Abstract
Symbiotic nitrogen fixation in legume nodules requires substantial energy investment from host plants, and soybean (Glycine max (L.) supernodulation mutants show stunting and yield penalties due to overconsumption of carbon sources. We obtained soybean mutants differing in their nodulation ability, among which rhizobially induced cle1a/2a (ric1a/2a) has a moderate increase in nodule number, balanced carbon allocation, and enhanced carbon and nitrogen acquisition. In multi-year and multi-site field trials in China, two ric1a/2a lines had improved grain yield, protein content and sustained oil content, demonstrating that gene editing towards optimal nodulation improves soybean yield and quality.


Similar content being viewed by others
Data availability
The accession numbers and gene IDs are available in Supplementary Tables 1–6. The raw data for the RNA-seq experiment are available in Supplementary Tables 1–6. The genome editing lines and plasmids generated are available from the corresponding authors on request, while adhering to the regulatory policy of genome-editing crops and soybean germplasm from the Ministry of Agriculture and Rural Affairs of the People’s Republic of China. Source data are provided with this paper.
References
van Dijk, M., Morley, T., Rau, M. L. et al. A meta-analysis of projected global food demand and population at risk of hunger for the period 2010–2050. Nat. Food 2, 494–501 (2021). https://doi.org/10.1038/s43016-021-00322-9
Patil, G. et al. Molecular mapping and genomics of soybean seed protein: a review and perspective for the future. Theor. Appl. Genet. 130, 1975–1991 (2017).
Mahmoud, A. A. et al. Effect of six decades of selective breeding on soybean protein composition and quality: a biochemical and molecular analysis. J. Agric. Food Chem. 54, 3916–3922 (2006).
Jhu, M.-Y. & Oldroyd, G. E. D. Dancing to a different tune, can we switch from chemical to biological nitrogen fixation for sustainable food security? PLoS Biol. 21, e3001982 (2023).
Searle, I. R. et al. Long-distance signaling in nodulation directed by a CLAVATA1-like receptor kinase. Science 299, 109–112 (2003).
Nishida, H. & Suzaki, T. Two negative regulatory systems of root nodule symbiosis: how are symbiotic benefits and costs balanced? Plant Cell Physiol. 59, 1733–1738 (2018).
Ferguson, B. J. et al. Legume nodulation: the host controls the party. Plant Cell Environ. 42, 41–51 (2019).
Zipfel, C. & Oldroyd, G. E. D. Plant signalling in symbiosis and immunity. Nature 543, 328–336 (2017).
Mortier, V. et al. CLE peptides control Medicago truncatula nodulation locally and systemically. Plant Physiol. 153, 222–237 (2010).
Reid, D. E., Ferguson, B. J. & Gresshoff, P. M. Inoculation- and nitrate-induced CLE peptides of soybean control NARK-dependent nodule formation. Mol. Plant Microbe Interact. 24, 606–618 (2011).
Kennedy, P., Leonforte, A. & Butsch, M. in Biological Nitrogen Fixation (ed. de Bruijn, F.J.) Ch. 106 (John Wiley & Sons, 2015).
Sayuri, T. & Takuji, O. in Advances in Biology and Ecology of Nitrogen Fixation (ed. Takuji, O.) Ch. 4 (IntechOpen, 2014).
Ferguson, B. J. in A Comprehensive Survey of International Soybean Research (ed. Board, J.) Ch. 2 (IntechOpen, 2013).
Carroll, B. J., McNeil, D. L. & Gresshoff, P. M. A supernodulation and nitrate-tolerant symbiotic (nts) soybean mutant. Plant Physiol. 78, 34–40 (1985).
Hungria, M. & Mendes, I. C. in Biological Nitrogen Fixation (ed. de Bruijn, F. J.) Ch.99 (John Wiley & Sons, 2015).
Herridge, D. & Rose, I. Breeding for enhanced nitrogen fixation in crop legumes. Field Crops Res. 65, 229–248 (2000).
Zeffa, D. M. et al. Effects of plant growth-promoting rhizobacteria on co-inoculation with Bradyrhizobium in soybean crop: a meta-analysis of studies from 1987 to 2018. PeerJ 8, e7905 (2020).
Fu, M., Sun, J., Li, X., Guan, Y. & Xie, F. Asymmetric redundancy of soybean Nodule Inception (NIN) genes in root nodule symbiosis. Plant Physiol. 188, 477–489 (2022).
Bai, M. et al. Generation of a multiplex mutagenesis population via pooled CRISPR–Cas9 in soya bean. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13239 (2019).
Bacanamwo, M. & Harper, J. E. Response of a hypernodulating soybean mutant to increased photosynthate supply. Plant Sci. 124, 119–129 (1997).
Yang, N. et al. Metabolomics reveals distinct carbon and nitrogen metabolic responses to magnesium deficiency in leaves and roots of soybean [Glycine max (Linn.) Merr.]. Front. Plant Sci. https://doi.org/10.3389/fpls.2017.02091 (2017).
Qin, J. et al. Identification of candidate genes and genomic selection for seed protein in soybean breeding pipeline. Front. Plant Sci. https://doi.org/10.3389/fpls.2022.882732 (2022).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Tian, T. et al. agriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res. 45, W122–W129 (2017).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408 (2001).
Shearer, G. B. & Kohl, D. H. N2-fixation in field settings: estimations based on natural 15N abundance. Aust. J. Plant Physiol. 13, 699–756 (1986).
Fischinger, S. A. & Schulze, J. The importance of nodule CO2 fixation for the efficiency of symbiotic nitrogen fixation in pea at vegetative growth and during pod formation. J. Exp. Bot. 61, 2281–2291 (2010).
Yang, J. et al. An improved method for the identification of soybean resistance to Phytophthora sojae applied to germplasm resources from the Huanghuaihai and Dongbei regions of China. Plant Dis. 104, 408–413 (2020).
Jin, T. et al. R(SC3) K of soybean cv. Kefeng No.1 confers resistance to soybean mosaic virus by interacting with the viral protein P3. J. Integr. Plant Biol. 65, 838–853 (2023).
GraphPad Prism v.8.0.0 for Windows (GraphPad Software Inc., 1999); www.graphpad.com
R: A language and environment for statistical computing (R Core Team, 2013).
Acknowledgements
We thank L. Yan from the Institute of Cereal and Oil Crops, Hebei Academy of Agricultural and Forestry Sciences, for help with field-trial design. We thank X. Li from the Root Biology Center, Fujian Agriculture and Forestry University, for assistance with the determination of soil chemical properties. We thank K. Li at the National Center for Soybean Improvement, Nanjing Agricultural University, for the SMV resistance tests. We thank X. Wang at the Key Laboratory of Soybean Disease and Pest Control, Ministry of Agriculture and Rural Affairs, for testing the resistance to Phytophthora. This work was funded by the National Key Research and Development Program of China (grant no. 2022YFD1201501 to F.K.), the Fujian Agriculture and Forestry University Scientific Research Project for Prominent Talents (grant no. Kxjq21010 to X. Zhong), the Natural Science Foundation of Hebei Province (grant no. C2020301020 to X.S.), the China Agriculture Research System of MOF and MARA (grant no. CARS-04-PS06 to C. Yang) and the Chinese Academy of Sciences Project for Young Scientists in Basic Research (grant no. YSBR-011 to E.W.).
Author information
Authors and Affiliations
Contributions
X. Zhong and J.W. performed most of the experiments, analysed the data and contributed to the manuscript writing. X.S. contributed to the field trials. M.B. and C. Yuan generated the different nodulation mutants. M.B. prepared the figures. C. Yuan, N.W. and X. Zhu performed functional validation of mutated RIC1a/2a genes. X.W. analysed the transcriptomic data. H.K. performed the soybean transformation. J.S., X.H., X.L. and W.Y. contributed to the physiological measurements of ric1a/2a mutants. C. Yang contributed to the design and analysis of the field trials. F.K., E.W. and Y.G. designed the experiments, wrote the paper together with X. Zhong and J.W., and conducted project administration and funding acquisition.
Corresponding authors
Ethics declarations
Competing interests
FAFU is the applicant on Chinese patent application no. 202410056923.2 with Y.G., E.W., X. Zhong and J.W. as co-inventors (all co-inventors are first or corresponding authors in the manuscript). The application was filed in January 2024, and status is submitted. The patent covered the applications of combined mutation of RIC genes to improve soybean yield. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Stig Andersen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genotype of nodulation mutants used in this study.
a, Mutation type of nin-4m, ric1b/2b, ric1a/2a-1, ric1a/2a-2, ric-6m, and nark. Pink indicates no mutations. Grey indicates the genotype of mutations. D, number of base pairs deleted; I, number of base pairs inserted, compared with the WT sequence. b, Expression of target genes in each mutant by RT–qPCR (n = 3). c, Predicted off-targets and Sanger sequencing for potential mutations. The central black lines in the dot plots represent the median in (b).
Extended Data Fig. 2 ric1a/2a plants are indistinguishable from HC-6 without inoculation or at high-N conditions.
a,b, Plant morphology (a) and whole-plant dry weight (b) of nin-4m, ric1b/2b, ric1a/2a-1, ric-6m,nark, and HC-6 in hydroponic culture without inoculation at 40 days. c–e, Plant morphology (c), whole-plant dry weight (d) and nodule number (e) of nin-4m, ric1b/2b, ric1a/2a-1, ric-6m, nark and HC-6 in hydroponic culture under high-N conditions (5 mM) with inoculation at 40 days. At high N, although ric1b/2b (9.6 ± 2.9), ric1a/2a (12.8± 4.0), ric-6m (17.0 ±3.0) and nark (28.4 ± 8.0) formed more nodules than HC-6 (8.3 ± 2.7), nodule number was inhibited and the nodule size was too small to score weight, thus were deemed non-functional. One-way ANOVA was performed in b, c and d. Different letters indicate statistically significant differences. The central black lines in the dot plots represent the median.
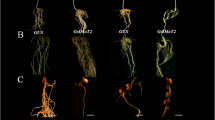
Extended Data Fig. 3 Over-expression of mutant RIC1a and RIC2a genes in transgenic composite plants does not alter nodule number.
a, b Alignment of wild-type and mutant RIC1a/2a amino-acid sequences. c, d Plant morphology (c) and nodule numbers (d) of transgenic composite plants expressing an empty-vector control and over-expressing wild-type RIC1a and RIC2a or mutated RIC1a and RIC2a genes present in the characterised mutants. One-way ANOVA was performed in d. Different letters indicate statistically significant differences. The central black lines in the dot plots represent the median.
Extended Data Fig. 4 15N isotope labelling of hydroponic culture for calculation of the contribution of biological N fixation to total N content in whole plants.
a, b Absolute isotopic 15N abundance (a) and atom % 15N (b) of HC-6 and ric1a/2a-1 plants grown with or without Rhizobia inoculation in a K15NO3-labelling assay (n ≥ 14). Mann-Whitney testing was performed. The central black lines in the dot plots represent the median.
Extended Data Fig. 5 Transcriptomic analysis of HC-6 and ric1a/2a-1 leaves.
a, Heat map of differential expression in HC-6 versus ric1a/2a-1 leaves. b, KEGG analysis of up-regulated genes in HC-6 versus ric1a/2a-1 leaves.
Extended Data Fig. 6 Agronomic traits of field-grown HC-6 and ric1a/2a-1.
a–e, Plant height (a), branch number (b), pod number (c), seed number (d), and 100-seed weight (e) of HC-6 and ric1a/2a-1 and ric1a/2a-2 in 2022 in Fuzhou and Shijiazhuang trials (n ≥ 14). f, Nodule number of HC-6 and ric1a/2a-1 mutants at flowering stage in 2022 in the Fuzhou field. Student t-tests were performed. The central black lines in the dot plots represent the median.
Extended Data Fig. 7 Flowering time of nodulation mutants under laboratory conditions.
a,b, Flowering time of nin-4m, ric1b/2b, ric1a/2a-1, ric-6m, nark, and wild-type HC-6 plants grown in hydroponic culture at low-N (0.5 mM, a) or high-N conditions (5 mM, b) with inoculation in a growth chamber with 14 h illumination and 10 h darkness period. One-way ANOVA was performed. Different letters indicate statistically significant differences. The central black lines in the dot plots represent the median.
Extended Data Fig. 8 Disease resistance of ric1a/2a mutants.
a, Resistance level of ric1a/2a-1, ric1a/2a-2 and HC-6 seedlings to different Phytophthora sojae strains (psMC1, PS4, USAR2, Ps41-1, and PsJS2). R, resistance. IR, intermediate resistance. b, Resistance level of ric1a/2a-1, ric1a/2a-2 and HC-6 seedlings to different strains of soybean mosaic virus (SC3, SC7). The number of samples showing different disease severity was shown. M: mosaic.MN: mosaic and necrosis.
Extended Data Fig. 9 Protein and oil yield of field-grown HC-6 and ric1a/2a mutants.
a,b, Calculated protein (a, b and c) and oil (d, e and f) yield per plant (2021) or per plot (2022 and 2023) in all locations. The hypothetical protein and oil production per plant or plot was calculated by multiplying protein/oil content by grain-yield per plant or plot. Student t-test was performed. The central black lines in the dot plots represent the median.
Extended Data Fig. 10 Natural-abundance estimation of the contribution of N fixation in HC-6 and ric1a/2a seeds.
a, b Natural-abundance atom %15N (a) and 15N/14N ratio (b) of HC-6 and ric1a/2a-1 seeds harvested in 2022 in Fuzhou. Mann-Whitney testing was performed. The central black lines in the dot plots represent the median.
Supplementary information
Supplementary Tables 1–6
Supplementary Tables 1–6.
Source data
Source Data Figs. 1 and 2 and Extended Data Figs. 1–4, 6, 7, 9 and 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhong, X., Wang, J., Shi, X. et al. Genetically optimizing soybean nodulation improves yield and protein content. Nat. Plants 10, 736–742 (2024). https://doi.org/10.1038/s41477-024-01696-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-024-01696-x
- Springer Nature Limited