Abstract
DNA-damaging agents exploit increased genomic instability, a hallmark of cancer. Recently, inhibitors targeting the DNA damage response (DDR) pathways, such as PARP inhibitors, have also shown promising therapeutic potential. However, not all tumors respond well to these treatments, suggesting additional determinants of response are required. Schlafen 11 (SLFN11), a putative DNA/RNA helicase that induces irreversible replication block, is emerging as an important regulator of cellular response to DNA damage. Preclinical and emerging clinical trial data suggest that SLFN11 is a predictive biomarker of response to a wide range of therapeutics that cause DNA damage including platinum salts and topoisomerase I/II inhibitors, as well as PARP inhibitors, which has raised exciting possibilities for its clinical application. In this article, we review the function, prevalence, and clinical testing of SLFN11 in tumor biopsy samples and circulating tumor cells. We discuss mounting evidence of SLFN11 as a key predictive biomarker for a wide range of cancer therapeutics and as a prognostic marker across several cancer types. Furthermore, we discuss emerging areas of investigation such as epigenetic reactivation of SLFN11 and its role in activating immune response. We then provide perspectives on open questions and future directions in studying this important biomarker.
Similar content being viewed by others
Introduction
Genomic instability and resistance to cell death are important hallmarks of cancer [1]. For decades, the cornerstones of cancer treatment have included direct DNA-damaging chemotherapies (such as platinum salts, alkylating agents, antimetabolites, topoisomerase I/II inhibitors) and radiation therapy, with varying degrees of response in unselected patients. More recently, small molecule inhibitors that directly target DNA damage response (DDR) have garnered significant interest. A poster child of this approach is the use of poly(ADP-ribose) polymerase inhibitors (PARPi), particularly in cancers harboring BRCA1 or BRCA2 (BRCA1/2) mutations, which are key proteins in the repair of double-stranded DNA breaks through homologous recombination repair (HRR) [2].
The Schlafen (SLFN; in German Schlafen means “sleeping”) family of genes were first described in 1998 as growth regulatory genes that affect thymocyte development and inhibit cell growth [3]. SLFN has ten known mouse isoforms (SLFN1, 1 L, 2, 3, 4, 5, 8, 9, 10, and 14) [4,5,6], and five human isoforms (SLFN5, 11, 12, 13, and 14) [6,7,8,9,10,11]. SLFN11 was recently identified as a biomarker predicting response to a wide range of DNA-damaging agents and PARPi in preclinical settings, across multiple cancer types including small cell lung cancer (SCLC) [12, 13], sarcoma [14], breast cancer [15], ovarian cancer [11, 16], colorectal cancer (CRC) [17], gastric cancer [18], and mesothelioma [19]. This finding has raised exciting possibilities for the use of SLFN11 as a biomarker in clinical applications. In this article, we review the function, prevalence, and clinical testing of SLFN11. We also review the current evidence of SLFN11 as a biomarker across different cancer types, with an emphasis on SCLC. Last, we discuss SLFN11 regulation and strategies to re-express SLFN11, and its role in activating an immune response.
SLFN11 as a key regulator in DNA damage response
DDR is a collective term for the various cellular pathways activated in response to DNA damage, which lead to downstream events such as cell-cycle arrest and DNA repair, with deficient DDR resulting in cell death. Key differentiating features of DDR in cancer compared with normal cells include increased endogenous DNA damage due to higher levels of mutations and proliferation, dysregulation of key proteins in DDR pathways such as BRCA1/2 and PARP, and increased replication stress [20]. Replication stress can be caused by intrinsic pressures such as the loss of retinoblastoma protein (Rb) causing premature entry into S-phase [21, 22], or extrinsically from the effects of DNA-damaging agents.
In response to replication stress, ATR/CHK1-mediated DDR is activated, resulting in slowing or stalling of replication fork progression and uncoupling of DNA polymerases from the replisome helicase [23]. Recently, SLFN11 has been identified as a key protein recruited in replication stress that induces an irreversible replication block and cell death [11, 24,25,26]. SLFN11 protein has an N-terminus nuclease domain and a C-terminus helicase domain that contains ATP binding Walker A and B motifs and a replication protein A (RPA)-binding region [25,26,27]. As an early response to replication stress, SLFN11 is recruited to the chromatin and stalled replication forks through its binding to RPA, which coats exposed single-stranded DNA at stalled forks. SLFN11 then interacts with the replicative helicase MCM3 (minichromosome maintenance complex component 3) as well as DHX9 (DExH-box helicase 9) [26, 28]. SLFN11 also causes the opening of chromatin near the replication initiation sites, thus blocking replication fork progression [26]. In addition, SLFN11 also regulates a genomic response to replication stress by globally enhancing chromatin accessibility at promoter regions and selectively activating transcription of immediate early genes, which is thought to further contribute to SLFN11-mediated DDR response [29]. A recent study shed more insight into how SLFN11 causes an irreversible replication block. During DNA damage, SLFN11 was shown to bind to DDB1-CUL4CDT2 E3 ubiquitin via its C-terminus ATPase domain and promote the degradation of CDT1, a key replication licensing factor [30, 31]. Without CDT1, replication origin firing is blocked, resulting in irreversible replication arrest [30, 32]. Another recent publication by the same group showed SLFN11 protected cells from unfolded protein response and endoplasmic reticulum stress by binding to protein folding and translation initiation factors [33]. Cells expressing SLFN11 had lower level of polyubiquitylation and reduced protein aggregation; while SLFN11-deficient cells were highly dependent on protein chaperonin and folding genes, and were sensitive to a ubiquitin-activating enzyme UBA1 inhibitor TAK-243.
SLFN11 has also been shown to induce inhibition of translation of ATR through cleavage of specific type II tRNAs through its N-terminus nuclease activity [34]. Therefore, SLFN11 acts as a master regulator of DDR to replication stress, independent of ATR [26, 31]. Accordingly, cells deficient in SLFN11 are reliant on ATR/CHK1-mediated replication stress response and DNA repair [25, 26, 31]. ATR elicits only a transient replication block in response to DNA damage, allowing cells to repair the DNA damage and recover replication, contributing to chemoresistance. Consequently, ATR and CHK1 inhibitors have been shown to sensitize otherwise resistant SLFN11-low cells to chemotherapy and PARPi [25, 31]. In addition to a replication block, reduced RPA loading and impaired HRR-mediated DNA repair have also been observed in SLFN11-proficient cells as a late response to DNA damage [24]. This is suggested to be secondary to a total replication block preventing generation of sister chromatids for homologous recombination [26]. Recently, SLFN11 has also been shown to bind a ribosomal protein (RPS4X) and block mTOR pathway activation [35], suggesting crosstalk with other signaling pathways to regulate tumor progression and perhaps DDR. Therefore, current evidence suggests that SLFN11 enhances sensitivity to DNA-damaging agents through several molecular mechanisms (Fig. 1).
SLFN11 also triggers a series of molecular events as listed. When SLFN11 is absent, ATR is recruited to RPA, and triggers ATR/CHK1-mediated DNA repair, hence the replication fork is repaired and replication resumes. IEGs immediate early genes, ER endoplasmic reticulum, HRR homologous recombination repair. Created with Biorender.com.
SLFN11 clinical testing
An immunohistochemistry (IHC) assay for the detection of SLFN11 has been used as a reliable method to quantify expression of SLFN11, as reported in several cancer types [36,37,38,39]. SLFN11 IHC consistently shows prominent nuclear localization [13, 25, 40] and based on our experience, tumors that express SLFN11 typically have diffuse moderate to strong nuclear staining for SLFN11 by IHC, while tumors that do not express SLFN11 have a distinct absence of staining. In practice, this effectively translates to tumoral nuclear staining of any intensity to be positive and complete loss of SLFN11 to be negative. The H-score, which is a function of percentage of positive cells and the intensity of cells stained, is a good marker for SLFN11 positivity (defined as H-score ≥1) (Fig. 2a). In addition to IHC, RNA sequencing (RNA-seq) and methylome analyses can be useful to assess SLFN11 expression in clinical samples. Having more than one expression assay may be valuable to score tumors with full confidence. Quantifying SLFN11 expression in circulating tumor cells (CTCs) using an immunofluorescence assay is also under development. In one study with prostate cancer patients, SLFN11 CTCs showed 85.7% (six of seven patients) concordance with SLFN11 mRNA expression of matched metastatic tumor biopsies [41]. We also previously demonstrated the unique advantage of tracking SLFN11 expression in CTCs longitudinally under different treatment conditions, in a non-invasive manner [42, 43].
a SLFN11 expression in human tumor tissue by immunohistochemistry (IHC). a: Positive for SLNF11 in SCLC: diffuse strong tumoral nuclear labeling. b: Negative for SLFN11 in SCLC: tumor cells with complete absence of SLFN11 labeling, note endothelial cells with nuclear labeling which can be used as a positive internal control. c: Normal lung: scattered lung macrophages with strong nuclear labeling, pneumocytes lining alveoli spaces is negative for SLFN11. SCLC: small cell lung cancer. (IHC Images from MD Anderson pathology department, courtesy of Dr. Junya Fujimoto. Images have not been published before. b SLFN11 mRNA expression in TCGA pan-cancer dataset.
Prevalence of SLFN11 in tumors
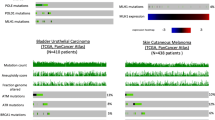
Previous reports have consistently shown that about 50% of all cancer cell lines express SLFN11 protein [7, 11, 26]. When comparing SLFN11 transcript levels in cell lines (by mRNA expression) with patient-derived xenograft models (PDXs; by IHC) from the same cancer tissue of origin, some concordance but significant variations were noted [44]. For example, high expression levels of SLFN11 were noted in both cell lines and in PDXs of bladder cancer, while for renal cell carcinoma higher SLFN11 expression was noted in cell lines than in PDXs. This discrepancy is not surprising as in vitro tumor cells grow in a very different setting than in vivo PDX models. Nonetheless, in a cohort of breast cancer PDXs, SLFN11 transcript and protein levels were highly correlated, which suggests that these assessment methods equally reflect levels in the tissue [44]. In 51 SCLC cell lines, SLFN11 protein and mRNA expression were found to be bimodally distributed, and also highly correlated with SLFN11 expression by IHC in PDXs [13]. Similarly, in a cohort of 47 patients with relapsed SCLC, ~50% of tumors were SLFN11 positive by IHC (H-score > =1) [38].
Notably, in an analysis of pan-cancer data from The Cancer Genome Atlas (TCGA) in 2012, SLFN11 mRNA expression was found to be highest in Ewing sarcoma, across 40 cancer types [7]. In a clinical cohort of 20 patients with Ewing sarcoma, SLFN11 was expressed in 95% of the cases [45]. About 85% of Ewing sarcomas harbor EWS-FLI, a fusion protein that encodes an oncogenic ETS transcription factor [46]. EWS-FLI1 has shown to bind to the ETS consensus site on the SLFN11 promoter and mediate SLFN11 expression, which sensitizes Ewing sarcoma cells to camptothecin and to PARPi plus temozolomide combinations [47]. The co-expression of FLI1 and SLFN11 has also been observed in other cancers including colon, breast, and prostate cancer and leukemias [47, 48]. In contrast, SLFN11 was only expressed in 25% of primary prostate cancers and 45% of metastatic castration-resistant prostate cancers in a cohort of 197 patients [41]. We also analyzed SLFN11 mRNA expression in TCGA pan-cancer data recently, which showed highest SLFN11 expression in mesothelioma, renal cell carcinoma, and sarcoma (Fig. 2b). As SLFN11 is highly prevalent among different cancer types and easy to measure by IHC, it can be rapidly translated to the clinical setting as a biomarker.
SLFN11 as a predictive biomarker for DNA-damaging agents and PARPi in multiple cancers
In preclinical studies, SLFN11 expression predicts sensitivity to many DNA-damaging agents including platinum salts (e.g. cisplatin, carboplatin, oxaliplatin), topoisomerase I inhibitors (topotecan, irinotecan, camptothecin, indotecan), topoisomerase II inhibitors (etoposide, doxorubicin, and epirubicin), alkylating agents (cyclophosphamide, temozolomide), antimetabolites (5-fluorouracil, gemcitabine, cytarabine, hydroxyurea), and PARPi (olaparib, veliparib, talazoparib, niraparib) [11, 13, 14, 18, 25, 28, 49, 50], which is not unexpected given its key function in DDR. Conversely, the loss of SLFN11 expression leads to resistance to these agents [7, 11, 28]. These observations are seen across a variety of cancer cell lines and PDXs, including SCLC [13], sarcoma [14], breast cancer [15], ovarian cancer [11, 16], CRC [17, 51], gastric cancer [18], and mesothelioma [19]. Furthermore, sensitivity to PARPi in SLFN11 positive tumors is found to be independent of BRCA1/2 mutations and HRR deficiency in preclinical models [13, 25], supporting clinical relevance beyond just tumors with inherent DDR mutations/HRR defects.
The PARP family of proteins plays critical roles in DNA repair through various DDR pathways. Targeting of PARP family proteins with PARPi disrupts DDR, and lead to eventual cell death. HRR deficient cells (e.g. with BRCA1/2 mutations) show greater reliance on PARP activity to maintain cell survival, thus are particularly vulnerable to PARPi. The use of PARPi in BRCA mutant breast cancer was the first reported example of a targeted therapy exploiting synthetic lethality to kill cancer cells [52]. Further studies have found that “PARP trapping”, a property of PARPi that “traps” PARP1/2 at the site of DNA damage and prevents DNA repair, may be important for the synergistic effect with SLFN11-mediated cell death [25]. The degree of PARP trapping varies among PARPi, with talazoparib being the strongest and veliparib the weakest [25, 53,54,55,56]. Therefore, the predictive strength of SLFN11 may differ among PARPi depending on the varying degree of PARP trapping.
In a recent study analyzing SLFN11 expression by IHC and testing the effect of combination treatment strategies of DNA-damaging chemotherapy with DDR inhibitors in different cancer types, the authors found chemotherapy (specifically gemcitabine) in combination with some DDR inhibitors, such as ATR, WEE1, or CHK1 inhibitors, re-sensitized SLFN11-low cancers [25, 44, 57]. Surprisingly, the study showed the PARPi olaparib had limited impact on SLFN11 in a cohort of breast cancer PDXs, contradicting previous observations in other cancer types [37, 58]. These results highlight the need to prospectively test treatment strategies with SLFN11 quantification in specific tumor types to confirm its predictive biomarker status.
SLFN11 as a prognostic biomarker in multiple cancers
The prognostic value of SLFN11 has been demonstrated in several cancer types including hepatocellular carcinoma (HCC) [35], CRC [36, 51, 59], breast cancer [15, 60, 61] and prostate cancer [41]. In a cohort of 3278 patients with CRC treated with 5-fluorouracil, leucovorin and irinotecan (FOLFIRI), high SLFN11 expression in patients with tumors with microsatellite instability (MSI) was associated with a better outcome compared to those with low SLFN11 expression [59]. However, in patients with microsatellite-stable (MSS) tumors, the opposite was observed. In another study, KRAS-wild-type CRC patients with high SLFN11 expression had significantly better overall survival (OS) compared to those with low SLFN11 expression [36]. These observations suggest a potential prognostic role of SLFN11 in addition to MSI and KRAS status in CRC.
Gene expression microarray profiling data from 7737 breast cancer cases were used to investigate the transcriptional landscape of SLFN11 in breast cancer [60, 61]. SLFN11 gene expression showed strong positive association with immune processes as well as tumor-infiltrating lymphocyte markers (CD3 and CD8). High SLFN11 gene expression was also independently associated with better prognosis [61]. In another retrospective analysis of 250 patients, patients with triple-negative breast cancer showed longer metastasis-free survival and OS in patients with higher tumor SLFN11 expression by IHC [15].
In a study of 182 paired tumor and non-tumor liver tissues from patients with HCC, SLFN11 expression was lower in about 75% of tumor tissues compared with non-tumor tissues [35]. Low SLFN11 expression correlated with high levels of alpha-fetoprotein, large tumor size, presence of microvascular invasion, and advanced stage. Survival analyses of this cohort following treatment with hepatectomy revealed longer OS and a lower recurrence rate in patients with tumors that had higher expression of SLFN11.
SLFN11 as a promising biomarker in SCLC
SLFN11 has recently emerged as a promising predictive biomarker in SCLC, a recalcitrant cancer lacking specific therapeutic targets and biomarkers. In preclinical studies using cell lines and PDXs, SLFN11 protein expression strongly predicted cisplatin and PARPi responses [13, 37, 62]. Our group and others have shown high SLFN11 expression to be associated with sensitivity to PARPi and cisplatin in SCLC cell lines and PDXs, while knock-down of SLFN11 induced resistance to these drugs [13, 37]. In a randomized Phase II trial of relapsed SCLC treated with the combination of temozolomide plus the PARPi veliparib or placebo, SLFN11-positive patients had significantly longer progression-free survival (PFS) (5.7 vs 3.6 mo, P = 0.009) and OS (12.2 vs.7.5 mo, P = 0.014) compared to SLFN11 negative patients in the temozolomide plus veliparib group [38]. However, no difference in PFS and OS was observed based on SLFN11 positivity in the temozolomide plus placebo group, indicating that SLFN11 expression is a predictive biomarker for veliparib, but not for temozolomide.
Based on the evidence that SLFN11 is a promising predictive biomarker for PARPi in SCLC, as well as the synergistic effects of PARP inhibition with anti-PD-L1 immunotherapy in preclinical models [63,64,65], a Phase II randomized trial of atezolizumab in combination with talazoparib versus atezolizumab alone in SCLC was developed by the SWOG Cancer Research Network (NCT04334941). In this ongoing trial, patients with extensive-stage SCLC will all receive frontline induction therapy with platinum-etoposide plus atezolizumab, while being prospectively screened for SLFN11 expression by IHC. Patients with SLFN11 positive tumors will be eligible to be randomized to one of the two arms in the maintenance phase: atezolizumab with or without talazoparib. As aforementioned, talazoparib has the strongest “PARP trapping” property, a property of PARPi that is similar to that of a cytotoxic chemotherapy [53, 56], and this may confer additional benefit compared to other PARPi in SLFN11 positive patients [25]. The primary objective of the trial is to compare PFS between the two arms, with secondary objectives of comparing OS, objective response and frequency of adverse effects. This is the first trial using SLFN11 prospectively as a biomarker to select patients, and the result will confirm whether patients with SLFN11-positive SCLC derive benefit from PARPi in addition to standard immunotherapy in the maintenance setting.
Epigenetic regulation and dynamic expression of SLFN11
Current evidence suggests the main regulation of SLFN11 expression is at the epigenetic and transcriptional levels, rather than mutations in the SLFN11 gene [27, 62, 66]. SLFN11 expression is frequently lost in cancer cell lines due to epigenetic silencing by hypermethylation at the promoter region of SLFN11, rendering these cells intrinsically resistant to DNA-damaging agents and PARPi [13, 28, 40, 48]. Epigenetic regulation may also account for changes in SLFN11 expression related to treatment and the development of treatment resistance. In an epigenome-wide study of 66 human SCLC cell lines, increased promoter methylation of SLFN11 correlated with low or no SLFN11 expression, which was associated with resistance to DNA-damaging agents [67]. Similarly, in CRC cell lines and patient tumor samples, SLFN11 was found to be frequently methylated in the promoter region, which was associated with SLFN11 downregulation, tumor resistance to cisplatin and decreased 5-year survival [51].
SLFN11 expression has also been shown to be dynamic with treatment pressure. One study generated chemo-resistant SCLC PDXs by treating tumor-bearing mice with cisplatin and etoposide, and found that SLFN11 expression was significantly downregulated in the chemo-resistant models [40] In another study using gastric cancer cell lines, decrease in SLFN11 expression was observed with continuous treatment of oxaliplatin and resistance development, which was reversed by treating the cells with epigenetic modifiers 5-azacytadine or entinostat [18]. In a Phase II clinical trial of cediranib plus olaparib in patients with PARPi-relapsed high-grade serous ovarian cancer, comparison of paired baseline and on-treatment biopsies revealed downregulation of SLFN11 as one of the mechanisms of PARPi resistance [68]. In our analyses of CTCs taken from SCLC patients, SLFN11 protein expression also appeared to be decreased when patients were treated with platinum-based chemotherapy [42, 43]. However, another study showed that SLFN11 protein levels did not decrease following chemotherapy in a prostate cancer cell line [44]. These discrepancies highlight the need for further testing in tumor-specific prospective clinical trial setting, with longitudinal patient data on SLFN11 status.
A recent study showed SLFN11 expression to be differentially regulated during B-cell development [69]. The authors speculated the reason for this may be the need to suppress SLFN11-mediated cell death in the setting of rapid proliferation and somatic hypermutations of the immunoglobulins, a critical step in B-cell development that introduces mutations to generate antibody diversity. The study further showed that this suppression was partially achieved by epigenetic modification of SLFN11 and was reversible by EZH2 inhibitor tazemetostat or histone deacetylase (HDAC) inhibitor panobinostat, which rendered germinal center B cells (GCB) more susceptible to cytarabine treatment. This study provided a preclinical rationale for combining epigenetic modifiers to induce SLFN11 expression with cytarabine in GCB-derived lymphomas. Notably, this study also demonstrated the feasibility of a dual IHC assay of SLFN11 and cluster of differentiation markers to further delineate SLFN11 expression in different stages of B cells. This finding shows SLFN11 IHC can be integrated with other potential biomarkers.
Strategies to re-express SLFN11
In preclinical models, reversal of SLFN11 promoter methylation by epigenetic modifiers has led to increased SLFN11 expression and sensitization to DNA-damaging agents. For example, pre-treatment of SLFN11 hypermethylated breast cancer cell lines with the DNA methylation inhibitor 5-azacytadine increased sensitivity to platinum drugs [28]. In another study using SCLC PDXs, treatment with EZH2 inhibitors restored the expression of SLFN11 and chemosensitivity, via a decreased level of H3K27me3 (a histone modification placed by EZH2) [40]. Interestingly, in this study, treatment with 5-azacytadine did not lead to SLFN11 re-expression, in contrary to the previous studies [18, 28]. Another recent study demonstrated low SLFN11 expression was associated with resistance to SG3199, a cytotoxic warhead of an antibody-drug-conjugate; which was abrogated by treating with an EZH2 inhibitor [70]. In addition, class I HDAC inhibitors romidepsin and entinostat was shown to induce SLFN11 expression in cells without promoter methylation and worked synergistically with topoisomerase inhibitor camptothecin [48].
Based on these observations, a Phase I/II trial investigating the combination of EZH1/2 inhibitor DS-3201b and irinotecan in recurrent SCLC (NCT03879798) is ongoing. This trial is testing whether the EZH1/2 inhibitor leads to re-expression of SLFN11, therefore enhancing response to chemotherapy or overcoming chemotherapy resistance that may have been acquired in the context of SLFN11 being downregulated due to prior lines of therapy. More studies are needed to investigate potential synergistic effects between epigenetic modifiers of SLFN11 expression and cytotoxic agents (Fig. 3a).
a Known therapeutic vulnerabilities and strategies of inducing SLFN11-mediated cell death. Potential synergistic effects between cytotoxic agents and epigenetic silencers targeting SLFN11; and potential synergistic effects between DDA and DDR agents. b SLFN11 induces replication block and cell death. Cell death leads to innate immune pathway cGAS-STING activation, which leads to upregulation of IFNs and immune activation. SLFN11 is upregulated by IFNs and increases chromatin accessibility, which leads to activation of IEGs and enhances SLFN11-dependent replication block and immune response. SLFN11 expression can also increase IFNγ-mediated cytotoxic T cell killing. DDA DNA-damaging agents, IEG immediate early genes, IFN interferon. Created with Biorender.com.
SLFN11 and interferon-regulated immune activation
SLFN11 has been implicated in the activation of innate immune response. SLFN11 is recognized as one of the interferon (IFN)-stimulated genes [71]. IFNs are a group of signaling proteins essential to innate and adaptive immunity against pathogens, as well as immune surveillance against cancer [71, 72]. Our group previously showed that in tumor samples from treatment-naive SCLC patients, SLFN11 mRNA expression was positively correlated with type I IFN pathway genes, such as STAT1, JAK1, and IRF9, as well as immune targets, including PD-L1 [13]. Notably, IFNs can be activated in innate immune response via the cGAS-STING signaling pathway. The cGAS-STING pathway can be activated by cytosolic DNA from tumor cells, viruses, and bacteria, and triggers downstream infiltration of effector immune cells such as T cells and natural killer cells [73]. In murine models of SCLC, we previously identified cGAS-STING pathway activation as an important mechanism for synergistic tumor killing through PD-L1 blockade and DDR inhibition (either by targeting PARP or CHK1) [65]. In a Phase I/II trial of olaparib and temozolomide (OT) in combination in patients with previously treated SCLC, SCLC PDXs were generated and treated with OT to model clinical OT sensitivity and resistance, and candidate genes were identified as biomarkers to predict OT sensitivity [74, 75]. The authors found that inflammatory-response genes, specifically those induced by IFN and TGFβ, in addition to SLFN11, were able to predict OT sensitivity. The association between SLFN11 and IFN signaling suggests different measurements of a common mechanism of PARPi sensitivity.
Another report showed that under replication stress, SLFN11 increased global chromatin accessibility and activated a subset of immediate early genes (IEGs) [29]. IEGs can induce cell cycle arrest, and this mechanism likely amplifies the SLFN11-dependent replication block and activates innate immune response, in addition to the direct effect by SLFN11 binding at the replication fork [29].
Intriguingly, virology studies have shown that SLFN11 expression correlates with viral inhibition in a range of viruses, such as HIV, West Nile virus, and Zika virus [9, 76]. Moreover, infection of human cells by the flavivirus West Nile virus and HIV-1 upregulated the expression of SLFN11, via type I IFN-dependent and IFN-independent pathways, respectively [77]. This suggests a common pathway of innate immune activation from replication stress caused by viral infection or DNA-damaging agents.
SLFN11 expression has also been shown to regulate IFN-γ (the type II interferon)-mediated cytotoxic T cell attack. In a study using a whole genome screening of the haploid human cell line HAP1 to investigate modulators of tumor sensitivity to T cell killing, the authors surprisingly found SLFN11 as the single IFN-γ induced gene that mediated tumor cell sensitivity to T cell attack [78]. For example, IFN-γ-mediated T cell toxicity was abrogated by SLFN11 loss in HAP1 cells, while restoration of SLFN11 expression re-sensitized the SLFN11-deficient HAP1 cells to IFN-γ mediated T cell killing. The authors went on further to show that SLFN11 regulated IFN-γ-mediated toxicity without altering IFN-γ receptor signaling activity, but rather by influencing the functional outcome of the signaling.
The interplay between SLFN11 and IFNs suggests SLFN11 may be an important biomarker in the cancer-immunity cycle. Further studies are needed to elucidate the mechanism of SLFN11-mediated antitumor immune response and its role in predicting response to immunotherapy (Fig. 3b).
Open questions and future directions
Despite the abundance of preclinical evidence that SLFN11 expression is predictive of many chemotherapy agents and PARPi targeting DDR, no prospective trials using SLFN11 as a predictive biomarker have been completed. To our knowledge, the randomized Phase II trial of atezolizumab plus talazoparib versus atezolizumab alone in the maintenance setting for SLFN11-positive extensive-stage SCLC (NCT04334941) is the first clinical trial selecting patients with SLFN11-positive tumors. We eagerly await more clinical trials designed to directly and prospectively test SLFN11 as a predictive biomarker, given its broad implications across multiple agents and tumor types.
Although tumor SLFN11 IHC is a reliable and effective method to detect SLFN11 as a tumor biomarker, SLFN11 expression in other cells, particularly immune cells in the tumor microenvironment, also warrants consideration. In a recent study evaluating SLFN11 expression by IHC in about 700 malignant and adjacent non-malignant tissues, SLFN11 expression was found to be tissue specific and highly variable [39, 79]. The study found discrepancies in SLFN11 expression from these tissues compared to the TCGA RNA-seq dataset in several organs and indicated that this may be due to the strong expression of SLFN11 in the infiltrating immune cells present in the TCGA RNA-seq data. This study highlights the consideration of using IHC rather than RNA-seq in order to accurately reflect SLFN11 in patient tumor cells as opposed to immune cells. However, the significance of SLFN11 expression in tumor-infiltrating immune cells is currently unknown and may warrant further investigation.
Another important question about SLFN11 is how its expression changes in response to treatment and the implications of this in therapeutic resistance. As multiple studies have shown that SLFN11 levels are lower following platinum-based and PARPi therapies, and the loss of SLFN11 is associated with treatment resistance [13, 40, 51, 67, 68], serial monitoring of SLFN11 may be important to guide treatment decisions. For example, in SCLC and other cancers using platinum-based treatment, the decision to re-treat with platinum is typically driven by the observation of clinical relapse more than 3 months after the last platinum treatment (i.e. platinum-sensitive). Instead, SLFN11 IHC could be a better way to select patients who should be re-treated with platinum. This, however, would require fresh tissue at the time of relapse, as SLFN11 expression in archival tissue may not have predictive value at the time of relapse. Given the challenges associated with re-biopsies, a blood-based assay such as SLFN11 expression in CTCs may be an attractive alternative to an IHC assay. In blood samples from SCLC patients, SLFN11 protein expression by immunofluorescence could be reliably established in CTCs. Moreover, the SLFN11 expression level varies among patients who were treatment naïve, on platinum treatment, or platinum-relapsed; confirming the dynamic nature of this biomarker [42, 43]. However, SLFN11 detection by CTC requires further validation, including careful compression of SLFN11 expression in CTCs and in tumor samples collected at the same time to ensure they are reflecting the same SLFN11 expression level.
Conclusion
SLFN11 is an important guardian of the genome, suppressing DDR at the stressed replication fork, which leads to irreversible replication block and cell death. Mounting preclinical evidence suggest SLFN11 predicts response to a wide range of DNA-damaging agents including platinum, topoisomerase I/II inhibitors, alkylating agents, antimetabolites, and PARPi. In addition, retrospective clinical data suggest the prognostic and, more importantly, the predictive value of SLFN11 expression across several cancer types. These findings support further investigation of the potential of SLFN11 IHC or other SLFN11 assays as a predictive biomarker that could have clinical applications for matching patients to DNA-damaging chemotherapies (such as platinum drugs and topoisomerase inhibitors) or targeted therapies (such as PARPi). Prospective trials using SLFN11 as a predictive biomarker are needed, as well as effective strategies to monitor SLFN11 expression longitudinally in order to determine its dynamic changes in response to therapy, which may contribute to treatment resistance. Furthermore, the role of SLFN11 in activating innate and adaptive immune response warrants further investigation, especially in relation to treatment with immunotherapy.
Data availability
Not applicable.
References
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Lee JM, Ledermann JA, Kohn EC. PARP Inhibitors for BRCA1/2 mutation-associated and BRCA-like malignancies. Ann Oncol. 2014;25:32–40.
Schwarz DA, Katayama CD, Hedrick SM. Schlafen, a new family of growth regulatory genes that affect thymocyte development. Immunity. 1998;9:657–68.
Ferguson DA, Chiang JT, Richardson JA, Graff J. eXPRESSION: an in silico tool to predict patterns of gene expression. Gene Expr Patterns. 2005;5:619–28.
Neumann B, Zhao L, Murphy K, Gonda TJ. Subcellular localization of the Schlafen protein family. Biochem Biophys Res Commun. 2008;370:62–66.
Bustos O, Naik S, Ayers G, Casola C, Perez-Lamigueiro MA, Chippindale PT, et al. Evolution of the Schlafen genes, a gene family associated with embryonic lethality, meiotic drive, immune processes and orthopoxvirus virulence. Gene. 2009;447:1–11.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7.
Katsoulidis E, Mavrommatis E, Woodard J, Shields MA, Sassano A, Carayol N, et al. Role of interferon {alpha} (IFN{alpha})-inducible Schlafen-5 in regulation of anchorage-independent growth and invasion of malignant melanoma cells. J Biol Chem. 2010;285:40333–41.
Li M, Kao E, Gao X, Sandig H, Limmer K, Pavon-Eternod M, et al. Codon-usage-based inhibition of HIV protein synthesis by human schlafen 11. Nature. 2012;491:125–8.
Razzak M. Genetics: Schlafen 11 naturally blocks HIV. Nat Rev Urol. 2012;9:605.
Zoppoli G, Regairaz M, Leo E, Reinhold WC, Varma S, Ballestrero A, et al. Putative DNA/RNA helicase Schlafen-11 (SLFN11) sensitizes cancer cells to DNA-damaging agents. Proc Natl Acad Sci USA. 2012;109:15030–5.
De Ruysscher D, Lueza B, Le Pechoux C, Johnson DH, O’Brien M, Murray N, et al. Impact of thoracic radiotherapy timing in limited-stage small-cell lung cancer: usefulness of the individual patient data meta-analysis. Ann Oncol. 2016;27:1818–28.
Allison Stewart C, Tong P, Cardnell RJ, Sen T, Li L, Gay CM, et al. Dynamic variations in epithelial-to-mesenchymal transition (EMT), ATM, and SLFN11 govern response to PARP inhibitors and cisplatin in small cell lung cancer. Oncotarget. 2017;8:28575–87.
Kang MH, Wang J, Makena MR, Lee JS, Paz N, Hall CP, et al. Activity of MM-398, nanoliposomal irinotecan (nal-IRI), in Ewing’s family tumor xenografts is associated with high exposure of tumor to drug and high SLFN11 expression. Clin Cancer Res. 2015;21:1139–50.
Coussy F, El-Botty R, Chateau-Joubert S, Dahmani A, Montaudon E, Leboucher S, et al. BRCAness, SLFN11, and RB1 loss predict response to topoisomerase I inhibitors in triple-negative breast cancers. Sci Transl Med. 2020;12:eaax2625.
Shee K, Wells JD, Jiang A, Miller TW. Integrated pan-cancer gene expression and drug sensitivity analysis reveals SLFN11 mRNA as a solid tumor biomarker predictive of sensitivity to DNA-damaging chemotherapy. PLoS ONE. 2019;14:e0224267.
Tian L, Song S, Liu X, Wang Y, Xu X, Hu Y, et al. Schlafen-11 sensitizes colorectal carcinoma cells to irinotecan. Anticancer Drugs. 2014;25:1175–81.
Takashima T, Taniyama D, Sakamoto N, Yasumoto M, Asai R, Hattori T, et al. Schlafen 11 predicts response to platinum-based chemotherapy in gastric cancers. Br J Cancer. 2021. https://doi.org/10.1038/s41416-021-01364-3.
Rathkey D, Khanal M, Murai J, Zhang J, Sengupta M, Jiang Q, et al. Sensitivity of mesothelioma cells to PARP inhibitors is not dependent on BAP1 but is enhanced by temozolomide in cells with high-Schlafen 11 and low-O6-methylguanine-DNA methyltransferase expression. J Thorac Oncol. 2020;15:843–59.
O’Connor MJ. Targeting the DNA damage response in cancer. Mol Cell. 2015;60:547–60.
Bester AC, Roniger M, Oren YS, Im MM, Sarni D, Chaoat M, et al. Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell. 2011;145:435–46.
Buisson R, Boisvert JL, Benes CH, Zou L. Distinct but concerted roles of ATR, DNA-PK, and Chk1 in countering replication stress during S phase. Mol Cell. 2015;59:1011–24.
Byun TS, Pacek M, Yee MC, Walter JC, Cimprich KA. Functional uncoupling of MCM helicase and DNA polymerase activities activates the ATR-dependent checkpoint. Genes Dev. 2005;19:1040–52.
Mu Y, Lou J, Srivastava M, Zhao B, Feng XH, Liu T, et al. SLFN11 inhibits checkpoint maintenance and homologous recombination repair. EMBO Rep. 2016;17:94–109.
Murai J, Feng Y, Yu GK, Ru Y, Tang SW, Shen Y, et al. Resistance to PARP inhibitors by SLFN11 inactivation can be overcome by ATR inhibition. Oncotarget. 2016;7:76534–50.
Murai J, Tang SW, Leo E, Baechler SA, Redon CE, Zhang H, et al. SLFN11 blocks stressed replication forks independently of ATR. Mol Cell. 2018;69:371–84. e376
Murai J, Thomas A, Miettinen M, Pommier Y. Schlafen 11 (SLFN11), a restriction factor for replicative stress induced by DNA-targeting anti-cancer therapies. Pharmacol Ther. 2019;201:94–102.
Nogales V, Reinhold WC, Varma S, Martinez-Cardus A, Moutinho C, Moran S, et al. Epigenetic inactivation of the putative DNA/RNA helicase SLFN11 in human cancer confers resistance to platinum drugs. Oncotarget. 2016;7:3084–97.
Murai J, Zhang H, Pongor L, Tang SW, Jo U, Moribe F, et al. Chromatin remodeling and immediate early gene activation by SLFN11 in response to replication stress. Cell Rep. 2020;30:4137–51. e4136
Aladjem MI, Redon CE. Order from clutter: selective interactions at mammalian replication origins. Nat Rev Genet. 2017;18:101–16.
Jo U, Murai Y, Chakka S, Chen L, Cheng K, Murai J, et al. SLFN11 promotes CDT1 degradation by CUL4 in response to replicative DNA damage, while its absence leads to synthetic lethality with ATR/CHK1 inhibitors. Proc Natl Acad Sci USA. 2021;118.
Pozo PN, Cook JG. Regulation and function of Cdt1; a key factor in cell proliferation and genome stability. Genes. 2016;8:2.
Murai Y, Jo U, Murai J, Jenkins L, Huang SN, Chakka S, et al. SLFN11 inactivation induces proteotoxic stress and sensitizes cancer cells to ubiquitin activating enzyme inhibitor TAK-243. Cancer Res. 2021. https://doi.org/10.1158/0008-5472.CAN-20-2694.
Li M, Kao E, Malone D, Gao X, Wang JYJ, David M. DNA damage-induced cell death relies on SLFN11-dependent cleavage of distinct type II tRNAs. Nat Struct Mol Biol. 2018;25:1047–58.
Zhou C, Liu C, Liu W, Chen W, Yin Y, Li CW, et al. SLFN11 inhibits hepatocellular carcinoma tumorigenesis and metastasis by targeting RPS4X via mTOR pathway. Theranostics. 2020;10:4627–43.
Deng Y, Cai Y, Huang Y, Yang Z, Bai Y, Liu Y, et al. High SLFN11 expression predicts better survival for patients with KRAS exon 2 wild type colorectal cancer after treated with adjuvant oxaliplatin-based treatment. BMC Cancer. 2015;15:833.
Lok BH, Gardner EE, Schneeberger VE, Ni A, Desmeules P, Rekhtman N, et al. PARP inhibitor activity correlates with SLFN11 expression and demonstrates synergy with temozolomide in small cell lung cancer. Clin Cancer Res. 2017;23:523–35.
Pietanza MC, Waqar SN, Krug LM, Dowlati A, Hann CL, Chiappori A, et al. Randomized, double-blind, phase II study of temozolomide in combination with either veliparib or placebo in patients with relapsed-sensitive or refractory small-cell lung cancer. J Clin Oncol. 2018;36:2386–94.
Takashima T, Sakamoto N, Murai J, Taniyama D, Honma R, Ukai S, et al. Immunohistochemical analysis of SLFN11 expression uncovers potential non-responders to DNA-damaging agents overlooked by tissue RNA-seq. Virchows Arch. 2020. https://doi.org/10.1007/s00428-020-02840-6.
Gardner EE, Lok BH, Schneeberger VE, Desmeules P, Miles LA, Arnold PK, et al. Chemosensitive relapse in small cell lung cancer proceeds through an EZH2-SLFN11 axis. Cancer Cell. 2017;31:286–99.
Conteduca V, Ku SY, Puca L, Slade M, Fernandez L, Hess J, et al. SLFN11 expression in advanced prostate cancer and response to platinum-based chemotherapy. Mol Cancer Ther. 2020;19:1157–64.
Byers LA, Stewart CA, Gay CM, Heymach J, Fernandez L, Lu D, et al. SLFN11 and EZH2 protein expression and localization in circulating tumor cells to predict response or resistance to DNA damaging therapies in small cell lung cancerp[Abstract]. In: Proceedings of the 110th Annual Meeting of the American Association for Cancer Researc; 2019 Mar 29–Apr 3; Atlanta, GA, Philadelphia (PA): AACR; 2019. Abstract nr 2215.
Zhang B, Stewart CA, Gay CM, Wang Q, Cardnell R, Fujimoto J, et al. Detection of DNA replication blocker SLFN11 in tumor tissue and circulating tumor cells to predict platinum and PARP inhibitor response in small cell lung cancer [abstract]. In: Proceedings of the 112th Annual Meeting of the American Association for Cancer Research; 2021 Apr 10-15; Virtual meeting. Philadelphia (PA): AACR; 2019. Abstract nr 1876.
Winkler C, Armenia J, Jones GN, Tobalina L, Sale MJ, Petreus T, et al. SLFN11 informs on standard of care and novel treatments in a wide range of cancer models. Br J Cancer. 2021;124:951–62.
Federico SM, Pappo AS, Sahr N, Sykes A, Campagne O, Stewart CF, et al. A phase I trial of talazoparib and irinotecan with and without temozolomide in children and young adults with recurrent or refractory solid malignancies. Eur J Cancer. 2020;137:204–13.
Sankar S, Lessnick SL. Promiscuous partnerships in Ewing’s sarcoma. Cancer Genet. 2011;204:351–65.
Tang SW, Bilke S, Cao L, Murai J, Sousa FG, Yamade M, et al. SLFN11 is a transcriptional target of EWS-FLI1 and a determinant of drug response in Ewing sarcoma. Clin Cancer Res. 2015;21:4184–93.
Tang SW, Thomas A, Murai J, Trepel JB, Bates SE, Rajapakse VN, et al. Overcoming resistance to DNA-targeted agents by epigenetic activation of Schlafen 11 (SLFN11) expression with class i histone deacetylase inhibitors. Clin Cancer Res. 2018;24:1944–53.
Sousa FG, Matuo R, Tang SW, Rajapakse VN, Luna A, Sander C, et al. Alterations of DNA repair genes in the NCI-60 cell lines and their predictive value for anticancer drug activity. DNA Repair. 2015;28:107–15.
Marzi L, Szabova L, Gordon M, Weaver Ohler Z, Sharan SK, Beshiri ML, et al. The Indenoisoquinoline TOP1 inhibitors selectively target homologous recombination-deficient and Schlafen 11-positive cancer cells and synergize with olaparib. Clin Cancer Res. 2019;25:6206–16.
He T, Zhang M, Zheng R, Zheng S, Linghu E, Herman JG, et al. Methylation of SLFN11 is a marker of poor prognosis and cisplatin resistance in colorectal cancer. Epigenomics. 2017;9:849–62.
Lord CJ, Ashworth A. PARP inhibitors: synthetic lethality in the clinic. Science. 2017;355:1152–8.
Pilie PG, Gay CM, Byers LA, O’Connor MJ, Yap TA. PARP inhibitors: extending benefit beyond BRCA-mutant cancers. Clin Cancer Res. 2019;25:3759–71.
Murai J, Huang SY, Das BB, Renaud A, Zhang Y, Doroshow JH, et al. Trapping of PARP1 and PARP2 by clinical PARP inhibitors. Cancer Res. 2012;72:5588–99.
Pommier Y, O’Connor MJ, de Bono J. Laying a trap to kill cancer cells: PARP inhibitors and their mechanisms of action. Sci Transl Med. 2016;8:362ps317.
Murai J, Huang SY, Renaud A, Zhang Y, Ji J, Takeda S, et al. Stereospecific PARP trapping by BMN 673 and comparison with olaparib and rucaparib. Mol Cancer Ther. 2014;13:433–43.
Coleman N, Zhang B, Byers LA, Yap TA. The role of Schlafen 11 (SLFN11) as a predictive biomarker for targeting the DNA damage response. Br J Cancer. 2021;124:857–9.
van Erp AEM, van Houdt L, Hillebrandt-Roeffen MHS, van Bree N, Flucke UE, Mentzel T, et al. Olaparib and temozolomide in desmoplastic small round cell tumors: a promising combination in vitro and in vivo. J Cancer Res Clin Oncol. 2020;146:1659–70.
Ballestrero A, Bedognetti D, Ferraioli D, Franceschelli P, Labidi-Galy SI, Leo E, et al. Report on the first SLFN11 monothematic workshop: from function to role as a biomarker in cancer. J Transl Med. 2017;15:199.
Isnaldi E, Ferraioli D, Ferrando L, Brohee S, Ferrando F, Fregatti P, et al. Correction to: Schlafen-11 expression is associated with immune signatures and basal-like phenotype in breast cancer. Breast Cancer Res Treat. 2019;177:773.
Isnaldi E, Ferraioli D, Ferrando L, Brohee S, Ferrando F, Fregatti P, et al. Schlafen-11 expression is associated with immune signatures and basal-like phenotype in breast cancer. Breast Cancer Res Treat. 2019;177:335–43.
Tlemsani C, Pongor L, Elloumi F, Girard L, Huffman KE, Roper N, et al. SCLC-CellMiner: a resource for small cell lung cancer cell line genomics and pharmacology based on genomic signatures. Cell Rep. 2020;33:108296.
Jiao S, Xia W, Yamaguchi H, Wei Y, Chen MK, Hsu JM, et al. PARP inhibitor upregulates PD-L1 expression and enhances cancer-associated immunosuppression. Clin Cancer Res. 2017;23:3711–20.
Parkes EE, Walker SM, Taggart LE, McCabe N, Knight LA, Wilkinson R, et al. Activation of STING-dependent innate immune signaling by S-phase-specific DNA damage in breast cancer. J Natl Cancer Inst. 2017;109.
Sen T, Rodriguez BL, Chen L, Corte CMD, Morikawa N, Fujimoto J, et al. Targeting DNA damage response promotes antitumor immunity through STING-mediated T-cell activation in small cell lung cancer. Cancer Discov. 2019;9:646–61.
Rajapakse VN, Luna A, Yamade M, Loman L, Varma S, Sunshine M, et al. CellMinerCDB for integrative cross-database genomics and pharmacogenomics analyses of cancer cell lines. iScience. 2018;10:247–64.
Krushkal J, Silvers T, Reinhold WC, Sonkin D, Vural S, Connelly J, et al. Epigenome-wide DNA methylation analysis of small cell lung cancer cell lines suggests potential chemotherapy targets. Clin Epigenetics. 2020;12:93.
Lheureux S, Oaknin A, Garg S, Bruce JP, Madariaga A, Dhani NC, et al. EVOLVE: a multicenter open-label single-arm clinical and translational phase II trial of cediranib plus olaparib for ovarian cancer after PARP inhibition progression. Clin Cancer Res. 2020;26:4206–15.
Moribe F, Nishikori M, Takashima T, Taniyama D, Onishi N, Arima H, et al. Epigenetic suppression of SLFN11 in germinal center B-cells during B-cell development. PLoS ONE. 2021;16:e0237554.
Mao S, Chaerkady R, Yu W, D’Angelo G, Garcia A, Chen H, et al. Resistance to pyrrolobenzodiazepine dimers is associated with SLFN11 downregulation and can be reversed through inhibition of ATR. Mol Cancer Ther. 2021;20:541–52.
Mavrommatis E, Fish EN, Platanias LC. The schlafen family of proteins and their regulation by interferons. J Interferon Cytokine Res. 2013;33:206–10.
Stark GR, Kerr IM, Williams BR, Silverman RH, Schreiber RD. How cells respond to interferons. Annu Rev Biochem. 1998;67:227–64.
Jiang M, Chen P, Wang L, Li W, Chen B, Liu Y, et al. cGAS-STING, an important pathway in cancer immunotherapy. J Hematol Oncol. 2020;13:81.
Farago AF, Yeap BY, Stanzione M, Hung YP, Heist RS, Marcoux JP, et al. Combination olaparib and temozolomide in relapsed small-cell lung cancer. Cancer Discov. 2019;9:1372–87.
Pacheco JM, Byers LA. Temozolomide plus PARP inhibition in small-cell lung cancer: could patient-derived xenografts accelerate discovery of biomarker candidates? Cancer Discov. 2019;9:1340–2.
Valdez F, Salvador J, Palermo PM, Mohl JE, Hanley KA, Watts D, et al. Schlafen 11 restricts flavivirus replication. J Virol. 2019;93:e00104-19.
Borrego AR, Corona-Ayala C, Salvador JC, Valdez FC, Llano M. Gene expression regulation of the type I interferon-induced protein Schlafen 11. Faseb J. 2020;34:1.
Mezzadra R, de Bruijn M, Jae LT, Gomez-Eerland R, Duursma A, Scheeren FA, et al. SLFN11 can sensitize tumor cells towards IFN-gamma-mediated T cell killing. PLoS ONE. 2019;14:e0212053.
Buettner R. Awakening of SCHLAFEN 11 by immunohistochemistry: a new biomarker predicting response to chemotherapy. Virchows Arch. 2021;478:567–8.
Acknowledgements
We would like to acknowledge Ms. Sunita Patterson for manuscript editing, and Lixia Diao, PhD for bioinformatics support.
Funding
This work was supported by: The NIH/NCI CCSG P30-CA016672 (Bioinformatics Shared Resource); NIH/NCI R01-CA207295; The Andrew Sabin Family Fellowship; and through generous philanthropic contributions to The University of Texas MD Anderson Lung Cancer Moon Shot Program.
Author information
Authors and Affiliations
Contributions
BZ and LAB conceived the project. All authors contributed to the writing of the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent to publish
Not applicable.
Competing interests
Dr. Byers serves on advisory committees for AstraZeneca, AbbVie, GenMab, BergenBio, Pharma Mar SA, Sierra Oncology, Merck, Bristol-Myers Squibb, Genentech, and Pfizer and has research support from AbbVie, AstraZeneca, GenMab, Sierra Oncology, Tolero Pharmaceuticals. Dr. Wistuba reports grants and personal fees from Genentech/Roche, grants and personal fees from Bayer, grants and personal fees from Bristol-Myers Squibb, grants and personal fees from AstraZeneca/Medimmune, grants and personal fees from Pfizer, grants and personal fees from HTG Molecular, grants and personal fees from Merck, personal fees from GlaxoSmithKline, grants and personal fees from Guardant Health, personal fees from MSD, grants from Oncoplex, grants from DepArray, grants from Adaptive, grants from Adaptimmune, grants from EMD Serono, grants from Takeda, grants from Amgen, grants from Karus, grants from Johnson & Johnson, grants from Iovance, grants from 4D, grants from Novartis, grants from Oncocyte, grants from Akoya, personal fees from Flame, outside the submitted work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Zhang, B., Ramkumar, K., Cardnell, R.J. et al. A wake-up call for cancer DNA damage: the role of Schlafen 11 (SLFN11) across multiple cancers. Br J Cancer 125, 1333–1340 (2021). https://doi.org/10.1038/s41416-021-01476-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-021-01476-w
- Springer Nature Limited
This article is cited by
-
CRISPR–Cas9 potential for identifying novel therapeutic targets in muscle-invasive bladder cancer
Nature Reviews Urology (2024)
-
Schlafen 11 further sensitizes BRCA-deficient cells to PARP inhibitors through single-strand DNA gap accumulation behind replication forks
Oncogene (2024)
-
Targeting DNA Damage Response Deficiency in Thoracic Cancers
Drugs (2024)
-
PARP inhibitors: enhancing efficacy through rational combinations
British Journal of Cancer (2023)
-
Mouse Slfn8 and Slfn9 genes complement human cells lacking SLFN11 during the replication stress response
Communications Biology (2023)