Abstract
Information in the genome is not only encoded within sequence or epigenetic modifications, but is also found in how it folds in three-dimensional space. The formation of self-interacting genomic regions, named topologically associated domains (TADs), is known as a key feature of genome organization beyond the nucleosomal level. However, our understanding of the formation and function of TADs in plants is extremely limited. Here we show that the genome of Marchantia polymorpha, a member of a basal land plant lineage, exhibits TADs with epigenetic features similar to those of higher plants. By analysing various epigenetic marks across Marchantia TADs, we find that these regions generally represent interstitial heterochromatin and their borders are enriched with Marchantia transcription factor TCP1. We also identify a type of TAD that we name ‘TCP1-rich TAD’, in which genomic regions are highly accessible and are densely bound by TCP1 proteins. Transcription of TCP1 target genes differs on the basis gene location, and those in TCP1-rich TADs clearly show a lower expression level. In tcp1 mutant lines, neither TCP1-bound TAD borders nor TCP1-rich TADs display drastically altered chromatin organization patterns, suggesting that, in Marchantia, TCP1 is dispensable for TAD formation. However, we find that in tcp1 mutants, genes residing in TCP1-rich TADs have a greater extent of expression fold change as opposed to genes that do not belong to these TADs. Our results suggest that, besides standing as spatial chromatin-packing modules, plant TADs function as nuclear microcompartments associated with transcription factor activities.






Similar content being viewed by others
Data availability
Short read data of in situ Hi-C, ChIP–seq, ATAC–seq and RNA-seq are publicly available at NCBI Sequence Read Archive under accession number PRJNA597314.
Large datasets, including Hi-C matrices (2-kb bin size for individual chromosomes), integrated epigenetic marks, ATAC–seq and ChIP–seq track files in 200-bp bin size are available from figshare repository, which are accessible with the following link: https://figshare.com/articles/dataset/Marchantia_TCP_transcription_factor_activity_correlates_with_3D_chromatin_structure/11309657. All figures presented in this manuscript are associated with these data.
The following public datasets were downloaded for coexpression analysis (with their accession numbers from the NCBI Sequence Read Archive): 11-day thalli (DRR050343, DRR050344, DRR050345), Archegoniophore (DRR050351, DRR050352, DRR050353), Antheridiophore (DRR050346, DRR050347, DRR050348), Antheridia (DRR050349, DRR050350), apical cell (SRR1553294, SRR1553295, SRR1553296), 13d-Sporophyte (SRR1553297, SRR1553298, SRR1553299), Sporelings 0 hr (SRR4450262, SRR4450261, SRR4450260), 24hr-Sporeling (SRR4450266, SRR4450265, SRR4450259), 48hr-Sporeling (SRR4450268, SRR4450264, SRR4450263), 72hr-Sporeling (SRR4450267, SRR4450258, SRR4450257), 96hr-Sporeling (SRR4450256, SRR4450255, SRR4450254), mock-inoculated plants, 1dpi (SRR7977545, SRR7977546, SRR7977548), mock-inoculated plants, 2dpi (SRR7977547, SRR7977550, SRR7977549), mock-inoculated plants, 3dpi (SRR7977552, SRR7977551, SRR7977554), mock-inoculated plants, 4dpi (SRR7977553, SRR7977556, SRR7977555), 1-month thallus (SRR6685782, SRR6685783, SRR6685784), Tak-1-1_Mp (SRR7772758, SRR7772757, SRR7772756, SRR7772755), Tak-1-2_Mp (SRR7772759, SRR7772761, SRR7772760, SRR7772762), Tak-1-3_Mp (SRR7772763, SRR7772764, SRR7772765, SRR7772766), and Mp-mock (SRR5905098, SRR5905099, SRR5905100). Source data are provided with this paper.
Code availability
All scripts used for pattern analysis are available upon request.
References
Gibcus, J. H. & Dekker, J. The hierarchy of the 3D genome. Mol. Cell 49, 773–782 (2013).
Sexton, T. & Cavalli, G. The role of chromosome domains in shaping the functional genome. Cell 160, 1049–1059 (2015).
Zheng, H. & Xie, W. The role of 3D genome organization in development and cell differentiation. Nat. Rev. Mol. Cell Biol. 20, 535–550 (2019).
Gomez-Diaz, E. & Corces, V. G. Architectural proteins: regulators of 3D genome organization in cell fate. Trends Cell Biol. 24, 703–711 (2014).
Ong, C. T. & Corces, V. G. CTCF: an architectural protein bridging genome topology and function. Nat. Rev. Genet. 15, 234–246 (2014).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Bonev, B. & Cavalli, G. Organization and function of the 3D genome. Nat. Rev. Genet. 17, 772 (2016).
Merkenschlager, M. & Nora, E. P. CTCF and cohesin in genome folding and transcriptional gene regulation. Annu. Rev. Genomics Hum. Genet. 17, 17–43 (2016).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Szabo, Q., Bantignies, F. & Cavalli, G. Principles of genome folding into topologically associating domains. Sci. Adv. 5, eaaw1668 (2019).
Andrey, G. et al. A switch between topological domains underlies HoxD genes collinearity in mouse limbs. Science 340, 1234167 (2013).
Lupianez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Narendra, V. et al. CTCF establishes discrete functional chromatin domains at the Hox clusters during differentiation. Science 347, 1017–1021 (2015).
Eser, U. et al. Form and function of topologically associating genomic domains in budding yeast. Proc. Natl Acad. Sci. USA 114, E3061–E3070 (2017).
Pope, B. D. et al. Topologically associating domains are stable units of replication-timing regulation. Nature 515, 402–405 (2014).
Flavahan, W. A. et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529, 110–114 (2016).
Ramirez, F. et al. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun. 9, 189 (2018).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Zhan, Y. et al. Reciprocal insulation analysis of Hi-C data shows that TADs represent a functionally but not structurally privileged scale in the hierarchical folding of chromosomes. Genome Res. 27, 479–490 (2017).
Dixon, J. R., Gorkin, D. U. & Ren, B. Chromatin domains: the unit of chromosome organization. Mol. Cell 62, 668–680 (2016).
Symmons, O. et al. Functional and topological characteristics of mammalian regulatory domains. Genome Res. 24, 390–400 (2014).
Xie, T. et al. Biased gene retention during diploidization in Brassica linked to three-dimensional genome organization. Nat. Plants 5, 822–832 (2019).
Wang, C. et al. Genome-wide analysis of local chromatin packing in Arabidopsis thaliana. Genome Res. 25, 246–256 (2015).
Dong, P. et al. 3D chromatin architecture of large plant genomes determined by local A/B compartments. Mol. Plant 10, 1497–1509 (2017).
Liu, C., Cheng, Y. J., Wang, J. W. & Weigel, D. Prominent topologically associated domains differentiate global chromatin packing in rice from Arabidopsis. Nat. Plants 3, 742–748 (2017).
Wang, M. et al. Asymmetric subgenome selection and cis-regulatory divergence during cotton domestication. Nat. Genet. 49, 579–587 (2017).
Wang, M. et al. Evolutionary dynamics of 3D genome architecture following polyploidization in cotton. Nat. Plants 4, 90–97 (2018).
Concia, L. et al. Wheat chromatin architecture is organized in genome territories and transcription factories. Genome Biol. 21, 104 (2020).
Dogan, E. S. & Liu, C. Three-dimensional chromatin packing and positioning of plant genomes. Nat. Plants 4, 521–529 (2018).
Stam, M., Tark-Dame, M. & Fransz, P. 3D genome organization: a role for phase separation and loop extrusion? Curr. Opin. Plant Biol. 48, 36–46 (2019).
Bowman, J. L. et al. Insights into land plant evolution garnered from the Marchantia polymorpha genome. Cell 171, 287–304.e15 (2017).
Montgomery, S. A. et al. Chromatin organization in early land plants reveals an ancestral association between H3K27me3, transposons, and constitutive heterochromatin. Curr. Biol. 30, 573–588.e7 (2020).
Krom, N. & Ramakrishna, W. Comparative analysis of divergent and convergent gene pairs and their expression patterns in rice, Arabidopsis, and Populus. Plant Physiol. 147, 1763–1773 (2008).
Reimegard, J. et al. Genome-wide identification of physically clustered genes suggests chromatin-level co-regulation in male reproductive development in Arabidopsis thaliana. Nucleic Acids Res. 45, 3253–3265 (2017).
Ren, X. Y., Stiekema, W. J. & Nap, J. P. Local coexpression domains in the genome of rice show no microsynteny with Arabidopsis domains. Plant Mol. Biol. 65, 205–217 (2007).
Williams, E. J. & Bowles, D. J. Coexpression of neighboring genes in the genome of Arabidopsis thaliana. Genome Res. 14, 1060–1067 (2004).
Braccioli, L. & de Wit, E. CTCF: a Swiss-army knife for genome organization and transcription regulation. Essays Biochem. 63, 157–165 (2019).
Rowley, M. J. & Corces, V. G. Organizational principles of 3D genome architecture. Nat. Rev. Genet. 19, 789–800 (2018).
Danisman, S. TCP transcription factors at the interface between environmental challenges and the plant’s growth responses. Front. Plant Sci. 7, 1930 (2016).
Li, S. The Arabidopsis thaliana TCP transcription factors: a broadening horizon beyond development. Plant Signal. Behav. 10, e1044192 (2015).
Busch, A. & Zachgo, S. Flower symmetry evolution: towards understanding the abominable mystery of angiosperm radiation. Bioessays 31, 1181–1190 (2009).
Navaud, O., Dabos, P., Carnus, E., Tremousaygue, D. & Herve, C. TCP transcription factors predate the emergence of land plants. J. Mol. Evol. 65, 23–33 (2007).
Franco-Zorrilla, J. M. et al. DNA-binding specificities of plant transcription factors and their potential to define target genes. Proc. Natl Acad. Sci. USA 111, 2367–2372 (2014).
O’Malley, R. C. et al. Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 165, 1280–1292 (2016).
Kosugi, S. & Ohashi, Y. DNA binding and dimerization specificity and potential targets for the TCP protein family. Plant J. 30, 337–348 (2002).
Busch, A. et al. MpTCP1 controls cell proliferation and redox processes in Marchantia polymorpha. New Phytol. 224, 1627–1641 (2019).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015).
Rowley, M. J. et al. Evolutionarily conserved principles predict 3D chromatin organization. Mol. Cell 67, 837–852 (2017).
Birchler, J. A., Bhadra, U., Bhadra, M. P. & Auger, D. L. Dosage-dependent gene regulation in multicellular eukaryotes: implications for dosage compensation, aneuploid syndromes, and quantitative traits. Dev. Biol. 234, 275–288 (2001).
Perales, M. et al. Threshold-dependent transcriptional discrimination underlies stem cell homeostasis. Proc. Natl Acad. Sci. USA 113, E6298–E6306 (2016).
Oldfield, C. J. & Dunker, A. K. Intrinsically disordered proteins and intrinsically disordered protein regions. Annu. Rev. Biochem. 83, 553–584 (2014).
Kubota, A. et al. TCP4-dependent induction of CONSTANS transcription requires GIGANTEA in photoperiodic flowering in Arabidopsis. PLoS Genet. 13, e1006856 (2017).
Mazur, M. J. et al. Arabidopsis TCP transcription factors interact with the SUMO conjugating machinery in nuclear foci. Front. Plant Sci. 8, 2043 (2017).
Mukhopadhyay, P. & Tyagi, A. K. OsTCP19 influences developmental and abiotic stress signaling by modulating ABI4-mediated pathways. Sci. Rep. 5, 9998 (2015).
Valsecchi, I. et al. The intrinsically disordered C-terminal region of Arabidopsis thaliana TCP8 transcription factor acts both as a transactivation and self-assembly domain. Mol. Biosyst. 9, 2282–2295 (2013).
Pontvianne, F. & Liu, C. Chromatin domains in space and their functional implications. Curr. Opin. Plant Biol. 54, 1–10 (2019).
Meszaros, B., Erdos, G. & Dosztanyi, Z. IUPred2A: context-dependent prediction of protein disorder as a function of redox state and protein binding. Nucleic Acids Res. 46, W329–W337 (2018).
Theune, M. L., Bloss, U., Brand, L. H., Ladwig, F. & Wanke, D. Phylogenetic analyses and GAGA-motif binding studies of BBR/BPC proteins lend to clues in GAGA-motif recognition and a regulatory role in brassinosteroid signaling. Front. Plant Sci. 10, 466 (2019).
Santi, L. et al. The GA octodinucleotide repeat binding factor BBR participates in the transcriptional regulation of the homeobox gene Bkn3. Plant J. 34, 813–826 (2003).
Ishizaki, K., Chiyoda, S., Yamato, K. T. & Kohchi, T. Agrobacterium-mediated transformation of the haploid liverwort Marchantia polymorpha L., an emerging model for plant biology. Plant Cell Physiol. 49, 1084–1091 (2008).
Sugano, S. S. et al. CRISPR/Cas9-mediated targeted mutagenesis in the liverwort Marchantia polymorpha L. Plant Cell Physiol. 55, 475–481 (2014).
Ishizaki, K. et al. Development of gateway binary vector series with four different selection markers for the liverwort Marchantia polymorpha. PLoS ONE 10, e0138876 (2015).
Liu, C. et al. Genome-wide analysis of chromatin packing in Arabidopsis thaliana at single-gene resolution. Genome Res. 26, 1057–1068 (2016).
Servant, N. et al. HiTC: exploration of high-throughput ‘C’ experiments. Bioinformatics 28, 2843–2844 (2012).
Schmid, M. W. et al. Extensive epigenetic reprogramming during the life cycle of Marchantia polymorpha. Genome Biol. 19, 9 (2018).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Hu, B. et al. Plant lamin-like proteins mediate chromatin tethering at the nuclear periphery. Genome Biol. 20, 87 (2019).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Contreras-Lopez, O., Moyano, T. C., Soto, D. C. & Gutierrez, R. A. Step-by-step construction of gene co-expression networks from high-throughput Arabidopsis RNA sequencing data. Methods Mol. Biol. 1761, 275–301 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Liu, T. Use Model-based Analysis of ChIP-Seq (MACS) to analyze short reads generated by sequencing protein–DNA interactions in embryonic stem cells. Methods Mol. Biol. 1150, 81–95 (2014).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Lin, P. C. et al. Identification of miRNAs and their targets in the liverwort Marchantia polymorpha by integrating RNA-seq and degradome analyses. Plant Cell Physiol. 57, 339–358 (2016).
Acknowledgements
We thank S. Czemmel from the Center for Quantitative Biology (University of Tübingen) for their assistance with sequencing. We are grateful for inspiring discussions with members of the COST Action CA1612 INDEPTH network. We acknowledge computing support by the High Performance and Cloud Computing Group at the Zentrum für Datenverarbeitung of the University of Tübingen, the state of Baden-Württemberg through bwHPC and the German Research Foundation (DFG) through grant no. INST 37/935-1 FUGG. This work was supported by the Deutsche Forschungsgemeinschaft (LI 2862/4) and the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 757600).
Author information
Authors and Affiliations
Contributions
C.L. conceived and designed the experiments. E.S.K., N.W., N.F. and H.B. established and characterized transgenic lines. E.K. performed ChIP–seq, ATAC–seq and RNA-seq experiments. N.W. performed FISH and immunostaining experiments. Y.L. performed coexpression analysis. S.A.M. and F.B. performed epigenomic profiling. K.W.B. performed nuclei sorting. C.L., E.S.K. and S.L. performed Hi-C experiments. E.S.K. and C.L. wrote the manuscript with contributions from other authors. All authors read and accepted the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Stefan de Folter, Stefan Grob and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The Marchantia genome has different types of TADs.
a, Epigenetic marks across Marchantia TADs. ‘mCG-poor’ and ‘mCG-rich’ TADs are shown in green and brown curves, respectively. b, Comparison of gene expression according to gene locus location. Thalli transcriptome data was from (doi: 10.1093/pcp/pcw020). Boxplots from left to right: n = 16707, 2264, 972, 215 and 575. c, Chromatin accessibility across ‘mCG-poor’ (green) and ‘mCG-rich’ (brown) TADs. d, Clustering analysis of TADs according to histone marks. The epigenetic profiling of various histone marks was from our previous study (doi: 10.1016/j.cub.2019.12.015).
Extended Data Fig. 2 Co-expression analysis.
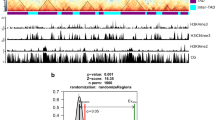
a, Clustering of transcriptome datasets used for calculating gene co-expression. The dendrogram shows hierarchical clustering based on Euclidean distance. b, Distribution of expression correlation coefficients as a function of distance. The distance of a given gene pair was determined according to their annotated TSSs. Boxplots from left to right: n = 6455, 9068, 9238, 9165, 8992, 8931, 8832, 9053, 8736, and 8793. c, d, TADs contain more co-expressed genes than expected. c, For all the gene pairs located in the same TAD, the fraction of co-expressed gene pairs (q < 0.05) was computed and indicated by a black dot at the top of the panel (highlighted with an arrow). The boxplot with twenty blue data points denotes results in which TADs were randomly assigned to the genome. The p-value is an empirical p-value calculated based on twenty simulations. d, Numbers of co-expressed and not co-expressed gene pairs in each round of shuffled TADs (simulation) are shown. As TADs are gene-poor (Fig. 1c), randomly shuffling TADs leads more genes (hence gene pairs) to overlap with TADs. The boxplots in b and c indicate the median (line within the box), the lower and upper quartiles (box), margined by the largest and smallest data points which are still within the interval of 1.5 times the interquartile range from the box (whiskers); outliers are not shown.
Extended Data Fig. 3 Motif analysis links TCP1 to TAD borders.
a, A phylogenetic tree of DNA binding domains of TCP proteins in Arabidopsis thaliana (green), Oryza sativa (yellow), and Marchantia polymorpha (blue). b, Alignment of the TCP DNA-binding domain sequences from two Marchantia TCP proteins and founder members of the TCP family. c-e, Motif analysis of Marchantia TAD borders. The scatter plot in c shows enrichment of motifs, which are recognized by various plant transcription factors (position weight matrices are according to the Arabidopsis DAP-seq dataset (doi: https://doi.org/10.1016/j.cell.2016.04.038)), at TAD boundary regions. The fold enrichment of a motif was calculated as the relative density of this motif in a 2 kb region overlapping with TAD borders compared to that in 20 kb region flanking TAD borders. The presence of motifs in query DNA sequences was determined by the ‘matchPWM’ function in the ‘Biostrings’ package in R, with the search stringency set to 85%. Red and black dots depict motifs recognized by class I and II TCP members, respectively. d, e, Analysis of TCP class I (d) and class II (e)consensus binding sites based on text search. Only sequences exactly matching the query motif were counted. Note that these consensus sequences, determined by Kosugi and Ohashi (doi: https://doi.org/10.1046/j.1365-313x.2002.01294.x), partially overlap.
Extended Data Fig. 4 Genome-wide identification of TCP1 target regions.
a, Verification of anti-TCP1 antibody for immunoprecipitation. TCP1:GFP fusion proteins under the control of 35S promoter were expressed transiently in Nicotiana benthamiana leaves. The presence of TCP1:GFP in each sample was examined by using anti-GFP antibody. Similar results were observed from two independent experiments. b, Snapshot showing reads distribution of ChIP-seq reads in different samples. tcp1 represents a TCP1 knock-out line (in Tak-1 background). See ‘methods’ for details of knock-out line generation. c, Venn diagram of genomic regions enriched in each biological replicate. Below this Venn diagram, the J(rep1, rep2) indicates the Jaccard index. d, Epigenetic marks across TCP1-bound chromatin regions (grey block).
Extended Data Fig. 5 Comparison of insulation scores of chromatin regions around wild-type and tcp1 mutant TAD borders.
a, Comparison of insulation scores in wild-type TAD borders with those in tcp1 no. 18-1 (blue plots) or in tcp1 no. 24-8 (green plots). See Fig. 2e legend for the definition of ‘TCP1-bound’ TAD borders. The titles above these plots, which indicate bin positions, are as those under boxplots in Fig. 2e. Assuming that TCP1 plays a structural role on TCP1-bound TAD borders, we expect that the removal of TCP1 results in specific changes in insulation scores of these regions compared to regions not bound by TCP1. The violin plots in this panel show distribution of changes in insulation scores in the mutant Hi-C maps. For each pair of comparison (that is, changes in insulation scores of TCP1-bound TAD borders vs. TCP1-free TAD borders), its p-value from the two-sided Mann-Whitney U test is given. To assess effect size, the cohen’s d (c’d) is also given below each p-value note. In general, the difference between two populations is considered ‘trivial’ or ‘negligible’ when the absolute value of cohen’s d is less than 0.2. b, Metagene plots showing chromatin contacts around TAD border regions. Pixels in the plots stand for 2-kb bins in the Hi-C matrices. For each plot, TAD borders are aligned and indicated with a dotted triangle. With careful inspection, we conclude that the differences of chromatin organization between TCP1-bound TAD borders and TCP1-free TAD borders are comparable in Tak-1 and tcp1, and loss-of-TCP1 does not led to drastic structural changes in TCP1-bound TAD boundaries.
Extended Data Fig. 6 Epigenetic and transcriptional profiling of TCP1-rich TADs.
Comparison of various epigenetic marks (a) and chromatin accessibility (b) between TCP1-rich TADs (blue curves) and the rest TADs (gray curves) belonging to the ‘mCG-poor’ category. c, d, Comparison of genes (c) and repeats (d) in different TADs. Same as those in panel b, the blue and gray curves denote TCP1-rich TADs and rest TADs in the ‘mCG-poor’ category, respectively. The brown curves denotes ‘mCG-rich’ TADs. Labels are the same as in Fig. 1b.
Extended Data Fig. 7 Changes in gene expression in relation to TCP1 binding, and the distribution of differentially expressed genes in tcp1 in relation to their location.
a, Changes in expression of genes bound by TCP1. Only genes with their gene bodies (defined as their transcribed region plus 0.5 kb flanking regions) overlapping with TCP1 ChIP-seq peaks are included in this plot. b, Distribution of gene expression changes in tcp1 mutants. All the genes from the genome are divided into four groups according to the extent to which they overlap with TCP1 ChIP-seq peaks. c, d, Differentially expressed genes bound (c) and not bound (d) by TCP1 are divided into different groups. The p value indicates two-sided Fisher’s exact test result. The term ‘regular TADs’ in these two panels refers to TADs that are not annotated as TCP-rich.
Extended Data Fig. 8 Motif analysis of Marchantia TAD borders and TADs.
a, Motif analysis of Marchantia TAD borders. This plot is the same as Extended Data Fig. 3c, but highlighting motifs of a few transcription factor families. b, Motif analysis of Marchantia mCG-poor and TCP1-rich TAD bodies. Fold enrichment was calculated as the ratio of motif density in TADs over that in 30 kb flanking genomic regions. Other than that, the motif search was performed as for Extended Data Fig. 3c.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6.
Supplementary Table 1
Number of Hi-C reads.
Supplementary Table 2
TADs annotation.
Supplementary Table 3
Motifs at TAD borders and within mCG-rich TADs.
Supplementary Table 4
TCP1 ChIP–seq peaks.
Supplementary Table 5
Motifs at TCP1-bound chromatin.
Supplementary Table 6
RNA-seq count table.
Supplementary Table 7
Motif position weight matrices.
Supplementary Table 8
ATAC–seq peaks.
Supplementary Table 9
FISH oligonucleotides.
Source data
Source Data Fig. 2
Statistical source data, related to Fig. 2.
Source Data Fig. 3
Statistical source data, related to Fig. 3.
Source Data Fig. 5
Statistical source data, related to Fig. 5.
Source Data Fig. 6
Statistical source data, related to Fig. 6.
Source Data Extended Data Fig. 2
Statistical source data, related to Extended Data Fig. 2.
Source Data Extended Data Fig. 4
Unprocessed western blots, related to Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Statistical source data, related to Extended Data Fig. 5.
Rights and permissions
About this article
Cite this article
Karaaslan, E.S., Wang, N., Faiß, N. et al. Marchantia TCP transcription factor activity correlates with three-dimensional chromatin structure. Nat. Plants 6, 1250–1261 (2020). https://doi.org/10.1038/s41477-020-00766-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-00766-0
- Springer Nature Limited
This article is cited by
-
The plant nuclear lamina disassembles to regulate genome folding in stress conditions
Nature Plants (2023)
-
The 3D architecture of the pepper genome and its relationship to function and evolution
Nature Communications (2022)
-
Homology-mediated inter-chromosomal interactions in hexaploid wheat lead to specific subgenome territories following polyploidization and introgression
Genome Biology (2021)
-
Charting the genomic landscape of seed-free plants
Nature Plants (2021)