Abstract
The role of oxytocin in social cognition has attracted tremendous interest in social neuroscience and psychiatry. Some studies have reported improvement in social symptoms following oxytocin treatment in autism spectrum disorders (ASD), while others point to endogenous factors influencing its efficiency and to mixed results in terms of long-term clinical benefits. Epigenetic modification to the oxytocin receptor gene (OXTR) in ASD could be an informative biomarker of treatment efficacy. Yet, little is known about the relationship between OXTR methylation, clinical severity, and brain function in ASD. Here, we investigated the relationship between OXTR methylation, ASD diagnosis (in N = 35 ASD and N = 64 neurotypical group), measures of social responsiveness, and resting-state functional connectivity (rsFC) between areas involved in social cognition and reward processing (in a subset of ASD, N = 30). Adults with ASD showed higher OXTR methylation levels in the intron 1 area compared with neurotypical subjects. This hypermethylation was related to clinical symptoms and to a hypoconnectivity between cortico-cortical areas involved in theory of mind. Methylation at a CpG site in the exon 1 area was positively related to social responsiveness deficits in ASD and to a hyperconnectivity between striatal and cortical brain areas. Taken together, these findings provide initial evidence for OXTR hypermethylation in the intron area as a potential biomarker for adults with ASD with less severe developmental communication deficits, but with impairments in theory of mind and self-awareness. Also, OXTR methylation in the exon 1 area could be a potential biomarker of sociability sensitive to life experiences.
Similar content being viewed by others
Introduction
Oxytocin (OXT) is a key modulator of the most intuitive and yet most complex socioemotional behaviors, including mother–infant attachment [1], pair bond formation [2, 3], consolation [4], and social recognition [5] across several species. By acting in the brain via the oxytocin receptor (OXTR) [6], which is primarily enriched in human subcortical reward and emotional regions [7], OXT affects social cognition by enhancing the salience of social cues and reward sensitivity to these cues [8, 9]. Intranasal OXT application also enhances resting-state functional connectivity (rsFC) between ventral striatal and frontal areas in humans [10, 11]. Pair bond formation in prairie voles, which is OXT-dependent, is associated with functional connectivity between the medial prefrontal cortex (mPFC) and the nucleus accumbens (NAcc) [12]. OXT is associated to various forms of social attachments and affects the activity and the connectivity of a social brain network involving NAcc, PFC, anterior cingulate cortex (ACC), anterior insula (AI), and amygdala [2, 13].
Autism spectrum disorder (ASD) is a neurodevelopmental disorder characterized by impairments in social interactions and communication, as well as repetitive behaviors and restricted interests (American Psychiatric Association, 2013). Based on Research Domain Criteria (RDoC), social deficits in ASD may be represented by two putative domains: (1) positive valence or reward responsiveness and (2) social processes which include facial processing and theory of mind capacities. Large multisite data sharing functional magnetic resonance imaging (fMRI) studies have shown that ASD is characterized by a general decrease in long-range cortico-cortical rsFC between areas involved in social cognition or mentalization, such as mPFC, posterior cingulate cortex (PCC), superior temporal sulcus (STS), and insular cortices [14, 15]; and increased connectivity between striatum and cortical areas [16]. Also, reduced neural activity in the NAcc and in the striatum has been documented in ASD in response to reward-related motivational tasks [17,18,19], highlighting motivational deficits in ASD.
DNA methylation of OXTR within the MT2 region (as shown in Fig. 1) has been shown to negatively correlate with OXTR transcription across tissues [20,21,22]. Variability in OXTR methylation in neurotypical subjects has been associated with differences in social cognition and brain response during social tasks [23,24,25]. In particular, blood oxygenation level dependent (BOLD) activity in brain regions involved in theory of mind and emotional processes (amygdala, ACC, and insula) have been related to the degree of OXTR methylation. Also, decreased functional connectivity between amygdala and other social brain areas were associated with increased OXTR methylation in neurotypical subjects [24]. These results suggest the utility of OXTR methylation as a biomarker of phenotypic variability in sociability.
CpG 16 (b.p. −989) showed significant increase in methylation in ASD as compared with neurotypical subjects (t(97) = −4.38; α = 0.0024; P < 0.0001). The schematic representation of the MT2 region in the bottom of the figure indicates that the Exon 1 includes CpG 1 to CpG 13 and that Intron 1 includes CpG 14 to CpG 27. Errors bars are standard errors.
Today, there is little information about the relationship of OXTR methylation and ASD diagnosis. A previous study has reported hypermethylation in the MT2 region in peripheral and in postmortem tissues in ASD that was associated with decreased OXTR mRNA expression in the brain [21]. However, research from other domains can provide hypotheses about potential links. In animals, low parental care early in life led to de novo OXTR DNA methylation in brain and the periphery, which was associated with a reduced OXTR expression in the NAcc [26]. This is in line with a recent study showing that maltreated children have higher OXTR methylation within the exon 1 area of the MT2 region (CpG 5,6) as compared with typically developed controls [27]. Also, increased methylation at CpG 5,6 was indirectly associated with insecure social attachment via brain volume alterations in the orbitofrontal cortex (OFC) [27]. ASD does not result from poor parental care or from early adversity or neglect. Nonetheless, the clinical phenotype of some individuals with ASD is characterized by lower social engagement, less attention to social cues and low reward sensitivity, resulting in reduced experience of positive social interactions. Our rationale is that this reduction in positive interactions might impact OXTR methylation at a CpG site (5,6) that is shown to be sensitive to extreme negative social experiences.
Despite the rapid growth of the OXT-based research in ASD, no study to date has examined associations between OXTR methylation, severity of ASD-related social symptoms and brain rsFC. Here, we investigated the relationship between salivary DNA OXTR methylation, ASD diagnosis, and symptoms severity in social responsiveness, and rsFC between key areas involved in social cognition and reward processing. Our primary hypothesis is that OXTR hypermethylation is associated with ASD symptomatology and with rsFC between brain regions that are linked to ASD social dysfunction and to OXT’s function in the brain. In an initial study, we tested this hypothesis by comparing methylation in the MT2 region of the OXTR in DNA extracted from salivary samples of 99 adult participants (35 ASD and 64 neurotypical adults) to locate CpG sites associated with ASD diagnosis. Based on the results from the first study, we focused on CpG 16 as a site of interest for subsequent analysis. We also selected CpG 5,6 as another site of interest, given that recent evidence linked its methylation to socially relevant early life experiences [27]. This makes CpG 5,6 an interesting candidate for reflecting phenotypical variability in sociability or social experiences in ASD (such as social avoidance) that are neurodevelopmental in nature. We therefore hypothesize that individuals with ASD who display lower social engagement or responsiveness will have higher OXTR methylation at CpG 5,6. In the second study, we collected rsFC data using fMRI from a subset of 30 adults with ASD. We performed hypothesis-based analysis to pinpoint the relationship between methylation of OXTR CpG sites of interest and rsFC between key brain regions involved in social cognition (STS and PCC) and reward processing (NAcc, vmPFC, amygdala, and ACC/AI). We predicted that OXTR methylation would relate to rsFC in these areas and that the severity of social symptoms would correlate with rsFC between areas involved in reward and social processing.
Materials and methods
Participants
ASD group
Forty adult male subjects with ASD (mean age = 27.02 ± 5.34; 70% Caucasian, 20% African American, 7.5% two or more races, 2.5% unknown) were recruited from the Emory Autism Center (EAC) of the Department of Psychiatry and Behavioral Sciences at the Emory School of Medicine. Subjects were initially part of a larger project, Autism Oxytocin Brain (AOB) trial (registered in clinicaltrials.gov), involving behavioral testing, brain imaging, and intranasal oxytocin administration that are not presented here. Subjects were first selected based on a DSM-5 ASD diagnosis, then confirmed by the Autism Diagnosis Interview-Revised (ADI-R) and the Autism Diagnostic Observation Schedule, 2nd edition (ADOS-2) criteria. Adults with ADHD were not excluded from the study. In addition to diagnostic tests, intellectual quotient (IQ) was also administered to these individuals during the screening session given that an IQ higher than 70 was part of the inclusion criteria. We used the Wechsler Adult Intelligence Scale (WAIS-II) (Table S1 for more details on demographic). Following screening at EAC, subjects participated in two separate studies.
Study 1 procedure
We examined the relationship between OXTR methylation, the Social Responsiveness Scale, 2nd edition (SRS-2) [28] and ASD diagnosis. SRS-2 was completed by parents. It is a widely used clinical measure that reflects severity of social deficits in terms of social motivation and awareness. We used total SRS-2 scores and subscale scores (social communication and interaction (SCI) and restricted and repetitive behaviors (RRB)) to correlate symptom severity with methylation levels. Salivary samples were collected from these subjects among other behavioral tests that are not part of this paper. Thirty-six ASD samples were processed for the DNA analysis of OXTR methylation and 35 ASD samples were included in the analysis (one sample did not have detectable rates of methylation and therefore was excluded from analysis). One subject withdrew from study 1 because of lack of interest.
Study 2 procedure
We examined the association between OXTR methylation, ASD symptom severity, and rsFC. Thirty-two males with ASD, a subset from study 1 (mean age = 28.97 ± 5.83, 84.37% Caucasian, 12.5% African American, 3.1% unknown), were included in a clinical trial (AOB) that was designed to study the effects of intranasal OXT on brain function during resting-state and social tests. The AOB had a within-subject randomized, placebo-controlled design. Thirty subjects with ASD were included in the fMRI analysis (because of technical problems with one subject data and one dropout due to anxiety related to MRI). For this paper, we only included data related to rsFC 40 min after placebo spray intake.
Comorbidity, adaptive functioning, and medication were assessed during this visit via a self-report demographic questionnaire (open ended). Comorbidity was assessed by a self-report demographic questionnaire and by the Symptom Checklist-90 Revised (SCL-90-R). SCL-90-R is a self-report instrument that evaluates a broad range of psychological problems and symptoms of psychopathology (see Table S2 for more details).
Neurotypical subjects
Seventy-four neurotypical adults (17 women, 57 men; mean age = 28.22 ± 6.96; 54.05% Caucasian, 24.32% African American, 13.51% Asiatic, 8.11% other and unknown) were recruited from Emory University and the general community by posting flyers and by using the advertisement company Teuteberg Incorporated (Wauwatosa, WI, USA). Only sixty-four were part of the final analysis. Ten subjects could not be included due to missing genetic data. Neurotypical subjects participated in the first study only. They provided salivary samples and completed the NEO Personality Inventory-Revised (NEO PI-R) [29], which examines subjects’ Big Five personality traits (extraversion, agreeableness, conscientiousness, neuroticism, and openness). IQ was also collected to match the ASD group in terms of cognitive capacities (see Table S1 for more details). We made sure that both groups have an average of total IQ within the normal range (85–115).
The first study was approved by the Emory Institutional Review Board (IRB#64623). The second study was approved by the Emory IRB (#93455). All subjects provided written informed consent before performing experimental procedures for each of the two studies (see Supplementary material for more details).
Although the AOB project was preregistered in clinical trials.gov (https://clinicaltrials.gov/ct2/show/NCT03033784), the studies in this paper were not preregistered as part of the AOB. They were conducted after submission of an additional grant and after supplemental funding were secured (see “Acknowledgments”). Our specific hypothesis regarding CpG 5,6 was added prior to analysis when the relevant association to social attachment and negative social experiences was demonstrated by our coauthors [27].
Behavioral outcome measures
In order to investigate the association between ASD symptom severity and OXTR methylation, we used SRS-2 as a primary outcome measure. SRS-2 total and it subscale scores (SCI and RRB) were considered in the analysis. We also focused the analysis on two CpG sites of interest: CpG 16 and CpG 5,6. We performed secondary exploratory analysis using the ADOS-2 calibrated severity scores and the scores of ADI-R to better characterize the association between OXTR methylation and ASD symptomatology.
In order to examine the association between OXTR methylation and sociability in neurotypical subjects, we used extraversion dimension from the NEO PI-R as a primary outcome given that it is considered the main dimension of sociability.
OXTR DNA methylation
Saliva samples were collected using Salivette® (SARSTEDT) and kept chilled on ice for up to 2 h before being stored at −80 °C until the day of separation. DNA was extracted from the Salivette® swab using the QIAamp mini kit (Qiagen, Hilden, Germany) and was quantified with PicoGreen® (Quant-iTTM PicoGreen® dsDNA Assay Kit, Thermo Fisher Scientific Inc., Waltham, MA). The average yield from 100 samples was 2.7 μg/swab. One microgram of DNA was treated with sodium bisulfite using the EpiTect Fast Bisulfite Kit (Qiagen, Hilden, Germany). The OXTR gene (chr3: 8792095 to 8811300; GRCh37/hg19 build) was interrogated using EpiTYPER (MassARRAY system; Agena Bioscience, San Diego, CA) according to the manufacturer’s instructions. We aimed to target a specific 406-bp region (chr3: 8810719–8811124) termed MT2 [22] located in a CpG island, as it is the regulatory region related to transcription of the gene. We selected the EpiTYPER platform to cover almost all CpG sites (21/27 CpGs in MT2) (see Supplementary material).
For each participant, the OXTR methylation ratios were retained and used as the criterion variable in subsequent statistical analyses. The reproducibility and sensitivity of the OXTR EpiTYPER assay product were assessed using commercially available standards (EpiTect control DNA, Qiagen) run in triplicate and measured as previously shown [30].
MRI acquisition
MR images were acquired on a 3T Siemens Prisma-Fit scanner with a 64-channel receiver array head + neck coil. BOLD contrast rsfMRI scans were acquired using a conventional EPI sequence with FOV = 192 mm, TR/TE/FA = 3000 ms/25 ms/90°; fifty-eight 2.4-mm-thick oblique slices; 1.5 mm × 1.5 mm in-plane resolution. There were 160 measurements in each rsfMRI scan (see Supplementary material for more details).
Data analysis
The rsfMRI data were analyzed with standard preprocessing pipeline including image distortion correction, slice-time correction, 3D volume registration, and spatial normalization to MNI152 template. The preprocessing pipeline (see Supplementary material for more details) included an added motion artifact reduction step conducted through the independent component analysis (ICA)-AROMA technique [31] (more details in Table S4 and Supplementary material). ICA was performed on temporally concatenated data of the whole subject cohort using the GIFT software [32]. Whole-brain group ICA spatial maps (SMs; with component strength expressed as t-scores), as well as corresponding IC time-courses (TCs) for each subject were obtained through back-projection [32]. The functional classification of the ICs were performed with image based meta-analyses conducted with the publicly available Neurosynth meta-analyses database [33]; utilizing the well-established correspondence between resting-state networks and task-related brain networks [34]. Functional network connectivity (FNC) were assessed between pairs of IC networks that can be unambiguously classified as representing specific brain functions or functional domains, through cross-correlation coefficient (CC) of their respective TCs.
Statistical analysis
In the first study, we performed independent t-tests on all the 21 fragments including 27 CpG sites of the MT2 region of the OXTR gene. We further performed multiple regressions on these CpG sites to control for factors of age and race. We added race as a covariate in our model given the increasing evidence showing race-specific alterations in DNA methylation [35, 36]. Future studies should look into race-specific differences in OXTR DNA methylation. The significance of these tests was controlled for multiple comparisons using Bonferroni corrections. Based on the results, we selected CpG 16 as primary site of interest. We also selected CpG 5,6 as a primary site of interest given its association to social attachment [27]. We used the generalized linear model (GLM) to examine these two CpG sites as predictors (added separately in two GLM models) of clinical severity of ASD symptoms, using the SRS-2 total score as the dependent measure. We added age as a covariate. We included main effects of each of the predictors and interaction between the predictors within the statistical model. We also performed exploratory analysis to examine the relationship between these CpG sites and ADOS-2 and ADI-R using correlation analysis. GLM was also applied for neurotypical controls with extraversion as the dependent measure and CpG 5,6 and CpG 16 as predictors. Age and gender were added as covariates in the GLM.
In the second study, for the brain correlates of the levels of methylation in ASD, we used GLM with CpG 5,6 and CpG 16 as dependent variables and rsFC in brain functional networks, whose dysfunctions are implicated in ASD, as predictors, in addition to age. For rsFC predictors, we included FNC between IC networks, whose SMs spanned areas involved in reward processing (IC4, IC5, IC16, and IC18), as well as between ICs which spanned areas involved in the theory of mind function (IC11 and IC20), from a total of 14 FNC networks (see Table S3). The GLM assesses the relationship between these FNCs (expressed in terms of z-transformed cross-correlations coefficients) and OXTR methylation in these CpG sites of interest. We performed a correlation between SRS-2, ADI-R scores, and the above FNC networks.
Results
OXTR methylation in ASD diagnosis
In the first study, to evaluate the association between ASD diagnosis and OXTR methylation in our sample (n = 99; Table 1), we compared the methylation of each CpG site between subjects with ASD and neurotypical subjects using t-tests (Table 1). We observed that the ASD group had significantly higher methylation at CpG 16 than did the neurotypical group (t(97) = −4.38; α = 0.0024; P < 0.0001; Fig. 1). We also performed multiple regressions to account for potential influence of sex and race on this finding. The analysis showed that ASD diagnosis was significantly associated with CpG 16 methylation (ß = 0.36, t = 3.71, P < 0.001), but age (ß = −1.05, t = −1.07, P = 0.29), and race (ß = −0.05, t = −0.51, P = 0.61) were not (Table 1). We also noticed that methylation at CpG 19.20 and CpG 21 became significantly different after corrections (t = −3.39, P = 0.001; t = −3.383, P = 0.001). Given that the neurotypical group differed from the ASD group in gender distribution, we conducted a secondary analysis with males only and this analysis did not change the results. Methylation at CpG 16 remained significantly higher in ASD compared with the neurotypical group (t = 3.379, P = 0.001).
Focused OXTR methylation analysis and ASD symptom severity
We focused the analysis on two sites of interest: (1) CpG 16 (as part of the intron 1 of MT2 region that was found to be significantly higher in ASD group in this study), (2) CpG 5,6 (part of the exon 1 of MT2 region) that was recently associated with deficits in social attachment [27]. The effect of OXTR methylation on ASD symptoms severity was analyzed with a GLM for both CpG sites (CpG 16 and CpG 5,6). Methylation at CpG 5,6 had a significant effect on social responsiveness total score (SRS-2) after correction for age (Fig. 2a; χ2 = 6.35, P < 0.05). It predicted SRS-2 subscale of SCI scores (χ2 = 7.62; P < 0.05) but did not predict scores of RRB (χ2 = 0.003; P = 0.96) after correcting for age. Methylation at CpG 16 was not found to be associated with the SRS-2 total score (χ2 = 0.86, P = 0.35) nor with it subscales (SCI (χ2 = 0.66, P = 0.417) and RRB (χ2 = 0.9, P = 0.34)).
The X axis represents the predicted estimated value of both predictors in relation to the dependent variable. a Positive correlation between methylation at CpG 5,6 (b.p. −1121, −1119) and total scores of SRS-2 (χ2 = 5.25, P < 0.05). b Negative correlation between methylation at CpG 16 (b.p. −989) and ADI-R total scores (χ2 = 4.92; P < 0.05). c Correlations between methylation at CpG 16 (b.p. −989) and ADI-R subcomponents (χ2 = 4.71; P < 0.05). SI social interaction, LC language and communication, RRB restricted and repetitive behavior. Dashed lines denote the 95% confidence interval.
We performed secondary exploratory analysis to examine the association between OXTR methylation, ADOS-2 and ADI-R scores, measures relevant to ASD diagnosis and developmental aptitudes. Methylation at CpG 5,6 did not correlate with the diagnostic measures (ADOS-2: r = −0.23, P = 0.177; ADI-R: r = 0.172, P = 0.346). Methylation at CpG 16 did not correlate with ADOS-2 scores, but did correlate with ADI-R total scores (r = −0.403, P < 0.05) (Fig. 2b). Methylation at CpG 16 did correlate with ADI-R subscales of social interaction (r = −0.405, P < 0.05) and language and communication (r = −0.35, P < 0.05), but not with the ADI-R subscale of restricted interests and repetitive behaviors (r = −0.036, P = 0.84) (Fig. 2c). Increased methylation at CpG 16 in adulthood seems to be related to less severe developmental and social communicative skills.
Focused OXTR methylation analysis and social personality traits in a neurotypical population
Here, we focused the analysis on the domain of extraversion of the NEO PI-r given that it is considered the domain of sociability. Using GLM for both CpG sites revealed no significant relationship between OXTR methylation and extraversion (CpG 5,6: χ2 = 1.264, P > 0.05; CpG 16: χ2 = 0.389, P > 0.05) after correcting for age, gender, and race. Main effects of each of these covariates as well as interactions between each of them and the CpG site of interest were entered in the model.
rsFC and DNA methylation of OXTR
Levels of methylation at CpG 16 and CpG 5,6 were negatively predicted by FNC between IC11 (which encompasses the theory of mind regions (e.g., STS)) and IC20 (which encompasses PCC) (CpG 16: ß = −0.404, χ2 = 15.03, P < 0.0001; CpG 5,6: ß = −0.11, χ2 = 8.97, P = 0.003), respectively (Fig. 3a).
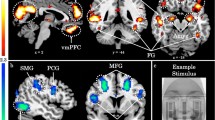
a Negative correlation between FNC between IC1 and IC20 and methylation levels at CpG 16 (b.p. −989) (General linear model, ß = -0.404, χ2 = 15.03, P < 0.0001). b Positive correlation between FNC between IC1 and IC20 and ADI-R total scores (r = 0.57, P < 0.003). The GICA-derived IC strength t-test maps have been thresholded at FDR q < 0.0001. Dashed lines denote the 95% confidence interval.
Also, the FNC between reward areas that includes the IC5 (that includes the ventral striatum) and IC16 (that includes vmPFC) positively predicted degree of methylation at CpG 5,6 (ß = 0.062, χ2 = 7.15, P < 0.009) (Fig. 4a) but did not predict levels of methylation at CpG 16 (ß = 0.052, χ2 = 0.401, P > 0.05).
a Positive correlation between IC5 (that includes ventral striatum) and IC16 (that includes ventromedial prefrontal cortex) and methylation levels at CpG 5,6 (b.p. −1121, −1119) of the oxytocin receptor gene (ß = 0.062, χ2 = 7.15, P < 0.009). b Negative correlation between IC5 (that includes ventral striatum) and IC18 (that includes anterior cingulate cortex and insula) and SRS-2 total scores (r = −0.41, P < 0.05). c Negative correlation between IC5 (that includes ventral striatum) and IC4 (that includes the amygdala) and SRS-2 total scores (r = −0.45, P < 0.05). The GICA-derived IC strength t-test maps have been thresholded at FDR q < 0.0001. Dashed lines denote the 95% confidence interval. vstriatum ventral striatum, vmPFC ventromedial prefrontal cortex, ACC anterior cingulate cortex, Amy amygdala.
Correlation between rsFC and clinical assessments
We observed a negative correlation between SRS-2 total scores and FNC between IC5 (which encompasses the reward areas (e.g., ventral striatum)) and IC18 (which encompasses the anterior cingulate cortex (ACC) and insula) (r = −0.41, P < 0.05) (Fig. 4b) as well as between IC5 (ventral striatum) and IC4 (which encompasses amygdala) (r = −0.45, P < 0.05) (Fig. 4c). SRS-2 total scores did not correlate with FNC between IC5 and IC16 nor between IC11 and IC20 (r = 0.006, P = 0.976; r = 0.005, P = 0.982, respectively).
In addition, we performed secondary analysis to examine the relationship between ADI-R and rsFC between theory of mind areas. We found a positive correlation between rsFC between STS and PCC and ADI-R total scores (r = 0.57, P < 0.003) (Fig. 3b), social interaction scores (r = 0.67, P < 0.0001) and communication scores (r = 0.52, P < 0.006), but not restricted interests and repetitive behaviors (r = −0.2, P = 0.32). Individuals who showed less early deficits in social communication (lower ADI-R) show less rsFC between areas involved in theory of mind. This reduction in rsFC and early social deficits was also found related to an increased methylation at CpG 16 (results described above).
Discussion
We investigated the relationship between OXTR methylation, ASD diagnosis, severity of social symptoms, and the relationship of OXTR methylation with brain network-level functional connectivity at rest in adults with ASD. We first showed that subjects with ASD have higher OXTR methylation at CpG 16 (within the intron 1 of MT2) compared with neurotypical subjects. While ASD did not significantly differ in levels of methylation at CpG 5,6 (within the exon 1 of MT2), compared with neurotypical subjects, methylation levels at this site correlated positively with total scores on the Social Responsiveness Scale, Second edition (SRS-2). It also correlated with subscale of SRS-2 of social communication. This indicates that increased levels of methylation correspond to greater deficits in social responsiveness. We also conducted secondary analysis using ADOS-2 and ADI-R scores and found that while there was no correlation with CpG 5,6, methylation at CpG 16 did correlate with ADI-R scores, and in particular with social interaction and communication scores. This CpG site did not correlate with extraverted personalities in neurotypical subjects. In terms of brain correlates of OXTR methylation and ASD symptom severity, we found that rsFC between regions involved in theory of mind and default mode (STS and PCC) is negatively correlated with both CpG sites of interest. Higher methylation at these sites corresponded to lower connectivity between STS and PCC. Also, a reduced connectivity was correlated with reduced developmental ASD symptoms (as measured by ADI-R). Methylation at CpG 5,6 was further correlated with rsFC between reward areas, which include the ventral striatum and the vmPFC (ACC and OFC). Finally, we found a negative correlation between the SRS-2 and connectivity between the reward processing and empathy networks, particularly between NAcc, ACC/AI, and amygdala. These results suggest that epigenetic variations in the OXT signaling pathway can be associated with ASD symptom severity in the domain of social responsiveness and repetitive behaviors, and atypical functional connectivity between networks involved in social cognition and reward processing in adults with ASD.
OXTR methylation and its relationship to ASD clinical phenotypes and rsFC have not been investigated. However, one early study showed OXTR hypermethylation within the MT2 intron 1 area (around CpG 19,20 at close proximity with our hypermethylated CpG 16 site) in children with ASD, which was associated with decreased OXTR mRNA [21]. In addition, hypermethylation of OXTR was found to be associated with autistic traits in children with OXTR rs53576 G-allele [37]. These studies together with our data presented here, suggest that hypermethylation within intron 1 of MT2 region is associated with ASD diagnosis. More replications in larger sample sizes are needed to confirm this hypermethylation and to examine its specificity to ASD diagnosis.
OXTR hypermethylation at CpG 16 seems to be associated to ASD diagnosis. We found hypermethylation at this site in ASD compared with the neurotypical group. However, within ASD subjects, higher methylation did not uniformly predict increased clinical deficits, but rather was associated with higher verbal functioning and more deficits in other domains. Methylation levels at CpG 16 were associated with ADI-R scores (related to developmental history of symptom severity in social and communication domain), with increased methylation predicting reduced early deficits in social communication. We also found a positive correlation between CpG 16 methylation and total verbal IQ (r = 0.384, P < 0.05), but not with performance IQ (r = 0.14, P = 0.44). One interpretation of this unexpected finding could be that OXTR hypermethylation at CpG 16 is depicting a subtype of ASD (within our sample of high-functioning ASD) that is characterized by less severe verbal and communication deficits, and more selective deficits in theory of mind and self-awareness (as described below). It may also be that this observed hypermethylation of OXTR in subjects with higher functioning is a compensatory mechanism that helps us to alleviate some of the social deficits in ASD. More studies are needed to investigate these speculations and to replicate our findings.
Methylation at CpG 5,6 within the exon 1 of MT2, which was not significantly hypermethylated in ASD, nonetheless predicted social responsiveness in these subjects. This is in line with our rationale that altered social behavior (reduced social reciprocity) may impact methylation levels at the same CpG site (5,6) that was shown to be affected by negative social interactions [27]. This also sheds light on the heterogeneous nature of ASD phenotype and that those who show enhanced social responsiveness, and probably increased reward sensitivity to social interactions, show lower levels of OXTR methylation at CpG 5,6. Its methylation did not correlate with ADOS-2 and ADI-R scores. This could be related to the fact that methylation at this site is more related to social motivation and reward sensitivity to social cues. Future replications with reward outcome measures are needed to better characterize its function. Methylation at CpG 5,6 may be more generally predictive of social experience, and lower sociability in ASD could translate into higher methylation and potentially downregulation of OXTR expression. This could also have an indirect effect on OXT release through dysfunction at the level of autoreceptors. Indeed, we have found previously [38] that adult males with ASD have significantly lower salivary OXT levels compared with neurotypical subjects. We were able to reconfirm this decrease in our present samples of ASD (data not shown here).
We did not find correlations between methylation at CpG 5,6 and extraverted personalities in neurotypical subjects. It is possible that methylation at this site correlates with other traits of personality or other measures of sociability, such as social motivation and reward processing. Future studies should incorporate measures of social reward to better assess the correlates of a neurotypical variability in methylation at CpG 5,6.
Our finding that OXTR hypermethylation is associated with reduced rsFC between areas involved in social cognition (such as STS and PCC) in ASD is in line with several neuroimaging studies in neurotypical subjects. OXTR hypermethylation was associated with enhanced neural activity to negative emotions in frontal areas of infants [39] and with BOLD activity in temporal areas and ACC when attributing intentionality and socioemotional content to geometric shapes [23]. OXTR hypermethylation is also associated with a reduction in functional coupling between social salience and attention networks in response social attention [25]. Also, STS and PCC, which are areas involved in social perception, theory of mind [40], and self-awareness [41], have been already reported to be deficient in ASD, including deficits in intrinsic functional connectivity in the default network [15, 42,43,44,45,46]. It is possible that this hypermethylation at CpG 16 is a biomarker of a particular subtype of ASD that is characterized by less severe verbal communication deficits, but with more pronounced deficits in theory of mind and self-awareness. In our sample, higher CpG 16 methylation was associated with higher verbal IQ and fewer early life deficits (as measured by ADI-R), but with reduced rsFC between theory of mind areas. Other authors have found that individuals with autism show stronger connectivity between PCC and temporal areas compared with neurotypical subjects [47], and that lower severity in verbal and nonverbal communication (measured by the ADI-R) are associated with less connectivity in the default network (including the PCC) [48]. In our sample, those who have less severe communication deficits early in life (but still have ASD) showed more pronounced deficits in self-awareness and theory of mind in adulthood (as depicted by lower functional connectivity). More studies are needed to better understand this finding and to investigate the potential dissociation between self-awareness, theory of mind capacities and verbal communication skills in ASD.
We also found that methylation at CpG 5,6 is positively correlated with rsFC between reward regions, and in particular between NAcc and vmPFC, a known network for pair bond formation [12]. However, this network also includes the connection between OFC and the NAcc core, which is associated with obsessive compulsive disorder symptomatology [49, 50]. Interestingly, here we found that subjects with above average OXTR methylation at CpG 5,6 (higher than 0.7) reported 15 words related to specific interests (digital design, coding, programming, toy trains, game developer, electronics and gadgets, Star Wars, astronomy, and railroad history), while subjects with below average OXTR methylation reported only two specific interests (camera editing and spotting details at high speed). However, the latter reported interests in more general topics that includes socialization, such as art, cooking, sports, reading people’s emotions, literature, and playing with friends. These findings are in line with previous reports of increased subcortical–cortical functional connectivity in ASD [51,52,53] and that striatal-cortical hyperconnectivity is associated with repetitive behaviors [54]. It is possible that a subtype of ASD with higher OXTR methylation within the exon 1 of MT2 region and frontostriatal hyperconnectivity display less social responsiveness, but more specific interests in nonsocial topics (such as trains and computers).
OXTR methylation did not directly correlate with areas involved in empathy (ACC/insula and amygdala). It is possible that this epigenetic modification is a better predictor of sociability per se than brain-based empathic responses. However, the connectivity between NAcc, ACC/insula, and amygdala, areas involved in reward, empathy [4, 55], and salience [56], did predict severity of clinical symptoms in ASD. This is in line with the motivation [57], empathy deficit hypothesis [58], and reward deficit in ASD [19]. We speculate that the coupling between reward and empathy brain network-based deficits could be a predictor of social deficits in ASD.
There are several limitations in our study. We acknowledge that replications with larger samples sizes and with larger diversity (children and adults, verbal and nonverbal capacities, male and female) would be crucial to confirm the findings and conclusions reported here. Also, replications across different psychiatric diagnosis are needed to better assess OXTR methylation function. Also, we were restricted to performing DNA methylation analysis from saliva rather than the brain. However, methylation of DNA extracted from saliva may have similar patterns to methylation observed in brain tissues [59], and peripheral OXTR methylation seems to correlate with brain OXTR methylation after parental neglect [26]. However, whether this is true for the human OXTR gene remains to be determined. Nevertheless, it is possible that peripheral OXTR methylation could be a biomarker relevant to brain physiology and treatment efficacy. Our peripheral marker may relate to brain methylation. However, caution should be taken when using any peripheral marker as a proxy for neuronal methylation. Examination of a publicly available dataset that assesses correlation between saliva, buccal cells, and brain (IMAGE-CpG (https://han-lab.org/methylation/default/imageCpG#)) suggests that only methylation of cg17285225 (CpG3 in our study) in buccal cells was positively correlated with methylation in the brain. However, this was the only CpG that was common between that dataset and the ones included in this study. Finally, individual variation in the proportion of epithelial to buccal cells in saliva can influence the observed methylation values. Thus, we believe that cellular heterogeneity in saliva DNA can be a limitation and that future studies should collect buccal swabs or quantify and adjust for buccal cell proportion in fresh saliva samples to minimize this heterogeneity.
We also acknowledge that the data and samples used in this study were collected as part of a larger preregistered clinical trial (AOB), and that the predictions/hypothesis related to methylation were not a part of the AOB but of a subsequent grant. Future replication studies with larger sample sizes should also include genotype × methylation interaction to better understand the specificity and genetic predispositions of these epigenetic markers. Another limitation of the study is the lack of direct early life measures of social responsiveness that could have better informed developmental trajectory of the clinical symptoms and its relationship to methylation levels. Longitudinal studies would help us to reveal whether some of these epigenetic modifications are acquired through social experiences and whether some of them are innate. We were limited in this study to the personality test in neurotypical subjects and did not have SRS-2 measurement for this group. Future studies should include SRS-2 and other measures of social motivation in neurotypical subjects to assess whether OXTR methylation is a biomarker of sociability. In summary, these findings should be interpreted in the context of the limitations and be viewed as hypothesis generating and preliminary to support further investigation.
In conclusion, using a combined imaging and epigenetic approach, we provide the first evidence that functional epigenetic modification within OXTR predicts symptoms severity in verbally capable adult males with ASD and that these modifications are associated with brain functional connectivity in areas involved in theory of mind, self/social-awareness, and reward. OXTR hypermethylation in the intron 1 area of MT2 was related to a less severe developmental phenotype of social communication and to a hypoconnectivity between areas involved in theory reward and self-awareness. In other terms, this site can be a potential biomarker of adults with ASD who have less severe verbal communication deficits, but who still have an intrinsic theory of mind deficits. OXTR hypermethylation could also be a compensatory mechanism that may help us to alleviate some of the social and communicative deficits in ASD. On the other hand, variability in OXTR methylation within the exon 1 area of MT2, along with hyperconnectivity between frontostriatal networks at rest, might be a biomarker of an ASD subtype characterized by lower social responsiveness and more participation in restricted interests and repetitive behaviors. We also provide compelling evidence that the reward and empathy networks uniquely predict clinical severity of social symptoms in ASD. Our results suggest that neural networks involved in reward processing and social responsiveness in ASD involve the OXT system, which may be impacted by epigenetic processes that could reflect environmental exposures that may be related to ASD symptoms. Given that methylation is reversible, modifying environmental contexts with social stimulations early in life might reverse the pattern of methylation for some patients, potentially leading to improved social functioning. These findings also provide a hypothetical assumption that while the intron 1 area of MT2 is possibly more reflective of ASD diagnosis (with hypermethylation in one of the sites), the exon 1 area of MT2 is a potential biomarker of social motivation and reward processing.
Funding and disclosure
This work was supported by NIH grants P50MH100023 to LJY, ORIP/OD P51OD011132 to YNPRC, and pilot grant from Center for Mind, Brain, and Culture (CMBC), Emory College of Arts and Sciences to EA. The pilot grant money from CMBC was specifically related to the study of OXTR methylation in ASD. EA was supported by NIH grants P50MH100023. Authors do not have competing financial interests in relation to the work described. The project was also supported in part by National Institutes of Health/National Center for Advancing Translational Sciences grant UL1 TR002378 and the Georgia Clinical and Translational Science Alliance (Georgia CTSA).
References
Rilling JK, Young LJ. The biology of mammalian parenting and its effect on offspring social development. Science. 2014;345:771–6.
Johnson ZV, Young LJ. Oxytocin and vasopressin neural networks: Implications for social behavioral diversity and translational neuroscience. Neurosci Biobehav Rev. 2017;76:87–98.
Walum H, Young LJ. The neural mechanisms and circuitry of the pair bond. Nat Rev Neurosci. 2018;19:643–54.
Burkett JP, Andari E, Johnson ZV, Curry DC, de Waal FB, Young LJ. Oxytocin-dependent consolation behavior in rodents. Science. 2016;351:375–8.
Ferguson JN, Aldag JM, Insel TR, Young LJ. Oxytocin in the medial amygdala is essential for social recognition in the mouse. J Neurosci. 2001;21:8278–85.
Jurek B, Neumann ID. The oxytocin receptor: from intracellular signaling to behavior. Physiol Rev. 2018;98:1805–908.
Quintana DS, Rokicki J, van der Meer D, Alnaes D, Kaufmann T, Cordova-Palomera A, et al. Oxytocin pathway gene networks in the human brain. Nat Commun. 2019;10:668.
Young LJ, Barrett CE. Neuroscience. Can oxytocin treat autism? Science. 2015;347:825–6.
Andari E, Hurlemann R, Young LJ. A precision medicine approach to oxytocin trials. Curr Top Behav Neurosci. 2018;35:559–90.
Bethlehem RAI, Lombardo MV, Lai MC, Auyeung B, Crockford SK, Deakin J, et al. Intranasal oxytocin enhances intrinsic corticostriatal functional connectivity in women. Transl Psychiatry. 2017;7:e1099.
Zhao Z, Ma X, Geng Y, Zhao W, Zhou F, Wang J, et al. Oxytocin differentially modulates specific dorsal and ventral striatal functional connections with frontal and cerebellar regions. Neuroimage. 2019;184:781–9.
Amadei EA, Johnson ZV, Jun Kwon Y, Shpiner AC, Saravanan V, Mays WD, et al. Dynamic corticostriatal activity biases social bonding in monogamous female prairie voles. Nature. 2017;546:297–301.
Wigton R, Radua J, Allen P, Averbeck B, Meyer-Lindenberg A, McGuire P, et al. Neurophysiological effects of acute oxytocin administration: systematic review and meta-analysis of placebo-controlled imaging studies. J Psychiatry Neurosci. 2015;40:E1–22.
Di Martino A, Yan CG, Li Q, Denio E, Castellanos FX, Alaerts K, et al. The autism brain imaging data exchange: towards a large-scale evaluation of the intrinsic brain architecture in autism. Mol Psychiatry. 2014;19:659–67.
Alaerts K, Woolley DG, Steyaert J, Di Martino A, Swinnen SP, Wenderoth N. Underconnectivity of the superior temporal sulcus predicts emotion recognition deficits in autism. Soc Cogn Affect Neurosci. 2014;9:1589–600.
Cerliani L, Mennes M, Thomas RM, Di Martino A, Thioux M, Keysers C. Increased functional connectivity between subcortical and cortical resting-state networks in autism spectrum disorder. JAMA Psychiatry. 2015;72:767–77.
Assaf M, Hyatt CJ, Wong CG, Johnson MR, Schultz RT, Hendler T, et al. Mentalizing and motivation neural function during social interactions in autism spectrum disorders. Neuroimage Clin. 2013;3:321–31.
Delmonte S, Balsters JH, McGrath J, Fitzgerald J, Brennan S, Fagan AJ, et al. Social and monetary reward processing in autism spectrum disorders. Mol Autism. 2012;3:7.
Scott-Van Zeeland AA, Dapretto M, Ghahremani DG, Poldrack RA, Bookheimer SY. Reward processing in autism. Autism Res. 2010;3:53–67.
Gouin JP, Zhou QQ, Booij L, Boivin M, Cote SM, Hebert M, et al. Associations among oxytocin receptor gene (OXTR) DNA methylation in adulthood, exposure to early life adversity, and childhood trajectories of anxiousness. Sci Rep. 2017;7:7446.
Gregory SG, Connelly JJ, Towers AJ, Johnson J, Biscocho D, Markunas CA, et al. Genomic and epigenetic evidence for oxytocin receptor deficiency in autism. BMC Med. 2009;7:62.
Kusui C, Kimura T, Ogita K, Nakamura H, Matsumura Y, Koyama M, et al. DNA methylation of the human oxytocin receptor gene promoter regulates tissue-specific gene suppression. Biochem Biophys Res Commun. 2001;289:681–6.
Jack A, Connelly JJ, Morris JP. DNA methylation of the oxytocin receptor gene predicts neural response to ambiguous social stimuli. Front Hum Neurosci. 2012;6:280.
Puglia MH, Lillard TS, Morris JP, Connelly JJ. Epigenetic modification of the oxytocin receptor gene influences the perception of anger and fear in the human brain. Proc Natl Acad Sci USA. 2015;112:3308–13.
Puglia MH, Connelly JJ, Morris JP. Epigenetic regulation of the oxytocin receptor is associated with neural response during selective social attention. Transl Psychiatry. 2018;8:116.
Perkeybile AM, Carter CS, Wroblewski KL, Puglia MH, Kenkel WM, Lillard TS, et al. Early nurture epigenetically tunes the oxytocin receptor. Psychoneuroendocrinology. 2019;99:128–36.
Fujisawa TX, Nishitani S, Takiguchi S, Shimada K, Smith AK, Tomoda A. Oxytocin receptor DNA methylation and alterations of brain volumes in maltreated children. Neuropsychopharmacology. 2019;44:2045–53.
Constantino JN, Gruber CP. The social responsiveness scale-second edition. Los Angeles: Western Psychological Services; 2012.
McCrae RR, Costa PT Jr, Martin TA. The NEO-PI-3: a more readable revised NEO personality inventory. J Pers Assess. 2005;84:261–70.
Kogan SM, Cho J, Beach SRH, Smith AK, Nishitani S. Oxytocin receptor gene methylation and substance use problems among young African American men. Drug Alcohol Depend. 2018;192:309–15.
Pruim RH, Mennes M, van Rooij D, Llera A, Buitelaar JK, Beckmann CF. ICA-AROMA: A robust ICA-based strategy for removing motion artifacts from fMRI data. Neuroimage. 2015;112:267–77.
Allen EA, Erhardt EB, Damaraju E, Gruner W, Segall JM, Silva RF, et al. A baseline for the multivariate comparison of resting-state networks. Front Syst Neurosci. 2011;5:2.
Yarkoni T, Poldrack RA, Nichols TE, Van Essen DC, Wager TD. Large-scale automated synthesis of human functional neuroimaging data. Nat Methods. 2011;8:665–70.
Smith SM, Fox PT, Miller KL, Glahn DC, Fox PM, Mackay CE, et al. Correspondence of the brain's functional architecture during activation and rest. Proc Natl Acad Sci USA. 2009;106:13040–5.
Chitrala KN, Hermandez DG, Nalls MA, Mode NA, Zonderman AB, Ezike N, et al. Race-specific alterations in DNA methylation among middle-aged African Americans and Whites with metabolic syndrome. Epigenetics. 2019;12:1–21.
Chan MA, Ciaccio CE, Gigliotti NM, Rezaiekhaligh M, Siedlik JA, Kennedy K, et al. DNA methylation levels associated with race and childhood asthma severity. J Asthma. 2017;54:825–32.
Rijlaarsdam J, van IMH, Verhulst FC, Jaddoe VW, Felix JF, Tiemeier H, et al. Prenatal stress exposure, oxytocin receptor gene (OXTR) methylation, and child autistic traits: The moderating role of OXTR rs53576 genotype. Autism Res. 2017;10:430–8.
Andari E, Duhamel JR, Zalla T, Herbrecht E, Leboyer M, Sirigu A. Promoting social behavior with oxytocin in high-functioning autism spectrum disorders. Proc Natl Acad Sci USA. 2010;107:4389–94.
Krol KM, Puglia MH, Morris JP, Connelly JJ, Grossmann T. Epigenetic modification of the oxytocin receptor gene is associated with emotion processing in the infant brain. Dev Cogn Neurosci. 2019;37:100648.
Zilbovicius M, Meresse I, Chabane N, Brunelle F, Samson Y, Boddaert N. Autism, the superior temporal sulcus and social perception. Trends Neurosci. 2006;29:359–66.
Buckner RL, Andrews-Hanna JR, Schacter DL. The brain's default network: anatomy, function, and relevance to disease. Ann N Y Acad Sci. 2008;1124:1–38.
Kana RK, Libero LE, Hu CP, Deshpande HD, Colburn JS. Functional brain networks and white matter underlying theory-of-mind in autism. Soc Cogn Affect Neurosci. 2014;9:98–105.
Mueller S, Keeser D, Samson AC, Kirsch V, Blautzik J, Grothe M, et al. Convergent findings of altered functional and structural brain connectivity in individuals with high functioning autism: a multimodal MRI Study. PLoS ONE. 2013;8:e67329.
Cherkassky VL, Kana RK, Keller TA, Just MA. Functional connectivity in a baseline resting-state network in autism. Neuroreport. 2006;17:1687–90.
Monk CS, Peltier SJ, Wiggins JL, Weng SJ, Carrasco M, Risi S, et al. Abnormalities of intrinsic functional connectivity in autism spectrum disorders. Neuroimage. 2009;47:764–72.
Assaf M, Jagannathan K, Calhoun VD, Miller L, Stevens MC, Sahl R, et al. Abnormal functional connectivity of default mode sub-networks in autism spectrum disorder patients. Neuroimage. 2010;53:247–56.
Monk CS, Peltier SJ, Wiggins JL, Weng SJ, Carrasco M, Risi S, Lord C. Abnormalities of intrinsic functional connectivity in autism spectrum disorders. Neuroimage. 2009;47:764–72.
Weng SJ, Wiggins JL, Peltier S, Carrasco M, Risi S, Lord C, Monk CS. Alterations of resting state functional connectivity in the default network in adolescents with autism spectrum disorders. Brain Res. 2010;1313:202.
Dong C, Yang Q, Liang J, Seger CA, Han H, Ning Y, et al. Impairment in the goal-directed corticostriatal learning system as a biomarker for obsessive-compulsive disorder. Psychol Med. 2019;5:1–11.
Harrison BJ, Soriano-Mas C, Pujol J, Ortiz H, Lopez-Sola M, Hernandez-Ribas R, et al. Altered corticostriatal functional connectivity in obsessive-compulsive disorder. Arch Gen Psychiatry. 2009;66:1189–200.
Di Martino A, Kelly C, Grzadzinski R, Zuo XN, Mennes M, Mairena MA, et al. Aberrant striatal functional connectivity in children with autism. Biol Psychiatry. 2011;69:847–56.
Di Martino A, Yan CG, Quigyang Li, Denio E, Castellanos FX, Alaerts K, et al. The autism brain imaging data exchange: towards large-scale evaluation of the intrinsic brain architecture in autism. Mol Psychiatry. 2014;19:659–67.
Picci G, Gotts SJ, Scherf KS. A theoretical rut: revisitng and critically evaluating the generalized under/over-connectivity hypothesis of autism. Dev Sci. 2016;19:524–49.
Abbott AE, Linke AC, Nair A, Jahedi A, Alba LA, Keown CL, et al. Repetitive behaviors in autism are linked to imbalance of corticostriatal connectivity: a functional connectivity MRI study. Soc Cogn Affect Neurosci. 2018;13:32–42.
Singer T. The neuronal basis and ontogeny of empathy and mind reading: review of literature and implications for future research. Neurosci Biobehav Rev. 2006;30:855–63.
Seeley WW, Menon V, Schatzberg AF, Keller J, Glover GH, Kenna H, et al. Dissociable intrinsic connectivity networks for salience processing and executive control. J Neurosci. 2007;27:2349–56.
Dawson G, Webb SJ, McPartland J. Understanding the nature of face processing impairment in autism: insights from behavioral and electrophysiological studies. Dev Neuropsychol. 2005;27:403–24.
Baron-Cohen S, Wheelwright S. The empathy quotient: an investigation of adults with Asperger syndrome or high functioning autism, and normal sex differences. J Autism Dev Disord. 2004;34:163–75.
Smith AK, Kilaru V, Klengel T, Mercer KB, Bradley B, Conneely KN, et al. DNA extracted from saliva for methylation studies of psychiatric traits: evidence tissue specificity and relatedness to brain. Am J Med Genet B Neuropsychiatr Genet. 2015;168B:36–44.
Acknowledgements
We acknowledge the help of Dr Thomas R. Ziegler, Dr Greg S. Martin, Ms Jane Clark, MS, Ms Debora A. Clem and the research nurses of the Georgia CTSA Clinical Research Center at Emory University Hospital. We thank the Center for Systems Imaging (CSI) and Dr John Oshinski for providing us free scans to perform training sessions for all subjects with autism as part of the Autism Oxytocin Brain project. We thank the Center for Mind, Brain, and Culture at Emory College of Arts and Sciences for giving EA a pilot grant to perform epigenetic analysis. We also thank Robert Smith III and Zhou Lei for their generous assistance during MRI scans. We thank Salvation Manadom and Stacy Ladd for their help. We thank Dr Matthew Joel Segall and Lauren Castriota who contributed on the clinical diagnostic assessments for the subjects with Autism. We thank Dr James K. Rilling for his generous support. We thank Dr. James Burkett and Alexander Burkett for their continuous support. We thank the students Morganne Reid and Nichole Andrea Evans for their assistance. We also thank Mrs Toni Thomas for her great help in recruitment efforts at the Emory Autism Center. Special thanks to families and all participants who are the essence of our work.
Author information
Authors and Affiliations
Contributions
EA conceived, designed the work, acquired and analyzed the data, and wrote the paper. SN played a role in the conception of the work, analyzed the data, played a role in interpreting the results, and revised the paper. GK analyzed the data, played a role in interpreting the results, and revised the paper. GC acquired and analyzed data and revised the paper. MM and OO acquired data relevant to this paper, played a role in interpreting the results and revised the paper. AS and JC played a role in the conception of the work, interpretation of the results, and revised the paper. LY revised the paper and played a role in the conception of the work and in the interpretation of the results.
Corresponding author
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Andari, E., Nishitani, S., Kaundinya, G. et al. Epigenetic modification of the oxytocin receptor gene: implications for autism symptom severity and brain functional connectivity. Neuropsychopharmacol. 45, 1150–1158 (2020). https://doi.org/10.1038/s41386-020-0610-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41386-020-0610-6
- Springer Nature Switzerland AG
This article is cited by
-
Intrapartum exposure to synthetic oxytocin, maternal BMI, and neurodevelopmental outcomes in children within the ECHO consortium
Journal of Neurodevelopmental Disorders (2024)
-
Intranasal oxytocin in a genetic animal model of autism
Molecular Psychiatry (2024)
-
Peripheral oxytocin levels are linked to hypothalamic gray matter volume in autistic adults: a cross-sectional secondary data analysis
Scientific Reports (2024)
-
Evaluation of the pooled sample method in Infinium MethylationEPIC BeadChip array by comparison with individual samples
Clinical Epigenetics (2023)
-
A targeted long-read sequencing approach questions the association of OXTR methylation with high-functioning autism
Clinical Epigenetics (2023)