Abstract
Bipolar disorder (BD) is one of the most heritable mental illnesses, but the elucidation of its genetic basis has proven to be a very challenging endeavor. Genome-Wide Association Studies (GWAS) have transformed our understanding of BD, providing the first reproducible evidence of specific genetic markers and a highly polygenic architecture that overlaps with that of schizophrenia, major depression, and other disorders. Individual GWAS markers appear to confer little risk, but common variants together account for about 25% of the heritability of BD. A few higher-risk associations have also been identified, such as a rare copy number variant on chromosome 16p11.2. Large scale next-generation sequencing studies are actively searching for other alleles that confer substantial risk. As our understanding of the genetics of BD improves, there is growing optimism that some clear biological pathways will emerge, providing a basis for future studies aimed at molecular diagnosis and novel therapeutics.
Similar content being viewed by others
Introduction
The genome-wide association studies (GWAS) era has transformed our understanding of bipolar disorder (BD). Ten years ago, BD was considered a distinct, highly heritable disorder for which genes of major effect had eluded detection by linkage studies but were expected to be found eventually. Now, numerous common genetic markers have been found by GWAS, none of which confers major risk for disease, and many of which overlap with markers associated with schizophrenia or major depression. A few higher-risk associations have also been identified, involving rare copy number variants (CNVs) that are usually not inherited. Now, BD can be regarded as a point on a spectrum of risk, ranging from major depression to schizophrenia. Despite this substantial progress, most of the inherited risk for BD remains unexplained, suggesting that there is still much to learn about the genetics of BD. In this review, we will summarize the key developments in BD genetics over the past decade and frame some open questions that will need to be addressed by future studies before we can fully realize the promise of “genomic medicine” in the diagnosis and treatment of BD.
The phenotype
Common
BD is among the most common of major mental illnesses, with prevalence estimates in the range of 1–4% [1]. However, since the diagnosis rests on reports of subjective symptoms that can be subtle, diagnosed cases probably represent the tip of an iceberg of very common disturbances in mood and behavior that blend imperceptibly into the clinical realm. Genetic studies have focused almost entirely on individuals who can be easily diagnosed by interview or are already in treatment, which undoubtedly provides an incomplete picture. Imagine trying to describe the genetics of hypertension by studying only stroke patients.
Varied clinical features
The genetic complexity of BD is belied by its complex and varied clinical presentation [2]. Although the first episode of major depression or mania typically begins between ages 18 and 24 [3], earlier or later onset cases are not rare. Episodes can be frequent or separated by many years, and some patients experience rapid cycling with a period of hours or days [4]. Comorbid anxiety [5, 6] and substance abuse [7, 8] are common, and psychotic features are often a component of mood episodes, particularly manias. Interepisode periods can be completely symptom-free or beset with chronic depressive or manic symptoms. Some people suffer only from manias, although this is uncommon [9]. Mixed states are frequent, as are periods of prolonged, treatment-resistant depression [2]. With such protean manifestations, it seems likely that what we now call BD may ultimately be resolved into dozens of biologically distinguishable disease entities.
Many studies have examined the familiality of clinical features in BD. Age at onset [10], psychotic symptoms [11, 12], frequency of manic and depressive episodes [13], and polarity (mania or depression) at onset [14] are all highly familial, while comorbid anxiety and substance abuse are less so [15]. Below we will address some of the genetic signals that may help explain these patterns.
High risk of suicide
Many studies have pointed to a high risk of suicide in BD [16,17,18,19,20]. On average, about 15% of people diagnosed with BD die of suicide [21], a number that has remained discouragingly stable for decades. Several small studies have reported that suicide may be especially common in some families with BD [18, 22, 23], suggesting specific genetic or shared environmental factors, but these have so far remained elusive.
Cycling as a distinct trait
Signs and symptoms of BD are so wide-ranging that they can be seen, in part, in just about every major psychiatric disorder. This makes for challenging differential diagnosis, one of the reasons that it has proven more difficult to accumulate very large samples of BD than schizophrenia, autism, or major depression. The one very distinctive trait seen in everyone with BD is cycling: episodic elevations and depressions of mood and behavior, separated by periods of relative or complete euthymia [4]. This is such a core feature of BD as currently conceived that we will probably not consider the genetics of BD to be solved until the genetic mechanism of cycling itself has been elucidated.
Response to lithium
Another relatively distinctive clinical feature of some people with BD is the response to lithium. Indeed about one-third of people diagnosed with BD will experience a dramatic improvement in the frequency and severity of mood episodes while receiving lithium, and another third with be at least somewhat improved [24]. Lithium is also the only drug shown to exert a protective effect against suicide in BD [17, 19, 20, 25]. No other major mental illness shows this kind of specific response to lithium, suggesting that genetic risk factors unique to BD are in some way related to the pharmacodynamics of lithium and that biologically meaningful subtypes of BD may be identifiable, at least in part, by response to lithium therapy. A few GWAS of lithium response have been published, but the results so far are divergent [26,27,28,29]. Some recent studies using cellular models lend support to the view that lithium-responsive BD carries a distinct neurobiological signature [30,31,32].
Genetic epidemiology
Before the era of molecular genetics, much of our etiologic understanding of BD rested upon the methods of genetic epidemiology. Family studies demonstrated that BD runs in families, with a 10–15% risk of mood disorder among first-degree relatives of people with BD, but could not distinguish the effects of shared environment from those of shared genes [33]. Twin studies showed that much of the shared familial risk could indeed be explained by shared genes, with heritability estimates on the order of 70–90% [33]. Adoption studies lent further support to a largely genetic etiology, since BD was elevated only in the biological parents of adult adoptees with the illness [33]. Despite the strong and consistent evidence in favor of a genetic etiology; however, segregation analyses could not find a clear, Mendelian pattern of transmission, tending instead to favor more complex models of inheritance [34].
Assortative mating
Assortative mating refers to nonrandom mating among individuals in a population [35]. People with similar phenotypes may be more likely to mate or may selectively avoid potential mates with other phenotypes. A number of studies over the past decades have demonstrated varying degrees of assortative mating in BD, with an increased rate of matings between individuals with BD and those with BD, major depression, alcoholism, or other phenotypes [35,36,37,38,39,40,41,42,43]. Recent, large population-based studies have found similar patterns of assortative mating across psychiatric and other traits, including height [44], activity level [45], emotional intelligence [46], and educational and social status [47].
Such substantial rates of assortative mating are likely to have a major impact on the genetic landscape of BD but are often not considered in studies of the disorder. Theoretically, assortative mating can lead to accumulation of risk alleles in subsequent generations, with consequent increases in rates or severity of illness across generations of a family, a phenomenon known as anticipation [48]. Assortative mating across traits can also induce genetic correlations and comorbidity between the traits in offspring, but these are not likely to persist in the face of random mating by subsequent generations [49]. Assortative mating does not appear to effect heritability estimates by twin studies but may contribute to underestimates of heritability by empirical relationship methods based on SNP arrays [50]. This is because individuals drawn from populations with nonrandom mating will tend to share more risk alleles than would be expected based on their overall genetic relatedness.
Risk loci
Initial searches for risk loci depended on a very limited set of genetic methods, chiefly genetic linkage analysis [14, 51, 52]. However, since linkage methods do not work well in the face of complex patterns of inheritance, linkage studies of BD failed to produce definitive, replicable findings [53]. A similar problem faced linkage studies of most other common, complex traits.
Candidate genes
In an attempt to overcome the limitations of linkage methods, many researchers tried to find genetic markers that were chosen on the basis of their proximity to genes that encoded proteins of known neurobiological importance, such as the serotonin transporter [54]. Unfortunately, this candidate gene strategy was largely unsuccessful. This is because the selection of candidate genes with a high-prior probability of involvement in BD proved to be quite difficult. Most candidate gene studies of BD also suffered from the same biases due to small sample size and undetected genetic mismatch between cases and controls that bedeviled other such studies of a variety of common traits [55]. While meta-analyses do tend to support a small contribution from at least a few well-studied candidates, including the serotonin transporter, SLC6A4 [56,57,58,59], d-amino acid oxidase, DAOA [58, 60,61,62], and brain-derived neurotrophic factor [58, 63,64,65,66,67,68,69,70], the most reliable association evidence has come from GWAS.
GWAS
Genome-wide association studies, wherein large numbers of genetic markers spanning the genome are tested for association with a trait, typically in large, case–control samples, have so far been the most successful strategy for identifying genetic variants associated with BD. Since the first BD GWAS appeared in 2007 [71], almost 20 such studies have been published. Most have focused on typical case definitions of bipolar I disorder [26, 72,73,74,75,76,77,78,79,80,81,82,83], but some have examined clinical subtypes such as schizoaffective disorder [84], bipolar II [85], or BD in the context of personality [86] or other traits. The most recent published GWAS, based on ~50 K cases, detected 30 genome-wide significant loci, of which 20 were newly identified [87].
Genome-wide significant loci reported to date are summarized in Table 1. As with most other common traits, risk loci are numerous, most of the lead SNPs are noncoding, and odds ratios are small (1.1–1.3). Although many of the loci have been implicated by several studies, only a few loci can be resolved to single genes [88, 89] based on current information, so it is still too early to make firm conclusions about specific risk genes underlying most GWAS loci. As functional genomic data accumulates, convergent findings are expected to point toward specific risk genes and pathways.
Convergent data so far highlight at least three genes. ANK3, located on chromosome 10q21.2, was one of the earliest genes to be implicated in BD by GWAS [72, 90,91,92,93]. Significant association has now been found between BD and SNPs near ANK3 by several studies, and several of those SNPs affect expression of ANK3 [90, 91, 94,95,96]. ANK3 encodes ankyrin B, a protein involved in axonal myelination, with expression in multiple tissues, especially brain [97]. Numerous alternative transcripts exist, suggesting a potential role for alternative splicing [98]. A conditional knock-out mouse displays cyclic changes in behavior that resemble BD and respond to treatment with lithium [99]. CACNA1C, located on chromosome 12p13, has also been implicated by genome-wide significant SNP associations in several studies of BD, along with schizophrenia and major depression; some of the associated SNPs are also associated with expression of CACNA1C in multiple tissues, including brain [73, 74, 87, 100,101,102,103]. The gene encodes an L-type voltage-gated ion channel with well-established roles in neuronal development and synaptic signaling. Heterozygous knockdown of the gene in mice alters a variety of behaviors thought to reflect mood, but without a clear syndromic resemblance to BD [102]. TRANK1, which resides on chromosome 3p22, has been implicated by genome-wide significant association with nearby SNPs in studies of BD and schizophrenia [75,76,77, 104, 105]. TRANK1 encodes a large, mostly uncharacterized protein, highly expressed in multiple tissues, especially brain, and may play a role in maintenance of the blood–brain barrier [106]. The expression of TRANK1 is increased by treatment with the mood stabilizer valproic acid, and cells carrying the risk allele show decreased expression of the gene and its protein [104]. Recent transcriptomic studies suggest that DCLK3 may be another gene in the same 3p22 GWAS locus that contributes to risk for both BD and schizophrenia [88, 107].
While each individual GWAS “hit” has only a small effect on risk, polygenic risk scores that combine the additive effects of many risk alleles (often hundreds or thousands) can index substantially more genetic risk by including variants that have so far escaped detection individually at genome-wide significance [108]. Recent studies that use the PRS strategy have shown that common variation accounts for about 25% of the total genetic risk for BD (less of the phenotypic variance), that PRS overlap substantially between BD and schizophrenia, and that PRS derived from large schizophrenia samples are associated with increased rates of psychotic symptoms and decreased response to lithium in BD [101, 105, 109].
Copy number variants (CNVs)
CNVs are stretches of DNA that occur in one (deleted), three (duplicated) or more copies on a chromosome, rather than the typical two copies expected in the diploid human genome. Initially discovered by use of hybridization or SNP array methods that could detect deletions and duplications too small to be found reliably by cytogenetic methods, large (30–1000 kb) CNVs have since been shown to play a major role in neurodevelopmental disorders [110,111,112,113,114,115,116] and some cases of schizophrenia [110, 117,118,119,120,121,122,123].
CNVs seem to play a smaller role in BD [124], but at least two CNVs have been associated with BD in large, case–control samples. The 650 kb duplication on chromosome 16p11.2 was initially described in a de novo study of schizophrenia [125] and was later detected as a de novo event in a proband with early-onset BD [126]. Genome-wide significant evidence of association with BD is based on a large meta-analysis of SNP array data, in which the duplication conferred an OR of 4.37 (95% CI: 2.12–9.00) [127]. This same study also found evidence of association with a deletion on 3q29, but this fell short of genome-wide significance [127]. Both of these CNVs have also been associated with schizophrenia, autism, and intellectual disability [128]. A reciprocal deletion in the 16p11.2 region is associated with autism and ID [129, 130]. One recent study found enrichment of genic CNVs in schizoaffective BD [131]. Taken together, these findings suggest that the genetic overlap between BD and schizophrenia extends beyond common, low-risk alleles to rare alleles of larger effect.
Most published CNV studies to date have relied on technologies that cannot reliably detect CNVs much below ~30 kb. As WGS and other technologies come to the fore, we will doubtless find very large numbers of smaller CNVs in the human genome. Many such smaller CNVs may also be associated with various neurodevelopmental and adult psychiatric disorders and may well be found to play an important role in BD in the future.
Single nucleotide variants (SNVs) and and small insertions/deletions (indels)
Next-generation sequencing (NGS) technology has enabled a search for rare single nucleotide and small insertion/deletion variants that are not represented in SNP arrays [132, 133]. Such studies may uncover alleles conferring greater risk than the common alleles detectable by GWAS, but the lower allele frequencies and large number of potential variants usually demand very large sample sizes, often larger than those needed for GWAS [134].
A few early NGS studies have been published in BD and several others are underway [135,136,137,138]. While the early studies lacked statistical power to demonstrate significant evidence of association after correction for multiple testing, as sample sizes grow significant findings may emerge. Ongoing consortia efforts that aim to achieve larger sample sizes through meta-analysis of multiple independent samples have perhaps the best likelihood of success. Studies that leverage the increased frequencies of otherwise rare alleles sometimes seen in unusual populations [134, 139, 140] may also succeed as sample sizes grow and sequencing technology improves.
Other studies have used NGS to sequence RNA expressed in brain tissue obtained post-mortem from people diagnosed with BD [107, 141, 142]. Such studies can identify diagnosis-associated changes in gene expression, inform efforts to fine-map GWAS loci to individual genes [143], and potentially reveal other transcriptomic events (such as alternative splicing [144]) that mediate risk of inherited genetic variants.
Pathways
One way to deal with the substantial genetic heterogeneity of illnesses like BD is to group implicated genes across studies into pathways or networks of functionally related genes. In this way, increased power to detect association may follow if different alleles in different genes converge at the level of gene sets. Several such pathway studies have been published, with little apparent agreement so far [85, 93, 145,146,147,148,149,150]. The multiplicity of implicated pathways and probably reflects genetic heterogeneity, the relatively small number of robust genetic associations found so far for BD, and the still-challenging problem of assigning common genetic markers found by GWAS to the appropriate gene or genes. Calcium signaling is probably the most supported pathway in BD to date. Calcium signaling has been implicated by animal and ex vivo models of BD [90, 151, 152]. The most compelling genetic evidence for this pathway in BD follows from the known function of the risk gene, CACNA1C [73, 102, 103, 153]. Lithium is also theorized to act by decreasing intracellular calcium signaling [154].
Pathways related to chronobiology and circadian rhythm have long been suspected to play a role in BD. Sleep disturbance is often reported by patients suffering from BD, and changes in sleep schedule (as in transmeridian travel) can provoke episodes in susceptible people [155,156,157]. Genes that influence entrainment of circadian rhythm to the light/dark cycle have been widely studied in BD, with some nominally significant findings [141, 158, 159], but none of these genes have so far been directly implicated by GWAS. Mutations of the CLOCK gene, a canonical gene in the circadian pathway, have been associated with mood disturbance and sleep disorders [160].
Mitochondrial dysfunction, with resulting disturbance in energy metabolism, has also long been theorized to play a role in BD. Patients with some known mitochondrial disorders also show increased rates of mood disturbances consistent with depression or BD [161, 162]. There is also some evidence of mitochondrial dysfunction in induced pluripotent stem cell (iPSC)-derived neurons from BD patients [163]. However, GWAS have failed to detect any significant association between mitochondrial DNA polymorphisms and BD [164].
The pathway analyses of genes implicated in the most recent BD GWAS highlight ion transport, neurotransmitter receptors, insulin secretion, and endocannabinoid signaling, which may provide novel targets for therapeutic development [87].
Genetic architecture
Heritability
Twin studies have consistently demonstrated that most of the individual difference in risk for BD is explained by inherited genetic factors. Studies that compare monozygotic with dizygotic twins have estimated values for narrow-sense heritability of about 70% [165]. Some concern has been raised that the traditional twin design may overestimate heritability under specific circumstances that violate model assumptions [166]. These include assumptions about unbiased ascertainment, equivalence of environments shared by MZ as compared to DZ twins, and potential gene-environment correlations [165]. (Gene–gene and gene–environment interactions, however important they may be in BD, do not contribute to narrow-sense heritability estimates [167]). Recent, population-based studies that do not depend on the same assumptions as twin studies have found very similar heritability estimates [168]. Thus, any overestimation of heritability in the earlier twin studies is likely to be small.
Recent methods allow estimates of heritability based on distant kinds of relatedness that may exist in large, case–control samples [169]. These methods rely on empirical estimates of relatedness derived from sharing of common alleles genotyped by SNP arrays. As has been observed for most common, complex disorders, the SNP-based heritability estimates for BD tend to range from around 25–45% [78, 170]. This “heritability gap” or “missing heritability” is not fully understood, but may reflect imprecision in the method, overestimates of heritability in twin studies (noted above), or a contribution of rare variants not captured on SNP arrays.
Models of etiology and risk
We still lack good models that can bring together genetic and other data heuristically. Four possibilities broadly consistent with the available data come to mind, but others are hard to rule out: (1) Two-hit model. Under this model, we imagine that classes of risk factors interact nonadditively to determine outcome, with combinations accounting for phenotypic distinctions [171]. For example, given two individuals with similar polygenic risk burden, one might develop BD while the other, exposed to a second hit from maternal influenza, develops schizophrenia. (2) Multifactorial threshold model. Under this model, there is a large but finite set of nonspecific genetic and other risk factors, whose total dosage determines specific phenotypes [172]. Thus, BD would occupy some intermediate space, with more risk factors than depression but fewer than schizophrenia. This is a more general version of the two-hit model and fits best when each risk factor has a small, additive effect on outcome. (3) Risk-resilience model. Under this model, genetic differences might confer risk or resilience, with the phenotypic outcome reflecting a delicate balance of harmful and protective factors [173, 174]. Thus, BD might result from genetic risk factors conferring, say, unstable mood, nearly balanced by stable temperament, and advantageous life circumstances. (4) Omnigenic Model. Under this model, almost all genetic differences contribute in some small way to risk (or resilience), while phenotypic outcomes are determined largely by which genes are involved and their relative importance in relevant cells and tissues [175]. Thus, BD might result from genetic risk factors that happen to impact genes that play an important role in cells that underlie neural circuits involved in regulation of mood and behavior.
It has been said that all models are wrong, but some are useful. Each of these models has supporters and critics. The two-hit model resonates with long-held theories of gene × environment interaction, but robust evidence of such interactions has proven elusive [176,177,178,179,180]. The Omnigenic Model has generated much recent debate, since it would seem to imply that larger and larger GWAS cannot alone solve complex traits. In any case, we clearly need more and better ways to incorporate nongenetic risk factors into models of etiology and risk prediction.
Genetic correlations
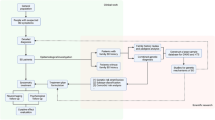
Genetic correlation refers to the degree to which two distinct traits share genetic influences (or more formally, the proportion of additive genetic variance—heritability—that is shared [167]). Traditionally, estimated through laborious twin and family studies, genetic correlation can now be estimated much more easily from overlapping sets of common SNPs genotyped in existing samples [181]. Such studies have so far revealed many expected and some unexpected genetic correlations with BD. In addition to the substantial genetic overlap with schizophrenia that was already apparent early in the GWAS era, significant genetic correlations are observed between bipolar and major depressive disorder [87, 182, 183], attention deficit hyperactivity disorder [184], neuroticism [185], and borderline personality disorder [86]. Small but significant genetic correlations have also emerged between BD and educational attainment [87], creativity [186], and leadership [187]. These findings lend support to the view that BD represents a point on a spectrum of genetic risk, with quantitative rather than categorical genetic differences underlying a range of common disorders of mood, perception, and cognition (Fig. 1).
Genetic and symptomatic relationships between bipolar and some other psychiatric disorders. Shared heritability of bipolar disorder (BD) with schizophrenia (Scz), attention deficit disorder (ADD), and major depressive disorder (MDD). Genetic correlation values were extracted from Ref. [181].
Pharmacogenetics
Pharmacogenetic studies aim to use genetic information to help match patients with the safest, most effective treatments. Several pharmacogenetic studies have been performed in patients with BD, but replicated findings have not yet emerged. This may reflect the fact that many past studies relied on a candidate gene design, while GWAS have not generally been able to achieve sample sizes large enough to detect variants of minor effect. The measurement of treatment response in BD brings additional challenges, since the episodic nature of the illness makes short-term assessments of outcome unreliable.
Some promising findings have nevertheless emerged from recent studies. The largest study to date, by the Consortium on Lithium Genetics, carried out a GWAS of lithium response in over 2000 individuals with BD who were treated with lithium and systematically rated for response. Significant association was detected with a set of genetic variants within a noncoding region on chromosome 21 [27]. Another recent GWAS compared lithium-responsive patients to healthy controls, revealing significant association with a SNP near SESTD1 [188]. The apparent lack of agreement between these two GWAS studies probably reflects limited power to detect small effects. One study in a highly selected set of Taiwanese claimed a locus of major effect [28], but several well-powered studies have failed to replicate this finding [29, 189,190,191]. As sample sizes grow, it seems likely that common loci influencing response to lithium or other drugs will be identified. Larger samples may also enable PRS derived from pharmacogenomic studies to illuminate pathways of drug response or help identify subgroups of patients most likely to respond to a specific treatment regimen.
In contrast to studies of treatment response, those focused on serious adverse events have detected strong and reproducible signals for drugs that are sometimes used in the treatment of BD. Patients exposed to carbamazepine occasionally develop serious adverse cutaneous reactions (ACR), such as Stevens–Johnson Syndrome. Genetic association studies initially carried out in people of Asian ancestry identified an HLA haplotype that conferred substantial risk of ACR after carbamazepine exposure [192]. Subsequent studies have confirmed this association also in patients of European ancestry [193], albeit with a different HLA haplotype. Other studies have identified additional, apparently independent HLA haplotypes that predispose to ACR after exposure to lamotrigine or phenytoin [194]. Based on these findings, HLA testing is advised in all patients being considered for carbamazepine and may also be informative for treatment decisions concerning other anticonvulsants [195].
Genetics of clinical subtypes
It has long been assumed that the clinical diversity of BD reflects, at least in part, differences in underlying risk alleles. Limited statistical power has so far forestalled a complete genetic dissection of the bipolar phenotype, but several studies have found suggestive evidence of genetic differences in bipolar cases with psychosis or catatonic features, and in cases with bipolar II disorder [84, 105, 196, 197]. One large study found a significant positive correlation between genetic risk for schizophrenia and psychotic episodes in patients with BD [84]. This same study detected significant heritability, as estimated from genome-wide SNP data, for psychotic features and suicide attempts in BD.
Ongoing studies aim to go beyond clinical symptoms to define subtypes of disease based on neuroimaging [198,199,200,201], neurocognitive tests [202, 203], and EEG patterns [201, 204, 205], as well as genetic markers. Such studies hold promise for a future nosology of bipolar (and other psychiatric) disorders that better reflects neurobiological disease entities.
Future directions
Cellular phenotyping
The generation of iPSCs from patients allows for in vitro evaluation of cell-autonomous traits that might be associated with clinical diagnosis [206, 207]. Cellular morphology, gene expression, and cellular functions are just some of the phenotypes that can be analyzed using iPSC-based cellular models. More complex models, such as 3D organoids, can explore more macroscopic interactions and might shed light on disorder-specific changes in brain circuitry. So far, only a few published studies have used iPSC derived from patients with BD [104, 151, 163, 208, 209], but several studies are underway. Initial results suggest some differences in neurons derived from patients with BD.
Reverse phenotyping
As we begin to identify genes that have a substantial influence on risk (either collectively, as with PRS, or individually, as with certain CNVs or rare variants), it may be instructive to study individuals who carry substantial risk but do not present in a psychiatric clinic. This approach, dubbed “reverse phenotyping” [210] or “genetics-first” [211, 212] has begun to bear fruit in studies of CNVs and aneuploidies that confer high risk for ASD or schizophrenia [116, 213,214,215]. These kinds of studies are needed for accurate estimates of penetrance [110, 114, 216, 217] and may also reveal an unheralded range of phenotypes related to identified genetic risk factors [218, 219]. Longitudinal studies of genetically high-risk individuals may also shed light on protective or resilience factors and could provide the basis for assessing the impact of primary prevention strategies.
Drug development
The path from the identification of risk alleles to the development of new drugs is complex and beyond the scope of this review. Readers interested in exploring this topic further should consult some recent reviews [220,221,222].
Clinical genetic testing
Genetic testing with utility for the diagnosis of BD or its treatment is not on the horizon right now. Too little of the risk is explained by current polygenic risk scores [170], and known pathogenic CNVs are so far quite rare in BD [124, 127]. However, some models suggest that PRS may ultimately prove useful in psychiatric diagnosis as GWAS samples reach sizes on the order of one million, at least for those individuals with the highest risk allele burdens [223, 224].
Genome-wide approaches help us navigate through the complex genetic landscape in an unbiased manner. However, multiple testing means that GWAS can only detect robust associations in large samples. Increasing the number of samples through involvement of different sample collection sites may improve power but can also introduce substantial genetic heterogeneity. This could be due to the innate genetic variability present across different populations and differences in ascertainment or clinical diagnosis by different research groups. This challenge highlights the need for further global-scale collaborations, standard practices of clinical assessment and phenotype characterization across different groups, and genome-scale modeling that can elucidate the biological impact of the many different risk alleles that are detected in large, population-based studies.
Conclusions
What emerges most clearly from molecular genetic findings over the past decade is a concept of BD that includes several features: (1) BD is a heterogeneous set of illnesses united by the core clinical feature of cyclic elevation in mood and activity, with substantial individual variation in depressive and psychotic symptoms; (2) there is strong sharing of weak, common genetic risk factors with schizophrenia and major depression; (3) high-risk alleles also exist, but they are rare and nonspecific, and there is so far no evidence for monogenic forms of BD.
As a disease entity, BD may resemble stroke or type II diabetes in the sense that several subclinical states create a meta-stable condition that periodically erupts in symptoms. For stroke, we understand that hypertension and cerebrovascular disease create vulnerabilities that may present periodically with paralysis, language, or cognitive deficits. And while there are rare, high-risk alleles that cause stroke, most of the genetic risk resides in large numbers of common alleles that each have a small impact on blood pressure, vascular health, and coagulability [225]. This analogy suggests that we need to identify the fundamental neurobiological processes that are most directly influenced by common risk alleles and we should expect that these processes are underway long before the first manic episode. The analogy further suggests that secondary preventive strategies will need to take aim at these underlying processes, probably beginning at or around the time of the first manic symptoms.
It remains to be seen whether genetic findings to date will continue to coalesce into clear neurobiological pathways. If they do, identification of new drug targets may be possible. The advent of cellular modeling through iPSC technology offers a new platform for screening large numbers of potential new drug treatments, but the success of this approach will depend heavily on the identification of robust cellular phenotypes that reflect at least some of same the genetic risk factors that predispose to bipolar or related disorders. Meanwhile, even if single genes of large effect remain elusive, it seems likely that polygenic approaches incorporating numerous common risk alleles will continue to be useful for research and may ultimately find modest applications in some clinical settings. We have finally made it through the first era of molecular genetics of BD, but the road to new methods of diagnosis and treatment may well remain long and uncertain.
References
Merikangas KR, Jin R, He J-P, Kessler RC, Lee S, Sampson NA, et al. Prevalence and correlates of bipolar spectrum disorder in the world mental health survey initiative. Arch Gen Psychiatry. 2011;68:241–51.
Oxford University Press. Manic-Depressive illness: bipolar disorders and recurrent depression. 2nd ed. Oxford, New York: Oxford University Press; 2007.
McMahon FJ, Stine OC, Chase GA, Meyers DA, Simpson SG, DePaulo JRJ. Influence of clinical subtype, sex, and lineality on age at onset of major affective disorder in a family sample. Am J Psychiatry. 1994;151:210–5.
Perlis RH, Ostacher MJ, Goldberg JF, Miklowitz DJ, Friedman E, Calabrese J, et al. Transition to mania during treatment of bipolar depression. Neuropsychopharmacol. 2010;35:2545–52.
Simon NM, Otto MW, Wisniewski SR, Fossey M, Sagduyu K, Frank E, et al. Anxiety disorder comorbidity in bipolar disorder patients: data from the first 500 participants in the Systematic Treatment Enhancement Program for Bipolar Disorder (STEP-BD). Am J Psychiatry. 2004;161:2222–9.
Deckersbach T, Peters AT, Sylvia L, Urdahl A, Magalhaes PVS, Otto MW, et al. Do comorbid anxiety disorders moderate the effects of psychotherapy for bipolar disorder? Results from STEP-BD. Am J Psychiatry. 2014;171:178–86.
Merikangas KR, Mehta RL, Molnar BE, Walters EE, Swendsen JD, Aguilar-Gaziola S, et al. Comorbidity of substance use disorders with mood and anxiety disorders: results of the International Consortium in Psychiatric Epidemiology. Addict Behav. 1998;23:893–907.
Kessler RC, Crum RM, Warner LA, Nelson CB, Schulenberg J, Anthony JC. Lifetime co-occurrence of DSM-III-R alcohol abuse and dependence with other psychiatric disorders in the National Comorbidity Survey. Arch Gen Psychiatry. 1997;54:313–21.
Reich T, Clayton PJ, Winokur G. Family history studies: V. The genetics of mania. Am J Psychiatry. 1969;125:1358–69.
Strober M. Relevance of early age-of-onset in genetic studies of bipolar affective disorder. J Am Acad Child Adolesc Psychiatry. 1992;31:606–10.
Goes FS, Zandi PP, Miao K, McMahon FJ, Steele J, Willour VL, et al. Mood-incongruent psychotic features in bipolar disorder: familial aggregation and suggestive linkage to 2p11-q14 and 13q21-33. Am J Psychiatry. 2007;164:236–47.
Potash JB, Chiu Y-F, MacKinnon DF, Miller EB, Simpson SG, McMahon FJ, et al. Familial aggregation of psychotic symptoms in a replication set of 69 bipolar disorder pedigrees. Am J Med Genet Part B. 2003;116B:90–7.
Fisfalen ME, Schulze TG, DePaulo JRJ, DeGroot LJ, Badner JA, McMahon FJ. Familial variation in episode frequency in bipolar affective disorder. Am J Psychiatry. 2005;162:1266–72.
Kassem L, Lopez V, Hedeker D, Steele J, Zandi P, McMahon FJ. Familiality of polarity at illness onset in bipolar affective disorder. Am J Psychiatry. 2006;163:1754–9.
Schulze TG, Hedeker D, Zandi P, Rietschel M, McMahon FJ. What is familial about familial bipolar disorder? Resemblance among relatives across a broad spectrum of phenotypic characteristics. Arch Gen Psychiatry. 2006;63:1368–76.
Coryell W, Kriener A, Butcher B, Nurnberger J, McMahon F, Berrettini W, et al. Risk factors for suicide in bipolar I disorder in two prospectively studied cohorts. J Affect Disord. 2016;190:1–5.
Cipriani A, Hawton K, Stockton S, Geddes JR. Lithium in the prevention of suicide in mood disorders: updated systematic review and meta-analysis. BMJ. 2013;346:f3646. https://doi.org/10.1136/bmj.f3646.
Willour VL, Zandi PP, Badner JA, Steele J, Miao K, Lopez V, et al. Attempted suicide in bipolar disorder pedigrees: evidence for linkage to 2p12. Biol Psychiatry. 2007;61:725–7.
Song J, Sjölander A, Joas E, Bergen SE, Runeson B, Larsson H, et al. Suicidal behavior during lithium and valproate treatment: a within-individual 8-year prospective study of 50,000 patients with bipolar disorder. Am J Psychiatry. 2017;174:795–802.
Cipriani A, Pretty H, Hawton K, Geddes JR. Lithium in the prevention of suicidal behavior and all-cause mortality in patients with mood disorders: a systematic review of randomized trials. Am J Psychiatry. 2005;162:1805–19.
Pompili M, Gonda X, Serafini G, Innamorati M, Sher L, Amore M, et al. Epidemiology of suicide in bipolar disorders: a systematic review of the literature. Bipolar Disord. 2013;15:457–90.
Potash JB, Kane HS, Chiu YF, Simpson SG, MacKinnon DF, McInnis MG, et al. Attempted suicide and alcoholism in bipolar disorder: clinical and familial relationships. Am J Psychiatry. 2000;157:2048–50.
Egeland JA, Sussex JN. Suicide and family loading for affective disorders. J Am Med Assoc. 1985;254:915–8.
Grof P, Muller-Oerlinghausen B. A critical appraisal of lithium’s efficacy and effectiveness: the last 60 years. Bipolar Disord. 2009;11(Suppl 2):10–9.
Smith KA, Cipriani A. Lithium and suicide in mood disorders: updated meta-review of the scientific literature. Bipolar Disord. 2017. https://doi.org/10.1111/bdi.12543.
Song J, Bergen SE, Di Florio A, Karlsson R, Charney A, Ruderfer DM, et al. Genome-wide association study identifies SESTD1 as a novel risk gene for lithium-responsive bipolar disorder. Mol Psychiatry. 2017;22:1223. https://doi.org/10.1038/mp.2016.246.
Hou L, Heilbronner U, Degenhardt F, Adli M, Akiyama K, Akula N, et al. Genetic variants associated with response to lithium treatment in bipolar disorder: a genome-wide association study. Lancet. 2016;387:1085–93.
Chen C-H, Lee C-S, Lee M-TM, Ouyang W-C, Chen C-C, Chong M-Y, et al. Variant GADL1 and response to lithium therapy in bipolar I disorder. N. Engl J Med. 2014;370:119–28.
Hou L, Heilbronner U, Rietschel M, Kato T, Kuo P-H, McMahon FJ, et al. Variant GADL1 and response to lithium in bipolar I disorder. N. Engl J Med. 2014;370:1857–9.
Wang JL, Shamah SM, Sun AX, Waldman ID, Haggarty SJ, Perlis RH. Label-free, live optical imaging of reprogrammed bipolar disorder patient-derived cells reveals a functional correlate of lithium responsiveness. Transl Psychiatry. 2014;4:e428. https://doi.org/10.1038/tp.2014.72.
Liang M-H, Wendland JR, Chuang D-M. Lithium inhibits Smad3/4 transactivation via increased CREB activity induced by enhanced PKA and AKT signaling. Mol Cell Neurosci. 2008;37:440–53.
Ferensztajn-Rochowiak E, Tarnowski M, Samochowiec J, Michalak M, Ratajczak MZ, Rybakowski JK. Increased mRNA expression of peripheral glial cell markers in bipolar disorder: the effect of long-term lithium treatment. Eur Neuropsychopharmacol. 2016;26:1516–21.
Smoller JW, Finn CT. Family, twin, and adoption studies of bipolar disorder. Am J Med Genet. 2003;123C:48–58.
Rice J, Cloninger CR, Reich T. Multifactorial inheritance with cultural transmission and assortative mating. I. Description and basic properties of the unitary models. Am J Hum Genet. 1978;30:618–43.
Merikangas KR, Spiker DG. Assortative mating among in-patients with primary affective disorder. Psychol Med. 1982;12:753–64.
Merikangas KR. Assortative mating for psychiatric disorders and psychological traits. Arch Gen Psychiatry. 1982;39:1173–80.
Mathews CA, Reus VI. Assortative mating in the affective disorders: a systematic review and meta-analysis. Compr Psychiatry. 2001;42:257–62.
Baron M, Mendlewicz J, Gruen R, Asnis L, Fieve RR. Assortative mating in affective disorders. J Affect Disord. 1981;3:167–71.
Maes HHM, Neale MC, Kendler KS, Hewitt JK, Silberg JL, Foley DL, et al. Assortative mating for major psychiatric diagnoses in two population-based samples. Psychol Med. 1998;28:1389–401.
Merikangas KR. Divorce and assortative mating among depressed patients. Am J Psychiatry. 1984;141:74–76.
Dunner DL, Fleiss JL, Addonizio G, Fieve RR. Assortative mating in primary affective disorder. Biol Psychiatry. 1976;11:43–51.
Krueger RF, Moffitt TE, Caspi A, Bleske A, Silva PA. Assortative mating for antisocial behavior: developmental and methodological implications. Behav Genet. 1998;28:173–86.
Gershon ES, Dunner DL, Sturt L, Goodwin FK. Assortative mating in the affective disorders. Biol Psychiatry. 1973;1:63–74.
Stulp G, Simons MJP, Grasman S, Pollet TV. Assortative mating for human height: a meta-analysis. Am J Hum Biol Off J Hum Biol Counc. 2017;29:1–10.
Montiglio P-O, Wey TW, Chang AT, Fogarty S, Sih A. Multiple mating reveals complex patterns of assortative mating by personality and body size. J Anim Ecol. 2016;85:125–35.
Smieja M, Stolarski M. Assortative mating for emotional intelligence. Curr Psychol. 2018;37:180–7.
Krzyzanowska M, Mascie-Taylor CGN. Educational and social class assortative mating in fertile British couples. Ann Hum Biol. 2014;41:561–7.
McInnis MG, McMahon FJ, Chase GA, Simpson SG, Ross CA, DePaulo JRJ. Anticipation in bipolar affective disorder. Am J Hum Genet. 1993;53:385–90.
de Jong S, Diniz MJA, Saloma A, Gadelha A, Santoro ML, Ota VK, et al. Applying polygenic risk scoring for psychiatric disorders to a large family with bipolar disorder and major depressive disorder. Commun Biol. 2018;1:163. https://doi.org/10.1038/s42003-018-0155-y.
Peyrot WJ, Robinson MR, Penninx BWJH, Wray NR. Exploring boundaries for the genetic consequences of assortative mating for psychiatric traits. JAMA Psychiatry. 2016;73:1189–95.
Grover D, Verma R, Goes FS, Mahon PLB, Gershon ES, McMahon FJ, et al. Family-based association of YWHAH in psychotic bipolar disorder. Am J Med Genet Part B. 2009;150B:977–83.
McInnis MG, Breschel TS, Margolis RL, Chellis J, MacKinnon DF, McMahon FJ, et al. Family-based association analysis of the hSKCa3 potassium channel gene in bipolar disorder. Mol Psychiatry. 1999;4:217–9.
Prathikanti S, McMahon FJ. Genome scans for susceptibility genes in bipolar affective disorder. Ann Med. 2001;33:257–62.
Judy JT, Seifuddin F, Mahon PB, Huo Y, Goes FS, Jancic D, et al. Association study of serotonin pathway genes in attempted suicide. Am J Med Genet Part B. 2012;159B:112–9.
Risch N, Merikangas K. The future of genetic studies of complex human diseases. Science. 1996;273:1516–7.
Kraft JB, Peters EJ, Slager SL, Jenkins GD, Reinalda MS, McGrath PJ, et al. Analysis of association between the serotonin transporter and antidepressant response in a large clinical sample. Biol Psychiatry. 2007;61:734–42.
Allen NC, Bagade S, McQueen MB, Ioannidis JPA, Kavvoura FK, Khoury MJ, et al. Systematic meta-analyses and field synopsis of genetic association studies in schizophrenia: the SzGene database. Nat Genet. 2008;40:827–34.
Gatt JM, Burton KLO, Williams LM, Schofield PR. Specific and common genes implicated across major mental disorders: a review of meta-analysis studies. J Psychiatr Res. 2015;60:1–13.
Hu X-Z, Rush AJ, Charney D, Wilson AF, Sorant AJM, Papanicolaou GJ, et al. Association between a functional serotonin transporter promoter polymorphism and citalopram treatment in adult outpatients with major depression. Arch Gen Psychiatry. 2007;64:783–92.
Schulze TG, Ohlraun S, Czerski PM, Schumacher J, Kassem L, Deschner M, et al. Genotype-phenotype studies in bipolar disorder showing association between the DAOA/G30 locus and persecutory delusions: a first step toward a molecular genetic classification of psychiatric phenotypes. Am J Psychiatry. 2005;162:2101–8.
Maheshwari M, Shi J, Badner JA, Skol A, Willour VL, Muzny DM, et al. Common and rare variants of DAOA in bipolar disorder. Am J Med Genet Part B. 2009;150B:960–6.
Detera-Wadleigh SD, Liu C, Maheshwari M, Cardona I, Corona W, Akula N, et al. Sequence variation in DOCK9 and heterogeneity in bipolar disorder. Psychiatr Genet. 2007;17:274–86.
Dreimuller N, Schlicht KF, Wagner S, Peetz D, Borysenko L, Hiemke C, et al. Early reactions of brain-derived neurotrophic factor in plasma (pBDNF) and outcome to acute antidepressant treatment in patients with Major Depression. Neuropharmacology. 2012;62:264–9.
Laje G, Perlis RH, Rush AJ, McMahon FJ. Pharmacogenetics studies in STAR*D: strengths, limitations, and results. Psychiatr Serv Wash DC. 2009;60:1446–57.
Liu L, Foroud T, Xuei X, Berrettini W, Byerley W, Coryell W, et al. Evidence of association between brain-derived neurotrophic factor gene and bipolar disorder. Psychiatr Genet. 2008;18:267–74.
Boulle F, Van den Hove DLA, Jakob SB, Rutten BP, Hamon M, Van Os J, et al. Epigenetic regulation of the BDNF gene: implications for psychiatric disorders. Mol Psychiatry. 2012;17:584–96.
Domschke K, Lawford B, Laje G, Berger K, Young R, Morris P, et al. Brain-derived neurotrophic factor (BDNF) gene: no major impact on antidepressant treatment response. Int J Neuropsychopharmacol. 2010;13:93–101.
Gao Y, Galante M, El-Mallakh J, Nurnberger JIJ, Delamere NA, Lei Z, et al. BDNF expression in lymphoblastoid cell lines carrying BDNF SNPs associated with bipolar disorder. Psychiatr Genet. 2012;22:253–5.
Duncan LE, Hutchison KE, Carey G, Craighead WE. Variation in brain-derived neurotrophic factor (BDNF) gene is associated with symptoms of depression. J Affect Disord. 2009;115:215–9.
Lopez JP, Mamdani F, Labonte B, Beaulieu MM, Yang JP, Berlim MT, et al. Epigenetic regulation of BDNF expression according to antidepressant response. Mol Psychiatry. 2013;18:398–9.
Burton PR, Clayton DG, Cardon LR, Craddock N, Deloukas P, Duncanson A, et al. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature. 2007;447:661–78.
Smith EN, Bloss CS, Badner JA, Barrett T, Belmonte PL, Berrettini W, et al. Genome-wide association study of bipolar disorder in European American and African American individuals. Mol Psychiatry. 2009;14:755–63.
Ferreira MAR, O’Donovan MC, Meng YA, Jones IR, Ruderfer DM, Jones L, et al. Collaborative genome-wide association analysis supports a role for ANK3 and CACNA1C in bipolar disorder. Nat Genet. 2008;40:1056–8.
Group PGCBDW. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat Genet. 2011;43:977–83.
Ikeda M, Takahashi A, Kamatani Y, Okahisa Y, Kunugi H, Mori N, et al. A genome-wide association study identifies two novel susceptibility loci and trans population polygenicity associated with bipolar disorder. Mol Psychiatry. 2018;23:639–47.
Chen DT, Jiang X, Akula N, Shugart YY, Wendland JR, Steele CJM, et al. Genome-wide association study meta-analysis of European and Asian-ancestry samples identifies three novel loci associated with bipolar disorder. Mol Psychiatry. 2013;18:195–205.
Muhleisen TW, Leber M, Schulze TG, Strohmaier J, Degenhardt F, Treutlein J, et al. Genome-wide association study reveals two new risk loci for bipolar disorder. Nat Commun. 2014;5:3339. https://doi.org/10.1038/ncomms4339.
Hou L, Bergen SE, Akula N, Song J, Hultman CM, Landén M, et al. Genome-wide association study of 40,000 individuals identifies two novel loci associated with bipolar disorder. Hum Mol Genet. 2016. https://doi.org/10.1093/hmg/ddw181.
Baum AE, Akula N, Cabanero M, Cardona I, Corona W, Klemens B, et al. A genome-wide association study implicates diacylglycerol kinase eta (DGKH) and several other genes in the etiology of bipolar disorder. Mol Psychiatry. 2008;13:197–207.
Xu W, Cohen-Woods S, Chen Q, Noor A, Knight J, Hosang G, et al. Genome-wide association study of bipolar disorder in Canadian and UK populations corroborates disease loci including SYNE1 and CSMD1. BMC Med Genet. 2014;15:2. https://doi.org/10.1186/1471-2350-15-2.
Smith EN, Koller DL, Panganiban C, Szelinger S, Zhang P, Badner JA, et al. Genome-wide association of bipolar disorder suggests an enrichment of replicable associations in regions near genes. PLoS Genet. 2011;7:e1002134. https://doi.org/10.1371/journal.pgen.1002134.
Bergen SE, O’dushlaine CT, Ripke S, Lee PH, Ruderfer DM, Akterin S, et al. Genome-wide association study in a Swedish population yields support for greater CNV and MHC involvement in schizophrenia compared with bipolar disorder. Mol Psychiatry. 2012;17:880–6.
Kuo PH, Chuang LC, Liu JR, Liu CM, Huang MC, Lin SK, et al. Identification of novel loci for bipolar I disorder in a multi-stage genome-wide association study. Prog Neuropsychopharmacol Biol Psychiatry. 2014;51:58–64.
Ruderfer DM, Ripke S, McQuillin A, Boocock J, Stahl EA, Pavlides JMW, et al. Genomic dissection of bipolar disorder and schizophrenia, including 28 subphenotypes. Cell. 2018;173:1705–15.e16.
Kao C-F, Chen H-W, Chen H-C, Yang J-H, Huang M-C, Chiu Y-H, et al. Identification of susceptible loci and enriched pathways for bipolar ii disorder using genome-wide association studies. Int J Neuropsychopharmacol. 2016. https://doi.org/10.1093/ijnp/pyw064.
Witt SH, Streit F, Jungkunz M, Frank J, Awasthi S, Reinbold CS, et al. Genome-wide association study of borderline personality disorder reveals genetic overlap with bipolar disorder, major depression and schizophrenia. Transl Psychiatry. 2017;7:e1155. https://doi.org/10.1038/tp.2017.115.
Stahl EA, Breen G, Forstner AJ, McQuillin A, Ripke S, Trubetskoy V, et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat Genet. 2019;51:793–803.
Huckins L, Dobbyn A, McFadden W, Wang W, Ruderfer D, Hoffman G, et al. Transcriptomic imputation of bipolar disorder and bipolar subtypes reveals 29 novel associated genes. BioRxiv. 2017:222786. https://doi.org/10.1101/222786.
Akula N, Marenco S, Johnson K, Feng N, Cross J, England B, et al. Deep transcriptome sequencing of subgenual anterior cingulate cortex reveals disorder-specific expression changes in major psychiatric disorders. BioRxiv. 2019:598649. https://doi.org/10.1101/598649.
Hayashi A, Le Gal K, Södersten K, Vizlin-Hodzic D, Ågren H, Funa K. Calcium-dependent intracellular signal pathways in primary cultured adipocytes and ANK3 gene variation in patients with bipolar disorder and healthy controls. Mol Psychiatry. 2015;20:931–40.
Rueckert EH, Barker D, Ruderfer D, Bergen SE, O’Dushlaine C, Luce CJ, et al. Cis-acting regulation of brain-specific ANK3 gene expression by a genetic variant associated with bipolar disorder. Mol Psychiatry. 2013;18:922–9.
Belmonte Mahon P, Pirooznia M, Goes FS, Seifuddin F, Steele J, Lee PH, et al. Genome-wide association analysis of age at onset and psychotic symptoms in bipolar disorder. Am J Med Genet Part B Neuropsychiatr Genet. 2011;156B:370–8.
Durak O, de Anda FC, Singh KK, Leussis MP, Petryshen TL, Sklar P, et al. Ankyrin-G regulates neurogenesis and Wnt signaling by altering the subcellular localization of beta-catenin. Mol Psychiatry. 2014. https://doi.org/10.1038/mp.2014.42.
Lippard ETC, Jensen KP, Wang F, Johnston JAY, Spencer L, Pittman B, et al. Effect of ANK3 variation on gray and white matter in bipolar disorder. Mol Psychiatry. 2017;22:1345–51.
Schulze TG, Detera-Wadleigh SD, Akula N, Gupta A, Kassem L, Steele J, et al. Two variants in Ankyrin 3 (ANK3) are independent genetic risk factors for bipolar disorder. Mol Psychiatry. 2009;14:487–91.
Lim CH, Zain SM, Reynolds GP, Zain MA, Roffeei SN, Zainal NZ, et al. Genetic association of LMAN2L gene in schizophrenia and bipolar disorder and its interaction with ANK3 gene polymorphism. Prog Neuropsychopharmacol Biol Psychiatry. 2014;54:157–62.
Hannon E, Lunnon K, Schalkwyk L, Mill J. Interindividual methylomic variation across blood, cortex, and cerebellum: implications for epigenetic studies of neurological and neuropsychiatric phenotypes. Epigenetics. 2015;10:1024–32.
Yamankurt G, Wu HC, McCarthy M, Cunha SR. Exon organization and novel alternative splicing of Ank3 in mouse heart. PLoS ONE 2015;10:e012817. https://doi.org/10.1371/journal.pone.0128177.
Zhu S, Cordner ZA, Xiong J, Chiu C-T, Artola A, Zuo Y, et al. Genetic disruption of ankyrin-G in adult mouse forebrain causes cortical synapse alteration and behavior reminiscent of bipolar disorder. Proc Natl Acad Sci USA. 2017;114:10479–84.
Moskvina V, Craddock N, Holmans P, Nikolov I, Pahwa JS, Green E, et al. Gene-wide analyses of genome-wide association data sets: evidence for multiple common risk alleles for schizophrenia and bipolar disorder and for overlap in genetic risk. Mol Psychiatry. 2009;14:252–60.
Cross Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet. 2013;381:1371–9.
Dao DT, Mahon PB, Cai X, Kovacsics CE, Blackwell RA, Arad M, et al. Mood disorder susceptibility gene CACNA1C modifies mood-related behaviors in mice and interacts with sex to influence behavior in mice and diagnosis in humans. Biol Psychiatry. 2010;68:801–10.
Gershon ES, Grennan K, Busnello J, Badner JA, Ovsiew F, Memon S, et al. A rare mutation of CACNA1C in a patient with bipolar disorder, and decreased gene expression associated with a bipolar-associated common SNP of CACNA1C in brain. Mol Psychiatry. 2014;19:890–4.
Jiang X, Detera-Wadleigh SD, Akula N, Mallon BS, Hou L, Xiao T, et al. Sodium valproate rescues expression of TRANK1 in iPSC-derived neural cells that carry a genetic variant associated with serious mental illness. Mol Psychiatry. 2019;24:613–24.
Ruderfer DM, Fanous AH, Ripke S, McQuillin A, Amdur RL, Consortium SWG of PG. et al. Polygenic dissection of diagnosis and clinical dimensions of bipolar disorder and schizophrenia. Mol Psychiatry. 2014;19:1017–24.
Schiavone S, Mhillaj E, Neri M, Morgese MG, Tucci P, Bove M, et al. Early loss of blood-brain barrier integrity precedes NOX2 elevation in the prefrontal cortex of an animal model of psychosis. Mol Neurobiol. 2017;54:2031–44.
Gandal MJ, Zhang P, Hadjimichael E, Walker RL, Chen C, Liu S, et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science. 2018;362:eaat8127. https://doi.org/10.1126/science.aat8127.
Purcell SM, Wray NR, Stone JL, Visscher PM, O’Donovan MC, Sullivan PF, et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature. 2009;460:748–52.
Schulze TG, Akula N, Breuer R, Steele J, Nalls MA, Singleton AB, et al. Molecular genetic overlap in bipolar disorder, schizophrenia, and major depressive disorder. World J Biol Psychiatry. 2014;15:200–8.
Kirov G, Rees E, Walters JT, Escott-Price V, Georgieva L, Richards AL, et al. The penetrance of copy number variations for schizophrenia and developmental delay. Biol Psychiatry. 2014;75:378–85.
Leppa VM, Kravitz SN, Martin CL, Andrieux J, Le Caignec C, Martin-Coignard D, et al. Rare inherited and de novo cnvs reveal complex contributions to asd risk in multiplex families. Am J Hum Genet. 2016;99:540–54.
Sanders SJ, Ercan-Sencicek AG, Hus V, Luo R, Murtha MT, Moreno-De-Luca D, et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron. 2011;70:863–85.
Williams NM, Zaharieva I, Martin A, Langley K, Mantripragada K, Fossdal R, et al. Rare chromosomal deletions and duplications in attention-deficit hyperactivity disorder: a genome-wide analysis. Lancet. 2010;376:1401–8.
Olsen L, Sparsø T, Weinsheimer SM, Dos Santos MBQ, Mazin W, Rosengren A, et al. Prevalence of rearrangements in the 22q11.2 region and population-based risk of neuropsychiatric and developmental disorders in a Danish population: a case-cohort study. Lancet Psychiatry. 2018;5:573–80.
Gilissen C, Hehir-Kwa JY, Thung DT, van de Vorst M, van Bon BW, Willemsen MH, et al. Genome sequencing identifies major causes of severe intellectual disability. Nature. 2014;511:344–7.
Pinto D, Pagnamenta AT, Klei L, Anney R, Merico D, Regan R, et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature. 2010;466:368–72.
Rippey C, Walsh T, Gulsuner S, Brodsky M, Nord AS, Gasperini M, et al. Formation of chimeric genes by copy-number variation as a mutational mechanism in schizophrenia. Am J Hum Genet. 2013;93:697–710.
Szatkiewicz JP, O’Dushlaine C, Chen G, Chambert K, Moran JL, Neale BM, et al. Copy number variation in schizophrenia in Sweden. Mol Psychiatry. 2014;19:762–73.
Yuan J, Hu J, Li Z, Zhang F, Zhou D, Jin C. A replication study of schizophrenia-related rare copy number variations in a Han Southern Chinese population. Hereditas. 2017;154:2. https://doi.org/10.1186/s41065-016-0025-x.
Bassett AS, Marshall CR, Lionel AC, Chow EW, Scherer SW. Copy number variations and risk for schizophrenia in 22q11.2 deletion syndrome. Hum Mol Genet. 2008;17:4045–53.
Gulsuner S, McClellan JM. Copy number variation in schizophrenia. Neuropsychopharmacol. 2015;40:252–4.
Ingason A, Rujescu D, Cichon S, Sigurdsson E, Sigmundsson T, Pietilainen OPH, et al. Copy number variations of chromosome 16p13.1 region associated with schizophrenia. Mol Psychiatry. 2011;16:17–25.
Ahn K, An SS, Shugart YY, Rapoport JL. Common polygenic variation and risk for childhood-onset schizophrenia. Mol Psychiatry. 2016;21:94–6.
Grozeva D, Kirov G, Ivanov D, Jones IR, Jones L, Green EK, et al. Rare copy number variants: a point of rarity in genetic risk for bipolar disorder and schizophrenia. Arch Gen Psychiatry. 2010;67:318–27.
McCarthy SE, Makarov V, Kirov G, Addington AM, McClellan J, Yoon S, et al. Microduplications of 16p11.2 are associated with schizophrenia. Nat Genet. 2009;41:1223–7.
Malhotra D, McCarthy S, Michaelson JJ, Vacic V, Burdick KE, Yoon S, et al. High frequencies of de novo CNVs in bipolar disorder and schizophrenia. Neuron. 2011;72:951–63.
Green EK, Rees E, Walters JTR, Smith KG, Forty L, Grozeva D, et al. Copy number variation in bipolar disorder. Mol Psychiatry. 2015;21:2189–93.
Kirov G. CNVs in neuropsychiatric disorders. Hum Mol Genet. 2015;24:R45–9.
Zufferey F, Sherr EH, Beckmann ND, Hanson E, Maillard AM, Hippolyte L, et al. A 600 kb deletion syndrome at 16p11.2 leads to energy imbalance and neuropsychiatric disorders. J Med Genet. 2012;49:660–8.
Guha S, Rees E, Darvasi A, Ivanov D, Ikeda M, Bergen SE, et al. Implication of a rare deletion at distal 16p11.2 in schizophrenia. JAMA Psychiatry. 2013;70:253–60.
Charney AW, Stahl EA, Green EK, Chen C-Y, Moran JL, Chambert K, et al. Contribution of rare copy number variants to bipolar disorder risk is limited to schizoaffective cases. Biol Psychiatry. 2019;86:110–9.
Gudbjartsson DF, Helgason H, Gudjonsson SA, Zink F, Oddson A, Gylfason A, et al. Large-scale whole-genome sequencing of the Icelandic population. Nat Genet. 2015. https://doi.org/10.1038/ng.3247.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164. https://doi.org/10.1093/nar/gkq603.
Zuk O, Schaffner SF, Samocha K, Do R, Hechter E, Kathiresan S, et al. Searching for missing heritability: designing rare variant association studies. Proc Natl Acad Sci USA. 2014;111:E455–64.
Ament SA, Szelinger S, Glusman G, Ashworth J, Hou L, Akula N, et al. Rare variants in neuronal excitability genes influence risk for bipolar disorder. Proc Natl Acad Sci USA. 2015;112:3576–81.
Goes FS, Pirooznia M, Parla JS, Kramer M, Ghiban E, Mavruk S, et al. Exome sequencing of familial bipolar disorder. JAMA Psychiatry 2016;73:590–7.
Kataoka M, Matoba N, Sawada T, Kazuno A-A, Ishiwata M, Fujii K, et al. Exome sequencing for bipolar disorder points to roles of de novo loss-of-function and protein-altering mutations. Mol Psychiatry. 2016;21:885–93.
Georgi B, Craig D, Kember RL, Liu W, Lindquist I, Nasser S, et al. Genomic view of bipolar disorder revealed by whole genome sequencing in a genetic isolate. PLoS Genet. 2014;10:e1004229. https://doi.org/10.1371/journal.pgen.1004229.
Hou L, Faraci G, Chen DT, Kassem L, Schulze TG, Shugart YY, et al. Amish revisited: next-generation sequencing studies of psychiatric disorders among the Plain people. Trends Genet. 2013;29:412–8.
Hou L, Kember RL, Roach JC, O’Connell JR, Craig DW, Bucan M, et al. A population-specific reference panel empowers genetic studies of Anabaptist populations. Sci Rep. 2017;7:6079. https://doi.org/10.1038/s41598-017-05445-3.
Akula N, Barb J, Jiang X, Wendland JR, Choi KH, Sen SK, et al. RNA-sequencing of the brain transcriptome implicates dysregulation of neuroplasticity, circadian rhythms and GTPase binding in bipolar disorder. Mol Psychiatry. 2014;19:1179–85.
Pacifico R, Davis RL. Transcriptome sequencing implicates dorsal striatum-specific gene network, immune response and energy metabolism pathways in bipolar disorder. Mol Psychiatry. 2017;22:441–9.
Akula N, Wendland JR, Choi KH, McMahon FJ. An integrative genomic study implicates the postsynaptic density in the pathogenesis of bipolar disorder. Neuropsychopharmacology. 2016;41:886–95.
Li YI, Geijn B, van de, Raj A, Knowles DA, Petti AA, Golan D, et al. RNA splicing is a primary link between genetic variation and disease. Science. 2016;352:600–4.
Pandey A, Davis NA, White BC, Pajewski NM, Savitz J, Drevets WC, et al. Epistasis network centrality analysis yields pathway replication across two GWAS cohorts for bipolar disorder. Transl Psychiatry. 2012;2:e154. https://doi.org/10.1038/tp.2012.80.
Chang S, Wang J, Zhang K, Wang J. Pathway-based analysis for genome-wide association study data of bipolar disorder provides new insights for genetic study. Protein Cell. 2015;6:912–5.
Zandi PP, Belmonte PL, Willour VL, Goes FS, Badner JA, Simpson SG, et al. Association study of Wnt signaling pathway genes in bipolar disorder. Arch Gen Psychiatry. 2008;65:785–93.
Berridge MJ. Calcium signaling and psychiatric disease: bipolar disorder and schizophrenia. Cell Tissue Res. 2014;357:477–92.
Nurnberger JIJ, Koller DL, Jung J, Edenberg HJ, Foroud T, Guella I, et al. Identification of pathways for bipolar disorder: a meta-analysis. JAMA Psychiatry. 2014;71:657–64.
Patel SD, Le-Niculescu H, Koller DL, Green SD, Lahiri DK, McMahon FJ, et al. Coming to grips with complex disorders: genetic risk prediction in bipolar disorder using panels of genes identified through convergent functional genomics. Am J Med Genet Part B. 2010;153B:850–77.
Chen HM, DeLong CJ, Bame M, Rajapakse I, Herron TJ, McInnis MG, et al. Transcripts involved in calcium signaling and telencephalic neuronal fate are altered in induced pluripotent stem cells from bipolar disorder patients. Transl Psychiatry. 2014;4:e375. https://doi.org/10.1038/tp.2014.12.
de Groot MWGDM, Dingemans MML, Rus KH, de Groot A, RHS Westerink. Characterization of calcium responses and electrical activity in differentiating mouse neural progenitor cells in vitro. Toxicol Sci J Soc Toxicol. 2014;137:428–35.
Yoshimizu T, Pan JQ, Mungenast AE, Madison JM, Su S, Ketterman J, et al. Functional implications of a psychiatric risk variant within CACNA1C in induced human neurons. Mol Psychiatry. 2014;20:162–9.
Schlecker C, Boehmerle W, Jeromin A, DeGray B, Varshney A, Sharma Y, et al. Neuronal calcium sensor-1 enhancement of InsP3 receptor activity is inhibited by therapeutic levels of lithium. J Clin Investig. 2006;116:1668–74.
Krane-Gartiser K, Steinan MK, Langsrud K, Vestvik V, Sand T, Fasmer OB, et al. Mood and motor activity in euthymic bipolar disorder with sleep disturbance. J Affect Disord. 2016;202:23–31.
Ng TH, Chung K-F, Ho FY-Y, Yeung W-F, Yung K-P, Lam T-H. Sleep–wake disturbance in interepisode bipolar disorder and high-risk individuals: a systematic review and meta-analysis. Sleep Med Rev. 2015;20:46–58.
Pagani L, Clair PAS, Teshiba TM, Fears SC, Araya C, Araya X, et al. Genetic contributions to circadian activity rhythm and sleep pattern phenotypes in pedigrees segregating for severe bipolar disorder. Proc Natl Acad Sci USA. 2016;113:E754–61.
Castro J, Zanini M, Gonçalves B, da SB, Coelho FMS, Bressan R, et al. Circadian rest–activity rhythm in individuals at risk for psychosis and bipolar disorder. Schizophr Res. 2015;168:50–5.
Geoffroy PA, Etain B, Lajnef M, Zerdazi E-H, Brichant‐Petitjean C, Heilbronner U, et al. Circadian genes and lithium response in bipolar disorders: associations with PPARGC1A (PGC‐1α) and RORA. Genes Brain Behav. 2016;15:660–8.
Shi J, Wittke-Thompson JK, Badner JA, Hattori E, Potash JB, Willour VL, et al. Clock genes may influence bipolar disorder susceptibility and dysfunctional circadian rhythm. Am J Med Genet Part B Neuropsychiatr Genet. 2008;147B:1047–55.
Mancuso M, Orsucci D, Ienco EC, Pini E, Choub A, Siciliano G. Psychiatric involvement in adult patients with mitochondrial disease. Neurol Sci. 2013;34:71–4.
Kasahara T, Ishiwata M, Kakiuchi C, Fuke S, Iwata N, Ozaki N, et al. Enrichment of deleterious variants of mitochondrial DNA polymerase gene (POLG1) in bipolar disorder. Psychiatry Clin Neurosci. 2017;71:518–29.
Mertens J, Wang Q-W, Kim Y, Yu DX, Pham S, Yang B, et al. Differential responses to lithium in hyperexcitable neurons from patients with bipolar disorder. Nature. 2015;527:95–9.
Sequeira A, Martin MV, Rollins B, Moon EA, Bunney WE, Macciardi F, et al. Mitochondrial mutations and polymorphisms in psychiatric disorders. Front Genet. 2012;3:103. https://doi.org/10.3389/fgene.2012.00103.
Craddock N, Jones I. Genetics of bipolar disorder. J Med Genet. 1999;36:585–94.
Zuk O, Hechter E, Sunyaev SR, Lander ES. The mystery of missing heritability: genetic interactions create phantom heritability. Proc Natl Acad Sci USA. 2012;109:1193–8.
Visscher PM, Hill WG, Wray NR. Heritability in the genomics era—concepts and misconceptions. Nat Rev Genet. 2008;9:255–66.
Song J, Bergen SE, Kuja-Halkola R, Larsson H, Landén M, Lichtenstein P. Bipolar disorder and its relation to major psychiatric disorders: a family-based study in the Swedish population. Bipolar Disord. 2014;17:184–93.
Yang J, Lee SH, Goddard ME, Visscher PM. GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet. 2011;88:76–82.
Stahl E, Forstner A, McQuillin A, Ripke S, Bipolar Disorder Working Group of the Psychiatric Genetics Consortium, Ophoff R. et al. Genomewide association study identifies 30 loci associated with bipolar disorder. Nat Genet. 2019;51:793–803.
Girirajan S, Rosenfeld JA, Cooper GM, Antonacci F, Siswara P, Itsara A, et al. A recurrent 16p12. 1 microdeletion supports a two-hit model for severe developmental delay. Nat Genet. 2010;42:203–9.
McGue M, Gottesman II, Rao DC. The transmission of schizophrenia under a multifactorial threshold model. Am J Hum Genet. 1983;35:1161–78.
Feder A, Nestler EJ, Charney DS. Psychobiology and molecular genetics of resilience. Nat Rev Neurosci. 2009;10:446–57.
McGrath LM, Cornelis MC, Lee PH, Robinson EB, Duncan LE, Barnett JH, et al. Genetic predictors of risk and resilience in psychiatric disorders: a cross-disorder genome-wide association study of functional impairment in major depressive disorder, bipolar disorder, and schizophrenia. Am J Med Genet Part B. 2013;162B:779–88.
Boyle EA, Li YI, Pritchard JK. An expanded view of complex traits: from polygenic to omnigenic. Cell. 2017;169:1177–86.
Hosang GM, Fisher HL, Cohen-Woods S, McGuffin P, Farmer AE. Stressful life events and catechol-O-methyl-transferase (COMT) gene in bipolar disorder. Depress Anxiety. 2017;34:419–26.
Oliveira J, Etain B, Lajnef M, Hamdani N, Bennabi M, Bengoufa D, et al. Combined effect of TLR2 gene polymorphism and early life stress on the age at onset of bipolar disorders. PloS ONE. 2015;10:e0119702. https://doi.org/10.1371/journal.pone.0119702.
Hosang GM, Uher R, Keers R, Cohen-Woods S, Craig I, Korszun A, et al. Stressful life events and the brain-derived neurotrophic factor gene in bipolar disorder. J Affect Disord. 2010;125:345–9.
Miller S, Hallmayer J, Wang PW, Hill SJ, Johnson SL, Ketter TA. Brain-derived neurotrophic factor val66met genotype and early life stress effects upon bipolar course. J Psychiatr Res. 2013;47:252–8.
Zeni CP, Mwangi B, Cao B, Hasan KM, Walss-Bass C, Zunta-Soares G, et al. Interaction between BDNF rs6265 Met allele and low family cohesion is associated with smaller left hippocampal volume in pediatric bipolar disorder. J Affect Disord. 2016;189:94–7.
Bulik-Sullivan B, Finucane HK, Anttila V, Gusev A, Day FR, Loh P-R, et al. An atlas of genetic correlations across human diseases and traits. Nat Genet. 2015;47:1236–41.
Middeldorp CM, de Moor MH, McGrath LM, Gordon SD, Blackwood DH, Costa PT, et al. The genetic association between personality and major depression or bipolar disorder. A polygenic score analysis using genome-wide association data. Transl Psychiatry. 2011;1:e50. https://doi.org/10.1038/tp.2011.45.
Huang J, Perlis RH, Lee PH, Rush AJ, Fava M, Sachs GS, et al. Cross-disorder genomewide analysis of schizophrenia, bipolar disorder, and depression. Am J Psychiatry. 2010;167:1254–63.
Weber H, Kittel-Schneider S, Gessner A, Domschke K, Neuner M, Jacob CP, et al. Cross-disorder analysis of bipolar risk genes: further evidence of DGKH as a risk gene for bipolar disorder, but also unipolar depression and adult ADHD. Neuropsychopharmacology. 2011;36:2076–85.
O’Brien HE, Hannon E, Hill MJ, Toste CC, Robertson MJ, Morgan JE, et al. Expression quantitative trait loci in the developing human brain and their enrichment in neuropsychiatric disorders. Genome Biol. 2018;19:194. https://doi.org/10.1186/s13059-018-1567-1.
Power RA, Steinberg S, Bjornsdottir G, Rietveld CA, Abdellaoui A, Nivard MM, et al. Polygenic risk scores for schizophrenia and bipolar disorder predict creativity. Nat Neurosci. 2015;18:953–5.
Kyaga S, Lichtenstein P, Boman M, Landén M. Bipolar disorder and leadership–a total population study. Acta Psychiatr Scand. 2015;131:111–9.
Song J, Bergen SE, Di Florio A, Karlsson R, Charney A, Ruderfer DM, et al. Genome-wide association study identifies SESTD1 as a novel risk gene for lithium-responsive bipolar disorder. Mol Psychiatry. 2016;21:1290–7.
Anghelescu I, Dettling M. Variant GADL1 and response to lithium in bipolar I disorder. N Engl J Med. 2014;370.
Ikeda M, Kondo K, Iwata N. Variant GADL1 and response to lithium in bipolar I disorder. N. Engl J Med. 2014;370:1856–7.
Lee CS, Cheng AT. Variant GADL1 and response to lithium in bipolar I disorder. N Engl J Med. 2014;370:1859–60.
Chung W-H, Hung S-I, Hong H-S, Hsih M-S, Yang L-C, Ho H-C, et al. Medical genetics: a marker for Stevens–Johnson syndrome. Nature. 2004;428:486.
McCormack M, Alfirevic A, Bourgeois S, Farrell JJ, Kasperavičiūtė D, Carrington M, et al. HLA-A* 3101 and carbamazepine-induced hypersensitivity reactions in Europeans. N Engl J Med. 2011;364:1134–43.
Li X, Yu K, Mei S, Huo J, Wang J, Zhu Y, et al. HLA-B*1502 increases the risk of phenytoin or lamotrigine induced stevens-johnson syndrome/toxic epidermal necrolysis: evidence from a meta-analysis of nine case-control studies. Drug Res Stuttg. 2014. https://doi.org/10.1055/s-0034-1375684. 28 May 2014.
Amstutz U, Shear NH, Rieder MJ, Hwang S, Fung V, Nakamura H, et al. Recommendations for HLA-B*15:02 and HLA-A*31:01 genetic testing to reduce the risk of carbamazepine-induced hypersensitivity reactions. Epilepsia. 2014;55:496–506.
Charney AW, Ruderfer DM, Stahl EA, Moran JL, Chambert K, Belliveau RA, et al. Evidence for genetic heterogeneity between clinical subtypes of bipolar disorder. Transl Psychiatry. 2017;7:e993. https://doi.org/10.1038/tp.2016.242.
Allardyce J, Leonenko G, Hamshere M, Pardinas A, Forty L, Knott S, et al. Association Between Schizophrenia-Related Polygenic Liability and the Occurrence and Level of Mood-Incongruent Psychotic Symptoms in Bipolar Disorder. JAMA Psychiatry. 2018;75:28–35.
Mathew I, Gardin TM, Tandon N, Eack S, Francis AN, Seidman LJ, et al. Medial temporal lobe structures and hippocampal subfields in psychotic disorders: findings from the Bipolar-Schizophrenia Network on Intermediate Phenotypes (B-SNIP) study. JAMA Psychiatry. 2014;71:769–77.
Emsell L, Leemans A, Langan C, Van Hecke W, Barker GJ, McCarthy P, et al. Limbic and callosal white matter changes in euthymic bipolar I disorder: an advanced diffusion magnetic resonance imaging tractography study. Biol Psychiatry. 2013;73:194–201.
Liu X, Akula N, Skup M, Brotman MA, Leibenluft E, McMahon FJ. A genome-wide association study of amygdala activation in youths with and without bipolar disorder. J Am Acad Child Adolesc Psychiatry. 2010;49:33–41.
Meda SA, Ruano G, Windemuth A, O’Neil K, Berwise C, Dunn SM, et al. Multivariate analysis reveals genetic associations of the resting default mode network in psychotic bipolar disorder and schizophrenia. Proc Natl Acad Sci USA. 2014;111:E2066–75.
Fears SC, Service SK, Kremeyer B, Araya C, Araya X, Bejarano J, et al. Multisystem component phenotypes of bipolar disorder for genetic investigations of extended pedigrees. JAMA Psychiatry. 2014;71:375–87.
Cardenas SA, Kassem L, Brotman MA, Leibenluft E, McMahon FJ. Neurocognitive functioning in euthymic patients with bipolar disorder and unaffected relatives: a review of the literature. Neurosci Biobehav Rev. 2016;69:193–215.
Glahn DC, Winkler AM, Kochunov P, Almasy L, Duggirala R, Carless MA, et al. Genetic control over the resting brain. Proc Natl Acad Sci USA. 2010;107:1223–8.
Tamminga CA, Pearlson GD, Stan AD, Gibbons RD, Padmanabhan J, Keshavan M, et al. Strategies for advancing disease definition using biomarkers and genetics: the bipolar and schizophrenia network for intermediate phenotypes. Biol Psychiatry Cogn Neurosci Neuroimaging. 2017;2:20–7.
Muffat J, Li Y, Jaenisch R. CNS disease models with human pluripotent stem cells in the CRISPR age. Curr Opin Cell Biol. 2016;43:96–103.
O’Shea KS, McInnis MG. Induced pluripotent stem cell (iPSC) models of bipolar disorder. Neuropsychopharmacol Publ Am Coll Neuropsychopharmacol. 2015;40:248–9.
O’shea KS, McInnis MG. Neurodevelopmental origins of bipolar disorder: iPSC models. Mol Cell Neurosci. 2016;73:63–83.
Stern S, Santos R, Marchetto MC, Mendes APD, Rouleau GA, Biesmans S, et al. Neurons derived from patients with bipolar disorder divide into intrinsically different sub-populations of neurons, predicting the patients’ responsiveness to lithium. Mol Psychiatry. 2017. https://doi.org/10.1038/mp.2016.260.
Schulze TG, McMahon FJ. Defining the phenotype in human genetic studies: forward genetics and reverse phenotyping. Hum Hered. 2004;58:131–8.
Stessman HA, Bernier R, Eichler EE. A genotype-first approach to defining the subtypes of a complex disease. Cell. 2014;156:872–7.
Raznahan A. Genetics-first approaches in biological psychiatry. Biol Psychiatry. 2018;84:234–5.
Geschwind DH, State MW. Gene hunting in autism spectrum disorder: on the path to precision medicine. Lancet Neurol. 2015;14:1109–20.
Stefansson H, Meyer-Lindenberg A, Steinberg S, Magnusdottir B, Morgen K, Arnarsdottir S, et al. CNVs conferring risk of autism or schizophrenia affect cognition in controls. Nature. 2014;505:361–6.
Raznahan A, Cutter W, Lalonde F, Robertson D, Daly E, Conway GS, et al. Cortical anatomy in human X monosomy. NeuroImage. 2010;49:2915–23.
Hoeffding LK, Trabjerg BB, Olsen L, Mazin W, Sparsø T, Vangkilde A, et al. Risk of psychiatric disorders among individuals with the 22q11. 2 deletion or duplication: a Danish nationwide, register-based study. JAMA Psychiatry. 2017;74:282–90.
Gur R, Yi J, Tang S, Calkins M, Moore T, Schmitt J, et al. Psychosis risk in 22q11. 2 deletion syndrome: findings from the Philadelphia sample. Eur Neuropsychopharmacol. 2017;27:S480.
Sahoo T, Theisen A, Rosenfeld JA, Lamb AN, Ravnan JB, Schultz RA, et al. Copy number variants of schizophrenia susceptibility loci are associated with a spectrum of speech and developmental delays and behavior problems. Genet Med. 2011;13:868–80.
D’Angelo D, Lebon S, Chen Q, Martin-Brevet S, Snyder LG, Hippolyte L, et al. Defining the effect of the 16p11.2 duplication on cognition, behavior, and medical comorbidities. JAMA Psychiatry. 2016;73:20–30.
Tobe BTD, Brandel MG, Nye JS, Snyder EY. Implications and limitations of cellular reprogramming for psychiatric drug development. Exp Mol Med. 2013;45:e59. https://doi.org/10.1038/emm.2013.124.
Schadt EE, Buchanan S, Brennand KJ, Merchant KM. Evolving toward a human-cell based and multiscale approach to drug discovery for CNS disorders. Front Pharmacol. 2014;5:252. https://doi.org/10.3389/fphar.2014.00252.
Breen G, Li Q, Roth BL, O’Donnell P, Didriksen M, Dolmetsch R, et al. Translating genome-wide association findings into new therapeutics for psychiatry. Nat Neurosci. 2016;19:1392–6.
Chatterjee N, Shi J, García-Closas M. Developing and evaluating polygenic risk prediction models for stratified disease prevention. Nat Rev Genet. 2016;17:392–406.
Zhang Y, Qi G, Park J-H, Chatterjee N. Estimation of complex effect-size distributions using summary-level statistics from genome-wide association studies across 32 complex traits. Nat Genet. 2018;50:1318–26.
Tan RYY, Markus HS. Genetics and genomics of stroke. In: Kumar D, Elliott P, editors. Cardiovascular genetics and genomics. Principles and clinical practice, pp 695–722. Cham: Springer International Publishing; 2018.
McMahon FJ, Akula N, Schulze TG, Muglia P, Tozzi F, Detera-Wadleigh SD, et al. Meta-analysis of genome-wide association data identifies a risk locus for major mood disorders on 3p21.1. Nat Genet. 2010;42:128–31.
Wang K-S, Liu X-F, Aragam N. A genome-wide meta-analysis identifies novel loci associated with schizophrenia and bipolar disorder. Schizophr Res. 2010;124:192–9.
Fabbri C, Serretti A. Pharmacogenetics of major depressive disorder: top genes and pathways toward clinical applications. Curr Psychiatry Rep. 2015;17:1–11.
Green EK, Grozeva D, Forty L, Gordon-Smith K, Russell E, Farmer A, et al. Association at SYNE1 in both bipolar disorder and recurrent major depression. Mol Psychiatry. 2013;18:614–7.
Kerner B, Lambert CG, Muthen BO. Genome-wide association study in bipolar patients stratified by co-morbidity. PLoS ONE. 2011;6:e28477. https://doi.org/10.1371/journal.pone.0028477.
Jiang Y, Zhang H. Propensity score-based nonparametric test revealing genetic variants underlying bipolar disorder. Genet Epidemiol. 2011;35:125–32.
Cichon S, Mühleisen TW, Degenhardt FA, Mattheisen M, Miró X, Strohmaier J, et al. Genome-wide association study identifies genetic variation in neurocan as a susceptibility factor for bipolar disorder. Am J Hum Genet. 2011;88:372–81.
Funding
This study was supported by the Intramural Research Program of the NIMH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gordovez, F.J.A., McMahon, F.J. The genetics of bipolar disorder. Mol Psychiatry 25, 544–559 (2020). https://doi.org/10.1038/s41380-019-0634-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-019-0634-7
- Springer Nature Limited
This article is cited by
-
A bipolar disorder-associated missense variant alters adenylyl cyclase 2 activity and promotes mania-like behavior
Molecular Psychiatry (2024)
-
Genetic regulation of human brain proteome reveals proteins implicated in psychiatric disorders
Molecular Psychiatry (2024)
-
Associations of altered leukocyte DDR1 promoter methylation and childhood trauma with bipolar disorder and suicidal behavior in euthymic patients
Molecular Psychiatry (2024)
-
Progress and Implications from Genetic Studies of Bipolar Disorder
Neuroscience Bulletin (2024)
-
Esplorando il legame genetico tra disfunzione tiroidea e disturbi psichiatrici comuni: una comorbidità ormonale specifica o una comorbidità autoimmune generale
L'Endocrinologo (2024)