158, median = 1,501, 3rd quartile = 35,501). We assigned a value of average temperature (BIO1) and precipitation (BIO12) to each geolocalized record (which we represented using the R package rworldmap51; Extended Data Fig. 2), using Worldclim data with a resolution of 10 min of a degree, available in the R package plotbiomes48. We then ascribed one biome to each record according to their values of temperature and precipitation (Extended Data Fig. 2). If a species was present in more than one biome, only biomes with a proportion of records greater than 5% of the total number of records were taken into account. Biome information for the 87 species that were not found in GBIF was retrieved from online descriptions (principally from http://www.plantsoftheworldonline.org/).
Construction of the global trait space
We identified the main axes of functional trait variation by performing PCAs on the log-transformed and scaled functional traits, using the subset of species with complete empirical trait measurements for (1) only the aboveground traits (2,630 species), (2) only the fine-root traits (748 species), and (3) all ten traits together (301 species). We used Horn’s parallel analysis in the R package paran52 to determine the dimensionality of these PCAs and applied a varimax rotation to the selected components to facilitate the interpretation of results (see Supplementary Methods 2 for detailed explanations about this procedure). We refer to these reduced and rotated spaces as ‘functional spaces’ from now on.
Examination of the aboveground, fine-root and all-traits spaces revealed that, while both the aboveground and fine-root spaces could be summarized with two components (in correspondence with previous results3,4; see Supplementary Note 1), the spaces built with the ten traits (both aboveground and fine-root traits; ‘total space’) had a dimensionality of four (see main text for discussion). A first inspection of the spaces suggested that the first two components of the total space corresponded very markedly with the aboveground space, whereas the fine-root space corresponded markedly to the third and fourth components. We examined this correspondence by estimating the correlation between distance matrices of the species that were common to all spaces (that is, 301 species with complete trait information for all traits) through three Procrustes tests: one considering the scores of species in the first and second components of the total space and the aboveground space, one considering the third and fourth components of the total space and the fine-root space, and one considering the scores of species in the aboveground space and the fine-root space. For this, we used the ‘procuste.rtest’ function from the R package ade4 (ref. 53). To assess the significance of the correlation, permutation tests (9,999 randomizations) based on Monte-Carlo simulations were generated. The two first Procrustes tests indicated a strong correspondence between the only aboveground and only fine-root spaces and the corresponding planes of the total space (Procrustes correlation between aboveground space and components 1 and 2 of the total space = 0.988, P = 0.0001; Procrustes correlation between fine-root space and components three and four of the total space = 0.982, P = 0.0001); consequently, we used only the total space in subsequent analyses. The third Procrustes test revealed a weak correspondence between the position of species in the only aboveground and only fine-root spaces (Procrustes correlation between the aboveground space and the fine-root space = 0.178, P = 0.0001).
We estimated the probabilistic distribution of the species within the functional space based on the complete dataset by performing multivariate kernel density estimations with the trait probability density (TPD) and ks R packages54,55,56,57. The kernel for each species was a multivariate normal distribution centred in the coordinates of the species in the functional space and bandwidth chosen using unconstrained bandwidth selectors from the ‘Hpi’ function in the ks package54,55,58. The aggregated kernels for all species in the functional space result in the TPD function56,59 of plants in the corresponding space (we created TPD functions for the aboveground plane, fine-root plane and total space). Although TPD functions are continuous, to perform operations with them, it is more practical to divide the functional space into a D-dimensional grid composed of many equal-sized cells (we divided the two-dimensional spaces into 40,000 cells, 200 per dimension, and the four-dimensional space into 810,000 cells, 30 per dimension). The value of the TPD function in a given point of the space reflects the density of species in that particular area of the space (that is, species with similar functional traits). For each of these spaces, we represented graphically the global TPD as well as the contours containing 50%, 60%, 70%, 80%, 90% and 99% of the total probability.
We compared the distribution of species within the aboveground plane, fine-root plane and total space with a null model considering that species are distributed following a multivariate normal distribution3,22. For this, for each part of the space (aboveground, fine root and total), we drew 499 samples of 301 simulated species (that is, as many species as in the dataset with complete empirical trait information) from multivariate normal distributions with the same mean and covariance matrix as the observed data. For each of these samples, we estimated a TPD function and measured functional richness (amount of space occupied by the set of species56,59,60,61) at decreasing probability thresholds (from 99% to 1% quantiles in 0.1% intervals). This way, we estimated a ‘profile’ of the probabilistic distributions of species, reflecting what proportion of the functional space is occupied at different probabilities. We also estimated the profile of the observed TPD functions (based on the 301 species). This analysis allows examination of how realized trait syndromes are constrained within the potential space of all combinations. A very high concentration of species in small portions of the space will show trajectories in which functional richness increases drastically as we increase the probability threshold, indicating that there is high redundancy at the global scale and vice versa. We also estimated functional divergence for the null models and the observed TPD functions. Functional divergence is an indicator of the degree to which the density of species in the functional trait space is distributed towards the extremes of the distribution56,60,61: higher divergence in the observed distribution than expected from the null model would indicate that the most prevalent combinations of functional traits are relatively far from the centre of the functional space, whereas lower divergence would indicate a higher aggregation of species in the centre of the functional space (compared with the null model). We compared the estimations of functional richness at different probability thresholds and functional divergence of the observed and simulated data by estimating a standardized effect size (SES = (observed value – mean(simulated values))/s.d.(simulated values)). SES values indicate how many s.d. units the observed value deviates from the mean of the observed values, with SES > 0 indicating that the observed value of a given metric is bigger than the average of the simulated values and vice versa.
Exploration of patterns within the global trait space
We then explored the distribution of different groups of species within the trait space. Specifically, we focused on examining whether there were differences in patterns between the aboveground (first and second components) and the fine-root (third and fourth components) planes. For this, we grouped species according to their woodiness (herbaceous versus woody species), the taxonomic family they belong to and the biomes they occur in. To examine these questions in a larger set of species than the 301 species with complete empirical information for all traits, we used the functional space based on the imputed dataset including 1,218 species.
We performed the same set of analyses for each of the grouping criteria (woodiness, families and biomes). First, we estimated how much of the total variation in the position of the species on the aboveground and fine-root planes as well as in the total space was explained by differences among groups. For this, we calculated the dissimilarities (using Euclidean distances) between all pairs of species considering the three aspects of the space (C1 and C2 for the aboveground plane, C3 and C4 for the belowground plane, and C1, C2, C3 and C4 for the total space), and analysed the dissimilarity matrices using PERMANOVA (R package vegan62). For woodiness and families, we made a PERMANOVA considering the group to which each species is ascribed as the explanatory variable. For biomes, since the same species can belong to different biomes, we performed 500 repetitions of a PERMANOVA in which, in each iteration, biomes were assigned to species with a probability that was proportional to the relative frequency of the species in the biomes, and used biome as the explanatory variable. This procedure is equivalent to performing a partitioning of functional diversity across scales63,64. When the explanatory variable (for example, the family) explains a large proportion of variation, it means that differences between groups account for most of the functional variability; conversely, when the explanatory variable explains a little variance, most of the total functional variation is due to differences among species within groups (that is, within woodiness levels, families or biomes).
Then, we selected groups that included at least 15 species and estimated their TPD functions on the aboveground and fine-root planes (567 woody species and 617 herbaceous species; 21 families; 8 biomes). We applied a quantile threshold of 99% to all TPD functions to reduce the potential effect of outliers on the estimation of the amount of functional space occupied by the different groups56,65,66. After thresholding, the TPD functions were rescaled, so that they again integrated to one across the functional space, and the probabilities expressed in terms of quantiles to ease interpretability of the results.
We estimated the dissimilarity between pairs of groups as 1 – overlap between their respective TPD functions56,59,66,67,68,69. Compared with methods that consider exclusively the boundaries of the distributions, such as hypervolumes or convex hulls70,71,72, probabilistic-based dissimilarities between TPD functions also consider the differences in density of species within the boundaries. This approach provides a more complete idea of what the differences between the functional spectra are, particularly in cases in which functional redundancy is high73,74,75,76. Given that a high proportion of the considered species might be clumped in particular areas of the considered space3, this methodological aspect can be particularly useful to detect differences in the occupation of functional spaces among groups of species. For families and biomes, we explored the relationships between the dissimilarities on the aboveground and fine-root planes by means of Mantel tests using the R package vegan62,77. Finally, and specifically for biomes, we examined the relationship between dissimilarity in their occupation of the trait space and climate dissimilarity. For this, for each combination of biome and species observed, we estimated the average mean annual temperature and precipitation values, by considering the GBIF records of each species present in the corresponding biome (see ‘Ascribing species to biomes’). Then, we averaged the species by biome averages within each biome to get biome-level mean annual temperature and precipitation values and built a matrix of climate dissimilarity between biomes using the Gower’s78 dissimilarity based on these averages. We explored the relationship between dissimilarity in the trait space and climate dissimilarity between pairs of biomes using Mantel tests.
We then estimated functional richness as the amount of functional space occupied by the 0.99 quantile of the corresponding TPD. Given that the functional richness of a group is positively related to the number of species that compose it, we performed null models to express functional richness independently from species richness79. We compared the observed functional richness value in each group with the values expected under a random species assignment; for this, for each group (for woody and non-woody species, for each family and for each biome), we randomly selected the same number of species from the whole dataset and estimated the TPD of this null assemblage. We estimated 500 null values of functional richness for each group using this procedure and used them to estimate a SES of functional richness for each group64,79. Values of SES smaller than 0 for a given group indicate that the amount of functional space occupied by the group in question is smaller than expected for that number of species, that is, that there is higher than expected functional redundancy among the species in the considered group. Consequently, we used the opposite values of functional richness SES (that is, multiplying them by −1) to make the interpretation of redundancy more straightforward. We examined whether the levels of functional redundancy of species within families and biomes were higher in the aboveground or the belowground parts of the spectrum using paired t-tests. We then analysed the relationship between the functional redundancy aboveground and belowground at the family and biome levels using major axis regressions.
Finally, specifically for biomes, we examined the relationship between SES of functional richness and NPP. Specifically, we estimated aboveground, belowground and total NPP using the formulas provided in ref. 80. These estimations are based on mean annual precipitation, which we estimated for each biome as the average of the mean annual precipitation values of each of the species present in the biome (BIO12; see ‘Ascribing species to biomes’). We examined these relationships by regressing the values of redundancy in biomes in each aspect of the functional space (aboveground and fine-root planes, and total space) against its corresponding value of NPP (for example, aboveground redundancy regressed against aboveground NPP).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The datasets generated and analysed during the current study are available in the Figshare repository: https://doi.org/10.6084/m9.figshare.13140146.
Code availability
The R code used in the current study is available in the Figshare repository: https://doi.org/10.6084/m9.figshare.13140146.
References
Grime, J. P. Plant Strategies, Vegetation Processes, and Ecosystem Properties (John Wiley and Sons, 2001).
Reich, P. B. et al. The evolution of plant functional variation: traits, spectra, and strategies. Int. J. Plant Sci. 164, S143–S164 (2003).
Díaz, S. et al. The global spectrum of plant form and function. Nature 529, 167–171 (2016).
Bergmann, J. et al. The fungal collaboration gradient dominates the root economics space in plants. Sci. Adv. 6, eaba3756 (2020).
Wright, I. J. et al. The worldwide leaf economics spectrum. Nature 428, 821–827 (2004).
Kattge, J. et al. TRY plant trait database — enhanced coverage and open access. Glob. Chang. Biol. 26, 119–188 (2020).
Iversen, C. M. et al. A global Fine-Root Ecology Database to address below-ground challenges in plant ecology. New Phytol. 215, 15–26 (2017).
Guerrero-Ramírez, N. R. et al. Global root traits (GRooT) database. Glob. Ecol. Biogeogr. 30, 25–37 (2021).
McCormack, M. L. et al. Redefining fine roots improves understanding of below-ground contributions to terrestrial biosphere processes. New Phytol. 207, 505–518 (2015).
Rasse, D. P., Rumpel, C. & Dignac, M. F. Is soil carbon mostly root carbon? Mechanisms for a specific stabilisation. Plant Soil 269, 341–356 (2005).
Eissenstat, D. M. Costs and benefits of constructing roots of small diameter. J. Plant Nutr. 15, 763–782 (1992).
Freschet, G. T., Cornelissen, J. H. C., van Logtestijn, R. S. P. & Aerts, R. Evidence of the ‘plant economics spectrum’ in a subarctic flora. J. Ecol. 98, 362–373 (2010).
Reich, P. B. The world-wide ‘fast–slow’ plant economics spectrum: a traits manifesto. J. Ecol. 102, 275–301 (2014).
Shen, Y. et al. Linking aboveground traits to root traits and local environment: implications of the plant economics spectrum. Front. Plant Sci. 10, 1412 (2019).
Kramer-Walter, K. R. et al. Root traits are multidimensional: specific root length is independent from root tissue density and the plant economic spectrum. J. Ecol. 104, 1299–1310 (2016).
Bergmann, J., Ryo, M., Prati, D., Hempel, S. & Rillig, M. C. Root traits are more than analogues of leaf traits: the case for diaspore mass. New Phytol. 216, 1130–1139 (2017).
Weemstra, M. et al. Towards a multidimensional root trait framework: a tree root review. New Phytol. 211, 1159–1169 (2016).
Ma, Z. et al. Evolutionary history resolves global organization of root functional traits. Nature 555, 94–97 (2018).
de la Riva, E. G. et al. Root traits across environmental gradients in Mediterranean woody communities: are they aligned along the root economics spectrum? Plant Soil 424, 35–48 (2018).
Craine, J. M., Lee, W. G., Bond, W. J., Williams, R. J. & Johnson, L. C. Environmental constraints on a global relationship among leaf and root traits of grasses. Ecology 86, 12–19 (2005).
Liese, R., Alings, K. & Meier, I. C. Root branching is a leading root trait of the plant economics spectrum in temperate trees. Front. Plant Sci. 8, 315 (2017).
Carmona, C. P. et al. Erosion of global functional diversity across the tree of life. Sci. Adv. 7, eabf2675 (2021).
Niklas, K. J. Modelling below- and above-ground biomass for non-woody and woody plants. Ann. Bot. 95, 315–321 (2005).
Liu, G. et al. Coordinated variation in leaf and root traits across multiple spatial scales in Chinese semi-arid and arid ecosystems. New Phytol. 188, 543–553 (2010).
Galland, T., Carmona, C. P., Götzenberger, L., Valencia, E. & de Bello, F. Are redundancy indices redundant? An evaluation based on parameterized simulations. Ecol. Indic. 116, 106488 (2020).
Valverde‐Barrantes, O. J., Maherali, H., Baraloto, C. & Blackwood, C. B. Independent evolutionary changes in fine‐root traits among main clades during the diversification of seed plants. New Phytol. 228, 541–553 (2020).
Cornwell, W. K. et al. Plant species traits are the predominant control on litter decomposition rates within biomes worldwide. Ecol. Lett. 11, 1065–1071 (2008).
Freschet, G. T. et al. Climate, soil and plant functional types as drivers of global fine-root trait variation. J. Ecol. 105, 1182–1196 (2017).
De Deyn, G. B. & Van der Putten, W. H. Linking aboveground and belowground diversity. Trends Ecol. Evol. 20, 625–633 (2005).
Pausas, J. G. & Bond, W. J. Humboldt and the reinvention of nature. J. Ecol. 107, 1031–1037 (2019).
Poorter, H. et al. Biomass allocation to leaves, stems and roots: meta-analyses of interspecific variation and environmental control. New Phytol. 193, 30–50 (2012).
Moora, M. Mycorrhizal traits and plant communities: perspectives for integration. J. Veg. Sci. 25, 1126–1132 (2014).
Freschet, G. T. et al. Root traits as drivers of plant and ecosystem functioning: current understanding, pitfalls and future research needs. New Phytol. https://doi.org/10.1111/nph.17072 (2021).
McCormack, M. L. & Iversen, C. M. Physical and functional constraints on viable belowground acquisition strategies. Front. Plant Sci. 10, 1215 (2019).
Wells, C. E. & Eissenstat, D. M. Beyond the roots of young seedlings: the influence of age and order on fine root physiology. J. Plant Growth Regul. 21, 324–334 (2002).
Zanne, A. E. et al. Three keys to the radiation of angiosperms into freezing environments. Nature 506, 89–92 (2014).
USDA. USDA PLANTS Database (accessed 3rd July 2020); https://plants.sc.egov.usda.gov
Engemann, K. et al. A plant growth form dataset for the New World. Ecology 97, 3243 (2016).
BGCI. GlobalTreeSearch online database (accessed 3rd July 2020); https://www.bgci.org/globaltree_search.php
The Plant List. The Plant List (accessed 17th February 2020); http://www.theplantlist.org
Cayuela, L., Macarro, I., Stein, A. & Oksanen, J. Taxonstand: Taxonomic Standardization of Plant Species Names. R package version 2.2. https://CRAN.R-project.org/package=Taxonstand (2019).
Stekhoven, D. J. & Buhlmann, P. MissForest—non-parametric missing value imputation for mixed-type data. Bioinformatics 28, 112–118 (2012).
Oliveira, B. F., Sheffers, B. R. & Costa, G. C. Decoupled erosion of amphibians’ phylogenetic and functional diversity due to extinction. Glob. Ecol. Biogeogr. 29, 309–319 (2020).
Penone, C. et al. Imputation of missing data in life-history trait datasets: which approach performs the best? Methods Ecol. Evol. 5, 961–970 (2014).
Jin, Y. & Qian, H. V.PhyloMaker: an R package that can generate very large phylogenies for vascular plants. Ecography 42, 1353–1359 (2019).
Smith, S. A. & Brown, J. W. Constructing a broadly inclusive seed plant phylogeny. Am. J. Bot. 105, 302–314 (2018).
Whittakker, R. H. Communities and Ecosystems (Macmillan, 1975).
Stefan, V. & Levin, S. plotbiomes: Plot Whittaker biomes with ggplot2. R package version 0.0.0.9001 https://github.com/valentinitnelav/plotbiomes (2021).
Ricklefs, R. E. The Economy of Nature (W. H. Freeman and Company, 2008).
GBIF. GBIF Occurrence Download (accessed 15 December 2019); https://doi.org/10.15468/dl.thlxph
South, A. rworldmap: a new R package for mapping global data. R J. 3, 35–43 (2011).
Dinno, A. paran: Horn’s Test of Principal Components/Factors. R package version 1.5.2. https://CRAN.R-project.org/package=paran (2018).
Dray, S. & Dufour, A.-B. The ade4 package: implementing the duality diagram for ecologists. J. Stat. Softw. https://doi.org/10.18637/jss.v022.i04 (2007).
Duong, T. ks: kernel density estimation and kernel discriminant analysis for multivariate data in R. J. Stat. Softw. https://doi.org/10.18637/jss.v021.i07 (2015).
Duong, T. ks: Kernel smoothing. R package version 1.11.5 https://CRAN.R-project.org/package=ks (2019).
Carmona, C. P., Bello, F., Mason, N. W. H. & Lepš, J. Trait probability density (TPD): measuring functional diversity across scales based on TPD with R. Ecology 100, e02876 (2019).
Carmona, C. P. TPD: methods for measuring functional diversity based on Trait Probability Density. R package version 1.1.0. https://CRAN.R-project.org/package=TPD (2019).
Duong, T. & Hazelton, M. L. Plug-in bandwidth matrices for bivariate kernel density estimation. J. Nonparametr. Stat. 15, 17–30 (2003).
Carmona, C. P., de Bello, F., Mason, N. W. H. & Lepš, J. Traits without borders: integrating functional diversity across scales. Trends Ecol. Evol. 31, 382–394 (2016).
Mason, N. W. H., Mouillot, D., Lee, W. G. & Wilson, J. B. Functional richness, functional evenness and functional divergence: the primary components of functional diversity. Oikos 111, 112–118 (2005).
Villéger, S., Mason, N. W. H. & Mouillot, D. New multidimensional functional diversity indices for a multifaceted framework in functional ecology. Ecology 89, 2290–2301 (2008).
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.5-5 https://CRAN.R-project.org/package=vegan (2019).
Carmona, C. P. et al. Taxonomical and functional diversity turnover in Mediterranean grasslands: interactions between grazing, habitat type and rainfall. J. Appl. Ecol. 49, 1084–1093 (2012).
Micó, E. et al. Contrasting functional structure of saproxylic beetle assemblages associated to different microhabitats. Sci. Rep. 10, 1520 (2020).
Blonder, B. et al. New approaches for delineating n-dimensional hypervolumes. Methods Ecol. Evol. 9, 305–319 (2018).
Carmona, C. P., de Bello, F., Mason, N. W. H. & Lepš, J. The density awakens: a reply to Blonder. Trends Ecol. Evol. 31, 667–669 (2016).
Mouillot, D. et al. Niche overlap estimates based on quantitative functional traits: a new family of non-parametric indices. Oecologia 145, 345–353 (2005).
de Bello, F., Carmona, C. P., Mason, N. W. H., Sebastià, M.-T. & Lepš, J. Which trait dissimilarity for functional diversity: trait means or trait overlap? J. Veg. Sci. 24, 807–819 (2013).
Traba, J., Iranzo, E. C., Carmona, C. P. & Malo, J. E. Realised niche changes in a native herbivore assemblage associated with the presence of livestock. Oikos 126, 1400–1409 (2017).
Cornwell, W. K., Schwilk, D. W. & Ackerly, D. D. A trait-based test for habitat filtering: Convex Hull Volume. Ecology 87, 1465–1471 (2006).
Blonder, B., Lamanna, C., Violle, C. & Enquist, B. J. The n-dimensional hypervolume. Glob. Ecol. Biogeogr. 23, 595–609 (2014).
Blonder, B. Hypervolume concepts in niche- and trait-based ecology. Ecography 41, 1441–1455 (2018).
Ricotta, C. et al. Measuring the functional redundancy of biological communities: a quantitative guide. Methods Ecol. Evol. 7, 1386–1395 (2016).
Mouillot, D. et al. Functional over-redundancy and high functional vulnerability in global fish faunas on tropical reefs. Proc. Natl. Acad. Sci. USA 111, 13757–13762 (2014).
Carmona, C. P., de Bello, F., Sasaki, T., Uchida, K. & Pärtel, M. Towards a common toolbox for rarity: a response to Violle et al. Trends Ecol. Evol. 32, 889–891 (2017).
Violle, C. et al. Functional rarity: the ecology of outliers. Trends Ecol. Evol. 32, 356–367 (2017).
Borcard, D., Legendre, P. & Drapeau, P. Partialling out the spatial component of ecological variation. Ecology 73, 1045–1055 (1992).
Gower, J. C. General coefficient of similarity and some of its properties. Biometrics 27, 857–871 (1971).
Carmona, C. P. et al. Agriculture intensification reduces plant taxonomic and functional diversity across European arable systems. Funct. Ecol. 34, 1448–1460 (2020).
Gherardi, L. A. & Sala, O. E. Global patterns and climatic controls of belowground net carbon fixation. Proc. Natl Acad. Sci. USA 117, 20038–20043 (2020).
Acknowledgements
This study has been supported by the TRY initiative on plant traits (http://www.try-db.org). The TRY initiative and database is hosted, developed and maintained by J. Kattge and G. Boenisch (Max Planck Institute for Biogeochemistry, Jena, Germany). TRY is currently supported by Future Earth/bioDISCOVERY and the German Centre for Integrative Biodiversity Research (iDiv) Halle–Jena–Leipzig. This study was financed by the Estonian Ministry of Education and Research (PSG293 for C.P.C., C.G.B., S.T. and R.T.; PRG609 for M.P. and R.T.; PSG505 for A.T.; and PRG1065 for M.M., M.Z. and C.G.B.), by the European Union through the European Regional Development Fund (Centre of Excellence EcolChange), the University of Tartu (PLTOM20903). S.D. received partial funding from CONICET, FONCyT and Universidad Nacional de Córdoba (Argentina), IAI SGP and the Newton Fund. Partial funding for A.D.M. is from the Natural Sciences and Engineering Research Council of Canada.
Author information
Authors and Affiliations
Contributions
C.P.C., C.G.B., A.T., S.T., M.M. and R.T. conceived the study. R.T. and C.P.C. collected and processed the trait data. A.T. and S.T. collected and processed the biogeographical and climate data. C.P.C. and A.T. analysed the data with input from all authors. C.G.B., S.T. and R.T. performed a literature search. C.P.C. wrote a first draft of the manuscript, assisted by C.G.B., A.T., S.T. and R.T. S.D., M.M., A.M., M.P. and M.Z. contributed to the design of analyses, interpretation of results and article writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Ian Wright and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Correlations between traits.
Pairwise correlations between the considered traits in the different datasets (black: full dataset with 1,719 species, blue: imputed dataset with 1,218 species, orange: complete dataset with 301 species). The lower-left triangle of the matrix contains scatterplots of traits (after log-10 transformation) showing the relationship (including regression lines) between each pair of traits. The diagonal includes a probability density function showing the distribution of each individual trait. The upper-right triangle includes the value of the correlation coefficients and, in the case of the full dataset, the number of species with empirical data for both traits (imputed and complete dataset always considered the same numbers of species). Lines for each dataset have different thickness to allow visualization of the correlation and probability density function even when there is high overlap between lines corresponding to different datasets.
Extended Data Fig. 2 Geographical, climatic and phylogenetic cover of the datasets.
a, Global map (Robinson projection) showing the occurrences (according to GBIF: http://www.gbif.org) of the species in the imputed dataset (1,218 species with empirical information for at least three aboveground and two fine-root traits). b, Number of species present in the major biomes47 in the imputed dataset (in parentheses, number of species in the complete dataset). c, Distribution of species across the phylogeny of vascular plants excluding ferns (Polypodiopsida) and lycopods (Lycopodiopsida) in the complete (301 species) and imputed datasets.
Extended Data Fig. 3 Individual aboveground and fine-root functional spaces.
Probabilistic distributions of the 2,630 and 748 species with complete empirical information for aboveground (a) and fine-root (b) traits in the functional spaces defined by a PCA on the corresponding traits followed by varimax rotation. The colour gradient (red-yellow-white) depicts different density of species in the defined space (red areas are more densely populated). Arrow length is proportional to the loadings of the considered traits in the resulting space. Aboveground traits are represented in green tones and fine-root traits in brown tones. Thick contour lines indicate the 0.5 and 0.99 quantiles, and thinner lines indicate quantiles 0.6, 0.7, 0.8 and 0.9.
Extended Data Fig. 4 Functional space using the complete dataset.
Probabilistic distributions of the 301 species with complete empirical information in the functional space defined by a PCA followed by varimax rotation based on both aboveground and fine-root traits. Each panel shows a combination of two of the four components that define the full plant spectrum. The colour gradient (red-yellow-white) depicts different density of species in the defined space (red areas are more densely populated). Arrow length is proportional to the loadings of the considered traits in the resulting space. Only those traits that had a loading of at least 0.3 in any of the represented components are shown to improve visualization (see loadings of all components in table S2). Aboveground traits are represented in green tones and fine-root traits in brown tones. Thick contour lines indicate the 0.5 and 0.99 quantiles, and thinner lines indicate quantiles 0.6, 0.7, 0.8 and 0.9.
Extended Data Fig. 5 Comparison of the occupation of the functional space with multivariate normal distributions.
Functional richness profile (amount of functional space occupied by quantiles of the functional spectra), difference in % of functional space occupied with respect to the null models, and functional divergence (representing the degree to which the density of species in the trait space is distributed towards the extremes of the distribution of species; right column) considering the first and second (a), third and fourth (b) and all components (c). In the functional profile plots (top of the left column in each case) green lines represent the mean, 2.5% and 97.5% quantiles of the functional richness profiles of null models (n = 499) representing multivariate normal distributions with equivalent parameters (means and standard deviations) than the observed data; orange lines represent the functional richness profile of the observed spectra. The values of functional richness for the 0.5 and 0.99 quantiles of all profiles are shown for comparison. The difference plots (bottom of the left column in each case), represent the percentage of functional space occupied by each quantile in relation to the mean of the null models; negative percentages mean that the considered quantile of the observed distribution occupies less space than the average of the null models, and vice versa. Each repetition of the null model (n = 499) is represented with a thin green line, whereas thicker green lines represent the mean, 2.5% and 97.5% quantiles of the 499 null models, and the orange line represents the observed distribution. The right column of each case represents the observed and null values of functional divergence; two-sided p values were estimated by confronting the value of the Standardize Effect Size (SES) with the cumulative normal distribution with mean = 0 and standard deviation = 1. The centre, bounds of box, and whiskers of the boxplot represent the median, 25th and 75th percentiles, and 1.5 times the interquartile range, respectively.
Extended Data Fig. 6 Functional space using the imputed dataset.
Probabilistic distributions of the 1,218 species with information for at least three aboveground and two fine-root traits (imputed dataset) in the functional space defined by a PCA followed by varimax rotation based on both aboveground and fine-root traits of the subset of species with complete empirical information. Each panel shows a combination of two of the four components that define the full plant spectrum. The colour gradient (red-yellow-white) depicts different density of species in the defined space (red areas are more densely populated). Arrow lengths are proportional to the loadings of the considered traits in the resulting space. Only those traits that had a loading of at least 0.4 in any of the represented components are shown to improve visualization (see loadings of all components in Extended Data Table 2). Aboveground traits are represented in green tones and fine-root traits in brown tones. Thick contour lines indicate the 0.5 and 0.99 quantiles, and thinner lines indicate quantiles 0.6, 0.7, 0.8 and 0.9.
Supplementary information
Supplementary Information
This file contains Supplementary Application 1, Supplementary Note 1, Supplementary Methods 1, Supplementary Methods 2 and Supplementary references.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Carmona, C.P., Bueno, C.G., Toussaint, A. et al. Fine-root traits in the global spectrum of plant form and function. Nature 597, 683–687 (2021). https://doi.org/10.1038/s41586-021-03871-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03871-y
- Springer Nature Limited
This article is cited by
-
Molecular-level carbon traits underlie the multidimensional fine root economics space
Nature Plants (2024)
-
Biodiversity and network complexity of rhizosphere soil microbiomes regulate the differentiation of Capsicum growth strategies
Plant and Soil (2024)
-
Divergent intra- and interspecific root order variability identifies a two-dimensional root economics spectrum
Plant and Soil (2024)
-
Coordination of leaf and root economic space in alpine coniferous forests on the Tibetan Plateau
Plant and Soil (2024)
-
Leaf phenotypic plasticity and integration balance plant adaptation to water table decline: a mesocosm experiment
Plant and Soil (2024)