Abstract
Oxoglutarate dehydrogenase-like (OGDHL) is considered to be the isoenzyme of oxyglutarate dehydrogenase (OGDH) in the OGDH complex, which degrades glucose and glutamate. OGDHL was reported to reprogram glutamine metabolism to suppress HCC progression in an enzyme-activity-dependent manner. However, the potential subcellular localization and non-canonical function of OGDHL is poorly understood. We investigated the expression of OGDHL and its effect on HCC progression. By employing a variety of molecular biology techniques, we revealed the underlying mechanism of OGDHL-induced DNA damage in HCC cells in vitro and in vivo. AAV loaded with OGDHL exerts therapeutic effect on mouse HCC and prolongs survival time. OGDHL induces DNA damage in HCC cells in vitro and in vivo. We also observed that OGDHL possesses nuclear localization in HCC cells and OGDHL-induced DNA damage was independent of its enzymatic activity. Mechanistically, it was demonstrated that OGDHL binds to CDK4 in the nucleus to inhibit the phosphorylation of CDK4 by CAK, which in turn attenuates E2F1 signaling. Inhibition of E2F1 signaling downregulates pyrimidine and purine synthesis, thereby inducing DNA damage through dNTP depletion. We clarified the nuclear localization of OGDHL and its non-canonical function to induce DNA damage, which demonstrated that OGDHL may serve as a select potential therapeutic target for HCC.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma is one of the most common and deadly malignant tumors in the world [1, 2], especially in Asia [3]. Metabolic reprogramming is one of the hallmarks of tumor cells [4], evoked by carcinogenic factors to ensure that tumor cells obtain sufficient nutrients from the external environment [5]. Unsurprisingly, whether glycolysis [6, 7], the tricarboxylic acid cycle [8, 9], the pentose phosphate pathway [10, 11], or amino acid metabolism [12,13,14] and lipid metabolism [15], mitochondria are at the core. Therefore, it is meaningful to focus on dysregulated metabolic enzymes in mitochondria of HCC cells.
OGDHL is downregulated in a variety of cancers [16,17,18] and exerts tumor suppressor effects through multiple mechanisms [19, 20]. OGDHL has been reported to inhibit HCC progression by reprogramming glutamine metabolism, in an enzymatic activity-dependent manner [20]. However, the regulatory effects of metabolic enzymes arise not only from their canonical activities, but also from their non-canonical or non-metabolic activities [21]. In preliminary experiments, we accidentally discovered that OGDHL, which was originally thought to be localized in mitochondria, has nuclear localization in HCC cells, according to which we speculated that OGDHL may retain potential non-canonical functions.
Obviously, this phenomenon is widespread. In details, Lactate dehydrogenase A (LDHA), whose tetrameric structure is altered by reactive oxygen species (ROS), then translocates into the nucleus and induces histone methylation by interacting with telomere silencing 1-like (DOT1L) [22]. Further, Glycolytic glyceraldehyde-3-phosphate dehydrogenase (GAPDH) directly binds to OCT-1 in the nucleus to promote H2B transcription in S phase [23]. In addition, nuclear fructose-1,6-bisphosphatase 2 (FBP2) interacts with c-Myc and inhibits c-Myc-dependent Transcription of mitochondrial transcription factor A (TFAM) to repress mitochondrial function [24]. Clearly, the presence of non-canonical functions of metabolic enzymes is often associated with their aberrant subcellular localization. Therefore, this study was designed to evaluate the non-canonical function of OGDHL and its application in the treatment of HCC.
We identified the downregulation of OGDHL in human and mouse HCC, explored its therapeutic potential using AAV loaded with OGDHL. We further determined the nuclear localization of OGDHL and its potential nuclear localization sequences. Functionally, we observed that OGDHL induced DNA damage in HCC cells in vitro and in vivo. More importantly, OGDHL-mediated DNA damage in HCC cells was independent on its enzymatic activity. Mechanistically, OGDHL in the nucleus interacts with CDK4, suppressing the phosphorylation of CDK4 by CAK, thereby inhibiting E2F1 signaling. E2F1 is a key protein involved in DNA synthesis [25, 26], cell proliferation, apoptosis [27], metabolism and cell cycle [28], and its function is regulated by RB1. Unphosphorylated RB1 binds to and mediates E2F1 transcriptional repression [29]. Attenuated E2F1 signaling hinders pyrimidine and purine biosynthesis, which leads to DNA damage in HCC cells through dNTP depletion. Overall, these findings clarify the tumor suppressor effect of OGDHL’s non-canonical function on HCC, and reveal that OGDHL may serve as a suitable therapeutic target for HCC.
Results
OGDHL possesses nuclear localization
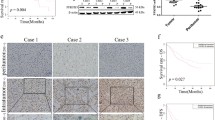
OGDHL was significantly down-regulated in clinical HCC samples (Fig. S1A, B), which also occurred in mouse HCC specimens (Fig. S1C, D).
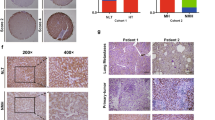
In the human cervical epithelial cell line ME180, OGDHL was observed in mitochondria [19]. Hep3B was selected for confocal immunofluorescence of OGDHL, and it was clear that a large fraction of OGDHL’s fluorescence coincided with the nucleus (Fig. 1A). This was also confirmed by subcellular fractionation of HCC cells (Fig. 1B). Further, a nuclear localization sequence(NLS) (560SKDKK564) for OGDHL was predicted [30] and validated. By replacing 3 lysines in NLS with alanines, we generated a nuclear-free form of OGDHL, annotated OGDHL/3KA (Fig. 1C, D). 293T cells were transfected with the WT and 3KA plasmids of OGDHL, Flag tag carried by the plasmids was used for immunofluorescence assay. Impressively, the 3KA mutation significantly restricted the nuclear localization of OGDHL (Fig. 1E). Additionally, we constructed AAV-OGDHL vector containing liver-specific transthyretin promoter and its control AAV vector. The Flag tag carried by AAV-OGDHL was used for more precise in vivo subcellular localization analysis. The staining results of mouse liver and clinical HCC samples further confirmed the fact that OGDHL possesses nuclear localization (Fig. 1F, G).
A Confocal images in Hep3B cells showed that OGDHL has nuclear localization. B Western blotting further revealed the nuclear localization of OGDHL in HCC cell lines. C Western blotting of WT and 3KA mutants of OGDHL in 293T cells. D The nuclear localization of the 3KA mutant of OGDHL was significantly attenuated. E Confocal images further revealed the subcellular localization of WT and 3KA mutants of OGDHL. F Immunofluorescence images after AAV-OGDHL treatment, and Flag tags were used to trace OGDHL. G Immunohistochemical staining results of OGDHL in clinical HCC samples.
Increased OGDHL induces DNA damage in HCC cells
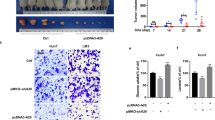
OGDHL inhibits the proliferation of HCC cells in vitro and in vivo (Fig. 2A–C). Subsequent KEGG enrichment analysis revealed marked downregulation of signals such as “DNA replication” and “pyrimidine metabolism” (Fig. 2D, E). Therefore, OGDHL may exert a tumor suppressor effect by mediating DNA damage in HCC cells. Comet assay (Fig. 2F) andγH2AX immunofluorescence confocal imaging (Fig. 2G) proved that OGDHL could induce DNA damage in HCC cells. Besides, elevated OGDHL downregulated FANCD2, which co-localizes with EdU representing active DNA synthesis (Fig. 2H). Activation of DNA damage response proteins such as ATM (pSer1981), ATR (pSer428), CHK1 (pSer317), CHK2 (pThr68) provides direct evidence for OGDHL-mediated DNA damage (Fig. 2I).
A Clonogenic assay in Huh7 cells, suggesting that upregulation of OGDHL inhibits HCC cell proliferation. **p < 0.01. B CCK8 assay in Huh7 cells also demonstrated that OGDHL inhibited HCC cell proliferation. *p < 0.05. C OGDHL could inhibit HCC cell growth in vivo. ***p < 0.001. D Results of KEGG enrichment analysis of transcriptome sequencing after overexpression of NC or OGDHL in Huh7 cells. These are up-regulated signals. E These are down-regulated signals. F The results of the comet assay suggested that OGDHL could induce DNA damage in HCC cells. **p < 0.01. G Confocal images of γH2AX in Huh7 cells further confirmed OGDHL-mediated DNA damage in HCC cells. *p < 0.05. H Confocal images indicated that the co-localization of FANCD2 foci with EdU in Huh7 cells was significantly reduced after OGDHL overexpression. *p < 0.05. I Western blotting revealed that upregulation of OGDHL in Huh7 cells triggered the activation of DDR proteins.
Hydrodynamic tail vein injection was adopted to construct a spontaneous HCC model in mice. AAV-OGDHL could also induce DNA damage in HCC cells (Fig. S2a) and activate DDR proteins (Fig. S2b) in vivo. However, our data indicated that OGDHL-3KA did not induce DDR proteins upregulation in HepG2, as did OGDHL-WT (Fig. S3).
Intranuclear OGDHL induces DNA damage, independent of its enzyme activity
To clarify the significance of OGDHL nuclear translocation on DNA damage of HCC cells, we constructed an OGDHL variant carrying MYC NLS signal peptide at N-terminal, OGDHL-NLS (Fig. 3A). The verification of its nuclear positioning efficiency has obtained satisfactory results (Fig. 3B, C). OGDHL-NLS can also induce DNA damage and inhibit HCC progression in vitro and in vivo (Fig. 3D, F, G), which is independent of its enzyme activity (Fig. 3E). The IHC of subcutaneous tumor further confirmed the efficient nuclear translocation of OGDHL-NLS (Fig. S4).
A Western blotting of WT and NLS mutants of OGDHL in HCCLM3 cells. B Confocal images in HCCLM3 cells showed that OGDHL-NLS has Strict nuclear enrichment. C Western blotting further revealed the nuclear localization of OGDHL-NLS in HCC cell lines. D Western blotting showed that OGDHL-NLS could also induce DNA damage in HCC cells. E OGDHL-NLS variant has no enzymatic activity in HCC cells. ***p < 0.001. F OGDHL-NLS can inhibit the growth of HCC cells in vivo. ***p < 0.001. G Representative images of immunohistochemical staining results of mouse HCC.
OGDHL mediates dNTP depletion in HCC cells
Since transcriptomics showed that OGDHL down-regulated nucleotide metabolism, we speculated that dNTP depletion might be the primary factor of OGDHL-mediated DNA damage in HCC cells. The MS results in HCC cell line confirmed our hypothesis (Fig. 4A, B). Further, the additional dNTPs compensated for the OGDHL-mediated upregulation of DDR proteins in HCC cell (Fig. S5). Moreover, OGDHL induced the decrease of other types of nucleotides in HCC cells. (Fig. S6).
Besides, We found that OGDHL could promote HCC cell apoptosis and G1 phase arrest (Fig. S7), as well as activate p53 signaling in vitro and in vivo (Fig. S8). Taken together, OGDHL mediates DNA damage in HCC cells by triggering dNTP depletion.
Nuclear OGDHL interacts with CDK4 through its Dehydrogenase, E1 component domain
To explore the interacting proteins of OGDHL, we searched on IntAct (https://www.ebi.ac.uk/intact/home), of which CDK4 caught our attention (Fig. 5A). Immunofluorescence colocalization assays in HCC cells indicated that OGDHL and CDK4 share a common and striking nuclear localization, revealing that they may interact in the nucleus (Fig. 5B). Exogenous co-immunoprecipitation experiments provide further confirmation (Fig. 5C, D). We also validated the direct interaction between OGDHL and CDK4 in LSPR assays. Human OGDHL protein has a relatively conserved domain structure: a small N-terminal domain, a Dehydrogenase, E1 component domain, a Transketolase-like, pyrimidine-binding domain and a C-terminal domain. CDK4 contains only one protein kinase domain (Fig. 5E). We found that OGDHL bound CDK4 through its Dehydrogenase, E1 component domain (Fig. 5F).
A We identified potential interacting proteins of OGDHL from IntAct(https://www.ebi.ac.uk/intact/search?query=id:Q9ULD0*#interactions). B The immunofluorescence confocal experiments of OGDHL and CDK4 in Hep3B cells suggested that the binding of the two may be mainly in the nucleus. C Results of exogenous co-immunoprecipitation assay in 293 T cells, IP: OGDHL-Flag, IB: CDK4-GFP. D Results of exogenous co-immunoprecipitation assay in 293T cells, IP: CDK4-GFP, IB: OGDHL-Flag. E OGDHL-GST captured on COOH chip can bind CDK4-MBP with an affinity constant of 39.4 nM as determined in a LSPR assay. F Schematic diagram of the protein domains of OGDHL and CDK4, and the specific domain that binds to CDK4.
OGDHL suppresses CAK-mediated phosphorylation of CDK4 at Thr172
Uncontrolled cell proliferation is a hallmark of cancer, and dysregulated cell cycle progression is common [4]. CDK4, a member of the Ser/Thr protein kinase family, is essential for progression in the G1 phase of the cell cycle. CDK4 has been shown to be responsible for the phosphorylation of the retinoblastoma gene product (Rb) [28, 31], in which CAK (MNAT1-CCNH-CDK7)-mediated phosphorylation of CDK4 at Thr172 is required [32, 33].
Elevated OGDHL in HCC cells greatly inhibited the phosphorylation of CDK4 at Thr172 (Fig. 6A, B). AAV-OGDHL also induces pCDK4 (Thr172) reduction in HCC cells in vivo (Fig. 6C, D). Moreover, IHC data from our clinical HCC samples revealed a negative correlation between OGDHL and pCDK4 (Thr172) at the protein level (Fig. 6E). To directly test whether OGDHL binding specifically blocks CDK4 phosphorylation, we constructed an in vitro CDK4 phosphorylation system. As expected, gradual addition of OGDHL protein to the in vitro CDK4 phosphorylation system inhibited the phosphorylation of CDK4 in a dose-dependent manner (Fig. 6F). Intranuclear OGDHL also induced the decrease of CDK4 phosphorylation in HCC cells (Fig. S9).
A The results of Western blotting indicated that overexpression of OGDHL in Huh7 resulted in decreased phosphorylation of CDK4. ***p < 0.001. B Overexpression of OGDHL in HepG2 also resulted in decreased phosphorylation of CDK4. *p < 0.05; **p < 0.01. C AAV-OGDHL induces downregulation of the phosphorylation level of CDK4 in HCC cells in vivo. *p < 0.05; **p < 0.01. D Representative images of IHC of HCC tissue from AAV-treated mice, detecting pCDK4. E Relative expression of OGDHL and pCDK4 in clinical HCC samples. F Results of in vitro phosphorylation assays. The loading amounts of OGDHL-GST were: 0, 0.8 ug, 1.6 ug, 2.4 ug, 3.2 ug. The loading amount of CDK4-MBP was 1ug. The loading amount of CAK complex was 1.5 ug. With the gradual increase of OGDHL-GST, the phosphorylation of CDK4 gradually decreased.
OGDHL inhibits the expression of rate-limiting enzymes of the nucleotide synthesis pathway by targeting CDK4-pRB-E2F1signaling
OGDHL was negatively associated with genes involved in pyrimidine metabolism at the mRNA level (Fig. 7A), although we focused on the rate-limiting ones (Fig. 7B). Overexpression of OGDHL in HCC cells not only reduced phosphorylated RB1, but also attenuated the rate-limiting enzymes in the nucleotide synthesis pathway (Fig. 7C–E).
A Heatmap showing the effect of OGDHL on genes involved in the de novo pyrimidine synthesis pathway in Huh7 cells. B A brief illustration of the de novo synthesis pathway of pyrimidine. C WB was used to identify the effect of OGDHL on the expression of pRB1 and the rate-limiting enzymes in the de novo pyrimidine and purine synthesis pathways in HCC cell lines. D AAV-OGDHL induced the down-regulation of pRB1 and rate-limiting enzymes in the de novo pyrimidine and purine synthesis pathways in vivo. E Relative protein quantification results. *p < 0.05; **p < 0.01; ***p < 0.001. F Knockdown of E2F1 modulates the expression of rate-limiting enzymes in the de novo synthesis pathway of pyrimidine and purine in HepG2. G Overexpression of E2F1 induces up-regulation of the rate-limiting enzymes in the de novo synthesis pathway of pyrimidine and purine in HepG2. H Effects of CDK4/6 inhibitors on the expression of pRB1 and rate-limiting enzymes in the de novo synthesis pathway of pyrimidine and purine in HepG2. I OGDHL inhibits the expression of rate-limiting enzymes in the de novo pyrimidine and purine biosynthesis pathway in a CDK4-dependent manner. J CHIP-PCR assay confirmed that E2F1 directly combined with the promoter regions of CAD and ADSL.
By knocking down or up-regulating E2F1 in HCC cells, we found that both nucleotide synthesis rate-limiting enzymes and p53 were affected (Fig. 7F, G). CDK4/6 inhibitor reduced the levels of pRB1 and the aforementioned rate-limiting enzymes in a dose-dependent manner (Fig. 7H). Moreover, we demonstrated that OGDHL-mediated downregulation of nucleotide synthesis rate-limiting enzymes is CDK4-dependent (Fig. 7I). We selected CAD and ADSL as representatives, and further CHIP-PCR confirmed the binding of E2F1 to their promoter regions (Fig. 7J).
OGDHL inhibits tumor growth and prolongs overall survival in vivo
More importantly, we assessed the effect of OGDHL on HCC progression and overall survival in mice in vivo (Fig. 8A). Each tumor-bearing mouse was injected with 7 × 10^10 virus particles through the tail vein, and sacrificed after 2 or 4 weeks. On the gross image of liver, the tumors of the mice in the AAV-OGDHL group were more regression (Fig. 8B). The tumor number and largest diameter were also smaller than those of AAV-NC group (Fig. 8C, D). Histologically, we further confirmed the in vivo inhibitory effect of AAV-OGDHL on HCC cells (Fig. 8E, F). Not only that, we found that AAV-OGDHL prolonged the overall survival time of tumor-bearing mice compared to the AAV-NC group (Fig. 8G). Taken together, our data suggest that OGDHL has a reliable in vivo therapeutic effect on HCC in vivo.
A Schematic representation of in vivo experiments to judge the effect of AAV-OGDHL treatment on tumor development and long-term survival of mice. B Representative images of mouse livers after 2 or 4 weeks of AAV-NC or AAV-OGDHL treatment. C Comparative data on the growth of liver tumors in mice after 2 weeks of AAV-NC or AAV-OGDHL treatment. *p < 0.05. D Comparative data on the growth of liver tumors in mice after 4 weeks of AAV-NC or AAV-OGDHL treatment. *p < 0.05. E Representative images of immunohistochemical staining results of mouse liver tumors after 2 weeks of AAV-NC or AAV-OGDHL treatment. F Quantitative results of immunohistochemical staining. *p < 0.05; **p < 0.01. G Kaplan–Meier survival analysis in mice.
Discussion
Tumor-specific promoter methylation is considered to be an important inducement for the down-regulation of OGDHL in various cancers [16, 17]. The expression of other components of OGDHC (Fig. S10A) in HCC was not as unambiguous as OGDHL. There was no significant difference between DLST and DLD in HCC cell lines and normal immortalized hepatocyte L02. Only OGDH, like OGDHL, was more abundant in L02 than in HCC cell lines (Fig. S10B, C). Curiously, OGDH showed no expression difference in TCGA-LIHC database (Fig. S10D). However, overexpression of OGDHL in Huh7 cells resulted in altered expression of OGDH and DLST (Fig. S10E). Theoretically, both the enzymatic and non-enzymatic functions of OGDHL can inhibit the biological behavior of HCC cells. However, whether the two are completely independent or intermingled is unknown.
A key observation of this study is the nuclear localization of OGDHL in HCC cells. Unlike the N-terminal signal peptides of proteins localized within the mitochondrial or endoplasmic reticulum, nuclear localization signals are always stable and may appear anywhere in the protein’s amino acid sequence [34]. Nuclear localization signals are generally divided into two categories: classical (cNLS) and non-classical (ncNLS) [30, 35]. cNLS possess one or two clusters of positively charged amino acid sequences, typically arginine (R), lysine (K) or proline (P) [36]. However, ncNLS does not have the above characteristics, and a large number of ncNLS do not have regular feature structures. Even, some proteins do not have NLS, but can use an independent nuclear transport mechanism to achieve entry into the nucleus, such as β-catenin [37]. The function and regulation of proteins are highly complex and unpredictable, precise and efficient methodological changes and in-depth exploration will bring light to this.
Interestingly, the subcellular localization of OGDHL in different tissues and cell lines varies greatly. OGDHL-deficient 293T cells were transfected with the OGDHL plasmid, and subsequent immunofluorescence images indicated that the signals were all observed in the nucleus, whereas in liver cells, they were mainly localized in the cytoplasm. Unexpectedly, the subcellular localization of OGDHL in HCC cell lines was also different. The intracellular translocation process of OGDHL should be regulated. How OGDHL translocation is regulated and whether subcellular shuttling of OGDHL implies better prognosis in HCC patients remains unclear and will be part of our future studies. Exploring the potential role of OGDHL in liver diseases such as NASH and liver failure is also of high priority.
Abnormal nuclear localization suggests that OGDHL has abundant and undefined capabilities. Not only mediating DNA damage, our pilot experiments observed that OGDHL enhances antitumor immunity in HCC. Oncolytic adenovirus is a popular and promising treatment for solid tumors. Oncolytic adenovirus expressing OGDHL may have stronger intratumoral proliferation and oncolytic ability. Further exploration and verification are needed.
In conclusion, Our data reveal that OGDHL triggers downregulation of E2F1 signaling by binding to CDK4 and inhibiting its phosphorylation, which in turn leads to reduced nucleotide synthesis, dNTP depletion, and subsequent DNA damage. Not only that, we also demonstrated that OGDHL-induced DNA damage in HCC cells, independent of its enzymatic activity, is a non-canonical function. In particular, our animal experiments suggest that OGDHL may serve as a potential therapeutic target for HCC.
Materials and methods
Cell culture
All cell lines used in this study were cultured in Dulbecco’s modified Eagle’s medium (Gibco, USA) containing 10% fetal bovine serum (Hyclone, USA). and 5% dual antibodies(Beyotime, China). A 37 °C, 5% CO2 incubator was used to provide a suitable culture environment. Contamination-free cells with normal growth status were used for subsequent detection and experiments.
Reagents and antibodies
jetPRIME® (Illkirch, France) was used for plasmid and siRNA transfection. dNTP Mixture (25 mM each)(Beyotime, China). Abemaciclib(Selleck, USA). The primary antibodies used in this study are listed in Table S1.
Vector construction
All plasmids for in vitro HCC cell experiments used in this study were synthesized by Wuhan GeneCreate Biological Engineering Co.Ltd and cloned after the CMV promoter of the pcDNA3.1(+) vector. Plasmids for protein purification and expression were synthesized and provided by GeneCodex (Wuhan, China).
Mouse models
pAAV-TBG-MCS-3×Flag and pAAV-TBG-OGDHL-3×Flag-tWPA, were constructed by OBiO Technology (Shanghai) Corp., Ltd. The 5-week-old C57BL/6 J mice were induced to develop a spontaneous HCC model by high-pressure hydrodynamic injection into the tail vein (30). Based on our laboratory’s observations, the combination regimen of NRas/cMYC/SB can induce overt HCC in C57BL/6 J mice at around 8 weeks. Therefore, after 8 weeks, each tumor-bearing mouse was injected with 7 × 10^10 virus particles of AAV-NC or AAV-OGDHL by tail vein injection. It should only be injected once to avoid an immune response against AAV. Samples were collected after 2 weeks and 4 weeks of AAV treatment, respectively, to determine the growth status of liver tumors in the two groups. In addition, some mice were used to measure survival time.
LC-MS/MS
HPLC-MS/MS (shimadzu LC20AD - API 3200MD TRAP) was used to detect intracellular dNTPs and NTP alterations. The liquid nitrogen quick-frozen cell samples were resuspended, centrifuged, diluted with methanol aqueous solution, and then directly tested on the machine. Observe the ion chromatogram and standard curve of each index, and obtain the experimental data.
Comets assay
Reagent Kit for Single Cell Gel Electrophresis Assay (elkbiotech,Wuhan) was used for comet assay to identify cellular DNA damage. According to the manufacturer’s instructions. Briefly, cells were washed with ice-cold PBS, centrifuged, and coated on slides of triple agarose gel. Pre-chilled Lysis Bufffer fully lyses cells. Place the slide in a horizontal electrophoresis tank. Pour into the freshly prepared alkaline electrophoresis buffer and place at room temperature for 20 ~ 60 min, in order to unwind the DNA under alkaline conditions and generate an alkali-labile segment, so that the DNA fragment can easily migrate in the electric field. Voltage 25 V, electrophoresis for 20 ~ 30 min, after neutralization and staining, observed under excitation light of 515 ~ 560 nm wavelength by fluorescence microscope.
Recombinant protein expression
Both GST-tagged and MBP-tagged recombinant proteins were expressed in E. coli strain BL21. Cells were harvested by centrifugation and lysed in ice-cold PBS containing 1% Triton X-100 and 1% w/v protease inhibitor. Lysates were purified by affinity chromatography. Elutes was concentrated and stored at −80 °Cfor further analysis. CAK complex (CDK7 & CCNH & MNAT1 Protein, Human, Recombinant (His Tag)) was purchased from SinoBiological (Beijing Yiqiao Shenzhou Technology Co., Ltd., China).
ChIP‐qPCR
ChIP assays were performed using the ChIP Assay Kit (Beyotime, China) following manufacturer’s instructions. Briefly, formaldehyde was added to the cell culture medium to sufficiently cross-link the target protein and genomic DNA, washed and lysed, and then sonicated to shear the genomic DNA. After centrifugation, the supernatant was collected and mixed with IgG and E2F1 antibodies, respectively, and mixed overnight at 4 °C with slow rotation or rocking. Then, after washing and de-crosslinking, it can be used for PCR detection of the target gene.
LSPR assay
Install the COOH chip according to the OpenSPRTM instrument standard operating procedure and start running at the maximum flow rate (150 µl/min) with the detection buffer PBS (pH 7.4). After the signal baseline has stabilized, the sample loop is flushed with buffer and emptied. Adjust the buffer flow rate to 20 µl/min. Load the EDC/NHS (1:1) solution to activate the chip. Diluted 200 µl of ligand protein was run for 4 min, and after binding stabilized, the sample loop was flushed with buffer. 200 µL of Blocking Solution was loaded, the sample loop was flushed with buffer, and evacuated with air. The baseline was observed for 5 minutes to ensure stability. The analyte was diluted with buffer, and the concentration was shown in the experimental results. The sample was loaded at 20 µl/min. The binding time of protein and ligand was 240 s; the natural dissociation was 360 s. Increase the flow rate to 150 µl/min and inject the appropriate regeneration buffer to remove the analyte. The analysis software used in this experiment was TraceDrawer (Ridgeview Instruments ab, Sweden), and the analysis method was the One To One analysis model.
Immunohistochemistry (IHC)
Briefly, the paraffin sections were dewaxed to water, placed in a retrieval box filled with citric acid antigen retrieval buffer (pH 6.0) for antigen retrieval in a microwave oven. Next, the sections were placed in 3% hydrogen peroxide solution, incubated at room temperature for 25 min in the dark, and then washed 3 times in PBS (pH 7.4) to block endogenous peroxidase. After serum blocking, the primary antibody can be added and incubated overnight at 4 °C in a humidified chamber. After secondary antibody incubation, DAB color development and counterstaining of nuclei, dehydration and mounting. Finally, the results were judged. The hematoxylin-stained nuclei were blue, and the positive expression of DAB was brown.
Western blot (WB)
Briefly, samples after quantification of BCA protein concentration were loaded on SDS-PAGE gel, and electrophoresis was performed at 80 V on the stacking gel and 120 V on the separating gel until bromophenol blue reached the lower edge of the gel plate. Cut the gel, transfer the membrane at a constant current of 300 mA, and adjust the transfer time according to the molecular weight of the target protein. The transferred membrane was blocked by adding blocking solution at room temperature for 1 h, removing the blocking solution, and adding the primary antibody diluted with primary antibody diluent at 4 °C overnight. The diluted primary antibody was recovered, washed three times with TBST, 5 min each time, added the diluted secondary antibody with secondary antibody diluent, incubated at room temperature for 30 min, and washed four times with TBST on a shaker at room temperature, 5 min each time. Finally, a freshly prepared ECL mixed solution (A:B = 1:1) was added dropwise to the protein side of the membrane, exposed in a dark room, and the exposure conditions were adjusted according to different light intensities, developing and fixing. The AlphaEaseFC software processing system analyzes the optical density value of the target band.
Statistical analyses
The western blots were analyzed with ImageJ software (National Institute of Health, USA) and the data from more than three independent experiments were averaged. Significance was determined with 2-tailed Student’s t test. All p < 0.05 values were considered significant. Kaplan-Meier analysis was used for survival studies. All recipient mice were age and gender-matched, therefore no formal randomization was performed.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Villanueva A. Hepatocellular Carcinoma. N. Engl J Med. 2019;380:1450–62.
Llovet JM, Villanueva A, Lachenmayer A, Finn RS. Advances in targeted therapies for hepatocellular carcinoma in the genomic era. Nat Rev Clin Oncol. 2015;12:408–24.
Zou H, Li M, Lei Q, Luo Z, Xue Y, Yao D, et al. Economic Burden and Quality of Life of Hepatocellular Carcinoma in Greater China: A Systematic Review. Front Public Health. 2022;10:801981.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Pavlova NN, Thompson CB. The Emerging Hallmarks of Cancer Metabolism. Cell Metab. 2016;23:27–47.
WARBURG O. On the origin of cancer cells. Science. 1956;123:309–14.
Seyfried TN, Shelton LM. Cancer as a metabolic disease. Nutr Metab (Lond). 2010;7:7.
Hui S, Ghergurovich JM, Morscher RJ, Jang C, Teng X, Lu W, et al. Glucose feeds the TCA cycle via circulating lactate. Nature. 2017;551:115–8.
Selak MA, Armour SM, MacKenzie ED, Boulahbel H, Watson DG, Mansfield KD, et al. Succinate links TCA cycle dysfunction to oncogenesis by inhibiting HIF-alpha prolyl hydroxylase. Cancer Cell. 2005;7:77–85.
Patra KC, Hay N. The pentose phosphate pathway and cancer. Trends Biochem Sci. 2014;39:347–54.
Deberardinis RJ, Sayed N, Ditsworth D, Thompson CB. Brick by brick: metabolism and tumor cell growth. Curr Opin Genet Dev. 2008;18:54–61.
Altman BJ, Stine ZE, Dang CV. From Krebs to clinic: glutamine metabolism to cancer therapy. Nat Rev Cancer. 2016;16:619–34.
Hensley CT, Wasti AT, DeBerardinis RJ. Glutamine and cancer: cell biology, physiology, and clinical opportunities. J Clin Invest. 2013;123:3678–84.
Jain M, Nilsson R, Sharma S, Madhusudhan N, Kitami T, Souza AL, et al. Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation. Science. 2012;336:1040–4.
Currie E, Schulze A, Zechner R, Walther TC, Farese RJ. Cellular fatty acid metabolism and cancer. Cell Metab. 2013;18:153–61.
Hoque MO, Kim MS, Ostrow KL, Liu J, Wisman GB, Park HL, et al. Genome-wide promoter analysis uncovers portions of the cancer methylome. Cancer Res. 2008;68:2661–70.
Ostrow KL, Park HL, Hoque MO, Kim MS, Liu J, Argani P, et al. Pharmacologic unmasking of epigenetically silenced genes in breast cancer. Clin Cancer Res. 2009;15:1184–91.
Jiao Y, Li Y, Fu Z, Hou L, Chen Q, Cai Y, et al. OGDHL Expression as a Prognostic Biomarker for Liver Cancer Patients. Dis Markers. 2019;2019:9037131.
Sen T, Sen N, Noordhuis MG, Ravi R, Wu TC, Ha PK, et al. OGDHL is a modifier of AKT-dependent signaling and NF-kappaB function. Plos One. 2012;7:e48770.
Dai W, Xu L, Yu X, Zhang G, Guo H, Liu H, et al. OGDHL silencing promotes hepatocellular carcinoma by reprogramming glutamine metabolism. J Hepatol. 2020;72:909–23.
Xu D, Shao F, Bian X, Meng Y, Liang T, Lu Z. The Evolving Landscape of Noncanonical Functions of Metabolic Enzymes in Cancer and Other Pathologies. Cell Metab. 2021;33:33–50.
Liu Y, Guo JZ, Liu Y, Wang K, Ding W, Wang H, et al. Nuclear lactate dehydrogenase A senses ROS to produce alpha-hydroxybutyrate for HPV-induced cervical tumor growth. Nat Commun. 2018;9:4429.
Zheng L, Roeder RG, Luo Y. S phase activation of the histone H2B promoter by OCA-S, a coactivator complex that contains GAPDH as a key component. Cell. 2003;114:255–66.
Huangyang P, Li F, Lee P, Nissim I, Weljie AM, Mancuso A, et al. Fructose-1,6-Bisphosphatase 2 Inhibits Sarcoma Progression by Restraining Mitochondrial Biogenesis. Cell Metab. 2020;31:174–88.
Chen HZ, Tsai SY, Leone G. Emerging roles of E2Fs in cancer: an exit from cell cycle control. Nat Rev Cancer. 2009;9:785–97.
DeGregori J, Kowalik T, Nevins JR. Cellular targets for activation by the E2F1 transcription factor include DNA synthesis- and G1/S-regulatory genes. Mol Cell Biol. 1995;15:4215–24.
Qin XQ, Livingston DM, Kaelin WJ, Adams PD. Deregulated transcription factor E2F-1 expression leads to S-phase entry and p53-mediated apoptosis. Proc Natl Acad Sci. 1994;91:10918–22.
Kent LN, Leone G. The broken cycle: E2F dysfunction in cancer. Nat Rev Cancer. 2019;19:326–38.
Helin K, Harlow E, Fattaey A. Inhibition of E2F-1 transactivation by direct binding of the retinoblastoma protein. Mol Cell Biol. 1993;13:6501–8.
Nguyen BA, Pogoutse A, Provart N, Moses AM. NLStradamus: a simple Hidden Markov Model for nuclear localization signal prediction. Bmc Bioinforma. 2009;10:202.
Sherr CJ, Beach D, Shapiro GI. Targeting CDK4 and CDK6: From Discovery to Therapy. Cancer Discov. 2016;6:353–67.
Kato JY, Matsuoka M, Strom DK, Sherr CJ. Regulation of cyclin D-dependent kinase 4 (cdk4) by cdk4-activating kinase. Mol Cell Biol. 1994;14:2713–21.
Matsuoka M, Kato JY, Fisher RP, Morgan DO, Sherr CJ. Activation of cyclin-dependent kinase 4 (cdk4) by mouse MO15-associated kinase. Mol Cell Biol. 1994;14:7265–75.
Martoglio B, Dobberstein B. Signal sequences: more than just greasy peptides. Trends Cell Biol. 1998;8:410–5.
Bradley KJ, Bowl MR, Williams SE, Ahmad BN, Partridge CJ, Patmanidi AL, et al. Parafibromin is a nuclear protein with a functional monopartite nuclear localization signal. Oncogene 2007;26:1213–21.
Willis AN, Dean SE, Habbouche JA, Kempers BT, Ludwig ML, Sayfie AD, et al. Nuclear localization signal sequence is required for VACM-1/CUL5-dependent regulation of cellular growth. Cell Tissue Res. 2017;368:105–14.
Sharma M, Jamieson C, Johnson M, Molloy MP, Henderson BR. Specific armadillo repeat sequences facilitate beta-catenin nuclear transport in live cells via direct binding to nucleoporins Nup62, Nup153, and RanBP2/Nup358. J Biol Chem. 2012;287:819–31.
Funding
This work was supported by grants from the National Natural Science Foundation of China (81871967, 82173129).
Author information
Authors and Affiliations
Contributions
Conceptualization: XJ, DY. Project administration: DY. Funding acquisition: DY; performed majority of the experiments and Writing original draft: XJ, YX, QL, YX and JH; Writing – review & editing: XJ, DY, SX; Investigation: CC, YW, LZ; Data curation: HL, YL, BL, JP; Supervision: DY.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics aprroval
The authors declare no competing interests. The research was approved by Experimental Animal Ethics Committee of Nanjing Hospital Affiliated to Nanjing Medical University (DWSY-2105599).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiang, X., Peng, J., Xie, Y. et al. Oxoglutarate dehydrogenase-like inhibits the progression of hepatocellular carcinoma by inducing DNA damage through non-canonical function. Cell Death Differ 30, 1931–1942 (2023). https://doi.org/10.1038/s41418-023-01186-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-023-01186-1
- Springer Nature Limited
This article is cited by
-
Development and Validation of a Carbohydrate Metabolism-Related Model for Predicting Prognosis and Immune Landscape in Hepatocellular Carcinoma Patients
Current Medical Science (2024)
-
Magnetic vagus nerve stimulation alleviates myocardial ischemia-reperfusion injury by the inhibition of pyroptosis through the M2AChR/OGDHL/ROS axis in rats
Journal of Nanobiotechnology (2023)