Abstract
Metazoan development requires the robust proliferation of progenitor cells, the identities of which are established by tightly controlled transcriptional networks1. As gene expression is globally inhibited during mitosis, the transcriptional programs that define cell identity must be restarted in each cell cycle2,3,4,5 but how this is accomplished is poorly understood. Here we identify a ubiquitin-dependent mechanism that integrates gene expression with cell division to preserve cell identity. We found that WDR5 and TBP, which bind active interphase promoters6,7, recruit the anaphase-promoting complex (APC/C) to specific transcription start sites during mitosis. This allows APC/C to decorate histones with ubiquitin chains branched at Lys11 and Lys48 (K11/K48-branched ubiquitin chains) that recruit p97 (also known as VCP) and the proteasome, which ensures the rapid expression of pluripotency genes in the next cell cycle. Mitotic exit and the re-initiation of transcription are thus controlled by a single regulator (APC/C), which provides a robust mechanism for maintaining cell identity throughout cell division.
Similar content being viewed by others
Main
The self-renewal of stem cells endows organisms with the capacity to establish or regenerate their many tissues, but the misregulation of self-renewal contributes to tumorigenesis, tissue degeneration or ageing8. Although tightly controlled transcriptional networks establish the identity of self-renewing stem cells during interphase1, changes in chromatin architecture and the activity of transcription factors restrict the synthesis of messenger RNA (mRNA) during mitosis9. Stem cells must therefore restart their gene-expression programs each time they enter a new cell cycle4,5, which is facilitated by promoter elements that remain unwound during mitosis2, hypersensitive to DNase I2,10, and accessible to RNA polymerase II and transcription factors such as the TATA-box binding protein TBP3,11,12,13. How dividing cells retain hallmarks of interphase transcription to preserve their identity is incompletely understood.
APC/C sustains stem cell identity
To understand how pluripotency is preserved through cell division, we fused green fluorescent protein (GFP) to the OCT4 (also known as POU5F1) locus of human embryonic stem (ES) cells. Diploid OCT4-GFP human ES cells responded to differentiation cues with an efficiency similar to that of their untagged counterparts (Extended Data Fig. 1a, b). Using lentiviral infection with pooled short hairpin (sh)RNAs, we depleted about 900 enzymes and effectors of ubiquitylation, which control cell division and differentiation14; propagated OCT4–GFP human ES cells in pluripotency medium, or briefly induced differentiation by neural conversion; and then deep-sequenced populations with low versus high levels of OCT4–GFP (Fig. 1a). shRNAs that decreased OCT4–GFP abundance in self-renewing human ES cells target pluripotency factors, whereas shRNAs that sustained OCT4–GFP expression upon neural conversion deplete proteins that are needed for robust differentiation.
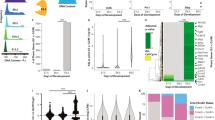
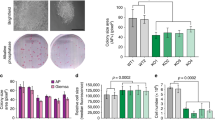
a, Schematic of the ultracomplex shRNA screen. hESC, human ES cell. b, shRNA screen identifies genes that are important for pluripotency. Each dot (n = 886 unique genes) represents the P value of a gene (two-sided Mann–Whitney U test, not corrected for multiple hypothesis testing), calculated from comparing the collection of shRNAs that target each gene to all negative-control shRNAs measured in each subpopulation (low versus high levels of OCT4–GFP). Orange, genes that encode enzymes or effectors of K11/K48 branched-chain synthesis; red, genes that encode deubiquitylases that oppose K11/K48-specific E3 ligases; blue, genes that encode DNA-repair enzymes; and green, positive controls. UFD2 is also known as UBE4B. Knockdown of genes indicated below and to the left of zero results in lower levels of OCT4–GFP; depletion of genes indicated above and to the right of zero maintains or increases the level of OCT4–GFP. c, Western blot of pluripotency markers upon APC/C-subunit knockdown in asynchronous H1 human ES cells. This experiment was performed five independent times with similar results. siCTRL, control small interfering (si)RNA; siAPC2, siCDC20, siUBE2C and siUBE2S denote siRNAs against APC2, CDC20, UBE2C and UBE2S, respectively. siUBE2C/S, siRNA against UBE2C and UBE2S. d, Interaction network of APC/C, WDR5 and USP44. Values listed in parentheses are total spectral counts of tryptic peptides of indicated proteins; values separated by a solidus denote proteins that coprecipitate with APC3 (left) or USP44 (right).
We recovered the positive-control OCT4, as well as known stem-cell E3 ligases such as DDB1, TRIM28 and UBR515,16,17, as pluripotency factors (Fig. 1b, Extended Data Fig. 1c). Consistent with the need for human ES cells to preserve genomic and proteomic integrity, we identified proteins involved in DNA repair (DDB1, RNF168 and USP7) and quality-control pathways (BAG6, HUWE1, PSMA1, PSMA6, UBR5 and UBXN7). Many of the enzymes of the latter pathways bind or produce K11/K48-branched ubiquitin chains18, which we confirmed in human ES cells (Extended Data Fig. 1d). Physiological pairs of E3 ligases and deubiquitylases (such as HUWE1 and USP7) clustered according to their opposing activities. Importantly, the APC2 subunit of APC/C was required for pluripotency, whereas the counteracting deubiquitylase USP4419 supported differentiation (Fig. 1b, Extended Data Fig. 1c, e). Other subunits of APC/C and APC/C-specific E2 enzymes scored as pluripotency factors, with P values that were slightly below our stringent screen cut-off (Extended Data Fig. 1c).
We confirmed that the depletion of subunits of APC/C, of the mitotic coactivator of APC/C (CDC20) or of APC/C-specific E2 enzymes inhibited human ES cell pluripotency, as revealed by decreased levels of OCT4 and NANOG (Fig. 1c, Extended Data Fig. 2a–c). Although less pronounced than its effects on protein levels, depletion of APC2 reduced the abundance of OCT4 and NANOG mRNA (Extended Data Fig. 2d). Human ES cells arrested in S phase and unable to enter mitosis did not require APC/C for pluripotency (Extended Data Fig. 2e), indicating that APC/C acts during cell division. However, it was unlikely that APC/C inhibition interfered with pluripotency simply by stalling mitotic progression, as loss of the APC/C-specific E2 enzyme UBE2C diminished OCT4 and NANOG levels without affecting the G2/M population (Fig. 1c, Extended Data Fig. 2f). Collectively, these findings indicated that the essential mitotic regulator APC/C also helps to preserve the stem-cell state, identifying APC/C as a strong candidate for maintaining cell identity through cell division.
APC/C works with WDR5 in human ES cells
We speculated that the identification of APC/C or USP44 substrate adaptors required for pluripotency might point to ubiquitylated proteins that preserve human ES cell identity. Using mass spectrometry, we found that USP44—in addition to known partners—also engaged WDR5, a chromatin-associated factor that binds methylated histone H3K4 at active interphase promoters6,7,20 (Fig. 1d). Endogenous APC/C also interacted with WDR5 during mitosis (Fig. 1d), which we confirmed by reciprocal purification of WDR5 (Extended Data Fig. 3a). In addition, mitotic WDR5 bound the transcription factor TFIID (which includes TBP), as well as chromatin remodellers INO80 and CHD1 (Extended Data Fig. 3a).
As with APC/C and TFIID–TBP21, depleting WDR5 diminished OCT4 and NANOG levels in human ES cells (Extended Data Fig. 3b). Human ES cells that are unable to enter mitosis did not require WDR5 for pluripotency (Extended Data Fig. 2e), which suggests that WDR5 acts during cell division. Consistently, the loss of WDR5 in human ES cells decreased the levels of K11-linked, as well as K11/K48-branched, ubiquitin chains—the mitotic products of APC/C18—to an extent similar to that seen after depletion of APC2 (Extended Data Fig. 3b). As in mouse ES cells20, loss of WDR5 did not affect mitotic duration (Extended Data Fig. 3c), but codepletion of WDR5 and APC2 caused human ES cells to die shortly after exiting mitosis (Extended Data Fig. 3d–g). These findings suggested that WDR5 cooperates with APC/C to ensure human ES cell identity and survival, and does not impinge on the role of APC/C in controlling cell division.
Reciprocal immunoprecipitations of endogenous proteins from somatic cells showed that APC/C, WDR5, and TBP engage each other during early mitosis, when APC/C binds CDC20 (Fig. 2a, b). A similar mitotic increase in the interaction between APC/C and WDR5 was seen in human ES cells (Extended Data Fig. 3h). Sequential affinity purifications revealed that APC/C, WDR5 and TBP were part of the same complex (Fig. 2c), the formation of which depended on WDR5 (Fig. 2d). In contrast to APC/C, WDR5 engaged USP44 also during interphase (Extended Data Fig. 3i).
a, Immunoprecipitation (IP) of endogenous APC3 from HeLa cells reveals that APC/C binds WDR5 and TBP in mitosis. Prometaphase HeLa cells were released into fresh medium to restart the cell cycle. This experiment was performed three independent times with similar results. b, Immunoprecipitation of endogenous WDR5 from HeLa cells confirms that WDR5 associates with APC/C subunits and TBP in mitosis. This experiment was performed three independent times with similar results. c, Sequential immunoprecipitations of APC/C in complex with Flag-tagged WDR5 from mitotic HEK293T cells reveal that APC/C–WDR5 and TBP form a ternary complex. Flag–WDR5 was first purified from prometaphase cells, and next purified with anti-APC3. This experiment was performed once. d, Endogenous APC3 immunoprecipitations from control versus WDR5-depleted human ES cells show that the association of APC/C with TBP is bridged through WDR5. This experiment was performed twice with similar results. siWDR5, siRNA against WDR5. Asterisk denotes nonspecific band. e, The approximately 20 Å resolution negative-stain electron-microscopy model corroborates the association of WDR5 with the catalytic core of APC/C. f, Flag–WDR5 purified from mitotic HeLa cells contains active APC/C. Flag-tagged wild-type (WT) WDR5 or Flag–WDR5(ΔWIN) were purified from mitotic HeLa cells, and incubated with E1, UBE2C, UBE2S, ubiquitin, ATP and 35S-labelled geminin. This experiment was performed two independent times with similar results. Autorad., autoradiography.
WDR5 uses distinct surfaces to recognize WDR5-binding motifs (WBMs) and WDR5-interacting (WIN) motifs6. Disrupting the ability of WDR5 to bind WIN motifs (WDR5(ΔWIN)) blocked the association of WDR5 with APC/C and USP44, but not with TBP (Extended Data Figs. 3i, 4a–c). Accordingly, the compound MM-102—which targets the site on WDR5 that binds the WIN motif22—prevented WDR5 from binding APC/C (Extended Data Fig. 4d), and WDR5(ΔWIN) did not sustain human ES cell pluripotency (Extended Data Fig. 4e). The ability of WDR5 to detect WBMs is not required for APC/C recognition, but is needed to bind TBP (Extended Data Fig. 4a, c).
Crosslinking experiments revealed that WDR5, but not WDR5(ΔWIN), binds APC/C close to CDC20 and the catalytic site that is composed of APC2 and APC11 (Extended Data Fig. 5a). Using in vitro translation, we identified APC2 as a specific binding partner of WDR5 (Extended Data Fig. 5b, c). We confirmed these findings by negative-stain electron microscopy, which showed that WDR5 is situated near CDC20 and docks against APC2 and APC11 (Fig. 2e).
Despite the proximity of WDR5 to the active site of APC/C, we could not detect APC/C-dependent ubiquitylation of WDR5 nor did excess WDR5 prevent the modification of APC/C substrates (Extended Data Fig. 6a, b). Instead, mitotic WDR5 complexes—which contain APC/C (Fig. 2b, Extended Data Fig. 3a)—supported the in vitro ubiquitylation of canonical APC/C substrates (Fig. 2f, Extended Data Fig. 6c). Mitotic WDR5 also coprecipitated K11-linked chains produced in cells (Extended Data Fig. 6d), which was dependent upon UBE2S (Extended Data Fig. 6e). We conclude that WDR5 binds active APC/C without being ubiquitylated itself, which suggests that WDR5 is a coadaptor that delivers APC/C to specific (probably chromatin-bound) substrates.
APC/C–WDR5 polyubiquitylates histones
To identify substrates of the APC/C–WDR5 complex, we used an approach that was previously established for SCF E3 ligases23. We fused WDR5 to the ubiquitin-binding domains of HHR23B or UBQLN2, which detect K11/K48-branched chains produced by APC/C18, and purified both constructs under conditions of low or high APC/C activity. Ubiquitylated substrates were expected to be trapped by both fusions in cells with active APC/C. These experiments identified histones as likely APC/C–WDR5 substrates (Fig. 3a).
a, Mass spectrometry of WDR5–HHR23B and WDR5–UBQLN2 traps identifies histones as candidate substrates. Traps were affinity-purified from prometaphase (PM) or anaphase (ANA) HeLa cells with low or high APC/C activity, respectively. TSC, total spectral counts. b, APC/C–CDC20 purified from mitotic HeLa S3 cells ubiquitylates recombinant human (Homo sapiens, Hs) H2A–H2B dimers. This experiment was performed four independent times with similar results. c, APC/C–WDR5 ubiquitylates H2B in polynucleosomes (nuc.) purified from HeLa cells and is inhibited by the APC/C inhibitor EMI1. This experiment was performed three independent times with similar results. d, APC/C–WDR5 ubiquitylates multiple Lys residues in histones, as seen with Lys-free ubiquitin (K0). This experiment was performed two independent times with similar results. e, APC/C-dependent ubiquitylation of H2B requires CDC20 in vitro. APC/C was purified from mitotic HeLa cells depleted of CDC20 or WDR5. This experiment was performed once. f, Ubiquitylation of H2B by APC/C is dependent on UBE2C and UBE2S, and inhibited by EMI1. This experiment was performed two independent times with similar results. g, Endogenous H2B is modified with K11/K48-branched chains, as seen by denaturing purification from synchronized HeLa cells. This experiment was performed three independent times with similar results. STLC, S-trityl-l-cysteine. h, Mitotic K11/K48 modification of endogenous H2B in human ES cells is dependent on UBE2C and UBE2S. This experiment was performed two independent times with similar results. i, Proteasome inhibition stabilizes mitotic K11/K48-modified H2B in H1 human ES cells. This experiment was performed two independent times with similar results. CFZ, carfilzomib.
In vitro reconstitution using human histone H2A–H2B dimers and H3–H4 tetramers, or Xenopus laevis H2A–H2B dimers and octamers, revealed efficient APC/C-dependent ubiquitylation of H2A, H2B and H3, but not of H4 (Fig. 3b, Extended Data Fig. 7a–c). H2A–H2B dimers, octamers and polynucleosomes were also strongly ubiquitylated by WDR5-bound APC/C and by endogenous APC/C purified from human ES cells (Fig. 3c, Extended Data Fig. 7b–d). Histone polyubiquitylation occurred at multiple sites (Fig. 3d), including K120 of H2B—the monoubiquitylation of which leads to transcriptional activation, and is negatively regulated by USP4424.
In contrast to mitotic APC/C, APC/C obtained from asynchronous or S-phase cells did not modify histones (Extended Data Fig. 7e). APC/C-dependent polyubiquitylation of histones was also blocked by the depletion of CDC20 (the mitotic coactivator of APC/C), by the addition of the APC/C inhibitor EMI1 or mutation of the K11 of ubiquitin (Fig. 3e, f, Extended Data Fig. 7f, g). H2B ubiquitylation was outcompeted by a canonical APC/C substrate, but less so by a D-box mutant substrate (Extended Data Fig. 7h), which indicates that histones are recognized by the D-box coreceptor composed of CDC20 and APC1025.
Denaturing purifications of K11/K48-branched chains revealed abundant ubiquitylation of endogenous H2B during early mitosis, at a time when CDC20 is decorated with such conjugates (Fig. 3g). Underscoring the role of APC/C, H2B modification with K11/K48-linked chains was strongly reduced by UBE2C and UBE2S depletion (Fig. 3h). Ubiquitylated H2B accumulated upon proteasome inhibition (Fig. 3i, Extended Data Fig. 7i), consistent with K11/K48-branched conjugates targeting proteins for degradation18,26. We conclude that APC/C–WDR5 modifies multiple histones with K11/K48-branched ubiquitin chains during mitosis.
APC/C acts at transcription start sites
As total histone levels did not drop during mitotic exit (Extended Data Fig. 8a), we hypothesized that APC/C–WDR5 targets histones at select chromosome locations. To identify this population, we performed genome-wide micrococcal-nuclease chromatin immunoprecipitation with sequencing (MNase ChIP–seq) analysis of K11-linked chains, WDR5 and TBP in prometaphase human ES cells. Because the vast majority of K11 linkages are assembled during mitosis by APC/C18,27, tracking this type of chain enabled us to monitor APC/C even if it interacted with its targets only transiently. MNase was used, as sonication fragmented polymeric ubiquitin chains and reduced the specific ChIP–seq signal (Extended Data Fig. 8b).
Notably, K11-linked and K11/K48-branched chains (that is, active APC/C) accumulated at specific genes in mitotic human ES cells that were co-occupied by WDR5 and TBP (Fig. 4a, b, Extended Data Fig. 8c–e). Chromatin-bound K11-linked chains were abundant during early mitosis (when APC/C is activated by CDC20), but were undetectable during late G1 or early S phase, when APC/C is inactive (Fig. 4c–e). By contrast, WDR5 and TBP were found at these promoters throughout the cell cycle (Fig. 4b). Depletion of CDC20, UBE2S or WDR5, and chemical inhibition of WDR5, strongly reduced K11-linked chains at APC/C–WDR5 target genes (Fig. 4f, Extended Data Fig. 8f, g). By heterologous expression of CDC20 and WDR5, we showed that mitotic APC/C–WDR5 also associated with specific genes in somatic cells (Extended Data Fig. 8h).
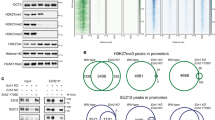
a, Genome browser track of E2F3. MNase ChIP–seq of indicated antibodies were performed from mitotic H1 human ES cells. b, K11 is deposited at select TSSs co-occupied by WDR5 in human ES cells. Heat map of co-occupied genes at TSSs from MNase ChIP–seq experiments of indicated antibodies. H1 human ES cells were collected after STLC treatment (mitosis) and after an 8-h release (late G1/S phase). CPM, counts per million. c, Genome browser track of E2F3 from MNase ChIP–seq of anti-K11 in human ES cells throughout a mitotic release. d, Flow cytometry analysis of H1 human ES cells upon mitotic synchronization and release into fresh medium (top). Metagene analysis of K11- and WDR5-occupied TSSs (middle). Heat map of individual K11- and WDR5-occupied TSSs from anti-K11 MNase ChIP–seq experiments throughout a mitotic release (bottom). e, Anti-K11 MNase ChIP–qPCR validates MNase ChIP–seq findings that K11 is deposited only during mitosis in H1 human ES cells. The same extract used in c was used for this experiment. f, Depletion of CDC20 or WDR5 causes robust depletion of K11 chains at select TSSs. g, MNase ChIP–seq from HUES64 human ES cells reveals that endogenous targets of APC/C–WDR5 are strongly enriched in binding sites for MYC, OCT4 and NANOG. h, Loss of APC/C–WDR5 function interferes with expression of genes marked with K11-linked chains in H1 human ES cells. Poly(A)-selected RNA was purified from asynchronous H1 human ES cells transfected with control siRNA or siRNA against WDR5 for 48 h and subjected to RNA sequencing. TPM, transcripts per million. i, Real-time qPCR analysis of nascent RNA reveals APC/C–WDR5 target genes are reactivated upon mitotic exit dependent on WDR5. Mitotic H1 human ES cells were treated with or without 50 μM MM102 and supplemented with 20 μM Z-VAD-FMK. Cells were released into fresh medium containing DMSO or 50 μM MM102. Real-time qPCR experiments were performed with oligonucleotides spanning intron–exon junctions. Values represent the mean of independent replicates ± s.e.m. (n = 3 for t = 15 min, n = 4 for t = 30, 60 and 480 min and n = 5 for t = 0, 120 and 240 min).
The majority of APC/C–WDR5 target sites were within 100 base pairs of the transcription start site (TSS); this location contains TBP-binding sites, as we confirmed for select targets by ChIP with quantitative PCR (ChIP–qPCR) (Extended Data Fig. 8i, j). Gene ontology (GO) analyses revealed that most APC/C–WDR5 target genes encode proteins that are involved in ribosome function (GO: 0003735, P = 1.2 × 10−56) and mRNA translation (GO: 0006413, P = 2.2 × 10−59). These genes are among the very first to be expressed upon mitotic exit4, dependent upon WDR5 and MYC6,28. Accordingly, APC/C–WDR5 target genes were strongly bound by the stem-cell transcription factors MYC, OCT4 and NANOG (Fig. 4g, Extended Data Fig. 8k), whereas transcription factors linked to differentiation did not accumulate at these sites (Extended Data Fig. 9). When we compared the set of APC/C–WDR5 target genes from HEK293T cells with gene-expression profiles, we noticed strong overlaps with human ES cell lines (Extended Data Fig. 10a).
Given the enrichment of APC/C–WDR5 at the TSSs of pluripotency genes and the requirement for this complex for self-renewal, we asked whether APC/C–WDR5 controls the transcription of its target genes. Notably, depletion of WDR5 strongly downregulated only those genes that were marked by K11-linked chains, WDR5 and TBP during mitosis (Fig. 4h, Extended Data Fig. 10b, c). Real-time qPCR analyses of nascent mRNAs using oligonucleotides that span intron–exon junctions showed that APC/C–WDR5 target genes were expressed immediately upon mitotic exit, dependent on WDR5 (Fig. 4i, Extended Data Fig. 10d). APC/C–WDR5 target genes are expressed at high levels (Extended Data Fig. 10e), and hence, particularly reliant on rapid reactivation after mitosis. Polyubiquitylation by APC/C–WDR5 therefore promotes early postmitotic expression of genes controlled by stem cell transcription factors.
APC/C recruits p97 and the proteasome
Consistent with K11/K48-branched chains recruiting the cellular degradation machinery18,26, the p97 adaptor UBXN7 and proteasome subunits scored in our screen (Fig. 1b). The p97–UBXN7 complex captured K11/K48-modified H2B in vitro (Extended Data Fig. 10f) and strongly bound K11/K48-ubiquitylated H2B in cells (Extended Data Fig. 10g). Moreover, p97 inhibition by NMS-873 caused the same strong increase in K11/K48-ubiquitylation of H2B as seen with proteasome inhibition (Extended Data Fig. 10h). Both MNase ChIP–seq and ChIP–qPCR experiments revealed that p97 and the proteasome were required for the loss of ubiquitylated proteins from the TSSs of APC/C–WDR5 target genes upon mitotic exit (Extended Data Fig. 10i, j). These findings suggest that APC/C–WDR5 might act by destabilizing histones at specific TSSs during mitosis.
Discussion
Our findings reveal a mechanism for how cell identity is preserved through cell division (Extended Data Fig. 10k). WDR5 and TBP bind promoters of genes transcribed in interphase. When cells enter mitosis, WDR5 and TBP remain associated with their targets but, instead of recruiting RNA polymerase II, they deliver APC/C to TSSs demarcated by the pluripotency factors MYC, OCT4 and NANOG. At these TSSs, APC/C decorates histones with K11/K48-branched chains, which attract p97 and the proteasome. We propose that subsequent histone degradation opens the TSSs for the rapid postmitotic expression of pluripotency genes. As it also triggers mitotic exit29, APC/C therefore tightly coordinates cell division and gene-expression regulation.
The newly identified cofactor WDR5 binds APC/C through the same surface as it uses to engage the MLL1 methyltransferase, another regulator of postmitotic gene expression30. Histone methylation might strengthen the interaction of WDR5 with promoters, which could facilitate subsequent recruitment of APC/C. WDR5 also engages OCT4, MYC and TFIID–TBP, all of which bind APC/C–WDR5 target genes and have vital roles in mitotic bookmarking. WDR5 thus appears to orchestrate distinct steps in the regulation of mitotic gene expression by mediating the recruitment of transcription factors, histone methylation and nucleosome destabilization.
Partial APC/C inhibition in neural progenitors triggered cell differentiation similar to that noted upon loss of APC/C–WDR5 in human ES cells31. Conversely, cellular reprogramming and somatic-cell nuclear transfer are more efficient during mitosis32,33, at times that coincide with APC/C–WDR5-dependent histone ubiquitylation. This further implies a role for APC/C–WDR5 in pluripotency control, which comes with practical implications: if APC/C–WDR5 acts in cancer stem cells as in human ES cells, combinations of APC/C and WDR5 inhibitors might impede the self-renewal of disease-driving cell populations and should be tested for their efficiency in cancer therapy.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized and investigators were not blinded to allocation during experiments and outcome assessment.
Mammalian cell culture
Human embryonic kidney (HEK)293T and HeLa cells were maintained in DMEM plus 10% fetal bovine serum. Plasmid transfections were performed using polyethylenimine (PEI) at a 1:3 ratio of DNA (in μg) to PEI (in μl at a 1 mg ml−1 stock concentration). siRNA transfections were performed using 40 nM of indicated siRNAs and a 1:400 dilution of RNAiMAX transfection reagent (Thermo Fisher, 13778150). Lentiviruses were produced in HEK293T cells by cotransfection of lentiviral and packaging plasmids using Lipofectamine 2000 transfection reagent (Thermo, 11668027). Viruses were collected 48 h after transfection, concentrated using the Lenti-X concentrator (Takara, 631232), aliquoted, and stored at −80 °C for later use. HEK293T cells were purchased directly from the Berkeley Cell Culture Facility (authenticated by short tandem repeat analysis). HeLa cells were not authenticated.
Human ES cells (WiCell, WA01/H1) were grown in mTeSR1 medium (StemCell Technologies, 85850) on human-ES-cell-qualified Matrigel-coated plates (Corning, 354277) with daily medium change. H1 cells were passaged by collagenase (StemCell Technologies, 07909) for routine maintenance or accutase (StemCell Technologies, 07920) for siRNA transfections, lentiviral infections or when single cells were required. For siRNA transfections, single-cell suspensions of H1 cells were generated by accutase treatment and 2–5 × 105 cells were seeded on a Matrigel-coated well of a 6-well plate with 1.8 ml of mTeSR1 containing 10 μM of Y-27632 (StemCell Technologies, 72308) and a 0.2 ml mixture of indicated siRNAs (at a final concentration of 40 nM) and a 1:400 dilution of RNAiMAX transfection reagent buffered in Opti-MEM. For lentiviral infections, single-cell suspensions of H1 cells were generated by accutase treatment and 1.5–3 × 105 cells were seeded on a Matrigel-coated well of a 6-well plate with 2 ml of mTeSR1 containing 10 μM of Y-27632, polybrene (at a final concentration of 6 μg ml−1), and lentiviruses produced from HEK293T cells for 2 h. The medium was immediately exchanged with 2 ml of fresh mTeSR1 containing 10 μM of Y-27632 only. Human ES cells were drug-selected 24–48 h after infection. H1 cells were positive for OCT4 and NANOG expression and karyotype analysis showed no chromosomal anomalies.
All cell lines were routinely tested for mycoplasma contamination using the MycoAlert mycoplasma detection kit (Lonza, LT07-218). All cell lines tested negative for mycoplasma.
Generation of OCT4-eGFP-P2A-PURO R human ES cells
The OCT4 locus was targeted for gene editing in H1 cells by TALE nucleases as previously described34. An in-frame fusion, consisting of enhanced GFP (eGFP) followed by the self-cleaving P2A peptide and the puromycin resistance gene (puromycin N-acetyltransferase), was generated at the C terminus of the OCT4 locus. In brief, single-cell suspensions of H1 cells were generated by accutase treatment and 1 × 107 cells were resuspended in ice-cold 1 × PBS with 40 μg of the DONOR plasmid and 5 μg each of the TALEN plasmids (T4 and T8). Cells were electroporated in a 0.4-cm cuvette at 250 V and 500 μF with the Gene Pulser II eletroporating system (Bio-Rad). Electroporated cells were immediately resuspended in mTeSR1, washed to remove lysed debris and seeded on 2 Matrigel-coated 15-cm plates in mTeSR1 containing 10 μM of Y-27632. H1 cells were selected for 10–14 days with puromycin (at a final concentration of 0.5 μg mg−1) 72 h after electroporation. Colonies were manually scored and transferred to fresh plates. A single allele of the OCT4 locus was fused with the eGFP-P2A-PUROR cassette as verified by Southern blot analysis (data not shown). Karyotype analysis was performed by WiCell.
Neural conversion of human ES cells
Neural induction of human ES cells were performed as previously described35, using STEMdiff Neural Induction Medium (StemCell Technologies, 05839). Single-cell suspensions of H1 cells were generated by accutase treatment and 1.5 × 106 cells were seeded in a well of 6-well plate with 4 ml of STEMdiff neural induction medium containing 10 μM Y-27632. Cells were treated with daily medium changes, and collected when indicated.
Ultracomplex shRNA screen
The shRNA library was constructed as previously described36. In brief, the shRNA library was divided into four sublibraries, cloned into lentiviral expression vectors and transfected into HEK293T cells with TransIT-293 transfection reagent (Mirus, MIR 2700) for virus production. Human ES cells were infected with lentiviruses overnight and cultured in mTeSR1 for six days or in mTeSR1 for six days followed by STEMdiff neural induction medium for one day. Human ES cells were then sorted by fluorescence-activated cell sorting using an INFLUX cell sorter (BD) at the Flow Cytometry Core Facility at UC Berkeley. Cells were sorted on the basis of the strength of their GFP expression into three populations. Sequencing libraries were prepared from sorted cells as previously described36, sequenced on a HiSeq 2000 (Illumina) and analysed using previously described scripts36.
Cell synchronization
HeLa cells were first synchronized in S phase by addition of thymidine (at a final concentration of 2 mM) for 24 h. S-phase cells were washed with 1 × PBS to remove excess thymidine and released into fresh medium (DMEM/10% FBS) for 3 h. To arrest cells in prometaphase, released cells were treated with STLC (Sigma, 164739) (at a final concentration of 5 μM) for 12–14 h. Finally, prometaphase cells were collected by vigorous pipetting, washed with 1× PBS and used for downstream applications, including immunoprecipitation assays and/or western blot analyses, or frozen in liquid nitrogen and stored at −80 °C for later use. For cell-cycle studies, prometaphase cells were released into fresh medium and collected at the indicated time points. For drug inhibition studies, cells were released into medium containing 2 μM carfilzomib (Selleck, PR-171), 20 μM (R)-MG132 (Cayman, 13697) and/or 10 μM NMS-873 (Sigma, SML1128) for indicated times. For depletion studies, HeLa cells were transfected with 40 nM of indicated siRNAs and a 1:400 dilution of RNAiMAX transfection reagent (Thermo Fisher, 13778150) 24 h before synchronization.
Mitotic enrichment of HEK293T cells and H1 cells was achieved by adding STLC (at a final concentration of 5 μM) to the culture medium for 14–16 h.
Purification of APC/C and APC/C–WDR5 complexes
Human APC/C and APC/C–WDR5 complexes were purified from HeLa extracts synchronized in prometaphase (see ‘Cell synchronization’). To purify APC/C–WDR5, HeLa cells were first PEI-transfected with 5 μg of pCMV 3×Flag–WDR5 (per 15-cm plate) for 24 h before synchronization. Collected prometaphase pellets were lysed in lysis buffer (20 mM HEPES, pH 7.4, 5 mM KCl, 150 mM NaCl, 1.5 mM MgCl2, 0.1% Nonidet P-40, 1 × cOmplete protease inhibitor cocktail (Roche, 04693159001) and 1 μl of benzonase (Millipore, 70746) per 15-cm plate). Detergent lysed cells were then subjected to a high-speed spin (20,000g) to remove cellular debris and the clarified extract was precleared with protein G-agarose resin (Roche, 11719416001). APC/C was purified with anti-CDC27 antibody (sc-9972, SCBT) precoupled to protein G-agarose resin for 3 h at 4 °C, and APC/C–WDR5 was purified with anti-Flag M2 affinity resin (Sigma, A2220) for 1.5 h at 4 °C. APC/C-coupled beads were washed 5× with lysis buffer (minus inhibitors and benzonase) before use.
Purification of recombinant proteins
WDR5 and WDR5(ΔWIN) were cloned into a pMAL expression vector containing a C-terminal 6×His tag and expressed in Escherichia coli BL21-CodonPlus (DE3)RIL cells. Transformed cells were grown at 37 °C to an optical density at 600 nm (OD600) of 0.5 in LB broth containing 100 μg ml−1 ampicillin, 34 μg ml−1 chloramphenicol and 0.2% glucose, chilled on ice for 30 min, induced with 100 μM isopropyl β-d-1-thiogalactopyranoside (IPTG) for 6 h at 16 °C, and collected by centrifugation. Collected cells were resuspended with lysis buffer (20 mM HEPES, pH 7.4, 300 mM NaCl, 2 mM 2-mercaptoethanol (BME), 1 mM EDTA, 10% glycerol, 0.2 mg ml−1 lysozyme, 1 mM phenylmethylsulfonyl fluoride (PMSF) and 0.1% Triton X-100), incubated on ice for 30 min, sonicated and clarified by high-speed centrifugation. The clarified extract was supplemented with 20 mM imidazole and bound to Ni-NTA resin (Qiagen, R90110) (2 ml of slurry per 1 l of bacterial culture) for 1 h at 4 °C. The resin was then washed 5× with wash buffer (20 mM HEPES, pH 7.4, 300 mM NaCl, 2 mM BME, 1 mM EDTA, 10% glycerol and 20 mM imidazole) and eluted 2 × with elution buffer (20 mM HEPES, pH 7.4, 300 mM NaCl, 2 mM BME, 1 mM EDTA, 10% glycerol and 300 mM imidazole). The elutions were pooled, dialysed overnight in dialysis buffer (20 mM HEPES, pH 7.4, 300 mM NaCl, 2 mM BME, 1 mM EDTA and 10% glycerol), concentrated, aliquoted, snap-frozen in liquid nitrogen and stored at −80 °C for later use.
Securin and its variants were cloned into a pET28 expression vector containing an N-terminal 6×His tag followed by a TEV-protease cleavage site and expressed in LOBSTR BL21(DE3)-RIL cells. Transformed cells were grown at 37 °C to an OD600 of 0.5 in LB broth containing 100 μg ml−1 ampicillin and 34 μg ml−1 chloramphenicol, chilled on ice for 30 min and induced with 100 μM IPTG for 14–16 h at 16 °C. Induced cells were centrifuged, resuspended in lysis buffer (20 mM HEPES, pH 7.4, 300 mM NaCl, 2 mM BME, 10% glycerol, 0.2 mg ml−1 lysozyme, 1 mM PMSF and 0.1% Triton X-100), incubated on ice for 30 min, sonicated and clarified by high-speed centrifugation. The clarified extract was supplemented with 20 mM imidazole and bound to Ni-NTA resin (2 ml of slurry per 1 l of culture) for 1 h at 4 °C. The resin was then washed 5× with wash buffer (20 mM HEPES, pH 7.4, 300 mM NaCl, 2 mM BME, 10% glycerol, 0.1% Triton X-100 and 20 mM imidazole) and eluted by TEV cleavage. The eluate was desalted using a PD10 column, concentrated, aliquoted, snap-frozen and stored at −80 °C for later use.
p97 was cloned into a pMAL expression vector and expressed in BL21-CodonPlus (DE3)RIL cells. Transformed cells were grown at 37 °C to an OD600 of 0.5 in LB broth containing 100 μg ml−1 ampicillin and 34 μg ml−1 chloramphenicol, chilled on ice for 30 min and induced with 0.5 mM IPTG overnight at 18 °C. Induced cells were centrifuged, resuspended in lysis buffer (20 mM Tris 7.4, 300 mM NaCl, 5% glycerol, 0.2 mg ml−1 lysozyme, 1 mM PMSF and 0.1% Triton X-100), incubated on ice for 30 min, sonicated and clarified by high-speed centrifugation. The clarified extract was bound to amylose resin (NEB, E8021) (2 ml of slurry per 1 l of culture) for 45 min at 4 °C. The resin was then washed 3× with 1× PBS, resuspended in 1× PBS containing 2 mM DTT, and stored at 4 °C for up to 1 month. Recombinant 6×His–p47 (also known as NSFL1C) and 6×His–UBXN7 were purified using previously described methods18.
In vitro transcription and translation
All in vitro synthesized substrates were cloned under the SP6 promoter. The corresponding plasmids can be found in Supplementary Table 1. 35S-labelled substrates were generated by incubating 3 μl (400 ng) of plasmid DNA in 20 μl of rabbit reticulocyte lysate (Promega, L2080) supplemented with 2 μl of 35S-Met (PerkinElmer, NEG009H001MC) for 1 h at 30 °C. Reactions were terminated by rapid dilution with 1× PBS. 35S-labelled substrates were used for in vitro ubiquitylation assays and/or MBP binding studies.
In vitro ubiquitylation
In vitro ubiquitylation assays were performed in a 10 μl reaction volume: 0.25 μl of 10 μM E1 (250 nM final), 1 μl of 10 μM UBE2C (1 μM final), 1 μl of 10 μM UBE2S (1 μM final), 1 μl of 10 mg ml−1 ubiquitin (1 mg ml−1 final) (Boston Biochem, U-100H), 1 μl of 100 mM DTT, 1.5 μl of energy mix (150 mM creatine phosphate, 20 mM ATP, 20 mM MgCl2, 2 mM EGTA, pH to 7.5 with KOH), 2.25 μl of 1 × PBS, 1 μl of 10 × ubiquitylation assay buffer (250 mM Tris 7.5, 500 mM NaCl, and 100 mM MgCl2) and 3 μl of substrate (in vitro translated or recombinant) were premixed and added to 5 μl of APC/C- or APC/C–WDR5-purified bed resin (see ‘Purification of APC/C and APC/C–WDR5 complexes’). Reactions were performed at 30 °C with shaking for 30 min, unless noted otherwise. Reactions were stopped by adding 2× urea sample buffer and resolved on SDS-acrylamide gels. E1, UBE2C and UBE2S were purified as previously described26. Recombinant human H2A–H2B dimers (NEB, M2508S), recombinant X. laevis H2A–H2B dimers and octamers, recombinant human H3–H4 tetramers (NEB, M2509S), or purified human nucleosomes (EpiCypher, 16-0003) were used at a final concentration of 500 nM.
MBP binding studies
For in vitro transcription and translation binding assays, 10 μl of 35S-labelled substrate was diluted down to 400 μl with prechilled 1× PBS containing 0.1% Nonidet P-40 and mixed with 2 μl of 1 mg ml−1 of MBP-fused bait (see ‘Purification of recombinant proteins’) and 8 μl of amylose slurry (NEB, E8021). The binding was performed for 2 h at 4 °C with mixing, and the amylose resin was subsequently washed 3× with 1× PBS. The bound prey was eluted with 2× urea sample buffer, resolved on an SDS-acrylamide gel and visualized by a Typhoon scanner.
For coadaptor-bound p97 binding studies, coadaptor-bound p97 resin was made by mixing 0.1 ml of p97-coupled amylose slurry (see ‘Purification of recombinant proteins’) with 0.2 ml of recombinant 6×His–p47 or 6×His–UBXN7 and 0.3 ml of 1× PBS containing 4 mM DTT for 45 min at 4 °C. The resin was washed 3× with 1× PBS containing 2 mM DTT and stored at 4 °C for up to 2 weeks. Ubiquitylated H2A–H2B dimers (see ‘In vitro ubiquitylation’) were added to 6 μl of coadaptor-bound p97 slurry brought up in 0.6 ml of 1× PBS, incubated for 20 min at 4 °C, washed 5× with 1× PBS, eluted with 2× urea sample buffer and resolved on an SDS-acrylamide gel.
Crosslinking studies
APC/C complexes were first purified from HeLa cells synchronized in prometaphase. Before crosslinking, a 200 μM working stock of the sulfhydryl-reactive and homobifunctional crosslinker 1,4-bismaleimidobutane (BMB) was prepared in DMSO and a 20 μM solution of recombinant MBP–WDR5 was pretreated with tris(2-carboxyethyl)phosphine (TCEP) (at a final concentration of 1 mM) in a 20 μl reaction volume. Ten microlitres of purified APC/C slurry (see ‘Purification of APC/C and APC/C–WDR5 complexes’) was mixed with TCEP-treated MBP–WDR5 (at a final concentration of 2 μM) and BMB (at a final concentration of 20 μM) and incubated for 30 min at 22 °C with shaking. Reactions were stopped by adding 2× urea sample buffer and resolved on SDS-acrylamide gels.
K11/K48 denaturing immunoprecipitations
Denaturing K11/K48-linked ubiquitin immunoprecipitations were performed from cells arrested in prometaphase. Three 15-cm plates of confluent cells were collected and lysed in equal pellet volume with urea lysis buffer (20 mM Tris 7.5, 135 mM NaCl, 10% glycerol, 8 M urea, 1% Triton X-100, 5 μM carfilzomib (Selleck, PR-171), 10 mM N-ethylmaleimide (NEM), 1 × phosSTOP (Roche, 4906837001) and 1 × cOmplete protease inhibitor cocktail (Roche, 04693159001)), rotated for 1 h at room temperature, sonicated with a microtip sonicator (15 pulses at 50 amps), diluted 2-fold in dilution buffer (20 mM Tris 7.5, 135 mM NaCl, 10% glycerol, 5 μM carfilzomib, 10 mM NEM, 1 × phosSTOP and 1 × cOmplete protease inhibitor cocktail) and clarified for 5 min at low speed (2,400g). Clarified extracts were incubated with 20 μg of anti-K11/K48 bispecific ubiquitin antibody or control normal mouse IgG and 40 μl of protein G-agarose slurry for 3 h at room temperature. Beads were washed 10× with dilution buffer, eluted with 2× urea sample buffer, and resolved on SDS-acrylamide gels.
Mass spectrometry
Mass spectrometry was performed on immunoprecipitates prepared from HEK293T cells. In brief, 20 15-cm plates of HEK293T cells were PEI-transfected (if indicated), grown to confluence, synchronized (if indicated), collected and lysed in lysis buffer (20 mM HEPES, pH 7.4, 5 mM KCl, 150 mM NaCl, 1.5 mM MgCl2, 0.1% Nonidet P-40 and 1 × cOmplete protease inhibitor cocktail). Lysed extracts were clarified by high-speed centrifugation, precleared with protein G-agarose slurry and bound to indicated antibodies pre-coupled to protein G-agarose resin (for immunoprecipitations of endogenous proteins) or anti-Flag M2 affinity resin (for immunoprecipitations of overexpressed proteins). Immunoprecipitates were then washed and eluted 3× at 30 °C with 0.5 mg ml−1 of 3× Flag peptide (Sigma, F4799) buffered in 1× PBS plus 0.1% Triton X-100. Elutions were pooled and precipitated overnight at 4 °C with 20% trichloroacetic acid. Immunoprecipitates were then pelleted, washed 3× with an ice-cold acetone/0.1 N HCl solution, dried, resolubilized in 8 M urea buffered in 100 mM Tris 8.5, reduced with TCEP (at a final concentration of 5 mM) for 20 min, alkylated with iodoacetamide (at a final concentration of 10 mM) for 15 min, diluted 4-fold with 100 mM Tris 8.5, and digested with 0.5 mg ml−1 of trypsin supplemented with CaCl2 (at a final concentration of 1 mM) overnight at 37 °C. Trypsin-digested samples were submitted to the Vincent J. Coates Proteomics/Mass Spectrometry Laboratory at UC Berkeley for analysis. Peptides were processed using multidimensional protein identification technology (MudPIT) and identified using a LTQ XL linear ion trap mass spectrometer. To identify high-confidence interactors, CompPASS analysis of the query mass spectrometry result was performed against mass spectrometry results from unrelated Flag immunoprecipitates performed in our laboratory.
For TMT labelling, samples were prepared in the same manner as previously described37. Following trypsin digestion, however, samples were desalted using a C18 column (Agilent, A57203), dried overnight, resuspended in 80 μl of 200 mM HEPES, pH 8.0 and quantified using the Pierce Quantitative Colorimetric Peptide Assay kit (Pierce, 23275) on a microplate reader. Peptides were then normalized to equal masses in 100 μl volumes with 200 mM HEPES, pH 8. TMT labelling was performed using the TMTsixplex Isobaric Mass Tagging Kit (Thermo Fisher, 90066) per the manufacturer’s instruction. Labelled peptides were combined in equal volumes, desalted, dried and identified using a Fusion Lumos mass spectrometer by the Vincent J. Coates Proteomics/Mass Spectrometry Laboratory.
Immunofluorescence microscopy
For immunofluorescence analysis of neural inductions, H1 cells and H1 cells undergoing neural conversion were seeded on Matrigel-coated 96-well plates in mTeSR1 or STEMdiff neural induction medium plus 10 μM Y-27632 for 24 h, washed with 1× PBS plus 1 mM MgCl2 and 1 mM CaCl2, fixed with 4% paraformaldehyde buffered in 1× PBS for 15 min, permeabilized in 1× PBS plus 0.1% Triton X-100 for 10 min, blocked in 10% FBS plus 0.1% Triton X-100 for 30 min and stained with indicated antibodies diluted in 10% FBS plus 0.1% Triton X-100. Images were taken on an Opera Phenix High-Content Screening System (PerkinElmer) using a 40× air objective and processed using Harmony High Content Imaging and Analysis Software (PerkinElmer).
Live-cell imaging
H2B–mCherry expressing H1 cells were transfected with indicated siRNAs and seeded on Matrigel-coated 8-chamber microscopy slides (Lab-TelII, 155409). Twenty-four to forty-eight hours after transfection, cells were imaged every 3 min for 12–14 h using a Zeiss LSM 710 confocal microscope with 20× objective. Mitotic cells were identified manually.
Analysis of cell-cycle progression
For DNA content analysis, single-cell suspensions were generated with trypsin, fixed for 15 min with 4% paraformaldehyde buffered in 1× PBS, washed with 1× PBS and incubated with 2 μg ml−1 of Hoescht 33342 buffered in 1× PBS for 30 min at room temperature with gentle rocking. Single cells were filtered through a mesh strainer and analysed using an LSRFortessa flow cytometer (Becton Dickinson). Cytometry data were processed using the FlowCytometryTools Python package and custom-built Python scripts.
Sonication and ChIP-qPCR analysis
Cells were resuspended in 1× PBS and fixed at room temperature with 1% formaldehyde (Fisher, UN1198) for 10 min or with 1.5 mM ethylene glycol bis(succinimidyl succinate) (EGS) for 20 min followed by 1% formaldehyde for an additional 10 min. Crosslinking reactions were quenched with 125 mM glycine buffered in 1× PBS for 10 min. Crosslinked cells were washed twice with 1× PBS, collected, snap-frozen and stored at −80 °C for later use. Collected pellets were resuspended in sonication buffer (50 mM Tris 8.0, 10 mM EDTA, 1% SDS and 1 × cOmplete protease inhibitor cocktail), incubated on ice for 15 min and pelleted at 2,000g. Pellets were washed 4× with sonication buffer and sonicated in 12 × 24-mm tubes (Covaris, 520056) at 150 W (peak power) using an S220 ultrasonicator (Covaris) with a duty factor of 20 and 200 cycles per burst for 30–35 cycles (30 s on and 30 s off). Sonicated extracts were clarified by high-speed centrifugation, snap-frozen and stored at −80 °C for later use. ChIP extracts were diluted 10-fold in dilution buffer (20 mM Tris 8.0, 167 mM NaCl, 1 mM EDTA, 1% Triton X-100 and 1× cOmplete protease inhibitor cocktail), precleared with protein G/A-agarose resin and bound overnight to the indicated antibodies (Supplementary Table 2) at 4 °C. Antibodies were pulled down by addition of BSA-blocked protein G/A-agarose resin. Beads were washed twice with low salt wash buffer (20 mM Tris 8.0, 150 mM NaCl, 2 mM EDTA, 1% Triton X-100 and 0.1% SDS), twice with high salt wash buffer (20 mM Tris 8.0, 500 mM NaCl, 2 mM EDTA, 1% Triton X-100 and 0.1% SDS), once with LiCl buffer (20 mM Tris 8.0, 250 mM LiCl, 1 mM EDTA, 1% deoxycholate and 1% Nonidet P-40) and twice with 1× TE. Samples were eluted twice at 30 °C with 1% SDS buffered in 1× TE. Eluates were pooled, treated with RNase A and reverse-crosslinked overnight at 65 °C. Samples were then treated with proteinase K, phenol:chloroform extracted, isopropanol precipitated and eluted in 10 mM Tris 8. Resuspended samples were quantified using the KAPA SYBR FAST Universal kit (Kapa Biosystems, KK406) on a QuantStudio 6 Flex Real-Time PCR System (Applied Biosystems). ChIP–qPCR primers used in this study can be found in Supplementary Table 3.
Real-time qPCR analysis
For real-time qPCR analysis, total RNA was purified from cells using the NucleoSpin RNA kit (Macherey-Nagel, no. 740955) or with acid phenol and reverse-transcribed using the Maxima First Strand cDNA Synthesis kit (Thermo Fisher, K1671). Expression levels were quantified using the Luna Universal qPCR Master Mix (NEB, M3003) on a QuantStudio 6 Flex Real-Time PCR System (Applied Biosystems). Real-time qPCR primers used in this study can be found in Supplementary Table 3.
Sonication and ChIP–seq analysis
For sonication and ChIP–seq analysis, samples were prepared as described in ‘Sonication and ChIP–qPCR analysis’. Libraries were constructed by the Functional Genomics Laboratory at UC Berkeley, multiplexed and sequenced by the Vincent J. Coates Genomics Sequencing Laboratory at UC Berkeley on a HiSeq2500 or a HiSeq4000 (Illumina). Alignments for the paired-end or single-read sequencing runs were performed against the hg19 reference genome using Bowtie2. ChIP peaks were called with MACS14. Downstream analyses were performed using Bedtools and Deeptools.
MNase ChIP–seq sample preparation
For MNase ChIP–seq analysis, human ES cells were collected by accutase treatment, washed once with ice-cold 1× PBS and resuspended in 1 ml of 1× PBS. Single-cell suspensions were crosslinked with 1% formaldehyde for 10 min at room temperature, quenched with glycine (at a final concentration of 125 mM) for 2 min, washed with 1× PBS, snap-frozen in liquid nitrogen and stored at −80 °C for later use. Frozen pellets were resuspended in an equal pellet volume of lysis buffer (1% SDS, 10 mM EDTA, 50 mM Tris 8.0, 1 × cOmplete protease inhibitor cocktail and 1 × phosSTOP), incubated on ice for 10 min, diluted 4-fold with dilution buffer (1% Triton X-100, 150 mM NaCl, 20 mM Tris 8.0, 2.5 mM CaCl2, 1× cOmplete protease inhibitor cocktail and 1× phosSTOP), digested with 150 units of MNase (Worthington, LS004798) per 200 μl of pellet volume for 5 min at 37 °C, quenched with 6 mM EDTA and 6 mM EGTA, spun at 20,000g to remove debris, aliquoted, snap-frozen in liquid nitrogen and stored at −80 °C for later use. MNase-digested chromatin was precleared with protein-G dynabeads (Thermo, 10003D) and bound to indicated antibodies overnight at 4 °C. Antibodies were immunoprecipitated by addition of BSA-blocked protein-G dynabeads. Beads were washed twice with low salt wash buffer (20 mM Tris 8.0, 150 mM NaCl, 2 mM EDTA, 1% Triton X-100 and 0.1% SDS), twice with high salt wash buffer (20 mM Tris 8.0, 500 mM NaCl, 2 mM EDTA, 1% Triton X-100 and 0.1% SDS), once with LiCl buffer (20 mM Tris 8.0, 250 mM LiCl, 1 mM EDTA, 1% deoxycholate and 1% Nonidet P-40), and twice with 1× TE. Samples were eluted twice at 30 °C with 1% SDS buffered in 1× TE. Eluates were pooled, treated with RNase A and reverse-crosslinked overnight at 65 °C. Samples were then treated with proteinase K, phenol:chloroform extracted, isopropanol precipitated and eluted in 10 mM Tris 8.
MNase ChIP–seq library construction
Purified DNA (see ‘MNase ChIP–seq sample preparation’) was quantified using a Fragment Analyzer (Agilent). Twenty-five nanograms of purified DNA was resuspended up to 50 μl in water. Ten microlitres of T4 DNA ligase buffer (NEB, B0202), 4 μl of 10 mM dNTPs, 5 μl of T4 DNA polymerase (NEB, M0203), 1 μl of Klenow DNA polymerase (NEB, M0210), 5 μl of T4 DNA polynucleotide kinase (NEB, M0201) and 25 μl of water were added to the diluted input DNA and incubated at 25 °C for 30 min. Samples were purified with Ampure XP beads (Beckman, A36881) and resuspended in 32 μl of water. Five microlitres of buffer 2 (NEB, B7002), 1 μl of 10 mM dATP, 3 μl of Klenow fragment (NEB, M0212) and 9 μl of water were added to the end-repaired DNA and incubated at 37 °C for 30 min. Samples were purified with Ampure XP beads (Beckman, A36881) and resuspended in 23 μl of water. Five microlitres of Truseq Y adaptors for paired-end sequencing (custom-made), 5 μl of 10× ligase buffer (NEB, B0202), 1.5 μl of T4 DNA ligase (NEB, M0202) and 15.5 μl of water were added to the 3′-adenylated DNA and incubated at room temperature for 1 h. Samples were purified with Ampure XP beads (Beckman, A36881) and resuspended in 30 μl of water. Three microlitres of adaptor-ligated DNA was used for PCR amplification (KAPA HiFi master mix, KK201).
MNase ChIP–seq and analysis
MNase ChIP–seq samples (see ‘ MNase ChIP–seq library construction’) were multiplexed and sequenced by the Vincent J. Coates Genomics Sequencing Laboratory at UC Berkeley on a HiSeq4000 (Illumina). Alignments for the single-read sequencing runs were performed against the hg19 reference genome using Bowtie2. ChIP peaks were called with MACS14. Downstream analyses were performed using Bedtools and Deeptools.
RNA-sequencing sample preparation and analysis
Total RNA was purified from cells with TRIzol (Thermo, 15596026) and digested with TURBO DNase (Thermo, AM2238). Total RNA was poly(A)-selected and sequencing libraries were constructed using the KAPA mRNA HyperPrep kit (KK8580) by the Functional Genomics Laboratory at UC Berkeley. Libraries were sequenced by the Vincent J. Coates Genomics Sequencing Laboratory at UC Berkeley on a HiSeq4000 (Illumina). Gene-expression analysis was performed using Kallisto against hg19 as the reference genome.
Bioinformatics
Identified ChIP peaks were subjected to bioinformatic analyses. GO enrichment analyses were performed using DAVID 6.8 (https://david.ncifcrf.gov). Comparison with SAGE data was performed using the CGAP-SAGE feature of DAVID, a web-based application (https://david.ncifcrf.gov).
Purification of phosphomimetic APC/C–CDC20 with WDR5 for negative-stain electron microscopy
Recombinant APC/C–CDC20 containing glutamate mutations that mimic phosphorylation38 was purified as previously described39. In brief, APC/C and CDC20 were expressed independently in High Five insect cells (Thermo Fisher Scientific) and colysed by mixing and sonication. Cleared lysate was treated to tandem Strep- and GST-affinity chromatography selections for APC2 and APC16, respectively. GST elution fractions containing APC/C–CDC20 were combined with TEV protease, HRV14 3C protease and purified MBP–Flag–WDR5–His containing a TEV proteolytic site N-terminal of the Flag tag. This mixture was further purified through Flag affinity chromatography and eluted with antigenic peptides.
Negative-stain electron microscopy
For negative-stain electron-microscopy studies, 125 μg of purified APC/C–CDC20–WDR5 eluate from Flag immunoprecipitations was loaded onto a 10–40% glycerol gradient containing 50 mM HEPES pH 8.0, 200 mM NaCl and 2 mM MgCl2. For particle fixation by GraFix40, the gradient also contained 0.025% and 0.1% glutaraldehyde in the lighter and denser glycerol solution, respectively, creating an additional glutaraldehyde gradient from top to bottom (0.025–0.1%). Centrifugation was performed at 34,000 rpm in a TH-660 rotor (Thermo Fisher Scientific) for 15 h at 0 °C and the solution was subsequently fractionated. APC/C particles were allowed to adsorb on a thin film of carbon, transferred onto a plasma-cleaned lacey grid (LC200-CU, Electron Microscopy Services) and then stained for 2 min with a 4% (w/v) uranyl formate solution. Micrographs were collected on a FEI Titan Halo at 300 KV with a Falcon 2 direct detector (FEI) (MPI of Biochemistry). The nominal magnification was 45,000×, resulting in an image pixel size of 2.37 Å per pixel on the object scale and data were collected in a defocus range of 1.5–3.5 μM. Particles were autopicked using Relion41. The contrast transfer function parameters were determined using CTFFIND442. Using Relion, particles were extracted from micrographs and subjected to 2D classification. Inconsistent class averages were removed before 3D classification in Relion.
Structural modelling was performed using UCSF Chimera (1.13.1)43. To identify electron microscopy density corresponding to WDR5, the electron microscopy reconstruction of APC/C–CDC20–WDR5 obtained from 3D classification in Relion was superimposed with a prior map from an APC/C–CDC20–substrate complex (EMDB-3385, ref. 44) low-pass-filtered to a comparable resolution. Although the resolution precludes definitive structural modelling, it allowed approximate, global placement of the crystal structures of WDR545, along with the APC2 winged-helix box and APC11 RING domains39,46, which are known to be mobile and to adopt distinct orientations when bound to different APC/C partner proteins.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
All original data are available from the corresponding author on request. ChIP–seq and RNA-sequencing data have been deposited at the Gene Expression Omnibus, accession code GSE122298.
Code availability
Custom Python scripts are available from the corresponding author on request.
References
Young, R. A. Control of the embryonic stem cell state. Cell 144, 940–954 (2011).
Michelotti, E. F., Sanford, S. & Levens, D. Marking of active genes on mitotic chromosomes. Nature 388, 895–899 (1997).
Teves, S. S. et al. A stable mode of bookmarking by TBP recruits RNA polymerase II to mitotic chromosomes. eLife 7, e35621 (2018).
Palozola, K. C. et al. Mitotic transcription and waves of gene reactivation during mitotic exit. Science 358, 119–122 (2017).
Hsiung, C. C. et al. A hyperactive transcriptional state marks genome reactivation at the mitosis–G1 transition. Genes Dev. 30, 1423–1439 (2016).
Thomas, L. R. et al. Interaction with WDR5 promotes target gene recognition and tumorigenesis by MYC. Mol. Cell 58, 440–452 (2015).
Wysocka, J. et al. WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell 121, 859–872 (2005).
Keyes, B. E. & Fuchs, E. Stem cells: aging and transcriptional fingerprints. J. Cell Biol. 217, 79–92 (2018).
Prescott, D. M. & Bender, M. A. Synthesis of RNA and protein during mitosis in mammalian tissue culture cells. Exp. Cell Res. 26, 260–268 (1962).
Martínez-Balbás, M. A., Dey, A., Rabindran, S. K., Ozato, K. & Wu, C. Displacement of sequence-specific transcription factors from mitotic chromatin. Cell 83, 29–38 (1995).
Caravaca, J. M. et al. Bookmarking by specific and nonspecific binding of FoxA1 pioneer factor to mitotic chromosomes. Genes Dev. 27, 251–260 (2013).
Festuccia, N. et al. Mitotic binding of Esrrb marks key regulatory regions of the pluripotency network. Nat. Cell Biol. 18, 1139–1148 (2016).
Kadauke, S. et al. Tissue-specific mitotic bookmarking by hematopoietic transcription factor GATA1. Cell 150, 725–737 (2012).
Rape, M. Ubiquitylation at the crossroads of development and disease. Nat. Rev. Mol. Cell Biol. 19, 59–70 (2018).
Buckley, S. M. et al. Regulation of pluripotency and cellular reprogramming by the ubiquitin-proteasome system. Cell Stem Cell 11, 783–798 (2012).
Gao, J. et al. The CUL4–DDB1 ubiquitin ligase complex controls adult and embryonic stem cell differentiation and homeostasis. eLife 4, e07539 (2015).
Hu, G. et al. A genome-wide RNAi screen identifies a new transcriptional module required for self-renewal. Genes Dev. 23, 837–848 (2009).
Yau, R. G. et al. Assembly and function of heterotypic ubiquitin chains in cell-cycle and protein quality control. Cell 171, 918–933 (2017).
Stegmeier, F. et al. Anaphase initiation is regulated by antagonistic ubiquitination and deubiquitination activities. Nature 446, 876–881 (2007).
Ang, Y. S. et al. Wdr5 mediates self-renewal and reprogramming via the embryonic stem cell core transcriptional network. Cell 145, 183–197 (2011).
Pijnappel, W. W. et al. A central role for TFIID in the pluripotent transcription circuitry. Nature 495, 516–519 (2013).
Karatas, H. et al. High-affinity, small-molecule peptidomimetic inhibitors of MLL1/WDR5 protein–protein interaction. J. Am. Chem. Soc. 135, 669–682 (2013).
Mark, K. G., Loveless, T. B. & Toczyski, D. P. Isolation of ubiquitinated substrates by tandem affinity purification of E3 ligase-polyubiquitin-binding domain fusions (ligase traps). Nat. Protoc. 11, 291–301 (2016).
Fuchs, G. et al. RNF20 and USP44 regulate stem cell differentiation by modulating H2B monoubiquitylation. Mol. Cell 46, 662–673 (2012).
Chang, L. F., Zhang, Z., Yang, J., McLaughlin, S. H. & Barford, D. Molecular architecture and mechanism of the anaphase-promoting complex. Nature 513, 388–393 (2014).
Meyer, H. J. & Rape, M. Enhanced protein degradation by branched ubiquitin chains. Cell 157, 910–921 (2014).
Matsumoto, M. L. et al. K11-linked polyubiquitination in cell cycle control revealed by a K11 linkage-specific antibody. Mol. Cell 39, 477–484 (2010).
Aho, E. R. et al. Displacement of WDR5 from chromatin by a WIN site inhibitor with picomolar affinity. Cell Rep. 26, 2916–2928 (2019).
King, R. W. et al. A 20S complex containing CDC27 and CDC16 catalyzes the mitosis-specific conjugation of ubiquitin to cyclin B. Cell 81, 279–288 (1995).
Blobel, G. A. et al. A reconfigured pattern of MLL occupancy within mitotic chromatin promotes rapid transcriptional reactivation following mitotic exit. Mol. Cell 36, 970–983 (2009).
Pilaz, L. J. et al. Prolonged mitosis of neural progenitors alters cell fate in the developing Brain. Neuron 89, 83–99 (2016).
Halley-Stott, R. P., Jullien, J., Pasque, V. & Gurdon, J. Mitosis gives a brief window of opportunity for a change in gene transcription. PLoS Biol. 12, e1001914 (2014).
Egli, D., Birkhoff, G. & Eggan, K. Mediators of reprogramming: transcription factors and transitions through mitosis. Nat. Rev. Mol. Cell Biol. 9, 505–516 (2008).
Hockemeyer, D. et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat. Biotechnol. 29, 731–734 (2011).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Kampmann, M., Bassik, M. C. & Weissman, J. S. Functional genomics platform for pooled screening and generation of mammalian genetic interaction maps. Nat. Protoc. 9, 1825–1847 (2014).
McGourty, C. A. et al. Regulation of the CUL3 ubiquitin ligase by a calcium-dependent co-adaptor. Cell 167, 525–538 (2016).
Qiao, R. et al. Mechanism of APC/CCDC20 activation by mitotic phosphorylation. Proc. Natl Acad. Sci. USA 113, E2570–E2578 (2016).
Brown, N. G. et al. RING E3 mechanism for ubiquitin ligation to a disordered substrate visualized for human anaphase-promoting complex. Proc. Natl Acad. Sci. USA 112, 5272–5279 (2015).
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Zhang, S. et al. Molecular mechanism of APC/C activation by mitotic phosphorylation. Nature 533, 260–264 (2016).
Zhang, P., Lee, H., Brunzelle, J. S. & Couture, J. F. The plasticity of WDR5 peptide-binding cleft enables the binding of the SET1 family of histone methyltransferases. Nucleic Acids Res. 40, 4237–4246 (2012).
Brown, N. G. et al. Mechanism of polyubiquitination by human anaphase-promoting complex: RING repurposing for ubiquitin chain assembly. Mol. Cell 56, 246–260 (2014).
Tsankov, A. M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349 (2015).
Acknowledgements
We thank N. Ingolia, R. Tjian, B. Schulman, J. Schaletzky and all members of M.R.’s laboratory for advice, helpful discussions and comments on the manuscript; and M. Matsumoto and V. Dixit for generously supplying us with linkage-specific ubiquitin antibodies. E.O. was funded by the Jane Coffin Childs Memorial Fund for Medical Research and the Siebel Stem Cell Institute. K.G.M. was funded by the NIH F32 postdoctoral fellowship (F32GM120956). A.M. was funded by the American Italian Cancer foundation and the California Institute for Regenerative Medicine. M.R. is an Investigator of the Howard Hughes Medical Institute. This work was also funded by an NIH grant (RO1GM083064) awarded to M.R.
Author information
Authors and Affiliations
Contributions
E.O. performed work with stem cells (including knockdowns and infections), and performed and analysed the ultracomplex screen, cytometry-based studies, microscopy-based studies, qPCR-based studies and ChIP–seq and RNA-sequencing experiments. K.G.M. performed in vitro assays. E.O., K.G.M., A.M. and D.D.C. prepared samples for mass spectrometry analyses, performed immunoprecipitation experiments and maintained cell culture. E.R.W. and J.R.P. performed the cryo-EM of APC/C–WDR5. M.K. helped to analyse the ultracomplex screen. N.G. and C.Y.Z. prepared recombinant histones for in vitro assays. E.O., K.G.M. and M.R. interpreted the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.R. is a cofounder and consultant to Nurix Therapeutics, a biotechnology company working in the ubiquitin space.
Additional information
Peer review information Nature thanks William P. Tansey and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Ultracomplex shRNA screen identifies APC/C and USP44 as regulators of human ES cell biology.
a, Karyotype analysis of H1 OCT4–GFP cell line shows normal chromosome architecture. This line was karyotyped before performing the screen by a third party vendor (WiCell). Twenty cells were counted, 8 were analysed and 4 were karyotyped as normal. No clonal abnormalities were detected at the band resolution of 450–475. b, H1 OCT4–GFP cells undergo neural conversion with an efficiency similar to that of the unmodified parent line. This experiment was performed three independent times with similar results. c, Deep sequencing read counts (log2-transformed) for individual shRNAs (red dots) targeting the indicated gene from the screen in Fig. 1b. Grey dots represent negative-control shRNAs. P values (two-sided Mann–Whitney U test, not corrected for multiple hypothesis testing) are indicated for each gene. d, Mass spectrometry analysis shows that many quality-control enzymes associate with K11/K48-branched chains in human ES cells. H1 human ES cells were synchronized in mitosis before being subjected to affinity purification using K11/K48-bispecific antibodies under denaturing conditions. Values listed in brackets are total spectral counts of tryptic peptides for each protein. e, CRISPR–Cas9-edited USP44 H1 human ES cells show impaired rates of neural conversion. Expression of markers of pluripotency (OCT4), neural crest cells (SNAIL2) or neural progenitors (PAX6) were determined at indicated times of differentiation by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) and western blotting using specific antibodies. This experiment was performed two independent times with similar results.
Extended Data Fig. 2 Characterization of APC/C and the role of USP44 in pluripotency.
a, Western blot of OCT4 and NANOG upon APC2 knockdown in asynchronous H1 human ES cells. This experiment was performed two independent times with similar results. b, Western blot of OCT4 and NANOG upon knockdown of APC/C subunits in asynchronous H1 human ES cells. This experiment was performed three independent times with similar results. c, Real-time qPCR of OCT4 and NANOG upon APC2 knockdown in asynchronous H1 human ES cells (mean of n = 4 independent experiments, ± s.d.). d, Flow cytometry analysis of APC2 depletion in H1 OCT4–GFP human ES cells. H1 OCT4–GFP human ES cells were transfected with siRNA against APC2 for 48 h before cytometry analysis. This experiment was performed two independent times with similar results. e, Loss of the pluripotency marker OCT4 upon depletion of APC2 or WDR5 requires entry into mitosis. H1 human ES cells were transfected with indicated siRNAs for 36 h and treated with DMSO (asynchronous), 5 μM STLC (mitotic arrest) or 200 mM thymidine (S-phase arrest) for an additional 12 h before collection for western blot analysis. This experiment was performed three independent times with similar results. f, Flow cytometry analysis of asynchronous H1 human ES cells transfected with indicated siRNAs for 72 h. This experiment was performed three independent times with similar results.
Extended Data Fig. 3 APC/C and WDR5 are required for human ES cell survival.
a, Mass spectrometry analysis of Flag–WDR5 purified from mitotic HEK293T cells. Values listed in brackets are total spectral counts of tryptic peptides of indicated proteins. b, Depletion of WDR5 phenocopies the depletion of APC2 in H1 human ES cells. H1 cells were depleted with the indicated siRNAs for 72 h before collection for western blot analysis. This experiment was performed once. c, A cumulative fraction curve measuring the length of each metaphase-to-anaphase transition. n = 112 cells for control siRNA; n = 105 cells for siRNA against APC2; n = 106 cells for siRNA against WDR5; and n = 217 cells for siRNAs against both APC2 and WDR5. d, Depletion of APC2 or WDR5 causes cell death in H1 human ES cells. Cell death was measured by trypan blue staining of dead cells (mean of n = 4 independent experiments ± s.d.). e, Quantifying cell survival using chromosome catastrophe as a proxy for cell death. H1 human ES cells virally expressing H2B–mCherry were transfected with the indicated siRNAs for 24 h before imaging by confocal microscopy. n = 97 cells for control siRNA; n = 104 cells for siRNA against APC2; n = 90 cells for siRNA against WDR5; and n = 213 cells for siRNAs against both APC2 and WDR5. f, Sister cells die immediately following mitotic exit when depleted of APC2 and WDR5. H1 human ES cells virally expressing H2B–mCherry were transfected with siRNA against APC2 and/or siRNA against WDR5 for 24 h before imaging by confocal microscopy. The time of death, as defined by cells undergoing chromosome catastrophe, was measured for each sister (mean of n = 57 pairs of cells ± s.d.). g, Representative frames of live-cell imaging from four independent experiments (in minutes) tracking the nuclei of siRNA-depleted H1 human ES cells virally expressing H2B–mCherry. Arrows mark individual sister cells upon mitotic exit. Chromosome catastrophe was used a proxy for cell death (time points 198 and 342). h, Flag–WDR5 associates with APC/C in mitotic H1 human ES cells. Flag–WDR5 immunoprecipitations were performed on asynchronous H1 human ES cells (A) or H1 human ES cells arrested in mitosis (M). Bound proteins were determined by SDS–PAGE and western blotting. This experiment was performed two independent times with similar results. i, Overexpressed haemagglutinin (HA)-tagged USP44 associates with Flag–WDR5 in both asynchronous and mitotic HEK293T cells. MYC–WDR5 was used as the control vector. This experiment was performed three independent times with similar results.
Extended Data Fig. 4 WDR5 associates with APC/C and TBP on distinct surfaces.
a, The WIN-motif binding site on WDR5 is critical for APC/C engagement, whereas the surface that binds the WBM is dispensable. A secondary binding surface (4A) is also important for the association of WDR5 with APC/C. The surface on WDR5 that binds the WBM is critical for TFIID association, whereas the WIN-motif binding site is dispensable. HEK293T cells were transfected with the indicated Flag–WDR5 variants, and cells were synchronized in mitosis. Flag–WDR5 was affinity-purified, and bound proteins were determined by western blotting. This experiment was performed five independent times with similar results. b, Reciprocal immunoprecipitations show that APC/C binds WDR5 through its WIN-motif binding site. Endogenous APC/C was purified from HEK293T cells expressing the indicated Flag–WDR5 variants, and bound proteins were determined by SDS–PAGE and western blotting. This experiment was performed three independent times with similar results. c, Heat map of bait-normalized total spectral counts identified from Flag–WDR5-purified mass spectrometry experiments. HeLa cells were transfected with Flag–WDR5 for 24 h before mitotic synchronization. d, The WDR5 inhibitor MM-102 impairs the association of WDR5with APC/C. Mitotic HeLa S3 cells were released into MM-102 for 2 h before immunoprecipitation experiments. Under these conditions, MM-102 did not prevent the association of WDR5 with MLL and RBBP5. This experiment was performed two independent times with similar results. e, Expression of wild-type WDR5 but not WDR5(∆WIN) rescues the pluripotency defect caused by WDR5 depletion in H1 human ES cells. H1 human ES cells virally expressing siRNA-resistant WDR5 variants (WDR5 versus WDR5(∆WIN)) were depleted of endogenous WDR5 (W) or treated with control siRNA (C). Expression of OCT4 and NANOG was determined by western blotting. This experiment was performed once.
Extended Data Fig. 5 WDR5 binds near the catalytic core of APC/C.
a, WDR5 forms BMB-dependent crosslinks with APC2, APC11 and CDC20. APC/C–CDC20 was affinity-purified from prometaphase-arrested HeLa cells and incubated with recombinant WDR5 or WDR5(ΔWIN) before the addition of crosslinker. Crosslinked APC/C subunits were detected by SDS–PAGE and western blotting using specific antibodies. This experiment was performed two independent times with similar results. b, In vitro translation binding assays reveal that APC2 directly interacts with recombinant WDR5. MBP-tagged WDR5 or WDR5(ΔWIN) were immobilized on amylose resin and incubated with 35S-labelled APC/C subunits produced by in vitro transcription and translation. Bound proteins were detected by SDS–PAGE and autoradiography. APC3 did not synthesize by in vitro transcription and translation. This experiment was performed once for the full set of APC/C subunits. APC2 binding was validated three independent times. c, Quantification of autoradiography blot shown in b.
Extended Data Fig. 6 WDR5 associates with active APC/C.
a, APC/C does not ubiquitylate WDR5 in vitro. Recombinant WDR5 was incubated with active APC/C, E1, UBE2C, UBE2S and ubiquitin, and potential reaction products were detected by western blotting against WDR5. This experiment was performed once. b, APC/C-dependent ubiquitylation of geminin is outcompeted by recombinant securin (comp), a canonical substrate, but not by recombinant WDR5. Securin or WDR5 was added to APC/C-dependent geminin ubiquitylation reactions at the indicated concentrations, and various reaction products were detected using western blotting. Asterisks represent cross-reactive bands. This experiment was performed once. c, APC/C–WDR5-dependent ubiquitylation of geminin is inhibited by EMI1. WDR5 affinity purifications from mitotic HeLa cells were incubated with E1, the APC/C-specific E2 enzymes UBE2C and UBE2S, and ubiquitin. EMI1 was added at indicated concentrations, and reaction products were detected by western blotting using antibodies against geminin. This experiment was performed two independent times with similar results. d, Immunoprecipitation of Flag–WDR5 from mitotic HEK293T cells coprecipitates K11-linked ubiquitin chains. HEK293T cells arrested in prometaphase were released into fresh medium, and WDR5 was affinity-purified at the indicated time points. Bound proteins were detected by western blotting. This experiment was performed once. e, Depletion of UBE2S eliminates WDR5-associated K11-linked ubiquitin chains in mitotic HEK293T cells. This experiment was performed once.
Extended Data Fig. 7 Mitotic APC/C–WDR5 complexes are catalytically active.
a, APC/C–WDR5 ubiquitylates human H3 in H3–H4 tetramers in vitro. APC/C was affinity-purified from mitotic HeLa cells and incubated with E1, UBE2C, UBE2S, ubiquitin and human H3–H4 tetramers, as indicated. Reaction products were detected by western blotting using antibodies against H3 and H4. This experiment was performed once. b, APC/C ubiquitylation of H2B in X. laevis H2A–H2B histone dimers. APC/C–WDR5 was purified from mitotic HeLa cells by Flag–WDR5 affinity purification and incubated with E1, UBE2C, UBE2S, ubiquitin and X. laevis histone octamers. Ubiquitylation was detected by western blotting against ubiquitylated H2B. This experiment was performed three independent times with similar results. c, APC/C–WDR5 ubiquitylation of H2B in X. laevis H2A–H2B–H3–H4 histone octamers. Reactions were performed as described in b. This experiment was performed two independent times with similar results. d, APC/C purified from H1 human ES cells is competent to ubiquitylate human H2B. This experiment was performed two independent times with similar results. e, APC/C purified from mitotic, but not S-phase, extracts can ubiquitylate H2B in vitro. APC/C was purified from HeLa cells synchronized at the indicated cell-cycle stages and incubated with E1, UBE2C, UBE2S, ubiquitin and X. laevis H2A–H2B dimers. Histone ubiquitylation was detected by western blotting using antibodies against ubiquitylated H2B. This experiment was performed once. f, APC/C-dependent ubiquitylation of H2B requires K11 residue on ubiquitin for chain elongation. Ubiquitylation of H2A–H2B dimers by APC/C–WDR5 was performed as described in e, but with ubiquitin variants. This experiment was performed once. g, APC/C-dependent ubiquitylation of H2B requires both K11 and K48 on ubiquitin for synthesis of branched chains. This experiment was performed two independent times with similar results. h, Securin, a canonical APC/C substrate, outcompetes H2A–H2B dimers for APC/C-dependent ubiquitylation. The D-box motif (an APC/C–CDC20-specific degron) is required for full competition, whereas the KEN motif (an APC/C–CDH1-specific degron) is not. This experiment was performed two independent times with similar results. i, Polyubiquitylated H2B is degraded by the proteasome. K11/K48-branched chains were purified under denaturing conditions from mitotic HeLa cells either in the presence or absence of MG132, and modified H2B was detected using western blotting. Proteasome inhibition with MG132 was found to stabilize endogenous polyubiquitylated H2B. This experiment was performed four independent times with similar results.
Extended Data Fig. 8 ChIP–seq analyses of APC/C- and WDR5-occupied gene targets.
a, Overall histone levels do not change upon release from mitosis. H1 cells were synchronized in mitosis by STLC and released into fresh medium. Indicated proteins were monitored by western blotting. This experiment was performed two independent times with similar results. b, Comparison between ChIP–seq and MNase ChIP–seq against anti-K11 from mitotic H1 human ES cells reveals that sonication shears polymeric ubiquitin linkages. c, Venn diagram of anti-K11 and anti-WDR5 ChIP peaks that colocalize with TSSs from MNase ChIP–seq experiments. MNase ChIP–seq experiments were performed from mitotic H1 human ES cells. d, Heat map of MNase ChIP–seq data from mitotic H1 human ES cells. Cluster 1 includes sites that are co-occupied by K11 and WDR5 near TSSs (within 100 bp); cluster 2 includes sites that are co-occupied by K11 and WDR5 outside of TSSs; and cluster 3 includes sites occupied only by K11, regardless of colocalization with TSSs. e, ChIP–qPCR analysis of candidate targets using K11- or K11/K48-linkage specific ubiquitin antibodies from mitotic H1 human ES cells. Mean of independent replicates ± s.d. n = 3 for K11/K48; n = 5 for IGG and K11 (except n = 4 for PUM1). f, ChIP–qPCR analysis of mitotic H1 human ES cells shows that K11 linkages synthesized at candidate sites are dependent on UBE2S and WDR5. This experiment was performed once. g, WDR5 inhibition prevents K11-ubiquitin chain formation at APC/C–WDR5-bound TSSs. H1 human ES cells were treated with or without 50 μM MM102 during mitotic synchronization with STLC before anti-K11 MNase ChIP–seq. Heat map of all APC/C–WDR5-bound TSSs are shown. h, Heat map of ChIP–seq peaks of individual genes co-occupied by Flag–CDC20 and Flag–WDR5. ChIP–seq against anti-Flag was performed on mitotic HEK293T cells that overexpress Flag–CDC20 or Flag–WDR5. i, Spatial profile of PUM1 of factor occupancy by ChIP–qPCR. This experiment was performed once. j, Spatial profile of E2F3 of factor occupancy by ChIP–qPCR. This experiment was performed once. k, Heat map of MNase ChIP–seq data of transcription-factor binding. Previously published MNase ChIP–seq data were obtained47, and APC/C-bound sites were analysed as described in d.
Extended Data Fig. 9 Select transcription factors are found at APC/C–WDR5-bound sites.
Heat map of MNase ChIP–seq data of transcription-factor binding. Previously published MNase ChIP–seq data were obtained47, and APC/C-bound sites were analysed as follows: cluster 1 includes sites that are co-occupied by K11 and WDR5 near TSSs (within 100 bp); cluster 2 includes sites that are co-occupied by K11 and WDR5 outside of TSSs; and cluster 3 includes sites only occupied by K11, regardless of colocalization with TSSs.
Extended Data Fig. 10 Regulation of chromatin and transcription by APC/C–WDR5.
a, Comparison of genes co-occupied by Flag–CDC20 and Flag–WDR5 from mitotic HEK293T cells with known gene-expression profiles reveals a strong overlap with ES cell and medulloblastoma cancer cell lines. n = 1,628 genes were analysed (P values represent a one-sided Fisher’s exact test with Bonferroni correction). b, Loss of APC/C–WDR5 function interferes with the expression of genes marked with K11-linked ubiquitin chains in H1 human ES cells. Poly(A)-selected RNA was purified from asynchronous H1 human ES cells transfected with control siRNA or siRNA against WDR5 for 48 h, and subjected to RNA-sequencing analysis (a biological replicate of Fig. 4h). c, Transcript analysis of WDR5 depletion on APC/C–WDR5-dependent genes (from Fig. 4h and b). Box plots include the median TPM value (n = 90 genes) with quartile ranges Q1–Q3; top whiskers represent the 3rd quartile + 1.5× interquartile range; bottom whiskers represent the 1st quartile − 1.5× interquartile range. P values were calculated from comparing individual TPM values of APC/C–WDR5-regulated genes (n = 90) versus all transcripts (n = 18,791) using a two-sided Student’s t-test (unpaired). d, Real-time qPCR analysis of nascent RNA reveals APC/C–WDR5 target genes are reactivated upon mitotic exit and gene reactivation is dependent on WDR5. Initial screening from a single experiment. e, The RNA levels of genes regulated by APC/C–WDR5 do not change upon mitotic exit. RNA-sequencing analysis was performed on poly(A)-selected RNA purified from H1 human ES cells at the indicated cell-cycle stages. Box plots were derived as described in c (n = 90 genes). f, Ubiquitylated H2B preferentially associates with p97–UBXN7 in vitro. H2B was preubiquitylated by APC/C in vitro, and incubated with immobilized p97 or p97–UBXN7 complexes. Bound histone H2B was detected by western blotting. This experiment was performed three independent times with similar results. g, Flag–UBXN7 associates with polyubiquitylated H2B, p97 and K11/K48-linked branched ubiquitin chains in mitosis. Native Flag–UBXN7 immunoprecipitations were performed on mitotic HEK293T cells and bound proteins were detected by western blotting or Ponceau staining. This experiment was performed three independent times with similar results. h, H2B ubiquitylation is stabilized by p97 inhibition in cells. Denaturing K11/K48 immunoprecipitations were performed on H1 human ES cells synchronized in prometaphase or released into 10 μM NMS-873 for 2 h. This experiment was performed four independent times with similar results. i, p97 inhibition restores K11 deposition at sites regulated by APC/C–WDR5 upon mitotic exit. Anti-K11 MNase ChIP–seq was performed from H1 human ES cells synchronized in mitosis (0 h) or released into fresh medium without (2 h + DMSO) or with p97 inhibition (2 h + 10 μM NMS-873). j, Anti-K11 MNase ChIP–qPCR of candidate targets from mitotic H1 human ES cells (mean of n = 3 independent replicates ± s.d.). H1 human ES cells were synchronized in mitosis (0 h) and released into fresh medium for 2 h with the indicated drugs. k, Model of APC/C-dependent gene activation upon mitotic exit.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-2 showing the uncropped gel data and Supplementary Tables 1-3, showing lists of plasmids and antibodies used in the study as well as oligonucleotide sequences used for siRNA knockdown, ChIP-qPCR and RT-qPCR.
Rights and permissions
About this article
Cite this article
Oh, E., Mark, K.G., Mocciaro, A. et al. Gene expression and cell identity controlled by anaphase-promoting complex. Nature 579, 136–140 (2020). https://doi.org/10.1038/s41586-020-2034-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2034-1
- Springer Nature Limited
This article is cited by
-
Mitotic gene regulation by the N-MYC-WDR5-PDPK1 nexus
BMC Genomics (2024)
-
Mitotic bookmarking redundancy by nuclear receptors in pluripotent cells
Nature Structural & Molecular Biology (2024)
-
Structural snapshots along K48-linked ubiquitin chain formation by the HECT E3 UBR5
Nature Chemical Biology (2024)
-
Spatial control of the APC/C ensures the rapid degradation of cyclin B1
The EMBO Journal (2024)
-
ASPSCR1-TFE3 reprograms transcription by organizing enhancer loops around hexameric VCP/p97
Nature Communications (2024)